Abstract
Alterations in transforming growth factor-β signaling, due to a decrease in Smad2 and especially Smad4 expression, has primarily been reported in pancreatic and colorectal cancers, although loss of the chromosomal region 18q21.1, containing the loci of Smad2 and Smad4, is among the most frequent molecular alterations in cervical cancer. The aim of our study was to investigate whether decreased Smad2 and Smad4 protein expression in primary cervical cancers is associated with molecular alterations at 18q21.1, mutations in the functional domains of Smad2 and Smad4 or hypermethylation, and to assess the biological relevance of decreased Smad2 and Smad4 expression. Subsequently, Smad2, Smad4 and p21 protein expression was determined by immunohistochemistry in 117 primary cervical carcinomas, assembled in a tissue array. Smad signaling was shown to be associated with p21 mRNA expression. All the tumors expressed Smad2 or Smad4. Weak cytoplasmic Smad2 or weak cytoplasmic Smad4 expression could not be attributed to loss of heterozygosity at 18q21.1. Despite weak/moderate Smad2 expression and absent nuclear Smad4 expression, the coding regions of the functional MH1 and MH2 domains of Smad2 and Smad4 were unchanged, as assessed by sequence analysis. The Smad4 promoter region was unmethylated in tumor samples with weak/moderate cytoplasmic Smad4 expression. Remarkably, both weak cytoplasmic Smad4 expression and absent nuclear Smad4 expression significantly correlated with poor disease-free (P=0.003 and P=0.003, respectively) and overall 5-year survival (P=0.003 and P=0.010, respectively). Our findings support the hypothesis that Smad4 is a target molecule for functional inactivation in cervical cancer.
Similar content being viewed by others
Main
Cervical cancer is caused by persistent infection with high-risk human papillomavirus (HPV). In addition to infection with HPV, molecular alterations of tumor suppressor genes and/or oncogenes are imperative for the development of cervical cancer. A characteristic molecular alteration associated with malignant progression is the loss of responsiveness to transforming growth factor-β1 (TGF-β1)-induced cell growth inhibition.1, 2 TGF-β1 acts as a double-edged sword; early in tumorigenesis, it operates as a tumor suppressor by inhibiting epithelial cell proliferation and, later in tumorigenesis, it operates as an angiogenic factor and suppresses an efficient anti-tumor immune response. Loss of TGF-β1 responsiveness is thought to be due to alterations in the TGF-β1 signal transduction route.3 A mechanism by which loss of the tumor suppressor function of TGF-β can occur is through inactivation of Smad genes. As transcription factors, Smads are important for inducing the expression of cell cycle inhibitors, such as p15, p21 and p27, thereby facilitating TGF-β-mediated inhibition of cell growth.4
Loss of heterozygosity (LOH) at 18q, the locus of Smad2 and Smad4, is a common mechanism for inactivation of Smad4 in pancreatic tumors.5 In cervical carcinomas, LOH at 18q was observed in 10–37% of cases and has been associated with poor prognosis.6, 7 In addition to LOH, mutations in Smad4 in pancreatic cancer and colorectal cancer, resulting in loss of expression or function, have been frequently reported.5, 8 Also, Smad2 mutations have been found in colorectal cancer.9 For cervical cancer, Smad4 mutations were reported in cervical cancer cell lines10 and a Smad2 mutation in a cervical tumor has been described.11 In addition to LOH and gene mutations, other mechanisms such as CpG methylation of the Smad promoter may play a role in downregulation of Smad expression. Recently, Smad4 promoter methylation was reported in Barrett's esophagus and gastric carcinoma as a mechanism for decreased Smad4 expression.12, 13
The aim of our study was to investigate whether decreased Smad2 and Smad4 protein expression in primary cervical carcinomas was associated with molecular alterations at 18q21.1, mutations in the functional domains of Smad2 and Smad4 or hypermethylation, and to assess the biological relevance of decreased Smad2 and Smad4 expression.
For this purpose, we determined protein expression of Smad2, P-Smad2, Smad4 and p21 by immunohistochemistry on a tissue array of 117 primary cervical cancers. Subsequently, the association between Smad2 and Smad4 protein expression and molecular alterations was investigated by determining LOH status at 18q21.1 (Smad2 and Smad4 loci), the occurrence of mutations in the functional domains of Smad2 and Smad4 and CpG methylation of the Smad4 promoter. Furthermore, to assess the biological relevance of our findings, we correlated the experimental results with clinico-pathological parameters.
Materials and methods
Patient Material
From 117 patients with cervical squamous carcinoma who underwent a radical hysterectomy combined with pelvic lymph adenectomy between 1985 and 2000, formalin-fixed paraffin-embedded tissue blocks were retrieved from the Department of Pathology, Leiden University Medical Center. None of the patients had received radio- or chemotherapy before surgery. The 117 primary cervical carcinomas consisted of 90 squamous cell carcinomas, 19 adenocarcinomas and 8 adenosquamous carcinomas. The use of clinical material was approved by the institutional review board, according to the guidelines of the Dutch Federation of Medical Research Associations. The clinical records of the women, all treated by the same three gynecologic oncologists of the Department of Gynecology, Leiden University Medical Center, with a radical hysterectomy type III for carcinoma of the uterine cervix between 1985 and 2000, were reviewed. The slides of all tumors were examined, using conventional histological sections stained with hematoxylin and eosin, by a trained pathologist.
Immunohistochemistry
Immunohistochemistry on a tissue array was performed on 4 μm sections using aminopropylethoxysilane-coated slides. A representative tumor area was selected by an experienced pathologist. Subsequently, three cores of each tumor were included in the tissue array. Standard immunohistochemical staining was performed as described previously.14 In the case of immunohistochemistry with the Smad2, P-Smad2 and Smad4 antibodies, antigen retrieval was performed with 0.01 M citrate buffer. mAb directed against Smad4 (clone B8; dilution 1:200; Santa Cruz, Santa Cruz, CA, USA),15 P-Smad2 (clone 138D4, dilution 1:50; Cell Signaling Technology, Beverly, CA, USA)16 and p21 (clone EA10, dilution 1:200; Calbiochem, San Diego, CA, USA)17 and a polyclonal antibody against Smad2 (dilution 1:200; Santa Cruz)18 were used. The cellular infiltrate provided an internal positive control (for Smad2 and Smad4). Immunohistochemistry was scored as described previously.19 The intensity of cytoplasmic staining was scored as weak, moderate or strong at low magnification (× 100). In the case of nuclear staining, the percentage of positive tumor cells was determined and divided into three groups: nuclear staining in 0% of tumor cells (absent), nuclear staining in ≥0% of tumor cells and ≤50% of tumor cells (low) or nuclear staining in ≥50% of tumor cells (high). Expression was scored by two independent researchers without knowing the clinical outcome of patients.
Genomic DNA Isolation from Flow-Sorted Tumor Cell Subpopulations
Formalin-fixed, paraffin-embedded cervical carcinomas were trimmed if necessary to remove normal epithelium and cervical intraepithelial neoplasia. The remaining tumor tissue was cut into 50 μ M sections, dewaxed and pretreated with 10 mM sodium citrate buffer (pH 6,4) as antigen retrieval. Dissociation of tissue was performed in a mixture of 0.1% collagenase I-a and 0.1% dispase in RPMI medium without FCS and antibiotics for 30–120 min at 37°C.20 The cells were stained for keratin, vimentin and DNA as described previously.20 Anti-vimentin antibodies applied were 3B4 (2 μg/ml; DAKOCytomation, Glostrup, Denmark) and V9-2b (supernatant diluted 1:5). Monoclonal antibodies used against keratins included MNF116 (2 μg/ml; DAKOCytomation) and Pan-keratin AE1/AE3 (5 μg/ml; Chemicon International Inc., Temecula, CA, USA). The vimentin-negative keratin-positive (V−K+) fraction, which represents the epithelial cell subpopulation, and vimentin-positive keratin-negative (V+K−) fraction, consisting of normal cells, were flow-sorted using a FACS Vantage (BD Biosciences, San Jose, CA, USA). The number of sorted cells varied from 50 000 to 1 000 000 between samples. DNA was extracted as described previously.21
Microsatellite Analysis
Seven Hex-labeled primer pairs of microsatellite markers comprising D18s877, D18s65, D18s460, D18s1137, D18s1118, D18s1109 and D18s844 (Genome Database, http://www.gdb.org) at chromosome 18q were used. The nearest marker for Smad2 is D18S1137 at 0.2 Mb distance and for Smad4, D18S1118 at 2.6 Mb. DNA, isolated from sorted tumor cell subpopulations, was used as source material.20 PCR, electrophoresis and analysis were performed as described previously.22 A reduction of 50% of one allele, equating to a ratio of ≥2 or ≤0.5, was considered as LOH. A reduction of 25% or more (1.5–2 and 0.75–0.5) of one allele was considered as allelic imbalance.
Mutation Analysis
Primers designed for amplification of the complete coding region of MH1 and MH2 domains of Smad2 and Smad4, applicable for paraffin material as template, are listed in Supplementary Tables 1 and 2. The following PCR conditions were used: 10 min at 95°C, followed by 40 cycles of 30 s at 95°C, 30 s at 53°C and 30 s extension at 72°C. Sequencing was performed by the Leiden Genome Technology Center (Leiden, The Netherlands). The obtained sequences were compared with DNA sequences of autologous normal material and with the corresponding sequences from the Genome Database (http:/www.gdb.org).
Methylation Assay
For the detection of CpG methylation in cell cultures, cervical cancer cell lines and sorted tumor cell subpopulations, the bisulfite method was applied for DNA methylation analysis using the EZ DNA Methylation Kit (ZYMO Research, Orange, CA, USA) according to the manufacturer's protocol. Before conversion with the bisulfite method, DNA from sorted samples was purified and concentrated with Wizard DNA clean-up (Promega, Madison, WI, USA). DNA input for the conversion reaction was 500 ng/sample. Primers designed for PCR and sequencing of bisulfite-converted DNA, encompassing the promoter region and exon 1 of Smad4, were as follows: 5′-GGTTGGGTTAATAGTTATTTATA (FWD) and CTCTAAATCTAAACCTAAACCTA (REV), 5′-TAGGTTTAGGTTTAGATTTAGAG (FWD) and AATTTAACCCTTTTCCCCCC (REV), 5′-GGGGGAAAAGGGTTAAATT (FWD) and CTAAACACCTAACTCCCCTC (REV).
Statistical Analysis
Statistical analysis was performed using SPSS 12.0 (SPSS Inc., Chicago, IL, USA). Data were processed using the Chi-squared test. Kaplan–Meier survival curves were generated to assess differences in disease-free survival (defined as the time in months from surgery to relapse of the disease) or cumulative overall survival (defined as the time in months from surgery to death due to cervical cancer). A Cox regression was used for univariate and multivariate survival analyses. P<0.05 was considered statistically significant.
Results
Smad2, P-Smad2, Smad4 and P21 Protein Expression in Primary Cervical Carcinomas
Protein expression of Smad2, P-Smad2, Smad4 and p21 was evaluated by immunohistochemistry in a tissue array containing 117 primary cervical carcinomas. Because of its nuclear localization p21 was chosen as the target product for TGF-β signaling.
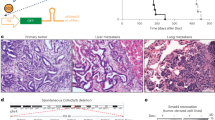
All tumors showed a homogeneous cytoplasmic staining pattern for Smad2 and Smad4 that varied in intensity (Figure 1 and Table 1). For the analysis of cytoplasmic staining results, the three categories from Table 1 were combined into two categories (weak vs moderate/strong). Weak cytoplasmic Smad2 staining intensity correlated significantly with weak cytoplasmic Smad4 staining intensity (P=0.050). All tumors showed nuclear staining for P-Smad2 (Table 1). Fifteen out of 111 tumors lacked nuclear staining for Smad4 (14%) and 15 out of 114 tumors lacked nuclear staining for p21 (13%). P-Smad2 consisted of two groups: a group with less than 50% of positive tumor cells (low) and a group with more than 50% of positive tumor cells (high). To analyze the results of the nuclear staining pattern, nuclear Smad4 and p21 were divided into a group without nuclear staining (absent) and a group with nuclear staining. The P-Smad2 nuclear staining pattern (1–50%) showed a significant correlation with the absence of p21 staining (P=0.011). Although not significant, association between absence of nuclear Smad4 and absence of p21 (P=0.051) was observed. In addition, a significant association between weak cytoplasmic Smad4 staining intensity and absent Smad4 nuclear staining was observed (P=0.001).
Expression of Smad2, P-Smad2, Smad4 and p21 proteins in primary cervical carcinomas. A tissue microarray of 117 cervical tumors was stained using immunohistochemistry as described in Materials and methods (× 125 magnification). (a) Smad2, strong cytoplasmic staining, (b) Smad2, moderate cytoplasmic staining, (c) Smad2, weak cytoplasmic staining, (d) positive nuclear staining of P-Smad2 in ≥50% of tumor cells, (d1) panel d in detail (× 400 magnification), (e) P-Smad2 nuclear staining in ≤50% of tumor cells, (e1) panel e in detail (× 400 magnification), (f) Smad4, positive nuclear staining in ≥50% of cells, (f1) panel f in detail (× 400 magnification), (g) Smad4, strong cytoplasmic intensity and absent nuclear staining, (g1) panel g in detail(× 400 magnification), (h) Smad4, moderate cytoplasmic intensity, (i) Smad4, weak cytoplasmic intensity, (j) p21 nuclear staining in ≥50% of tumor cells, (j1) panel j in detail (× 400 magnification), (k) p21 nuclear staining in ≥0 and ≤50% of tumor cells, (k1) panel k in detail (× 400 magnification), (l) absent p21 nuclear staining and (l1) panel l in detail (× 400 magnification).
LOH of the Smad2 and Smad4 Loci at 18q21.1
To establish the basis for weak Smad2 and weak Smad4 protein expression, we investigated whether this was due to haplo-insufficiency of the Smad2 or Smad4 genes caused by LOH. For LOH analysis, DNA was used from a flow-sorted tumor cell subpopulation (keratin positive, vimentin negative; K+V−) to prevent contamination by either stromal or inflammatory cells. LOH at 18q21.1 was determined in 46 tumors with low, moderate and strong expression of either Smad2 or cytoplasmic Smad4 (Figure 2b). High allelic imbalance ratios were frequently observed, often ranging from 2 to 10, representing complete LOH. Out of the 46 cases, 19 cases (41%) showed loss at markers flanking the Smad2 and Smad4 loci (allelic imbalance ≥1.5). Loss of the complete Smad2 and Smad4 loci was noticed in 14 cases (30%). Eleven out of these 14 cases seemed to have lost the complete chromosome arm. No significant association between Smad2 or Smad4 protein expression and LOH at 18q21.1 was apparent.
LOH at 18q21.1 in primary cervical carcinomas. (a) Microsatellite markers surrounding Smad2 and Smad4 genes at chromosome arm 18q. (b) LOH at 18q of 46 samples was measured using seven microsatellite markers as described in Materials and methods. For each tumor, Smad2 and cytoplasmic Smad4 staining intensity is depicted as w: weak, m: moderate and s: strong. A.I., allelic imbalance.
Mutation Analysis of the Functional MH1 and MH2 Domains of Smad2 and Smad4
Subsequently, we investigated whether mutations in the function-associated MH1 and MH2 domains of Smad2 or Smad4 were the underlying cause for their reduced expression. For this purpose, five tumors with a weak and five tumors with a moderate cytoplasmic Smad2 staining pattern were selected (all the tumors selected displayed less than 50% of P-Smad2-positive tumor cells). For Smad4, eight tumors with a weak and two tumors with a moderate cytoplasmic Smad4 staining pattern were selected (without nuclear Smad4 staining). PCR amplification and sequence analysis of DNA from the flow-sorted tumor cell subpopulation (V−K+) as well as DNA from the flow-sorted stromal cell/inflammatory cell subpopulation (V+K−) as a control were performed. No mutations in the MH1 and MH2 domains of Smad2 and Smad4 were observed (data not shown).
Methylation of the Promoter Region and Exon 1 of Smad4
Because of the lack of a significant association between Smad2 or Smad4 expression and LOH at 18q21.1, and the absence of mutations in the MH1 and MH2 domains of Smad2 and Smad4, we investigated whether methylation of the promoter region could explain the reduced protein expression. We focused on Smad4 promoter methylation, as it has been previously described as a tumor suppressor mechanism.12, 13 Four cervical cancer cell lines (HeLa, CaSki, CC-8 and CC-11) and four normal primary cervical epithelial cell cultures, both expressing the complete coding region of Smad4 (as determined by RT-PCR), were sequenced after bisulfite conversion to determine the CpG methylation status. Also, DNA from five flow-sorted tumor cell populations with weak and five flow-sorted tumor cell populations with moderate Smad4 cytoplasmic staining was analyzed for CpG methylation status. The region up to 300 bp upstream from the transcription start point, containing dense CpG islands, and 125 bp downstream from the transcription start, which includes exon 1, was sequenced (Supplementary Figure 1). None of the CpG sites investigated was methylated (data not shown).
Association between Smad2, Smad4 and P21 Expression and Clinico-pathological Parameters
Despite the unknown nature of decreased Smad2 and cytoplasmic Smad4 expression, we investigated whether the expression pattern of Smad2, P-Smad2, cytoplasmic Smad4, nuclear Smad4 and p21 correlated with clinico-pathological parameters (Table 2).
Weak cytoplasmic Smad4 staining intensity strongly correlated with the presence of positive lymph nodes (P=0.002) and recurrent disease (P=0.009). Absent nuclear Smad4 staining correlated with tumors ≥40 mm (P=0.020), infiltration depth ≥15 mm (P=0.036) and the presence of HPV18 (P=0.020). Both weak Smad2 expression and P-Smad2 expression (<50% of tumor cells) were not significantly associated with any of the clinico-pathological parameters tested.
Because of the significant correlation between a weak cytoplasmic Smad4 staining intensity and lymph node metastases and recurrent disease, we also assessed the association between the Smad protein expression patterns and disease-free and overall 5-year survival. Indeed, weak cytoplasmic Smad4 staining intensity correlated with both poor disease-free survival (P=0.003; Figure 3a) and poor overall survival (P=0.003; Figure 3b). Also the absence of nuclear Smad4 staining correlated with poor disease-free survival (P=0.010; Figure 3c) and poor overall survival (P=0.003; Figure 3d).
Association between cytoplasmic and nuclear Smad4 expression and survival in primary cervical carcinomas. Kaplan–Meier survival curves were generated to assess differences in disease-free survival (defined as the time in months from surgery to relapse of the disease) or cumulative overall survival (defined as the time in months from surgery to death due to cervical cancer) as described in Materials and methods. (a) Disease-free 5-year survival in patients with weak cytoplasmic Smad4 staining intensity (N=38) and moderate/strong intensity (N=63). (b) Overall survival in patients with weak cytoplasmic Smad4 intensity (N=40) and moderate/strong intensity (N=70) in primary cervical carcinomas. (c) Disease-free survival in patients with absent (N=13) and present (N=88) Smad4 nuclear staining. (d) Overall survival in patients with absent (N=15) and present (N=96) Smad4 nuclear staining.
Subsequently, we used the Cox proportional hazards model to calculate the predictive value for a shorter disease-free and overall 5-year survival of each of these parameters. Both weak cytoplasmic Smad4 expression and absent nuclear Smad4 expression were demonstrated to be prognostic factors for disease-free and overall 5-year survival (Table 3).
Furthermore, in a multivariate Cox analysis that included the four strongest prognostic factors according to univariate Cox analysis (lymph node status, tumor size, vaso-invasion and depth of infiltration),23 in addition to lymph node status, absent nuclear Smad4 staining was found to be the only independent prognostic factor for both disease-free (hazard ratio of 2.91) and overall 5-year survival (hazard ratio of 2.74).
Discussion
Loss of responsiveness to TGF-β-induced cell growth inhibition during cancer development is a feature of many tumors, including cervical cancer.1 Although a number of studies have investigated the expression status and alterations in the canonical TGF-β signaling (Smad) pathway, the role of Smad signaling in the progression of cervical cancer remains unclear.3, 10, 11, 24 In this study, we investigated whether decreased Smad2 and Smad4 protein expression in primary cervical cancers was associated with molecular alterations at 18q21.1, mutations in the functional domains of Smad2 and Smad4 or hypermethylation, and we assessed the biological relevance of decreased Smad2 and Smad4 expression.
Previously, Smad4 deficiency has been reported in 4 out of 13 cervical cancer cell lines.10 It should be noted that of these cell lines, one cell line was derived from a lymph node metastases (HT-3) and two other cell lines (C4-I and C4-II) were derived from the same patient. The origin of the fourth cell line was not specified. In contrast to Baldus et al,10 who reported a Smad4 deficiency in 10 out of 40 invasive cervical carcinomas, we did not observe tumors negative for Smad4 staining. There is no clear explanation for this difference. Baldus et al did not report the FIGO stage of the investigated tumors, whereas in our study only FIGO stage IB and IIA cervical carcinomas were investigated. Conceivably, differences in Smad4 expression may occur as late events during tumor progression. The expression of P-Smad2, nuclear Smad4 or p21 in the majority of the cervical carcinomas investigated suggests that the canonical TGF-β signal pathway is operative. However, in cases with a limited amount of cytoplasmic Smad4, the amount of Smad4 may be rate limiting.
Loss or decrease in Smad protein expression can be due to genetic alterations such as LOH and inactivating mutations.5 We found a relatively high frequency (41%) of LOH at 18q21.1, the locus of Smad2 and Smad4 genes, compared to other studies (10–37%).6, 7 The close proximity of the markers to the Smad2 and Smad4 loci in combination with the purity of the tumor samples may explain this increased frequency. For LOH analysis, we used DNA samples from flow-sorted cancer samples. The advantage of flow sorting over other techniques, such as laser microdissection, is the lack of contaminating stromal cells or cells from the immune system in the tumor sample. As LOH at 18q21.1 did not associate with weak Smad2 or weak Smad4 expression, genetic haplo-insufficiency of Smad2 and Smad4 as reported by Baldus et al10 is not supported by our data.
Only two studies have reported on mutations in the functional domains (MH1 and MH2) of Smad2 and Smad4 in cervical carcinoma.10, 11 Maliekal et al11 identified a missense mutation in the MH1 domain of Smad2 in a cervical tumor and reported lack of mRNA expression of the C-terminal part of the MH2 domains in six tumors. We did not detect mutations in the MH1 and MH2 domains of Smad2 in the tumor samples; therefore, it remains uncertain whether cervical tumors contain inactivating mutations in the MH2 domain. Nevertheless, we cannot exclude the presence of less frequently occurring mutations outside the MH1 and MH2 domains of Smad2 and Smad4.
Baldus et al10 reported deletions in part of the function-associated MH1 and MH2 domains of Smad4 in four cervical cancer cell lines. In agreement with Maliekal et al, in our study we did not observe mutations in MH1 and MH2 domains of Smad4 in cervical tumors. However, our approach does not exclude the presence of possible missense mutations. Our data suggest that genetic inactivation of Smad2 and Smad4 plays a minor role in cervical cancer.
Another mechanism by which Smad expression could be downregulated is promoter methylation. In our study, similar to the study of Wang et al13 in gastric carcinoma, we investigated both the promoter region and exon 1 of Smad4. No hypermethylation of the promoter region or exon 1 of Smad4 was observed in normal samples, cervical cancer cell lines and tumor samples. Also, in colorectal cancers, no hypermethylation of Smad4 was observed, whereas in Barrett's esophagus, a weak correlation between mRNA expression and promoter methylation was found.12, 25, 26 In gastric tumors, hypermethylation was rare. In this tumor type, a significant correlation was reported between hypermethylation and loss of protein expression.
Weak protein expression of Smad2 and Smad4 in cervical cancer could also be due to other regulatory processes, such as interaction of HPV16 E7 with Smad2, Smad3 and Smad4,27 degradation by a proteasome13 or activation of specific ubiquitin-Smad E3 ligases.28, 29
In our series, weak cytoplasmic Smad4 staining and the absence of Smad4 nuclear staining was associated with poor survival in cervical cancer patients. In a previous study, our group found an association between LOH at 18q and survival.7 Here, we did not find this association. The differences in the results could be explained by much closer proximity of the chosen markers to the Smad2 and Smad4 loci in the present study. In our study, weak nuclear Smad4 expression was found to be an independent prognostic factor for both disease-free and overall survival. Loss of nuclear Smad4 expression in gastric cancer also correlated with poor survival.13 In pancreatic cancer, Smad4 protein expression was an independent predictor for longer survival.30 In colorectal and breast cancers, loss of Smad4 protein expression correlated with negative outcome or the presence of lymph node metastasis.31, 32 These data emphasize the importance of the integrity of the canonical TGF-β pathway in carcinogenesis. Our findings support the hypothesis that Smad4 is a target molecule for functional inactivation in cervical cancer.
References
De Geest K, Bergman CA, Turyk ME, et al. Differential response of cervical intraepithelial and cervical carcinoma cell lines to transforming growth factor-beta 1. Gynecol Oncol 1994;55:376–385.
Kang SH, Won K, Chung HW, et al. Genetic integrity of transforming growth factor beta (TGF-beta) receptors in cervical carcinoma cell lines: loss of growth sensitivity but conserved transcriptional response to TGF-beta. Int J Cancer 1998;77:620–625.
Lee S, Cho YS, Shim C, et al. Aberrant expression of Smad4 results in resistance against the growth-inhibitory effect of transforming growth factor-beta in the SiHa human cervical carcinoma cell line. Int J Cancer 2001;94:500–507.
ten Dijke P, Goumans MJ, Itoh F, et al. Regulation of cell proliferation by Smad proteins. J Cell Physiol 2002;191:1–16.
Hahn SA, Schutte M, Hoque AT, et al. DPC4, a candidate tumor suppressor gene at human chromosome 18q21.1. Science 1996;271:350–353.
Harima Y, Sawada S, Nagata K, et al. Chromosome 6p21.2, 18q21.2 and human papilloma virus (HPV) DNA can predict prognosis of cervical cancer after radiotherapy. Int J Cancer 2001;96:286–296.
Kersemaekers AM, Kenter GG, Hermans J, et al. Allelic loss and prognosis in carcinoma of the uterine cervix. Int J Cancer 1998;79:411–417.
Fukushima T, Mashiko M, Takita K, et al. Mutational analysis of TGF-beta type II receptor, Smad2, Smad3, Smad4, Smad6 and Smad7 genes in colorectal cancer. J Exp Clin Cancer Res 2003;22:315–320.
Eppert K, Scherer SW, Ozcelik H, et al. MADR2 maps to 18q21 and encodes a TGFbeta-regulated MAD-related protein that is functionally mutated in colorectal carcinoma. Cell 1996;86:543–552.
Baldus SE, Schwarz E, Lohrey C, et al. Smad4 deficiency in cervical carcinoma cells. Oncogene 2005;24:810–819.
Maliekal TT, Antony ML, Nair A, et al. Loss of expression, and mutations of Smad 2 and Smad 4 in human cervical cancer. Oncogene 2003;22:4889–4897.
Onwuegbusi BA, Aitchison A, Chin SF, et al. Impaired transforming growth factor beta signalling in Barrett's carcinogenesis due to frequent SMAD4 inactivation. Gut 2006;55:764–774.
Wang LH, Kim SH, Lee JH, et al. Inactivation of SMAD4 tumor suppressor gene during gastric carcinoma progression. Clin Cancer Res 2007;13:102–110.
Hazelbag S, Gorter A, Kenter GG, et al. Transforming growth factor-beta1 induces tumor stroma and reduces tumor infiltrate in cervical cancer. Hum Pathol 2002;33:1193–1199.
Natsugoe S, Xiangming C, Matsumoto M, et al. Smad4 and transforming growth factor beta1 expression in patients with squamous cell carcinoma of the esophagus. Clin Cancer Res 2002;8:1838–1842.
Bruna A, Darken RS, Rojo F, et al. High TGFbeta–Smad activity confers poor prognosis in glioma patients and promotes cell proliferation depending on the methylation of the PDGF-B gene. Cancer Cell 2007;11:147–160.
Pruneri G, Pignataro L, Carboni N, et al. Clinical relevance of expression of the CIP/KIP cell-cycle inhibitors p21 and p27 in laryngeal cancer. J Clin Oncol 1999;17:3150–3159.
Torquati A, O′rear L, Longobardi L, et al. RUNX3 inhibits cell proliferation and induces apoptosis by reinstating transforming growth factor beta responsiveness in esophageal adenocarcinoma cells. Surgery 2004;136:310–316.
Ruiter DJ, Ferrier CM, van Muijen GN, et al. Quality control of immunohistochemical evaluation of tumour-associated plasminogen activators and related components. European BIOMED-1 Concerted Action on Clinical Relevance of Proteases in Tumour Invasion and Metastasis. Eur J Cancer 1998;34:1334–1340.
Corver WE, ter Haar NT, Dreef EJ, et al. High-resolution multi-parameter DNA flow cytometry enables detection of tumour and stromal cell subpopulations in paraffin-embedded tissues. J Pathol 2005;206:233–241.
Abeln EC, Corver WE, Kuipers-Dijkshoorn NJ, et al. Molecular genetic analysis of flow-sorted ovarian tumour cells: improved detection of loss of heterozygosity. Br J Cancer 1994;70:255–262.
Haven CJ, van Puijenbroek M, Karperien M, et al. Differential expression of the calcium sensing receptor and combined loss of chromosomes 1q and 11q in parathyroid carcinoma. J Pathol 2004;202:86–94.
Sevin BU, Lu Y, Bloch DA, et al. Surgically defined prognostic parameters in patients with early cervical carcinoma. A multivariate survival tree analysis. Cancer 1996;78:1438–1446.
Chen T, de Vries EG, Hollema H, et al. Structural alterations of transforming growth factor-beta receptor genes in human cervical carcinoma. Int J Cancer 1999;82:43–51.
Ando T, Sugai T, Habano W, et al. Analysis of SMAD4/DPC4 gene alterations in multiploid colorectal carcinomas. J Gastroenterol 2005;40:708–715.
Roth S, Laiho P, Salovaara R, et al. No SMAD4 hypermethylation in colorectal cancer. Br J Cancer 2000;83:1015–1019.
Lee DK, Kim BC, Kim IY, et al. The human papilloma virus E7 oncoprotein inhibits transforming growth factor-beta signaling by blocking binding of the Smad complex to its target sequence. J Biol Chem 2002;277:38557–38564.
Izzi L, Attisano L . Ubiquitin-dependent regulation of TGFbeta signaling in cancer. Neoplasia 2006;8:677–688.
Fukuchi M, Fukai Y, Masuda N, et al. High-level expression of the Smad ubiquitin ligase Smurf2 correlates with poor prognosis in patients with esophageal squamous cell carcinoma. Cancer Res 2002;62:7162–7165.
Tascilar M, Skinner HG, Rosty C, et al. The SMAD4 protein and prognosis of pancreatic ductal adenocarcinoma. Clin Cancer Res 2001;7:4115–4121.
Xie W, Mertens JC, Reiss DJ, et al. Alterations of Smad signaling in human breast carcinoma are associated with poor outcome: a tissue microarray study. Cancer Res 2002;62:497–505.
Xie W, Rimm DL, Lin Y, et al. Loss of Smad signaling in human colorectal cancer is associated with advanced disease and poor prognosis. Cancer J 2003;9:302–312.
Acknowledgements
We thank P Tholen for technical assistance with the optimization of Smad2 and Smad4 immunohistochemistry and mutation analysis. The study was supported by the Dutch Cancer Society, Grant RUL2001-2465.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/conflicts of interest
The authors state no conflicts of interest.
Supplementary Information accompanies the paper on Modern Pathology website (http://www.nature.com/modpathol)
Supplementary information
Rights and permissions
About this article
Cite this article
Kloth, J., Kenter, G., Spijker, H. et al. Expression of Smad2 and Smad4 in cervical cancer: absent nuclear Smad4 expression correlates with poor survival. Mod Pathol 21, 866–875 (2008). https://doi.org/10.1038/modpathol.2008.62
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/modpathol.2008.62
Keywords
This article is cited by
-
HPV-mediated regulation of SMAD4 modulates the DNA damage response in head and neck cancer
Journal of Experimental & Clinical Cancer Research (2022)
-
SMAD2 rs4940086 heterozygosity increases the risk of cervical cancer development among the women in Bangladesh
Molecular Biology Reports (2020)
-
Transforming growth factor-β1 in carcinogenesis, progression, and therapy in cervical cancer
Tumor Biology (2016)
-
microRNA Expression Pattern Modulates Temozolomide Response in GBM Tumors with Cancer Stem Cells
Cellular and Molecular Neurobiology (2014)
-
Partial loss of Smad signaling during in vitroprogression of HPV16-immortalized human keratinocytes
BMC Cancer (2013)