Abstract
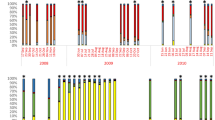
Genetic diversity is what selection acts on, thus shaping the adaptive potential of populations. We studied micro-evolutionary patterns of the key planktonic diatom Pseudo-nitzschia multistriata at a long-term sampling site over 2 consecutive years by genotyping isolates with 22 microsatellite markers. We show that both sex and vegetative growth interplay in shaping intraspecific diversity. We document a brief but massive demographic and clonal expansion driven by strains of the same mating type. The analysis of an extended data set (6 years) indicates that the genetic fingerprint of P. multistriata changed over time with a nonlinear pattern, with intermittent periods of weak and strong diversification related to the temporary predominance of clonal expansions over sexual recombination. These dynamics, rarely documented for phytoplankton, contribute to the understanding of bloom formation and of the mechanisms that drive microevolution in diatoms.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Arnaud-Haond S, Belkhir K . (2007). GENCLONE: a computer program to analyse genotypic data, test for clonality and describe spatial clonal organization. Mol Ecol Resour 7: 15–17.
Basu S, Patil S, Mapleson D, Russo M, Vitale L, Fevola C et al. (2017). Finding a partner in the ocean: molecular and evolutionary bases of the response to sexual cues in a planktonic diatom. New Phytol 215: 140–156.
Behrenfeld MJ, Boss E, Siegel DA, Shea DM . (2005). Carbon-based ocean productivity and phytoplankton physiology from space. Global Biogeochem Cycles 19: GB1006.
Bengtsson BO . (2003). Genetic variation in organisms with sexual and asexual reproduction. J Evol Biol 16: 189–199.
Berdalet E, Fleming LE, Gowen R, Davidson K, Hess P, Backer LC et al. (2016). Marine harmful algal blooms, human health and wellbeing: challenges and opportunities in the 21st century. J Mar Biol Assoc U K 96: 61–91.
D’Alelio D, Amato A, WHCF Kooistra, Procaccini G, Casotti R, Montresor M . (2009a). Internal transcribed spacer polymorphism in Pseudo-nitzschia multistriata (Bacillariophyceae) in the Gulf of Naples: recent divergence or intraspecific hybridization? Protist 160: 9–20.
D’Alelio D, Amato A, Luedeking A, Montresor M . (2009b). Sexual and vegetative phases in the planktonic diatom Pseudo-nitzschia multistriata. Harmful Algae 8: 225–232.
D’Alelio D, Gandolfi A . (2012). Recombination signals in the rpoC1 gene indicate gene-flow between Planktothrix (Cyanoprokaryota) species. J Phycol 48: 1424–1432.
D’Alelio D, Libralato S, Wyatt T, Ribera d’Alcalà M . (2016). Ecological-network models link diversity, structure and function in the plankton food-web. Sci Rep 6: 21806.
D’Alelio D, Ribera d’Alcalà M, Dubroca L, Sarno D, Zingone A, Montresor M . (2010). The time for sex: a biennial life cycle in a marine planktonic diatom. Limnol Oceanogr 55: 106–114.
D’Alelio D, Ruggiero MV . (2015). Interspecific plastidial recombination in the diatom genus Pseudo-nitzschia. J Phycol 51: 1024–1028.
D’Alelio D, Salmaso N, Gandolfi A . (2013). Frequent recombination shapes the epidemic population structure of Planktothrix (Cyanoprokaryota) in Italian subalpine lakes. J Phycol 49: 1107–1117.
De Bruin A, Ibelings BW, Rijkeboer M, Brehm M, van Donk E . (2004). Genetic variation in Asterionella formosa (Bacillariophyceae): is it linked to frequent epidemics of host-specific parasitic fungi? J Phycol 40: 823–830.
Dia A, Guillou L, Mauger S, Bigeard E, Marie D, Valero M et al. (2014). Spatiotemporal changes in the genetic diversity of harmful algal blooms caused by the toxic dinoflagellate Alexandrium minutum. Mol Ecol 23: 549–560.
Erdner DL, Richlen M, McCauley LAR, Anderson DM . (2011). Diversity and dynamics of a widespread bloom of the toxic dinoflagellate Alexandrium fundyense. PLoS One 6: e22965.
Godhe A, Kremp A, Montresor M . (2014). Genetic and microscopic evidence for sexual reproduction in the centric diatom Skeletonema marinoi. Protist 165: 401–416.
Godhe A, Sjöqvist C, Sildever S, Sefbom J, Harðardóttir S, Bertos-Fortis M et al. (2016). Physical barriers and environmental gradients cause spatial and temporal genetic differentiation of an extensive algal bloom. J Biogeogr 43: 1130–1142.
Gsell AS, de Senerpont Domis LN, Verhoeven KJF, van Donk E, Ibelings BW . (2013). Chytrid epidemics may increase genetic diversity of a diatom spring-bloom. ISME J 7: 2057–2059.
Guillard RRL . (1975). Culture of phytoplankton for feeding marine invertebrates. In: Culture of marine invertebrate animals. Springer, pp 29–60.
Härnström K, Ellegaard M, Andersen TJ, Godhe A . (2011). Hundred years of genetic structure in a sediment revived diatom population. Proc Natl Acad Sci USA 108: 4252–4257.
Haubold B, Hudson RR . (2000). LIAN 3.0: detecting linkage disequilibrium in multilocus data. Bioinformatics 16: 847.
Krueger-Hadfield SA, Balestreri C, Schroeder J, Highfield A, Helaouët P, Allum J et al. (2014). Genotyping an Emiliania huxleyi (prymnesiophyceae) bloom event in the North Sea reveals evidence of asexual reproduction. Biogeosciences 11: 5215–5234.
MacLeod A, Tweedie A, Welburn SC, Maudlin I, Turner CMR, Tait A . (2000). Minisatellite marker analysis of Trypanosoma brucei: Reconciliation of clonal, panmictic, and epidemic population genetic structures. Proc Natl Acad Sci USA 97: 13442–13447.
Maiden MCJ . (2006). Multilocus sequence typing of bacteria. Annu Rev Microbiol 60: 561–588.
McCabe RM, Hickey BM, Kudela RM, Lefebvre KA, Adams NG, Bill BD et al. (2016). An unprecedented coastwide toxic algal bloom linked to anomalous ocean conditions. Geophys Res Lett 43: 10,366–10,376.
Peakall ROD, Smouse PE . (2006). GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6: 288–295.
Pemberton TJ, DeGiorgio M, Rosenberg NA . (2013). Population structure in a comprehensive genomic data set on human microsatellite variation. G3 3: 891–907.
Raymond M, Rousset F . (1995). GENEPOP (Version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86: 248.
Rengefors K, Kremp A, Reusch TBH, Wood AM . (2017). Genetic diversity and evolution in eukaryotic phytoplankton: revelations from population genetic studies. J Plankton Res 39: 165–179.
Ribera d’Alcalà M, Conversano F, Corato F, Licandro P, Mangoni O, Marino D et al. (2004). Seasonal patterns in plankton communities in a pluriannual time series at a coastal Mediterranean site (Gulf of Naples): an attempt to discern recurrences and trends. Sci Mar 68: 65–83.
Richlen ML, Erdner DL, McCauley LAR, Liberal K, Anderson DM . (2012). Extensive genetic diversity and rapid population differentiation during blooms of Alexandrium fundyense (dinophyceae) in an isolated salt pond on cape cod, MA, USA. Ecol Evol 2: 2588–2599.
Rynearson TA, Armbrust EV . (2005). Maintenance of clonal diversity during a spring bloom of the centric diatom Ditylum brightwellii. Mol Ecol 14: 1631–1640.
Saravanan V, Godhe A . (2010). Genetic heterogeneity and physiological variation among seasonally separated clones of Skeletonema marinoi (Bacillariophyceae) in the Gullmar Fjord, Sweden. Eur J Phycol 45: 177–190.
Schurko AM, Logsdon JM, Eads BD . (2009a). Meiosis genes in Daphnia pulex and the role of parthenogenesis in genome evolution. BMC Evol Biol 9: 78.
Schurko AM, Neiman M, Logsdon JM . (2009b). Signs of sex: what we know and how we know it. Trends Ecol Evol 24: 208–217.
Smayda TJ . (1997). What is a bloom? Limnol Oceanogr 42: 1132–1136.
Smith JM, Smith NH, O’Rourke M, Spratt BG . (1993). How clonal are bacteria? Proc Natl Acad Sci USA 90: 4384–4388.
Speijer D, Lukeš J, Eliáš M . (2015). Sex is a ubiquitous, ancient, and inherent attribute of eukaryotic life. Proc Natl Acad Sci USA 112: 8827–8834.
Stoeckel S, Masson JP . (2014). The exact distributions of FISunder partial asexuality in small finite populations with mutation. PLoS One 9: e85228.
Tesson SVM, Borra M, Kooistra W, Procaccini G . (2011). Microsatellite primers in the planktonic diatom Pseudo-nitzschia multistriata (Bacillariophyceae). Am J Bot 98: E33–E35.
Tesson SVM, Legrand C, van Oosterhout C, Montresor M, Kooistra WHCF, Procaccini G . (2013). Mendelian inheritance pattern and high mutation rates of microsatellite alleles in the diatom Pseudo-nitzschia multistriata. Protist 164: 89–100.
Tesson SVM, Montresor M, Procaccini G, Kooistra WHCF . (2014). Temporal changes in population structure of a marine planktonic diatom. PLoS One 9: e114984.
Tibayrenc M, Ayala FJ . (2012). Reprodaductive clonality of pathogens: a perspective on pathogenic viruses, bacteria, fungi, and parasitic protozoa. Proc Natl Acad Sci USA 109: E3305–E3313.
Vanormelingen P, Evans KM, Mann DG, Lance S, Debeer AE, D’Hondt S et al. (2015). Genotypic diversity and differentiation among populations of two benthic freshwater diatoms as revealed by microsatellites. Mol Ecol 24: 4433–4448.
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P . (2004). MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4: 535–538.
von Dassow P, Montresor M . (2010). Unveiling the mysteries of phytoplankton life cycles: patterns and opportunities behind complexity. J Plankton Res 33: 3–12.
Weedall GD, Hall N . (2015). Sexual reproduction and genetic exchange in parasitic protists. Parasitology 142: S120–S127.
Wyatt T . (2012). Margalef’s mandala and phytoplankton bloom strategies. Deep Sea Res Part II Top Stud Oceanogr 101: 32–49.
Acknowledgements
We thank Carmen Minucci for technical help in genotyping; Eleonora Scalco and Christophe Legrand for providing additional data on mating types; Diana Sarno for providing cell abundance data for P. multistriata; Augusto Passarelli for providing physical-chemical data at LTER-MC; the Molecular Biology and Bioinformatic Unit for sequencing and the Monitoring and Environmental Data Unit (MEDA) for sampling. The study was funded by Flagship Project RITMARE—The Italian Research for the Sea—coordinated by the Italian National Research Council and funded by the Italian Ministry of Education, University and Research within the National Research Program 2011–2013; LV was supported by a PhD fellowship from Stazione Zoologica Anton Dohrn (SZN).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Ruggiero, M., D'Alelio, D., Ferrante, M. et al. Clonal expansion behind a marine diatom bloom. ISME J 12, 463–472 (2018). https://doi.org/10.1038/ismej.2017.181
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/ismej.2017.181
This article is cited by
-
An INDEL genomic approach to explore population diversity of phytoplankton
BMC Genomics (2024)
-
Multiannual patterns of genetic structure and mating type ratios highlight the complex bloom dynamics of a marine planktonic diatom
Scientific Reports (2024)
-
Machine learning identifies a strong association between warming and reduced primary productivity in an oligotrophic ocean gyre
Scientific Reports (2020)
-
Metabolomics analysis of Pseudomonas chlororaphis JK12 algicidal activity under aerobic and micro-aerobic culture condition
AMB Express (2018)
-
MRP3 is a sex determining gene in the diatom Pseudo-nitzschia multistriata
Nature Communications (2018)