Abstract
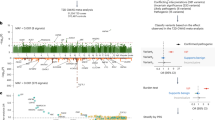
Several genetic loci (JAZF1, CDC123/CAMK1D, TSPAN8/LGR5, ADAMTS9, VEGFA and HHEX-IDE) were identified to be significantly related to the risk of type 2 diabetes and quantitative metabolic traits in European populations. Here, we aimed to evaluate the impacts of these novel loci on type 2 diabetes risk in a population-based case–control study of Han Chinese (1912 cases and 2041 controls). We genotyped 13 single-nucleotide polymorphisms (SNPs) in/near these genes and examined the differences in allele/genotype frequency between cases and controls. We found that both IDE rs11187007 and HHEX rs1111875 were associated with type 2 diabetes risk (for both variants: odds ratio (OR)=1.15, 95% confidence interval (CI) 1.04–1.28, P=0.009). In a meta-analysis where we pooled our data with the three previous studies conducted in East Asians, we found that the variants of JAZF1 rs864745 (1.09 (1.03–1.16); P=3.49 × 10−3) and TSPAN8/LGR5 rs7961581 (1.11(1.05–1.17); P=1.89 × 10−4) were significantly associated with type 2 diabetes risk. In addition, the meta-analysis (7207 cases and 8260 controls) also showed that HHEX rs1111875 did have effects on type 2 diabetes in Chinese population (OR=1.15(1.10–1.21); P=1.93 × 10−8). This large population-based study and meta-analysis further confirmed the modest effects of the JAZF1, TSPAN8/LGR5 and HHEX-IDE loci on type 2 diabetes in Chinese and other East Asians.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Prokopenko, I., McCarthy, M. I. & Lindgren, C. M. Type 2 diabetes: new genes, new understanding. Trends Genet. 24, 613–621 (2008).
Stancakova, A., Kuulasmaa, T., Paananen, J., Jackson, A. U., Bonnycastle, L. L., Collins, F. S. et al. Association of 18 confirmed susceptibility loci for type 2 diabetes with indices of insulin release, proinsulin conversion, and insulin sensitivity in 5327 nondiabetic Finnish men. Diabetes 58, 2129–2136 (2009).
Hu, C., Zhang, R., Wang, C., Wang, J., Ma, X., Lu, J. et al. PPARG, KCNJ11, CDKAL1, CDKN2A-CDKN2B, IDE-KIF11-HHEX, IGF2BP2 and SLC30A8 are associated with type 2 diabetes in a Chinese population. PLoS ONE 4, e7643 (2009).
Liu, Y., Zhou, D. Z., Zhang, D., Chen, Z., Zhao, T., Zhang, Z. et al. Variants in KCNQ1 are associated with susceptibility to type 2 diabetes in the population of mainland China. Diabetologia 52, 1315–1321 (2009).
Tan, J. T., Ng, D. P., Nurbaya, S., Ye, S., Lim, X. L., Leong, H. et al. Polymorphisms identified through genome-wide association studies and their associations with type 2 diabetes in Chinese, Malays, and Asian-Indians in Singapore. J. Clin. Endocrinol. Metab. 95, 390–397 (2009).
Wu, Y., Li, H., Loos, R. J., Yu, Z., Ye, X., Chen, L. et al. Common variants in CDKAL1, CDKN2A/B, IGF2BP2, SLC30A8, and HHEX/IDE genes are associated with type 2 diabetes and impaired fasting glucose in a Chinese Han population. Diabetes 57, 2834–2842 (2008).
Zeggini, E., Scott, L. J., Saxena, R., Voight, B. F., Marchini, J. L., Hu, T. et al. Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat. Genet. 40, 638–645 (2008).
Boesgaard, T. W., Gjesing, A. P., Grarup, N., Rutanen, J., Jansson, P. A., Hribal, M. L. et al. Variant near ADAMTS9 known to associate with type 2 diabetes is related to insulin resistance in offspring of type 2 diabetes patients—EUGENE2 study. PLoS ONE 4, e7236 (2009).
Grarup, N., Andersen, G., Krarup, N. T., Albrechtsen, A., Schmitz, O., Jorgensen, T. et al. Association testing of novel type 2 diabetes risk alleles in the JAZF1, CDC123/CAMK1D, TSPAN8, THADA, ADAMTS9, and NOTCH2 loci with insulin release, insulin sensitivity, and obesity in a population-based sample of 4516 glucose-tolerant middle-aged Danes. Diabetes 57, 2534–2540 (2008).
Simonis-Bik, A. M., Nijpels, G., van Haeften, T. W., Houwing-Duistermaat, J. J., Boomsma, D. I., Reiling, E. et al. Gene variants in the novel type 2 diabetes loci CDC123/CAMK1D, THADA, ADAMTS9, BCL11A and MTNR1B affect different aspects of pancreatic beta cell function. Diabetes 59, 293–301 (2009).
Omori, S., Tanaka, Y., Horikoshi, M., Takahashi, A., Hara, K., Hirose, H. et al. Replication study for the association of new meta-analysis-derived risk loci with susceptibility to type 2 diabetes in 6244 Japanese individuals. Diabetologia. 52, 1554–1560 (2009).
Sanghera, D. K., Been, L., Ortega, L., Wander, G. S., Mehra, N. K., Aston, C. E. et al. Testing the association of novel meta-analysis-derived diabetes risk genes with type II diabetes and related metabolic traits in Asian Indian Sikhs. J. Hum. Genet. 54, 162–168 (2009).
Takeuchi, F., Serizawa, M., Yamamoto, K., Fujisawa, T., Nakashima, E., Ohnaka, K. et al. Confirmation of multiple risk loci and genetic impacts by a genome-wide association study of type 2 diabetes in the japanese population. Diabetes 58, 1690–1699 (2009).
Meigs, J. B., Panhuysen, C. I., Myers, R. H., Wilson, P. W. & Cupples, L. A. A genome-wide scan for loci linked to plasma levels of glucose and HbA(1c) in a community-based sample of Caucasian pedigrees: the Framingham Offspring Study. Diabetes 51, 833–840 (2002).
Saxena, R., Voight, B. F., Lyssenko, V., Burtt, N. P., de Bakker, P. I., Chen, H. et al. Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science (New York, N.Y.) 316, 1331–1336 (2007).
Scott, L. J., Mohlke, K. L., Bonnycastle, L. L., Willer, C. J., Li, Y., Duren, W. L. et al. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science (New York, N.Y.) 316, 1341–1345 (2007).
Sladek, R., Rocheleau, G., Rung, J., Dina, C., Shen, L., Serre, D. et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445, 881–885 (2007).
WTCCC. Genome-wide association study of 14,000 cases of seven common diseases and 3000 shared controls. Nature 447, 661–678 (2007).
van Vliet-Ostaptchouk, J. V., Onland-Moret, N. C., van Haeften, T. W., Franke, L., Elbers, C. C., Shiri-Sverdlov, R. et al. HHEX gene polymorphisms are associated with type 2 diabetes in the Dutch Breda cohort. Eur. J. Hum. Genet. 16, 652–656 (2008).
Liu, Y., Yu, L., Zhang, D., Chen, Z., Zhou, D. Z., Zhao, T. et al. Positive association between variations in CDKAL1 and type 2 diabetes in Han Chinese individuals. Diabetologia 51, 2134–2137 (2008).
Zhou, D., Zhang, D., Liu, Y., Zhao, T., Chen, Z., Liu, Z. et al. The E23K variation in the KCNJ11 gene is associated with type 2 diabetes in Chinese and East Asian population. J. Hum. Genet. 54, 433–435 (2009).
Groves, C. J., Wiltshire, S., Smedley, D., Owen, K. R., Frayling, T. M., Walker, M. et al. Association and haplotype analysis of the insulin-degrading enzyme (IDE) gene, a strong positional and biological candidate for type 2 diabetes susceptibility. Diabetes 52, 1300–1305 (2003).
Gu, H. F., Efendic, S., Nordman, S., Ostenson, C. G., Brismar, K., Brookes, A. J. et al. Quantitative trait loci near the insulin-degrading enzyme (IDE) gene contribute to variation in plasma insulin levels. Diabetes 53, 2137–2142 (2004).
Mueller, J. C., Riemenschneider, M., Schoepfer-Wendels, A., Gohlke, H., Konta, L., Friedrich, P. et al. Weak independent association signals between IDE polymorphisms, Alzheimer's disease and cognitive measures. Neurobiol. Aging 28, 727–734 (2007).
Steinthorsdottir, V., Thorleifsson, G., Reynisdottir, I., Benediktsson, R., Jonsdottir, T., Walters, G. B. et al. A variant in CDKAL1 influences insulin response and risk of type 2 diabetes. Nat. Genet. 39, 770–775 (2007).
Zeggini, E., Weedon, M. N., Lindgren, C. M., Frayling, T. M., Elliott, K. S., Lango, H. et al. Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science (New York, NY) 316, 1336–1341 (2007).
Shi, Y. Y. & He, L. SHEsis, a powerful software platform for analyses of linkage disequilibrium, haplotype construction, and genetic association at polymorphism loci. Cell Res. 15, 97–98 (2005).
Ng, M. C., Park, K. S., Oh, B., Tam, C. H., Cho, Y. M., Shin, H. D. et al. Implication of genetic variants near TCF7L2, SLC30A8, HHEX, CDKAL1, CDKN2A/B, IGF2BP2, and FTO in type 2 diabetes and obesity in 6,719 Asians. Diabetes 57, 2226–2233 (2008).
Florez, J. C., Jablonski, K. A., McAteer, J., Sandhu, M. S., Wareham, N. J., Barroso, I. et al. Testing of diabetes-associated WFS1 polymorphisms in the diabetes prevention program. Diabetologia 51, 451–457 (2008).
Farris, W., Mansourian, S., Chang, Y., Lindsley, L., Eckman, E. A., Frosch, M. P. et al. Insulin-degrading enzyme regulates the levels of insulin, amyloid beta-protein, and the beta-amyloid precursor protein intracellular domain in vivo. Proc. Natl Acad. Sci. U.S.A. 100, 4162–4167 (2003).
Karamohamed, S., Demissie, S., Volcjak, J., Liu, C., Heard-Costa, N., Liu, J. et al. Polymorphisms in the insulin-degrading enzyme gene are associated with type 2 diabetes in men from the NHLBI Framingham Heart Study. Diabetes 52, 1562–1567 (2003).
Cauchi, S., Proenca, C., Choquet, H., Gaget, S., De Graeve, F., Marre, M. et al. Analysis of novel risk loci for type 2 diabetes in a general French population: the D.E.S.I.R. study. J. Mol. Med. (Berlin, Germany) 86, 341–348 (2008).
Omori, S., Tanaka, Y., Takahashi, A., Hirose, H., Kashiwagi, A., Kaku, K. et al. Association of CDKAL1, IGF2BP2, CDKN2A/B, HHEX, SLC30A8, and KCNJ11 with susceptibility to type 2 diabetes in a Japanese population. Diabetes 57, 791–795 (2008).
Pivovarova, O., Nikiforova, V. J., Pfeiffer, A. F. & Rudovich, N. The influence of genetic variations in HHEX gene on insulin metabolism in the German MESYBEPO cohort. Diabetes Metab. Res. Rev. 25, 156–162 (2009).
Collins, L. L., Lee, Y. F., Heinlein, C. A., Liu, N. C., Chen, Y. T., Shyr, C. R. et al. Growth retardation and abnormal maternal behavior in mice lacking testicular orphan nuclear receptor 4. Proc. Natl Acad. Sci. U.S.A. 101, 15058–15063 (2004).
Acknowledgements
We thank the individuals who participated in the present study. This work was supported by grants from the National Key Technology R&D Program (2006BAI05A05), the S973 Program (2006CB910601, 2007CB947300, 2010CB529600), 863 Program (2006AA02A407), the National Natural Science Foundation of China (30972529), the Opening Project of Shanghai Key Laboratory of Complex Prescription (07DZ22917), the Shanghai–Unilever Research and Development Fund (06SU07007), the Shanghai Leading Academic Discipline Project (B205), the Chinese Nutrition Society (05015), the Shanghai Municipality Science & Technology Commission (05JC14090) and the Chinese Academy of Sciences (KSCX2-YW-R-01).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Zhou, Dz., Liu, Y., Zhang, D. et al. Variations in/nearby genes coding for JAZF1, TSPAN8/LGR5 and HHEX-IDE and risk of type 2 diabetes in Han Chinese. J Hum Genet 55, 810–815 (2010). https://doi.org/10.1038/jhg.2010.117
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/jhg.2010.117
Keywords
This article is cited by
-
TMT-based proteomic and bioinformatic analyses of human granulosa cells from obese and normal-weight female subjects
Reproductive Biology and Endocrinology (2021)
-
Effect of smoking on the association of HHEX (rs5015480) with diabetes among Korean women and heavy smoking men
BMC Medical Genetics (2018)
-
Association between 28 single nucleotide polymorphisms and type 2 diabetes mellitus in the Kazakh population: a case-control study
BMC Medical Genetics (2017)
-
Association of JAZF1 and TSPAN8/LGR5 variants in relation to type 2 diabetes mellitus in a Saudi population
Diabetology & Metabolic Syndrome (2015)


