Abstract
The double homeobox 4 gene (DUX4) and its centromeric paralogue, DUX4C, reside in the subtelomeric region of chromosome 4 and have been implicated in facioscapulohumeral muscular dystrophy (FSHD) and cancers. However, the high sequence similarity between these genes, together with the widespread presence of DUX4-like paralogues across the human genome, has hindered accurate genotyping and expression profiling using short-read sequencing. To elucidate the genetic architecture and potential disease-associated functions of DUX4 and DUX4C, we first identified two distinct DUX4C haplotypes with expression quantitative trait effects—DUX4C-4qα and DUX4C-4qβ. We then integrated them with known DUX4 haplotypes to generate a reference genome, D4Ref-T2T, using long-read sequencing. Haplotype analysis indicated strong linkage disequilibrium between DUX4C and DUX4 haplotypes (r2 = 0.86). We further characterized full-length DUX4C mRNA isoforms and established a corresponding transcriptome reference. Applying these resources to breast tumor, FSHD, and lymphoblastoid cell line datasets revealed that DUX4 is expressed in both breast tumor and FSHD tissues, whereas DUX4C shows expression only in breast tumors. Differential gene expression and Gene Ontology enrichment analyses further suggested that DUX4C expression is associated with activation of pathways involved in leukocyte differentiation, chemotaxis, and cell migration.
Similar content being viewed by others
Introduction
The double homeobox 4 (DUX4) gene resides in the subtelomeric region of human chromosome 4, embedded within a macrosatellite array composed of approximately 1–100 copies of 3.3-kb D4Z4 tandem repeats. When the number of repeats contracts to ≤10 and a functional polyadenylation signal (PAS) is present within the terminal unit, mature DUX4 transcripts are produced, enabling expression of full-length DUX4 protein, which has been implicated in facioscapulohumeral muscular dystrophy (FSHD) and several cancer types [1, 2].
The complex structure of the DUX4 locus has been redefined in the recent telomere-to-telomere (T2T) project [3], and upstream of the macrosatellite array lies an additional repeat in the T2T-CHM13v2.0 genome. This repeat encodes the centromeric paralogue DUX4C (also known as DUX4L9), which has also been associated with FSHD pathology [4]. However, the extremely high sequence similarity between DUX4 and DUX4C, together with incomplete annotation of DUX4C transcripts, has made it difficult to distinguish these genes using short-read sequencing. These limitations have hindered efforts to define DUX4C-specific functions and to clarify its relationship with the telomeric gene DUX4 (hereafter DUX4T).
To overcome these challenges, we combined genomic and transcriptomic analyses using both short- and long-read sequencing technologies. This integrative framework aimed to: (i) delineate the major haplotypes of DUX4C and DUX4T through systematic analysis of publicly available short-read datasets and prior literature; (ii) reconstruct complete haplotype sequences using long-read whole-genome (LWGS) data to generate a haplotype-resolved reference suitable for accurate genotyping from short-read whole-genome (SWGS) data, while also evaluating the genetic relationship between DUX4C and DUX4T; and (iii) characterize transcript diversity using long-read RNA sequencing to develop an expanded transcript reference. Leveraging these resources, we further investigated the roles and regulatory mechanisms of DUX4C and DUX4T across multiple biological contexts, including breast tumors, FSHD, and interferon-α2 (IFNα2)–stimulated lymphoblastoid cell lines (LCLs).
Materials and methods
DNA and RNA sample preparation
Japanese LCLs, derived from B lymphocytes, were obtained from the NIGMS Human Genetic Cell Repository at the Coriell Institute. Cells were cultured in RPMI-1640 medium supplemented with 15% fetal bovine serum and 1% penicillin–streptomycin. For stimulation experiments, LCLs were treated with 50 ng/mL recombinant human IFNα2 (BioLegend) for 6 h, while untreated cells served as controls. Approximately 1 × 10⁷ cells were collected for each extraction.
Genomic DNA was isolated using the DNeasy Blood & Tissue Kit (QIAGEN), and total RNA was extracted using the RNeasy Mini Kit (QIAGEN), following the manufacturer’s instructions. Nucleic acid concentration and purity were assessed with a NanoDrop spectrophotometer (Thermo Fisher Scientific), and RNA integrity was evaluated using an Agilent 2100 Bioanalyzer (RNA 6000 Nano Kit). Only RNA samples with RNA integrity number (RIN) > 9.0 were used for downstream analyses.
Nanopore long-read sequencing
Genomic DNA libraries from LCLs (NA18943 and NA18948) were prepared using the SQK-LSK109 ligation sequencing kit (Oxford Nanopore Technologies) and sequenced on a PromethION platform equipped with FLO-PRO001 flow cells according to the manufacturer’s guidelines. Basecalling was performed using Guppy v6.4.1 in super-accurate mode.
For long-read transcriptome sequencing, cDNA libraries from IFNα2-stimulated NA18943 were prepared using the SQK-PCB114.24 cDNA-PCR Barcoding Kit V14 (Oxford Nanopore Technologies) and sequenced on PromethION PRO114M flow cells. Basecalling was performed using Dorado v0.9.8 in super-accurate mode.
Sequencing data acquisition
RNA-seq datasets for FSHD and control samples—including LCLs (n = 18), myoblasts (n = 6), and myotubes (n = 6)—were retrieved from Gene Expression Omnibus (GEO) accession GSE153523 [5]. Breast tumor and normal tissue RNA-seq data were obtained from GEO accession GSE233242, including Luminal A (n = 29), Luminal B (n = 3), triple-negative breast cancer (TNBC, n = 9), HER2-positive tumors (n = 2), and normal tissue controls (n = 43) [6].
SWGS data from Japanese LCLs (n = 104) were obtained from the 1000 Genomes Project for DUX4C and DUX4T genotyping. RNA-seq data from a subset of these Japanese LCLs, including IFNα2-stimulated (n = 94) and non-stimulated (n = 20) samples, were retrieved from the DDBJ Sequence Read Archive (SRA; accession DRA016395) [7]. Long-read RNA-seq data generated in this study have been deposited in the DDBJ Sequence Read Archive (DRA) under BioProject accession number PRJDB39925.
Haplotype analysis of DUX4C region
Expression quantitative trait loci (eQTL) data for DUX4L9 (ENSG00000224807.5) in LCLs were obtained from the GTEx Analysis Release V10 (dbGaP accession phs000424.v10.p2) on 08/01/2025. Variants with a normalized effect size (NES) ≥ 0.5 were selected (20 variants in total). Corresponding genotypes for these variants in the European (n = 373) and Japanese (n = 94) populations were obtained from the 1000 Genomes Project [8]. Variants with a minor allele frequency (MAF) < 0.1 were excluded. Pairwise linkage disequilibrium (LD) was calculated using PLINK v1.90b6.16 [9], and LD heatmaps were visualized in Python using Seaborn. Haplotype phasing and frequency estimation were performed using Beagle v5.5 [10].
LWGS data from NA18943 and NA18948 were basecalled using Guppy and aligned to the GRCh38 reference genome with Minimap2 v2.30 under default settings [11]. DNA methylation states were inferred using DeepMod2 v0.3.0 with the r9.4.1 model [12]. For each variant, mean methylation levels were calculated within a ± 50 bp window centered on the position.
Simulated mapping of DUX4T breakpoint sequences
Simulated short-read datasets were generated from known DUX4T haplotype breakpoint sequences (Table S2a) using Sandy v0.24 with the parameters -c 30 -m 50 -e 0.01 -t single-end. Simulated reads were aligned to the CHM13v2.0 and GRCh38 reference genomes using BWA-MEM v0.7.19 with default parameters [13]. Alignments were sorted, and low-quality reads (MAPQ < 10) were filtered using Samtools v1.22.1.
Extraction and alignment of DUX4-like genes
DUX4-like (DUX4L) sequences in the CHM13v2.0 reference genome were identified by extracting regions annotated as “DUX4” from both the genome FASTA and corresponding GFF3 files. Coding regions were compared against canonical DUX4 domains defined in the DUX4 mRNA reference (NCBI protein accession: NP_149418.2). Based on the presence or absence of key domains, DUX4L sequences were classified into three groups: (i) those containing both homeobox domains (HD1 and HD2) and the transcriptional activation domain (TAD); (ii) those retaining HD1 and HD2 but lacking a TAD; and (iii) those lacking all three domains. Multiple sequence alignment was conducted using ApE v3.1.8.1 [14].
Acquisition of complete DUX4C and DUX4T haplotype sequences
We used LWGS data selected from Japanese LCL samples, including one homozygous sample for DUX4C-4qα and DUX4T-4qB [15], as well as another homozygous sample for DUX4C-4qβ and DUX4T-4qA, to extract the complete DUX4 sequences. The DUX4C sequence within the FRG1–DUX4L9–FRG2 region (chr4:193,282,466–193,399,128) was retrieved from the CHM13v2.0 T2T genome. DUX4T haplotype sequences were obtained from the NCBI Nucleotide Database, including DUX4T-4A161 (HM190178.1), DUX4T-10A166 (HM190186.1), and DUX4T-4B163 (HM190161.1). Because DUX4T-4qB and DUX4T-10qB share identical breakpoint sequences, making them indistinguishable for genotyping, we extracted a complete DUX4T-B sequence from NA18943 (DUX4T: 4qB/4qB; 10qA/10qA), which served as the reference sequence for both 4qB and 10qB DUX4T haplotypes.
From these sequences, 200-bp regions spanning the DUX4T breakpoints and adjacent 3′ segments were extracted as mapping references [15]. LWGS data from NA18943 and NA18948 were aligned using Winnowmap2 v2.03 with a k-mer size of 20 [16]. Reads with MAPQ < 10 were removed using Samtools. Breakpoint-containing reads corresponding to DUX4T-4qA (NA18948), DUX4T-10qA (NA18948), and DUX4T-B (NA18943) were identified, and 21,680-bp regions spanning the last two D4Z4 repeats through exon 7 of the DUX4 long isoform (GenBank: NR_137167.1) were extracted. D4Z4 tandem repeat arrays (chr4:193,282,466–193,558,261 and chr10:134,615,120–134,741,148 in CHM13v2.0) were masked, and the three extracted 21,680-bp segments, together with the DUX4C sequence, were integrated into the masked genome to construct a custom reference.
To refine haplotype accuracy, LWGS reads from NA18943 and NA18948 were re-aligned to the custom reference with Winnowmap2, followed by variant calling using PEPPER-Margin-DeepVariant v0.8 with default settings [17]. Variants passing quality thresholds were incorporated to generate finalized haplotype-resolved sequences: DUX4C-4qα, DUX4C-4qβ, DUX4T-4qA, DUX4T-10qA, and DUX4T-B.
D4Ref-T2T genome construction and sequencing data alignment
Sequencing-error–corrected 15-kb haplotype sequences for DUX4C-4qα, DUX4C-4qβ, DUX4T-4qA, DUX4T-10qA, and DUX4T-B were incorporated into the CHM13v2.0 genome after masking the D4Z4 tandem repeat arrays and endogenous DUX4 loci (chr4:193,376,884–193,391,884; chr4:193,395,464–193,555,598; chr10:134,615,120–134,738,536). This produced a custom genotyping reference, designated D4Ref-T2T. To validate the completeness and accuracy of each haplotype sequence, LWGS reads from NA18943 and NA18948 were aligned to D4Ref-T2T using Winnowmap2 (k-mer size 20). Reads with MAPQ < 10 were removed using Samtools, and alignments were inspected in IGV v2.19.7 [18].
The D4Ref-T2T genome was then indexed and used as the reference for SWGS alignment. Reads were mapped using BWA-MEM implemented in Parabricks v4.3.2-1 under default settings, followed by MAPQ-based filtering (MAPQ < 10) using Samtools.
Genotyping of DUX4C and DUX4T haplotypes
Haplotype-specific motifs were defined for both genes. For DUX4C, two 6-bp sequences spanning rs7696384 and rs7696390 in the 3′ UTR were used to distinguish the DUX4C-4qα and DUX4C-4qβ haplotypes. For DUX4T, the canonical 6-bp PAS motif was used to identify DUX4T-4qA, and analogous 6-bp sequences at syntenic positions were used to genotype DUX4T-10qA and DUX4T-B.
Coverage of each haplotype-specific motif was calculated as the mean read depth across the six nucleotide positions:
Genotypes were inferred by computing the relative proportion of each haplotype-specific motif. For DUX4C:
and for DUX4T:
Expected motif proportions for DUX4C genotypes are 1 for 4qα/4qα, 0.5 for 4qα/4qβ, and 0 for 4qβ/4qβ. Because experimental variation can shift observed values away from these theoretical ratios, we applied permissive thresholds: 0.75–1.0 for 4qα/4qα, 0.25–0.75 for 4qα/4qβ, and 0–0.25 for 4qβ/4qβ.
For chromosome 4 DUX4T, theoretical proportions are 0.5 for 4qA/4qA, 0.25 for 4qA/4qB, and 0 for 4qB/4qB. Practical threshold ranges were set as ≥0.375 for 4qA/4qA, 0.125–0.375 for 4qA/4qB, and 0–0.125 for 4qB/4qB. For chromosome 10 DUX4T, the same threshold scheme (≥0.375, 0.125–0.375, and 0–0.125) was used to classify 10qA/10qA, 10qA/10qB, and 10qB/10qB genotypes, respectively. Haplotype phasing, frequency estimation, and pairwise LD calculations were performed using Beagle.
Novel DUX4C isoform identification
Short-read RNA-seq data from Japanese LCLs stimulated with IFN-α2 for 6 h were aligned to the D4Ref-T2T reference genome using STAR v2.7.11b [19], with the following parameters: --outFilterMultimapNmax 10, --outFilterIntronMotifs None, --outSJfilterDistToOtherSJmin 0 0 0 0, and --twopassMode Basic.
Genome annotation was derived from the CHM13v2.0 GTF file but modified such that the DUX4L9 locus was updated to reflect DUX4C-4qα coordinates (12,742–14,000). Long-read RNA-seq data from IFNα2-stimulated NA18943 were mapped using FLAIR v2.2.0. Splice-junction correction was applied with flair correct using STAR-generated junctions (SJ.out.tab) and the modified GTF. Isoforms were reconstructed with flair collapse using the parameters --support 2, --filter comprehensive, and --no_gtf_end_adjustment, based on the same annotation [20]. Coding sequences and predicted amino acid sequences were generated with SQANTI3 v5.5.1 [21]. Candidate DUX4C isoforms were curated using the following criteria: a 5′ UTR longer than 50 bp; a FANTOM5 CAGE peak located within the 5′ UTR; an initiating methionine (M) codon; and the presence of a Kozak consensus sequence at the translation start site.
DUX4 expression, differential gene expression, and gene ontology enrichment analyses
The longer DUX4C isoform (DUX4C-v1) was added to the transcriptome reference (D4Trans-hg38), which was constructed on GENCODE Release 49 (GRCh38.p14) [22]. Short-read RNA-seq data were aligned to D4Trans-hg38 and quantified with Salmon v1.10.2 [23]. Transcript-level estimates from quant.sf files were converted to gene-level values, and all entries annotated as “DUX4” were combined to represent DUX4T expression using the comprehensive GTF annotation. Expression levels were normalized as transcripts per million (TPM), and differential expression analysis of DUX4C and DUX4T was conducted: breast tumors versus matched controls (n = 43 each), FSHD patients versus controls (n = 15 each), and IFNα2-stimulated LCLs versus unstimulated controls (n = 20 each). Statistical significance was assessed using the Mann–Whitney U test implemented in pandas and scipy.stats. Data visualization was carried out using seaborn and matplotlib.
Counts per million (CPM) for the DUX4C-v1 exon 2 region were obtained from samtools-derived coverage. Gene-level count matrices were imported from Salmon outputs using tximport in R. Differential gene expression (DEG) analysis was conducted with DESeq2 on IFNα2-stimulated LCLs, grouped into case samples (DUX4C: 4qα/4qα; DUX4T: 4qB/4qB; CPM > 1; n = 3) and control samples (DUX4C: 4qβ/4qβ; DUX4T: 4qA/4qA; CPM = 0; n = 3) [24]. Control samples were selected from the candidate samples (n = 9) at random using a custom Python script. Genes encoding immunoglobulins (IGH, IGK, and IGL) were removed before Gene Ontology (GO) enrichment. GO enrichment was performed with clusterProfiler using padj < 0.05 and log₂ fold change > 0.5 as thresholds. Enrichment results were processed and visualized with ggplot2 and dplyr, with annotations supplied by org.Hs.eg.db.
Ethics
This study was approved by the Ethics Committee of the Institute of Science, Tokyo (Approved No.: O2019-005-08).
Results
DUX4C haplotypes in human populations
To distinguish DUX4 paralogues, the centromeric DUX4 repeat on chromosome 4 was defined as DUX4C, the terminal DUX4 repeats on chromosomes 4 and 10 were classified collectively as DUX4T, and the intervening array was designated as the D4Z4 tandem repeats (Fig. 1a). The GTEx project previously identified multiple single-nucleotide polymorphisms (SNPs) acting as eQTLs for DUX4C (DUX4L9) in LCLs from European individuals. To explore the underlying genetic structure of this locus, we examined haplotype organization across populations. eQTLs with NES ≥ 0.5 were selected from GTEx (20 variants; Fig. 1a; Table S1a). LD patterns in European and Japanese populations revealed two distinct haplotype blocks in each population as grouped by the LD index (r2 > 0.8) (Fig. 1b, c; Table S1e, g).
Genomic loci and haplotype architecture of DUX4C and DUX4T. a DUX4C (DUX4L9) is located between FRG1 and FRG2 on chromosome 4, whereas DUX4T resides within the terminal D4Z4 repeats on chromosomes 4 and 10. SNPs are shown as colored dots; DUX4C-defining SNPs are indicated in red (genotyping SNPs) and blue. SNPs located in the 3′ region of DUX4C correspond to cCREs. PAS and PAS-like motifs of DUX4T are shown as green bars, and SNP in DUX4-10qA is shown as black dot. b Pairwise LD (r2) matrix of DUX4L9 eQTLs in the European population. The same SNPs and color scheme as in (a) are used to indicate functional annotation. Symbols “●” and “▲” denote block 1 and block 2, respectively. c Pairwise LD (r2) matrix of DUX4L9 eQTLs in the Japanese population. Symbols “□” and “△” denote block 1 and block 2, respectively. d Haplotype frequency distribution of DUX4C-4qα and DUX4C-4qβ in the European population. e Haplotype frequency distribution of DUX4C-4qα and DUX4C-4qβ in the Japanese population
The 3′ region of DUX4C is enriched for cis-regulatory elements (cCREs) across multiple tissues and cell types, as annotated by ENCODE [25], and exhibits hypomethylation in LCLs (methylation rate < 0.2; Table S1b, c). Based on these features, we selected five eQTL SNPs with potential regulatory relevance (rs7696384, rs7696390, rs7654952, rs2006696, and rs1985683). These variants showed strong LD (r2 > 0.95) in both populations and were suitable markers for defining functional haplotypes. Haplotype inference using these variants identified two major haplotypes—T-A-T-A-G and G-G-G-G-C—corresponding to DUX4C-4qα and DUX4C-4qβ, respectively. Their frequencies were similar between populations (DUX4C-4qα: 58% in Europeans, 64% in Japanese; DUX4C-4qβ: 38% in Europeans, 35% in Japanese) (Fig. 1d, e; Table S1d, f), supporting their use as robust genetic markers for the DUX4C locus. The DUX4C haplotype sequences in the GRCh38 and CHM13v2.0 assemblies are identical except for their SNP alleles, with GRCh38 containing DUX4C‑4qβ and CHM13v2.0 containing DUX4C‑4qα.
Comparative analysis of DUX4T haplotypes integrated in reference genomes
DUX4T exhibits multiple haplotypes, among which DUX4T-4qA has been linked to disease susceptibility [2, 15]. In the GRCh38 reference genome, an assembly gap of approximately 82 kb is present downstream of the D4Z4 tandem repeats and upstream of DUX4T on chromosome 4 (Fig. 2b). This missing sequence was later resolved in the CHM13v2.0 assembly using long-read sequencing.
Comparison of GRCh38 and CHM13v2.0 assemblies and DUX4 family genes. a Breakpoint sequences for DUX4T haplotypes, including a 42-bp repeat and 38-bp flanking sequence. SNPs relative to DUX4T-4qA161 are indicated in red. Breakpoint sequences of DUX4T-4qB and DUX4T-10qB, highlighted in yellow, show substantial divergence from those of DUX4T-4qA, DUX4T-10qA, and DUX4T-4qC. Simulated single-end short reads were generated separately for group 1 and group 2 sequences. b Simulated reads mapped to GRCh38 and CHM13v2.0, with group 1 shown in black and group 2 in yellow. c Comparative sequences of DUX4 family genes annotated in the CHM13v2.0 assembly. d Detailed alignment of DUX4 family genes (from top to bottom: DUX4T; chr4-DUX4-D4Z4; chr10-DUX4-D4Z4; and DUX4C (DUX4L9)). Variants relative to DUX4T are shown in red
To evaluate the accuracy of these assemblies, we performed simulated mapping with DUX4T-specific breakpoint sequences. Eighty–base pair junction sequences spanning the transition between the final D4Z4 repeat and the 3′ flanking region—unique to each DUX4T haplotype [15]—were used to generate simulated single-end short reads, which were aligned to both GRCh38 and CHM13v2.0 (Fig. 2a, b; Table S2a).
Mapping results showed that in GRCh38, 56 reads corresponding to group 1 haplotypes (4qA, 4qC, and 10qA) aligned to the DUX4T region (chr4:190,175,561–190,175,579; 3′ side of the assembly gap), whereas 48 reads from group 2 haplotypes (4qB and 10qB) mapped to the final D4Z4 repeat at the 5′ side of the gap (Fig. 2b; Table S2b, c). In contrast, in CHM13v2.0, 36 group 1 reads aligned precisely to chr4:193,543,366–193,543,392 (Table S2d), whereas group 2 reads failed to map. These findings suggest that GRCh38 likely mis-integrated DUX4T-4qA and DUX4T-4qB haplotypes on chromosome 4, whereas CHM13v2.0 accurately represents the DUX4T-4qA haplotype alone, providing a more precise and haplotype-resolved assembly.
Comparative analysis of DUX4-like paralogues in CHM13v2.0 genome
DUX4-like paralogues (DUX4Ls) are dispersed across multiple chromosomes, with intact open reading frames (ORFs) restricted to the D4Z4 tandem repeats on chromosomes 4 and 10 (Fig. 1) [26]. Due to their high sequence similarity to DUX4C and DUX4T, these paralogues can complicate genotyping and isoform identification, especially when using short-read sequencing data. To elucidate their genetic characteristics, we extracted 96 DUX4-like sequences from the CHM13v2.0 assembly for comparative analysis.
Sequence alignment revealed that DUX4Ls embedded within the D4Z4 arrays on chromosomes 4 and 10—together with DUX4T—retain intact ORFs encompassing both HD1 and HD2 and TADs (Fig. 2c). In contrast, DUX4Ls located outside the D4Z4 regions (e.g., on chromosomes 9 and 18) harbor disruptive SNPs within their coding sequences, resulting in frameshifts or premature stop codons. The sole exception was DUX4C (annotated as DUX4L9), which retains both HDs but lacks the C-terminal TAD (Fig. 2c, d). These observations indicate that, in the CHM13v2.0 genome, only DUX4-like genes embedded within the D4Z4 tandem repeat arrays maintain full-length ORFs, whereas DUX4C contains an ORF that lacks the canonical TAD sequence observed in DUX4T. In contrast, DUX4-like loci located outside these arrays have become pseudogenized through accumulated mutations.
Genotyping of DUX4C and DUX4T using the D4Ref-T2T reference
To establish a comprehensive reference for DUX4 genotyping, we first extracted complete haplotype sequences from LWGS data of NA18943 (DUX4C: 4qα/4qα; DUX4T: 4qB/4qB; 10qA/10qA) and NA18948 (DUX4C: 4qβ/4qβ; DUX4T: 4qA/4qA; 10qA/10qA). Consensus haplotype sequences were generated by applying sample-specific variants to the raw haplotypes (DUX4C-4qα, DUX4T-4qB, and DUX4T-10qA from NA18943; DUX4C-4qβ and DUX4T-4qA from NA18948). Haplotype-specific SNPs were validated for DUX4C, and breakpoint sequences were confirmed for DUX4T within each complete haplotype. The 80-bp sequences (breakpoint) of DUX4C-4qα and DUX4C-4qβ, spanning from the repeat region to the 3′ region, were extracted (Table S2e). Simulated reads were generated from these sequences and aligned to the GRCh38 and CHM13v2.0 assemblies using the same strategy as for DUX4T. In both assemblies, reads from both haplotypes mapped to a single locus corresponding to the DUX4C locus (Table S2f–i).
To minimize ambiguous alignment of reads derived from DUX4C and DUX4T to the highly repetitive D4Z4 tandem arrays, all DUX4L sequences within the D4Z4 arrays on chromosomes 4 and 10 were masked in the CHM13v2.0 assembly. Verified haplotype sequences were then incorporated as accessory contigs, generating a DUX4-specific telomere-to-telomere reference genome, termed D4Ref-T2T (Fig. 3a). Remapping of NA18943 and NA18948 LWGS reads confirmed complete haplotype coverage consistent with the known genotypes of both samples (Fig. S1).
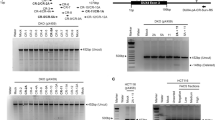
D4Ref-T2T genome architecture and relative base-depth at DUX4 haplotypes. a DUX4 and D4Z4 tandem repeat regions in CHM13v2.0 were masked, and complete DUX4C and DUX4T haplotype sequences were inserted as accessory contigs. b–e Relative base-depths calculated from SWGS data of NA18943 and NA18948. Haplotype-specific motifs are indicated by red lines. Panels show mapping results for b DUX4C-4qα, c DUX4C-4qβ, d DUX4T-4qA, and e DUX4T-B
A genotyping framework for DUX4C and DUX4T was established using SWGS data aligned to D4Ref-T2T. Relative base-depth—defined as per-base coverage normalized to the genome-wide average—was used to assess mapping specificity, with values approaching 1 indicating uniquely mapped, non-redundant alignments. Haplotype-informative positions included SNPs distinguishing DUX4C-4qα from DUX4C-4qβ, the functional PAS specific to DUX4T-4qA, and corresponding 6-bp sequences at syntenic positions of DUX4T-10qA and DUX4T-B, all located within high-specificity regions (Fig. 3b–e).
Genotypes were inferred by calculating the proportion of each haplotype-specific motif relative to the summed coverage of all motifs (Table S3a, b). In the Japanese cohort, D4Ref-T2T–based genotypes of DUX4T matched those from the previous study using simple sequence length polymorphism analysis and Southern blotting [15] in 41 of 45 cases. Among the four unmatched samples, we classified NA18951 and NA19005 as 4qB/4qB rather than 4qA/4qB, based on our results. This decision was supported by the observed proportions of the DUX4T-4qA motif, which were 0 and 0.02 (a single read), respectively. The single 4qA read was manually inspected and determined to be a mismatched read originating from 4qB, indicating a 4qB/4qB genotype. In NA19003, the presence of DUX4T-10qB may have biased the result of 4qB. DUX4T genotyping of chromosome 10 identified two samples—including NA19003—classified as 10qA/10qB. In NA19003, the proportions of DUX4T-4qA (0.38) and DUX4T-10qA (0.31) supported assignment as 4qA/4qA and 10qA/10qB genotypes, which were inconsistent with the previous result of 4qA/4qB and 10qA/10qA. The remaining sample, NA18995, was classified as 4qA/4qA and 10qA/10qA, as both DUX4T-4qA and DUX4T-10qA showed high proportions (0.38 and 0.40). This made it difficult to assign the relatively lower proportion of the DUX4T-B allele (0.23) to either 4qB or 10qB, although its genotype in the previous study was 4qA/4qB and 10qA/10qA. Due to the ambiguous genotyping results observed in NA19003 and NA18995, we excluded these samples, along with two additionally genotyped samples (NA19002 and NA19009), from the subsequent analyses, as they exhibited high proportions of DUX4T-4qA (≥0.375) and DUX4T-B (≥0.05), which may have led to misclassification of 4qA/4qB as 4qA/4qA.
Detection of DUX4C haplotype-derived isoforms and structural characterization
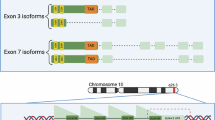
We hypothesized that novel DUX4C isoforms are expressed in IFNα2-stimulated LCLs. Long-read RNA-seq data from NA18943 and NA18948 were aligned to the D4Ref-T2T genome, revealing that DUX4C isoforms were detected only in NA18943. Isoform prediction for DUX4C was therefore performed using a combination of short-read and long-read RNA-seq data from a single IFNα2-stimulated LCL sample (NA18943). After filtering, two novel DUX4C isoforms with intact coding sequences were identified and compared with the previously reported DUX4C transcript (GenBank: AY500824.1) and DUX4L9 annotated in the CHM13v2.0 genome (Fig. 4a). The predicted isoforms contained a defined 5′ untranslated region (UTR; DUX4C-4qα: 14,000–14,174) and 3′ UTR (11,539–12,876), as well as two splice junctions at positions 11,976–12,212 and 11,976–12,219. In contrast, the previously reported DUX4C transcript and the DUX4L9 annotation lacked recorded 5′ and 3′ UTR structures. These features represent the first complete transcript structure reported for DUX4C, and the isoforms were confirmed to originate from the DUX4C-4qα haplotype.
Novel DUX4C isoforms, expression patterns of DUX4C and DUX4T, and functional analysis of DUX4C. a Novel isoforms DUX4C-v1 and DUX4C-v2 compared with previously published DUX4C and DUX4L9 transcripts. The CAGE-seq peak indicating the transcription start site is shown as a green triangle. b Domain architecture of DUX4C isoforms. HD1 and HD2 are shown in blue, and the IDR is shown in pink. Expression of DUX4C and DUX4T in different sample contexts: c breast tumor (43 cases and 43 controls), d FSHD patients (15 cases and 15 controls), and e IFNα2-stimulated LCLs (20 stimulated and 20 non-stimulated). Statistical significance was assessed using the Mann–Whitney U test (p < 0.05). f Haplotype frequency for DUX4C and DUX4T in the Japanese population. g, h GO enrichment analysis of DEGs between IFNα2-stimulated LCLs expressing DUX4C (genotype: DUX4C 4qα/4qα; DUX4T 4qB/4qB, n = 3) and those lacking DUX4C expression (genotype: DUX4C: 4qβ/4qβ; DUX4T: 4qA/4qA, n = 3). Biological Process (BP) of (g) cnetplot: gray nodes represent DEGs, orange nodes represent pathways, and orange node size reflects the number of associated DEGs. h Barplot: bars indicate enriched GO terms, color represents p-value, and length indicates the number of enriched DEGs
We then analyzed the predicted protein domain architecture of DUX4C (DUX4C-v1). The sequence aligned closely with the previously reported DUX4C mRNA (GenBank: AY500824.1) (Fig. 4a). While DUX4T harbors a C-terminal TAD, a structured domain essential for transcriptional activity, DUX4C lacks sequence conservation in this region. Instead, the C-terminus of DUX4C contains an intrinsically disordered region (IDR) (Fig. 4b). To identify potential interaction sites within this region, we used MoRFchibi to predict short Molecular Recognition Features (MoRFs)—binding-prone segments often found in intrinsically disordered sequences [27, 28]. A cluster of high-propensity MoRFs spanning residues 310 to the C-terminus was identified (MCW > 0.7) (Table S4a). We further used Eukaryotic Linear Motif (ELM) predictions to explore functional motifs potentially involved in signaling and regulation mediated by IDR. This analysis revealed several motifs within DUX4C IDR, including those associated with signal transduction (e.g., SH2 and PDZ-binding motifs), post-translational modifications (e.g., CK1 and GSK3 phosphorylation sites), and membrane trafficking (e.g., endocytic sorting signals) (Table S4b). These features suggest that the DUX4C IDR may function as a dynamic regulatory hub mediating protein–protein interactions and cellular signaling, although additional protein–protein interaction assays will be required to validate this possibility.
Expression dynamics of DUX4C and DUX4T across biological contexts, and a chemotactic role for DUX4C
To investigate potential functions of DUX4C, we constructed a customized transcriptome reference, D4Trans-hg38, by updating GENCODE Release 49 (GRCh38.p14) to replace the DUX4L9 entry with the full-length DUX4C-v1 transcript identified from our long-read RNA-seq data. Short-read RNA-seq datasets from diverse biological contexts—including human breast tumors (n = 43), FSHD patient samples (n = 15), and IFNα2-stimulated LCLs (n = 20)—were quantified against D4Trans-hg38. Both DUX4C and DUX4T were significantly upregulated in breast tumor tissues relative to normal breast samples (Fig. 4c). In contrast, only DUX4T exhibited significant upregulation in FSHD patient tissues, while DUX4C expression remained unchanged (Fig. 4d). Neither gene showed significant expression changes upon IFNα2 stimulation in LCLs (Fig. 4e). Interestingly, five samples in the FSHD control (unaffected family members of FSHD patients) exhibited elevated DUX4C expression (TPM > 1), suggesting that DUX4C may exert weaker myotoxic effects or potentially serve tissue-protective or reparative roles compared with DUX4T.
To further explore the relationship between genetic background and DUX4C function, we analyzed D4Ref-T2T–based haplotypes of DUX4C and DUX4T in the Japanese cohort (n = 100). We identified 123 4qα-4qB and 70 4qβ–4qA haplotypes, accounting for 61.5% and 35% of all haplotypes, respectively (Fig. 4f; Table S3c), where a strong LD was observed between these two loci (r2 = 0.86). These findings suggest a potential linkage between DUX4C-4qα and DUX4T-4qB, indicating that DUX4C expression may be modulated by DUX4T at the genetic level, possibly functioning in a compensatory or modulatory manner.
Given the potential haplotype effects of DUX4C, IFNα2-stimulated LCLs were stratified into case (DUX4C: 4qα/4qα; DUX4T: 4qB/4qB; CPM > 1; n = 3) and control (DUX4C: 4qβ/4qβ; DUX4T: 4qA/4qA; CPM = 0; n = 3) (Table S5a) for DEG and GO enrichment analyses, providing a limited yet exploratory assessment of DUX4C function. Upregulated genes (Table S5b) in the case (high-DUX4C expressing LCLs) were significantly enriched in pathways related to immune-related chemotactic processes, including “leukocyte chemotaxis,” “cell chemotaxis,” and “lymphocyte chemotaxis” (Fig. 4g, h; Table S5c), suggesting that DUX4C may modulate immune cell recruitment or migration in a haplotype-specific manner. This functional association is further supported by ELM-predicted linear motifs within the C-terminal IDR of DUX4C, which include interaction domains such as SH2, PDZ, and phosphorylation sites. These motifs are commonly found in signaling proteins involved in immune pathways, raising the possibility that DUX4C indirectly contributes to chemotactic signaling through dynamic protein–protein interactions or post-translational modifications.
Discussion
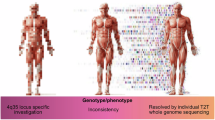
In this study, we developed a DUX4-specific reference genome (D4Ref-T2T) and transcriptome (D4Trans-hg38), enabling accurate genotyping and isoform identification of DUX4C and DUX4T. Comparison of genotyping results between CHM13v2.0 and D4Ref-T2T indicated that both references can reliably resolve DUX4C genotypes. In contrast, DUX4T-4qB lacks a complete reference sequence in CHM13v2.0 and diverges substantially from the DUX4T-4qA sequence represented. D4Ref-T2T addresses this limitation by providing a comprehensive reference that supports large-scale DUX4T genotyping using SWGS data.
A limitation of the current D4Ref-T2T genotyping framework is the observed discrepancy in four samples relative to previous reports, suggesting that PCR validation with DUX4T-specific primers may be necessary for definitive genotyping. Additionally, as previous studies have demonstrated that DUX4T expression is regulated by methylation within the D4Z4 tandem repeats [2], DUX4C expression may similarly be affected by local methylation. This underscores the importance of evaluating D4Z4 repeat structures to elucidate the functional roles of DUX4 haplotypes. However, evaluating these effects remains technically challenging: SWGS reads are substantially shorter than the ~3.3-kb D4Z4 repeat unit, and even with D4Ref-T2T, multi-mapping ambiguity persists within the repeat region (Fig. 3b–e). Consequently, investigating DNA methylation—particularly in a phased context—and its regulatory impact on DUX4C represents a critical avenue for future research.
Beyond establishing a genotyping framework, we identified previously uncharacterized full-length DUX4C isoforms in LCLs. Because DUX4C isoforms were determined from a single sample (NA18943), this analysis is limited in its ability to establish haplotype associations, and additional functional isoforms may remain undiscovered. We observed that in short-read RNA-seq data from IFNα2-stimulated LCL controls, a subset of samples with the DUX4C-4qβ/4qβ genotype exhibited low but detectable DUX4C expression (TPM < 0.15; Table S6b), suggesting the possible presence of DUX4C transcripts derived from the DUX4C-4qβ haplotype. However, in the absence of long-read RNA-seq data, the presence and structure of these transcripts could not be conclusively validated. In contrast, samples with the DUX4C: 4qα/4qα genotype consistently showed higher DUX4C expression levels (TPM ~ 2.1), with a statistically significant difference compared with the 4qβ/4qβ group (Mann–Whitney U test, p < 0.05; Table S6a–c). Taken together, these results suggest that the DUX4C-4qα haplotype may enhance DUX4C expression, rather than acting as a strict determinant of transcriptional activity.
Although DUX4C expression may be a consequence of a specific cellular state or other regulatory factors, we discuss the potential roles of DUX4C in signaling pathways. Functional analyses suggest that DUX4C harbors multiple phosphorylation-related motifs within its C-terminal region, including CK1 and GSK3β sites. These motifs are associated with signaling modules involved in β-catenin degradation and NFκB activation [29], both of which are central regulators of immune responses. Along with chemotaxis-related gene enrichment in DUX4C-expressing LCLs, these features suggest that DUX4C may promote immune activation. This contrasts with DUX4T, which suppresses interferon-stimulated gene expression [30]. Their opposing functions and distinct expression patterns point to haplotype structure as a potential mechanism balancing DUX4C and DUX4T activity.
Indeed, haplotype analysis of DUX4C and DUX4T revealed a strong LD between these two loci (r2 = 0.86) (Table S3c), although the subtelomeric regions of the genome are known to be susceptible to recombination. Two predominant haplotypes—4qα–4qB and 4qβ–4qA—accounted for 97% of all haplotype combinations. This skewed distribution raises the possibility that negative selection acts against the 4qα–4qA haplotype, which expresses both DUX4C and functional DUX4T, as well as against 4qβ–4qB, which expresses neither. As DUX4T is expressed during the early pre-implantation stage of human embryogenesis [31], the mutually exclusive expression of DUX4C and DUX4T, governed by two functional haplotypes, may play pivotal roles in human embryonic development.
In summary, our work establishes a framework for analyzing genes located in subtelomeric repeat regions and elucidates a potential mechanistic relationship between DUX4C and DUX4T in regulating immune activity. These findings lay the groundwork for future studies integrating genomic structure, methylation patterns, and transcript isoform diversity to better understand DUX4-related immune phenotypes.
References
Chew GL, Campbell AE, Neef ED, Sutliff NA, Shadle SC, Tapscott SJ, et al. DUX4 suppresses MHC Class I to promote cancer immune evasion and resistance to checkpoint blockade. Dev Cell. 2019;50:658–71.e7.
Lemmers RJLF, van der Vliet PJ, Klooster R, Sacconi S, Camaño P, Dauwerse JG, et al. A unifying genetic model for facioscapulohumeral muscular dystrophy. Science. 2010;329:1650–3.
Nurk S, Koren S, Rhie A, Rautiainen M, Bzikadze AV, Mikheenko A, et al. The complete sequence of a human genome. Science. 2022;376:44–53.
Ansseau E, Laoudj-Chenivesse D, Marcowycz A, Tassin A, Vanderplanck C, Sauvage S, et al. DUX4c is up-regulated in FSHD. It induces the MYF5 protein and human myoblast proliferation. PLoS ONE. 2009;4:e7482.
Banerji CRS, Panamarova M, Zammit PS. DUX4 expressing immortalized FSHD lymphoblastoid cells express genes elevated in FSHD muscle biopsies, correlating with the early stages of inflammation. Hum Mol Genet. 2020;29:2285–99.
Li SY, Hammarlund JA, Wu G, Lian JW, Howell SJ, Clarke RB, et al. Tumor circadian clock strength influences metastatic potential and predicts patient prognosis in luminal A breast cancer. Proc Natl Acad Sci USA. 2024;121:e2311854121.
Ueda MT, Inamo J, Miya F, Shimada M, Yamaguchi K, Kochi Y. Functional and dynamic profiling of transcript isoforms reveals essential roles of alternative splicing in interferon response. Cell Genom. 2024;4:100654.
Logsdon GA, Ebert P, Audano PA, Loftus M, Porubsky D, Ebler J, et al. Complex genetic variation in nearly complete human genomes. Nature. 2025;644:430–41.
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience. 2015;4:7.
Browning BL, Tian X, Zhou Y, Browning SR. Fast two-stage phasing of large-scale sequence data. Am J Hum Genet. 2021;108:1880–90.
Li H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics. 2018;34:3094–100.
Ahsan MU, Gouru A, Chan J, Zhou W, Wang K. A signal processing and deep learning framework for methylation detection using Oxford Nanopore sequencing. Nat Commun. 2024;15:1448.
Li H, Durbin R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics. 2010;26:589–95.
Davis MW, Jorgensen EM. ApE, A Plasmid Editor: a freely available DNA manipulation and visualization program. Front Bioinform. 2022;2:818619.
Lemmers, Vliet RJLF, van der PJ, Gaag, van der KJ, Zuniga, et al. Worldwide population analysis of the 4q and 10q subtelomeres identifies only four discrete interchromosomal sequence transfers in human evolution. Am J Hum Genet. 2010;86:364–77.
Jain C, Rhie A, Hansen NF, Koren S, Phillippy AM. Long-read mapping to repetitive reference sequences using Winnowmap2. Nat Methods. 2022;19:705–10.
Shafin K, Pesout T, Chang PC, Nattestad M, Kolesnikov A, Goel S, et al. Haplotype-aware variant calling with PEPPER-Margin-DeepVariant enables high accuracy in nanopore long-reads. Nat Methods. 2021;18:1322–32.
Robinson JT, Thorvaldsdóttir H, Winckler W, Guttman M, Lander ES, Getz G, et al. Integrative genomics viewer. Nat Biotechnol. 2011;29:24–6.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Tang AD, Felton C, Hrabeta-Robinson E, Volden R, Vollmers C, Brooks AN. Detecting haplotype-specific transcript variation in long reads with FLAIR2. Genome Biol. 2024;25:173.
Pardo-Palacios FJ, Arzalluz-Luque A, Kondratova L, Salguero P, Mestre-Tomás J, Amorín R, et al. SQANTI3: curation of long-read transcriptomes for accurate identification of known and novel isoforms. Nat Methods. 2024;21:793–7.
Mudge JM, Carbonell-Sala S, Diekhans M, Martinez JG, Hunt T, Jungreis I, et al. GENCODE 2025: reference gene annotation for human and mouse. Nucleic Acids Res. 2025;53:D966–75.
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C. Salmon: fast and bias-aware quantification of transcript expression using dual-phase inference. Nat Methods. 2017;14:417–9.
Soneson C, Love MI, Robinson MD. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Research. 2016;4:1521.
Moore JE, Purcaro MJ, Pratt HE, Epstein CB, Shoresh N, Adrian J, et al. Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature. 2020;583:699–710.
Clapp J, Mitchell LM, Bolland DJ, Fantes J, Corcoran AE, Scotting PJ, et al. Evolutionary conservation of a coding function for D4Z4, the tandem DNA repeat mutated in facioscapulohumeral muscular dystrophy. Am J Hum Genet. 2007;81:264–79.
Kumar M, Michael S, Alvarado-Valverde J, Zeke A, Lazar T, Glavina J, et al. ELM—the Eukaryotic Linear Motif resource—2024 update. Nucleic Acids Res. 2024;52:D442–55.
Malhis N, Jacobson M, Gsponer J. MoRFchibi SYSTEM: software tools for the identification of MoRFs in protein sequences. Nucleic Acids Res. 2016;44:W488–93.
Xue C, Chu Q, Shi Q, Zeng Y, Lu J, Li L. Wnt signaling pathways in biology and disease: mechanisms and therapeutic advances. Sig Transduct Target Ther. 2025;10:106.
Spens AE, Sutliff NA, Bennett SR, Campbell AE, Tapscott SJ. Human DUX4 and mouse Dux interact with STAT1 and broadly inhibit interferon-stimulated gene induction. eLife. 2023;12:e82057.
Vuoristo S, Bhagat S, Hydén-Granskog C, Yoshihara M, Gawriyski L, Jouhilahti EM, et al. DUX4 is a multifunctional factor priming human embryonic genome activation. iScience. 2022;25:104137.
Acknowledgements
The authors thank Ayako Ishii for technical assistance and Satomi Mitsuhashi for her initial research guidance. The authors are also grateful to Nao Nishida and Takayo Tsuchiura for their valuable experimental advice. The authors thank Dr. Richard J.L.F. Lemmers for providing the DUX4T genotype data. This work was supported by the Japan Society for the Promotion of Science (JSPS), Grant-in-Aid for Scientific Research (B) and (C) (grant numbers: 22H02597, 24K10050) from the MEXT of Japan. This work was also supported by grants from Nanken-Kyoten; Medical Research Center Initiative for High Depth Omics; the Kobayashi Foundation; and the Uehara Memorial Foundation.
Author information
Authors and Affiliations
Contributions
Conceptualization was carried out by ZZ, MTU, KY, and YK. All experiments and in silico analyses were performed by ZZ under the supervision of MTU, KY, and YK. ZZ drafted the original manuscript, and all authors contributed to its review and editing. Funding acquisition and overall supervision were provided by YK.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhuang, Z., Ueda, M.T., Yamaguchi, K. et al. A new integrated genetic and transcriptomic approach for investigating DUX4 and DUX4C. J Hum Genet (2026). https://doi.org/10.1038/s10038-025-01450-x
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s10038-025-01450-x