Abstract
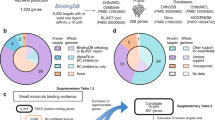
Malaria caused by Plasmodium falciparum is a disease that is responsible for 880,000 deaths per year worldwide. Vaccine development has proved difficult and resistance has emerged for most antimalarial drugs. To discover new antimalarial chemotypes, we have used a phenotypic forward chemical genetic approach to assay 309,474 chemicals. Here we disclose structures and biological activity of the entire library—many of which showed potent in vitro activity against drug-resistant P. falciparum strains—and detailed profiling of 172 representative candidates. A reverse chemical genetic study identified 19 new inhibitors of 4 validated drug targets and 15 novel binders among 61 malarial proteins. Phylochemogenetic profiling in several organisms revealed similarities between Toxoplasma gondii and mammalian cell lines and dissimilarities between P. falciparum and related protozoans. One exemplar compound displayed efficacy in a murine model. Our findings provide the scientific community with new starting points for malaria drug discovery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wongsrichanalai, C. & Meshnick, S. R. Declining artesunate-mefloquine efficacy against falciparum malaria on the Cambodia-Thailand border. Emerg. Infect. Dis. 14, 716–719 (2008)
Dondorp, A. M. et al. Artemisinin resistance in Plasmodium falciparum malaria. N. Engl. J. Med. 361, 455–467 (2009)
Ridley, R. G. Medical need, scientific opportunity and the drive for antimalarial drugs. Nature 415, 686–693 (2002)
Kissinger, J. C. et al. The Plasmodium genome database. Nature 419, 490–492 (2002)
Baniecki, M. L., Wirth, D. F. & Clardy, J. High-throughput Plasmodium falciparum growth assay for malaria drug discovery. Antimicrob. Agents Chemother. 51, 716–723 (2007)
Plouffe, D. et al. In silico activity profiling reveals the mechanism of action of antimalarials discovered in a high-throughput screen. Proc. Natl Acad. Sci. USA 105, 9059–9064 (2008)
Weisman, J. L. et al. Searching for new antimalarial therapeutics amongst known drugs. Chem. Biol. Drug Des. 67, 409–416 (2006)
Wells, T. N., Alonso, P. L. & Gutteridge, W. E. New medicines to improve control and contribute to the eradication of malaria. Nature Rev. Drug Discov. 8, 879–891 (2009)
Munos, B. Can open-source R&D reinvigorate drug research? Nature Rev. Drug Discov. 5, 723–729 (2006)
Shelat, A. A. & Guy, R. K. Scaffold composition and biological relevance of screening libraries. Nature Chem. Biol. 3, 442–446 (2007)
Shelat, A. A. & Guy, R. K. The interdependence between screening methods and screening libraries. Curr. Opin. Chem. Biol. 11, 244–251 (2007)
Smilkstein, M. et al. Simple and inexpensive fluorescence-based technique for high-throughput antimalarial drug screening. Antimicrob. Agents Chemother. 48, 1803–1806 (2004)
Cibulskis, R. E. et al. Estimating trends in the burden of malaria at country level. Am. J. Trop. Med. Hyg. 77, (suppl. 6)133–135 (2007)
Gujjar, R. et al. Identification of a metabolically stable triazolopyrimidine-based dihydroorotate dehydrogenase inhibitor with antimalarial activity in mice. J. Med. Chem. 52, 1864–1872 (2009)
Patel, V. et al. Identification and characterization of small molecule inhibitors of Plasmodium falciparum dihydroorotate dehydrogenase. J. Biol. Chem. 283, 35078–35085 (2008)
Weissbuch, I. & Leiserowitz, L. Interplay between malaria, crystalline hemozoin formation, and antimalarial drug action and design. Chem. Rev. 108, 4899–4914 (2008)
Pisciotta, J. M. et al. The role of neutral lipid nanospheres in Plasmodium falciparum haem crystallization. Biochem. J. 402, 197–204 (2007)
Sijwali, P. S. & Rosenthal, P. J. Gene disruption confirms a critical role for the cysteine protease falcipain-2 in hemoglobin hydrolysis by Plasmodium falciparum . Proc. Natl Acad. Sci. USA 101, 4384–4389 (2004)
Sijwali, P. S., Koo, J., Singh, N. & Rosenthal, P. J. Gene disruptions demonstrate independent roles for the four falcipain cysteine proteases of Plasmodium falciparum . Mol. Biochem. Parasitol. 150, 96–106 (2006)
Crowther, G. J. et al. Buffer optimization of thermal melt assays of Plasmodium proteins for detection of small-molecule ligands. J. Biomol. Screen. 14, 700–707 (2009)
Witola, W. H. et al. Disruption of the Plasmodium falciparum PfPMT gene results in a complete loss of phosphatidylcholine biosynthesis via the serine-decarboxylase-phosphoethanolamine-methyltransferase pathway and severe growth and survival defects. J. Biol. Chem. 283, 27636–27643 (2008)
Krnajski, Z. et al. Thioredoxin reductase is essential for the survival of Plasmodium falciparum erythrocytic stages. J. Biol. Chem. 277, 25970–25975 (2002)
McFadden, G. I. & Roos, D. S. Apicomplexan plastids as drug targets. Trends Microbiol. 7, 328–333 (1999)
Reynolds, M. G. & Roos, D. S. A biochemical and genetic model for parasite resistance to antifolates. Toxoplasma gondii provides insights into pyrimethamine and cycloguanil resistance in Plasmodium falciparum . J. Biol. Chem. 273, 3461–3469 (1998)
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 46, 3–26 (2001)
Shenai, B. R. et al. Structure-activity relationships for inhibition of cysteine protease activity and development of Plasmodium falciparum by peptidyl vinyl sulfones. Antimicrob. Agents Chemother. 47, 154–160 (2003)
Ritz, C. & Streibig, J. C. Bioassay analysis using R. J. Stat. Softw. 12, 22 (2005)
Acknowledgements
This work was supported by the American Lebanese Syrian Associated Charities (ALSAC) and St Jude Children’s Research Hospital (SJCRH, R.K.G.), the Medicines for Malaria Venture (W.C.V.V. and V.M.A.), National Institute of Allergy and Infectious Diseases (AI772682 (P.H.D.), AI075517 (R.K.G.), AI067921 (W.C.V.V.) and AI080625 (W.C.V.V.), AI28724 (D.S.R.), AI53862 (J.L.D.), AI35707 (P.J.R.), AI053680 (M.A.P. and P.K.R.), AI075594 (M.A.P., P.K.R. and I.B.), AI082617 (P.K.R.) and AI045774 (D.J.S.)), the National Cancer Institute (CA78039 (J.S.L.)), the Welch Foundation (I-1257 (M.A.P.)), the Doris Duke Charitable Foundation (P.J.R.), and the Ellison Medical Foundation (D.S.R.). We acknowledge A. B. Vaidya for providing the parasite strain D10_yDHOD. We acknowledge M. Sigal for assistance in the early leads project coordination, the SJCRH High Throughput Screening Center, particularly J. Cui; the SJCRH Lead Discovery Informatics Center, and the SJCRH High Throughput Analytical Chemistry Center, particularly C. Nelson and A. Lemoff; at UW, F. Buckner, W. Hol and A. Napuli (AI067921, W. Hol); S. Wei and W. Hao in the UT Southwestern HTS Center; and the Australian Red Cross Blood Service for the provision of O+ erythrocytes to Griffith University.
Author information
Authors and Affiliations
Contributions
W.A.G. and R.K.G. designed and coordinated the project. A.A.S. wrote the algorithms for the data analysis and generated the figures. Assays were conceived, performed and analysed by W.A.G. and D.B. (P. falciparum phenotypic screen), M.C. (human cell lines), D.C.S. (T. brucei), P.H.D. and D.S.R. (T. gondii), J.S.L. and E.R.S. (L. major), A.K.T. and D.J.S. (haemozoin inhibition), G.J.C. and W.C.V.V. (thermal melt experiments), M.A.P., P.K.R., F.E.M. and I.B. (PfDHOD), J.W.F. and P.K.R. (P. falciparum dihydrofolate reductase), J.G. and P.J.R. (PfFP-2), I.F. and M.K.R. (cytochrome bc1), J.C. (P. falciparum mutant drug sensitivity). E.B.W., S.D., J.L.D. and V.M.A. (independent antimalarial in vitro experiments), F.Z. (in vitro pharmacokinetics), M.B.J.D., M.S.M., I.A.-B. and S.F. (in vivo pharmacokinetics and efficacy), I.B. (coordination of technology development and network development), S.C. and P.L.M. (re-synthesis). W.A.G., A.A.S. and R.K.G. wrote the manuscript. All authors contributed to the design of the experiments and the preparation of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information (download PDF )
This file contains Supplementary Information comprising: Compound Library Screened; Parasite and cell-based assay methods; Enzymatic and protein assays; Data processing and screening results; Multiple-lab cross-validation study; Chemical structure network graph; Additional mechanistic studies; Detailed analysis of three early lead compounds; Other high-value compounds; Data availability; Supplementary Tables S1-S8, Supplementary Figures S1-S5 with legends and References. The Supplementary Information file was replaced on 24 June 2010. (PDF 952 kb)
Supplementary Data (download XLS )
This file contains Structural information and Primary screening for 1536 compounds, Screening data from 228 compounds for the Bland-Altman analysis, Calculated medicinal chemistry properties, Cytotoxic activity, Screen sensitivity and Differential activity for 172 compounds, Raw data from the Hemozoin polymerization inhibition assay, summary of activity of 172 compounds in the thermal melt assays and Raw data and calculated Kd for thermal melt assay hits. (XLS 5090 kb)
Rights and permissions
About this article
Cite this article
Guiguemde, W., Shelat, A., Bouck, D. et al. Chemical genetics of Plasmodium falciparum. Nature 465, 311–315 (2010). https://doi.org/10.1038/nature09099
Received:
Accepted:
Issue date:
DOI: https://doi.org/10.1038/nature09099
This article is cited by
-
The past, present and future of anti-malarial medicines
Malaria Journal (2019)
-
Advances in omics-based methods to identify novel targets for malaria and other parasitic protozoan infections
Genome Medicine (2019)
-
Two successful decades of Swiss collaborations to develop new anti-malarials
Malaria Journal (2019)
-
High-throughput screening of the Plasmodium falciparum cGMP-dependent protein kinase identified a thiazole scaffold which kills erythrocytic and sexual stage parasites
Scientific Reports (2019)
-
High Throughput and Computational Repurposing for Neglected Diseases
Pharmaceutical Research (2019)