Abstract
Palm-leaf manuscripts (PLMs), essential Asian cultural artifacts, necessitate accurate species identification for their preservation. Traditional destructive analysis or manual inspection prone to errors hampers large-scale conservation efforts. To address this, we present PLNet, a lightweight framework integrating Lion-optimized EfficientNetB2 with domain-specific augmentation strategies. Lion optimizer enhances training stability, while task-driven augmentation simulates manuscript degradation. PLNet achieves exceptional performance on a diverse test set: 99.07% accuracy and 99.21% F1-score, outperforming ResNet50, DenseNet169, and YOLOv11-cls. Crucially, PLNet operates with high efficiency, utilizing only 7.7 million parameters and processing a folio in 0.31 s. To enable practical application, we developed PLNet-GUI for large-scale identification. Validated on 142,679 PLMs from 8 countries, the framework demonstrates robust real-world applicability, reveals regional species distribution patterns, and establishes a method shift in heritage conservation by replacing destructive practices with AI-driven preservation.
Similar content being viewed by others
Introduction
Palm-leaf manuscripts (PLMs), serving as primary carriers of historical, cultural, and scientific knowledge across South and Southeast Asia for over two millennia, represent an irreplaceable component of global documentary heritage1. These manuscripts, primarily crafted from the leaves of Corypha umbraculifera and Borassus flabellifer palms, document diverse fields ranging from astronomy to philosophy, offering unique insights into pre-modern civilizations2. With an estimated one million PLMs extant worldwide, concentrated in India, Myanmar, Thailand, and China, their preservation is critically urgent3,4,5. However, accelerated degradation due to aging, environmental factors, and improper storage threatens this fragile heritage, demanding immediate conservation interventions6,7.
A critical prerequisite for effective conservation is the precise identification of the botanical species used in each manuscript. Species knowledge directly impacts traceability studies, material compatibility in restoration treatments, and preservation strategy formulation8,9,10. Traditional identification relies on two approaches: manual inspection and anatomical analysis. Manual inspection, although non-destructive, suffers from high inter-observer variability due to subjective assessment of visual traits (e.g., texture, color)11,12,13. Anatomical analysis, though accurate (leveraging distinct histological symmetries, dorsiventral in Corypha umbraculifera vs. iso-bilateral in Borassus flabellifer; Fig. S1), fundamentally violates heritage ethics14,15,16. This method requires destructive sampling (5–10 cm2 loss per folio) involving slicing, chemical staining, and microscopy, causing irreversible damage17,18. The tension between accurate identification and preservation integrity highlights an imperative need for scalable, objective, and non-destructive alternatives.
Recent digitization initiatives (e.g., British Library, Potala Palace) have generated >200,000 high-resolution PLM images, enabling computational solutions19,20. Early computational techniques for plant leaf classification combined hand-crafted features (e.g., morphology, color, texture descriptors) with classifiers such as neural networks21,22, SVM23,24, random forest25, and k-nearest neighbor26. However, these methods fail to generalize to PLMs due to unique challenges, including surface abrasions, ink residues, and uneven degradation obscure discriminative features absent in fresh/herbarium specimens. Deep learning approaches (e.g., Modified GoogleNet27, Dual-path CNN28, SWP-LeafNET29) automate feature extraction but face limitations including high computational costs, architectural complexity, cascading latency, and critically a focus on pristine specimens rather than degraded historical materials. Among these, EfficientNet30 offers a promising balance of accuracy and efficiency via compound scaling of depth, width, and resolution. Nevertheless, its direct application to PLMs remains suboptimal, as surface degradation and variable digitization quality compromise feature discriminability.
To bridge this gap, we propose PLNet, a novel deep learning framework designed specifically for non-destructive, rapid, and robust identification of PLM plant species. Our approach integrates a Lion-optimized EfficientNetB2 architecture with task-specific data augmentation strategies. Validated on a geographically diverse dataset spanning eight countries, PLNet achieves 99.07% accuracy at 0.31 s per folio, outperforming state-of-the-art models (ResNet50, DenseNet169, YOLOv11-cls) while using only 7.7 million parameters. We further deploy this framework via PLNet-GUI, an open-source tool enabling large-scale analysis. Applied to 142,679 manuscripts, PLNet reveals distinct regional species distribution patterns, demonstrating its dual utility in conservation science and historical studies. This work establishes a solution shift from destructive sampling to AI-driven preservation, safeguarding humanity’s fragile written legacy.
Methods
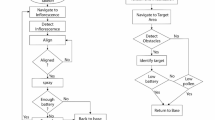
PLNet consists of three main components: dataset preparation, PLNet model construction, and PLM species prediction (Fig. 1a).
a The workflow of PLNet. b Dataset preparation. c The architecture of PLNet based on EfficientNet. d PLNet-based PLM species prediction.
Dataset preparation
The construction of the PLM dataset prioritized geographical diversity, preservation condition variability, and species balance to ensure robust model generalization. The dataset preparation process is shown in Fig. 1b. A total of 4004 images of PLM were sourced from the Potala Palace and the British Library’s open-access collections (https://www.bl.uk/), covering manuscripts from 8 countries. Then, 345 images of Corypha umbraculifera and Borassus flabellifer from Zhanjiang, China, were captured under controlled conditions (SONY IMX682 camera, 3456 × 4560 resolution) to establish baseline morphological features. The combined dataset of 4349 images was stratified by species and geographical origin, then split into training (80%, 3478 images) and validation (20%, 871 images) sets. The training set was used for model training, and the validation set was used for hyperparameter tuning. To rigorously evaluate generalization, a separate test set was constructed by selecting 1 image per manuscript volume from 642 volumes across 8 Asian repositories. This curation ensured zero content overlap with the training/validation data, resulting in a test set of 382 Corypha umbraculifera and 260 Borassus flabellifer images. This fully independent test set was used to evaluate the generalization ability of the model and compare performance differences between different models. Table S1 shows the quantities of the training, validation, and test sets, as well as their respective source distributions. Fig. S2 shows typical PLMs crafted from Corypha umbraculifera and Borassus flabellifer in the dataset.
The background of the digital images of PLM was first removed to eliminate its impact on model training. These images were then cropped into patches, allowing the CNN to focus more on the local details of the PLMs. Subsequently, the PLM boundary was obtained. A certain margin was left on each side of a PLM to ensure that the sub-image encompassed the entire manuscript. Patches with more than 20% background pixels were filtered out because they contained insufficient effective features and could potentially affect model training.
We implemented task-specific data augmentation using the Albumentations library31 to address unique challenges in PLM imagery. During training, each image underwent real-time transformations, including rotation, scaling, cropping, flipping, and color adjustments, dynamically tailored to simulate manuscript degradation patterns. These operations were optimized to amplify biologically discriminative features (e.g., vein structures, edge characteristics), ensuring augmented data aligned with species identification objectives. This approach enhanced training set diversity while mitigating overfitting and improving generalization. Detailed augmentation parameters are provided in Table S2.
Architecture of PLNet
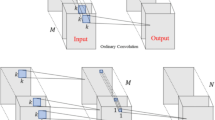
The core architecture of PLNet (Fig. 1c) builds upon EfficientNetB230, selected after rigorous evaluation of the EfficientNet family (B0-B7). As demonstrated in Fig. S3, EfficientNetB2 achieved peak test accuracy, the primary metric for real-world deployment, while balancing computational efficiency. PLNet maintains the foundational structure of EfficientNetB2, which consists of seven sequential blocks, each with multiple Mobile Inverted Bottleneck Convolution (MBConv) modules, including variants MBConv1 and MBConv6. Detailed schematics are provided in Fig. S4. The original 1000-class output layer was replaced with a binary classification head (2-unit layer + softmax activation). A dropout layer (rate=0.3) was inserted prior to the output to suppress overfitting. The model was initialized using EfficientNetB2 weights pre-trained on ImageNet. Full fine-tuning was performed on the patches of the PLM training dataset to adapt domain-specific features.
To address high inter-class similarity in PLM images, we employed label-smoothed cross-entropy loss32,33. For binary classification, the loss is defined as:
where y is the true label (0 or 1) and \(\widehat{y}\) is the predicted probability of the class label being 1.
We adopted the Lion optimizer34 due to its superior hyperparameter robustness and convergence stability. Unlike Adam-based methods requiring second-moment estimates, Lion uses a sign-based update rule:
\({w}_{t+1}\) is the weight at the next iteration, \({w}_{t}\) is the current weight, lr is a predefined step size, \({\rm{sign}}({m}_{t})\) is the sign of the momentum at the current iteration, \({m}_{t}\) is the current momentum, β is the momentum decay factor, \({m}_{t-1}\) is the momentum from the previous iteration, and \({g}_{t}\) is the gradient at the current iteration. This design minimizes sensitivity to learning rate and weight decay (validated in Fig. 2), crucial for distributed training. Lion achieved 2.5% higher accuracy than AdamW on ImageNet34, aligning with our need for efficient large-batch optimization. The binary output layer is intentionally designed as a pluggable component, enabling future expansion to multi-species identification (e.g., incorporating additional palm taxa) without structural overhaul.
Comparison of Adam and Lion optimizer loss with varying a learning rates and b weight decays.
Species identification for PLMs
During the prediction phase, a single PLM image is first divided into multiple non-overlapping patches, each measuring 224 × 224 pixels. Each patch is processed individually by the PLNet classifier to derive the classification logits for the two species, Borassus flabellifer and Corypha umbraculifera. To form a single, robust prediction for the entire manuscript, the logits from all constituent patches are averaged. The class with the highest average logit is then selected as the final identification result. This aggregation strategy was deliberately chosen to ensure that the model considers features from the entire manuscript surface, making the final prediction resilient to localized artifacts such as ink residue, stains, or physical damage that might corrupt individual patches and mislead the classifier. Averaging logits, which represent the unnormalized confidence of the model, yields higher accuracy on the test set compared to averaging probabilities, as shown in Table S3. Therefore, averaging logits provides a more nuanced, reliable, and accurate final decision for the PLNet framework. All performance metrics reported in this study are evaluated at this final PLM level to accurately reflect the model’s real-world utility in classifying entire manuscript folios.
Evaluation metrics
The metrics were used to quantitatively evaluate the identification performance, including accuracy, precision, sensitivity, and F1-score. The formulas for these evaluation metrics are provided below:
TP represents the number of images accurately identified as Borassus flabellifer, while TN signifies the number of images correctly identified as Corypha umbraculifera. Conversely, FN indicates the number of images incorrectly attributed to Corypha umbraculifera, and FP indicates the number of images incorrectly classified as Borassus flabellifer. These metrics can be obtained from a confusion matrix. Furthermore, the confusion matrix is a structured representation that compares the actual PLM labels with the model’s predictions. These metrics offer comprehensive evaluations of the model’s generalizability.
Uncertainty quantification
To quantify the model’s confidence in its predictions at the manuscript level, we calculate the Shannon entropy35 of the final probability distribution. This metric serves as a powerful indicator of classification uncertainty. The process begins after the patch-level logits are averaged to produce a final logit vector for the entire manuscript. This vector is then normalized into a probability distribution, denoted as P, using the softmax function. The Shannon entropy is subsequently calculated using the formula:
where P(i) is the probability of the i-th class. A low entropy value, close to 0, signifies a high-confidence prediction where one class has a very high probability, whereas a high entropy value, near 1 in a binary case, indicates high uncertainty, with probabilities distributed more evenly between classes. This allows the framework to flag low-confidence predictions for subsequent human review.
Guided GradCAM
In deep learning, models often make decisions in ways that are not directly observable. Guided GradCAM is an advanced visualization technique that merges the concepts of Guided Backpropagation (GBP) and Gradient-weighted Class Activation Mapping (GradCAM) to provide high-resolution, class-discriminative visual explanations for decisions made by CNNs. This method was proposed36 to address the limitations of CAM and to provide a more detailed understanding of model decisions.
The formula for Guided GradCAM involves an element-wise product of GBP and GradCAM attributions. The GradCAM attributions are computed with respect to a convolutional layer and are upsampled to match the input size. The GBP is a modified backpropagation technique that only permits positive gradients to pass through ReLUs, offering fine-grained visualizations. The mathematical representation of GradCAM is as follows:
where \({L}_{{\rm{GradCAM}}}^{c}\) is the GradCAM localization map for class (c), \({\alpha }_{k}^{c}\) represents the neuron importance weights, and \({A}^{k}\) are the forward activation maps of a convolutional layer.
Guided GradCAM synthesizes its results by merging the visual cues from GradCAM with the detailed guidance from GBP. This fusion is achieved by multiplying the specific elements of GBP attribution with a GradCAM visual map on a one-to-one basis, known as the Hadamard product. This method enhances the clarity and sharpness of the resulting attributions. For Guided GradCAM, the final visualization is obtained by:
where \(\odot\) denotes element-wise multiplication, and G is the guided backpropagation gradient map.
Results
Hyperparameter optimization and model training
To benchmark the performance of the PLNet architecture, we compared it against three representative models, including ResNet5037, DenseNet16938, and YOLO11x-cls39. ResNet50 serves as the baseline model, given its foundational role in deep learning. DenseNet169 and YOLO11x-cls are considered state-of-the-art models for the feature reuse capability and high-efficiency, respectively.
All models were implemented in Python 3.11.5 with PyTorch 2.2.0. Hyperparameter optimization and model training were conducted on a workstation equipped with an Intel(R) Xeon(R) W5-3435X processor, 512 GB of RAM, and two NVIDIA RTX 6000 Ada 48 G graphics cards for acceleration.
We conducted Bayesian optimization40 over 20 trials to determine optimal hyperparameters for all models (Table S4). The process of optimizing the initial learning rate, ranging from 10-7 to 10-5, and weight decay, from 0.01 to 1. A learning rate scheduler utilizing cosine decay was employed to adjust the learning rate throughout the training process dynamically. Additionally, to prevent overfitting, early stopping was implemented with a threshold of 5 epochs without improvement. Considering the balance of GPU memory and Lion optimizer efficiency, the batch size was established at 256.
The outcomes of hyperparameter optimization and the validation accuracy for ResNet50, DenseNet169, YOLO11x-cls, and PLNet are depicted in Fig. S5 and Table S5. On the validation set, PLNet achieved the highest accuracy of 99.31%, outperforming all competitors by >0.23%, while demonstrating robust convergence during optimization.
Comparison of lion and adam optimizers
We systematically evaluated the robustness of the Lion optimizer34 against the widely adopted Adam by analyzing validation loss sensitivity to hyperparameters (Fig. 2). Figure 2a shows the best validation loss as a function of the learning rate. The Lion optimizer consistently achieves lower validation loss than Adam across all tested learning rates (10−8–10−3). Both optimizers attained optimal performance near 10−4, but Lion consistently reaches a lower minimum loss, demonstrating its efficiency. The difference in robustness is most evident in Fig. 2b. Lion maintains near-constant low loss ( < 0.05) across weight decay values spanning 10−6 to 10−1. In contrast, the Adam optimizer is highly sensitive to weight decay. It performs well only at lower values (10−6 to 10−4), and the loss increases sharply after a weight decay of 10−3. The Lion optimizer demonstrates significant insensitivity to hyperparameters, ensuring stable convergence even with suboptimal hyperparameter settings. This capability substantially reduces optimization costs while guaranteeing reliable model performance. This robustness is critical for heritage informatics, where computational resources and tuning expertise may be constrained. The superior stability and efficiency of Lion validate its integration into PLNet.
Evaluation of model performance
All performance metrics were evaluated at the manuscript level, where the final classification output for each complete PLM image was derived by aggregating patch-level predictions. This approach ensures that reported results reflect real-world operational conditions. As illustrated in Fig. 3a, PLNet demonstrated superior performance across all key metrics on the independent test set, achieving the highest accuracy (99.07%), sensitivity (99.21%), and F1-score (99.21%) among the benchmarked models. While ResNet50 attained a marginally higher precision (99.47%), this advantage was negated by its substantially larger parameter footprint (23.51 million parameters vs. PLNet’s 7.70 million) and slower inference speed (0.40 s per folio vs. PLNet’s 0.31 s), as quantified in Fig. 3b and Tables S6–S7. PLNet exhibited exceptional efficiency without compromising accuracy. It operated 1.8× faster than DenseNet169 (0.31 s/folio vs. 0.48 s/folio) while maintaining near-perfect classification performance. The modern YOLOv11x-cls emerged as a competitive alternative, achieving a high F1-score of 98.68%. However, PLNet consistently outperformed it across all metrics, utilizing nearly four times fewer parameters (7.70 million vs. 29.64 million) and delivering faster inference. This efficiency stems from EfficientNet’s compound scaling methodology, which optimally balances network depth, width, and input resolution against computational constraints.
a Performance comparison of ResNet50, DenseNet169, YOLO11x-cls, and PLNet in terms of accuracy, precision, sensitivity, and F1-score. b Radar chart depicting the comparative metrics of ResNet50, DenseNet169, YOLO11x-cls, and PLNet in terms of the training time, inference speed, parameter numbers, and model size. c Confusion matrix of ResNet50, DenseNet169, YOLO11x-cls, and PLNet.
The confusion matrix analysis (Fig. 3c) further validated PLNet’s robustness under real-world degradation conditions. It correctly identified 257 out of 260 Borassus flabellifer samples (98.8% recall) and 379 out of 382 Corypha umbraculifera samples (99.2% recall), with merely three misclassifications per species. Such balanced performance fully demonstrates PLNet’s capability to discern subtle interspecies morphological features despite manuscript deterioration.
To assess predictive reliability, we quantified classification uncertainty using Shannon entropy. The results revealed a strong inverse correlation between entropy and accuracy. Correctly classified manuscripts exhibited exceptionally low entropy (0.0299), while errors corresponded to high uncertainty (0.7627; Table S8). This enables automated flagging of low-confidence predictions for expert review, enhancing the framework’s utility in conservation workflows.
In summary, PLNet achieves an optimal equilibrium between accuracy (99.07%), efficiency (0.31 s/folio), and reliability, establishing itself as an indispensable tool for large-scale non-destructive PLM analysis. Its compact architecture, coupled with self-diagnostic capability through entropy monitoring, addresses critical needs in cultural heritage preservation.
Model interpretability
Although deep neural networks often function as “black boxes”, understanding their decision rationale is paramount in scientific applications. To illuminate PLNet’s classification logic, we employed Guided Grad-CAM, a state-of-the-art visualization technique that fuses gradient-weighted class activation maps with guided backpropagation. This method generates high-resolution heatmaps (Fig. 4) by aggregating channel-wise gradients from shallow convolutional layers and overlaying them onto input images through element-wise multiplication. The resulting saliency maps pinpoint pixel regions that critically influence species determinations, thereby bridging artificial intelligence with botanical expertise.
a Original images of Corypha umbraculifera. b Interpretable explanation based on Guided Grad-CAM for Corypha umbraculifera. c Original images of Borassus flabellifer. d Interpretable explanation based on Guided Grad-CAM for Borassus flabellifer.
Crucially, these visualizations reveal that PLNet prioritizes morphologically discriminative traits for species identification, such as vein architecture, marginal features, and surface artifacts. Corypha umbraculifera exhibits parallel secondary veins (Fig. 4b), whereas Borassus flabellifer displays reticulate tertiary veins (Fig. 4d). The serrated edges of Corypha umbraculifera versus the smooth margins of Borassus flabellifer serve as key discriminators. Furthermore, distinct tool-induced engraving patterns emerge on each species due to their differing leaf material properties, providing another morphological distinction.
It is worth noting that these visualizations generated from shallow layers (Fig. 4b, d) provide biologically interpretable insights, in contrast to those from deeper layers, which encode more abstract and less localized representations. This alignment between model attention and botanical taxonomy confirms PLNet’s capacity to replicate expert reasoning, prioritizing taxonomically significant features without compromising computational efficiency. The framework thus achieves a critical dual objective, operational speed essential for large-scale digitization projects, and interpretability necessary for scholarly validation in heritage science.
PLNet-GUI: bridging AI and heritage conservation
To empower cultural heritage practitioners with minimal technical expertise, we developed PLNet-GUI, an intuitive cross-platform interface (Windows/macOS/Linux) that operationalizes non-destructive species identification. Built on Python 3.11 and PyQt5 5.15, this software integrates into digitization workflows through three core modules (Fig. 5). Single-Folio Analysis: Users load individual manuscript images via an embedded file browser (Fig. 5a). Real-time processing ( <1 second) displays species probabilities alongside confidence scores (Fig. 5b), enabling instant verification during conservation assessments. Batch Processing Mode: Toggled via a dedicated interface (Fig. 5d), this module processes entire manuscript collections, displaying aggregated results in a sortable table. Conservators can rapidly compare hundreds of samples while filtering by species, confidence, or origin, which is critical for large-scale provenance studies. Data Curation & Reporting: Export functionalities generate structured CSV files with species IDs, confidence metrics, and entropy-based uncertainty flags, and publication-ready PDF reports containing statistical summaries (e.g., species and confidence distribution). This automates documentation for conservation archives and scholarly publications. The interface design prioritizes accessibility, zero-code operation and drag-and-drop functionality. By combining the computational efficiency of PLNet (0.31 s/folio) with the GUI development technique, PLNet-GUI transforms cutting-edge AI into a practical tool for museums, libraries, and field conservation teams engaged in palm-leaf manuscript preservation.
a Load PLM images. b Identification result. c Save data and generate reports. d Display the batch identification result.
Application in PLMs across different countries
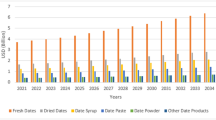
Leveraging the non-destructive identification capability of PLNet, we analyzed 142,679 PLMs from eight Asian countries. This continental-scale investigation uncovered striking geographical patterns in palm species utilization (Fig. 6), offering new insights into historical craft traditions and ecological adaptations.
Percentage distribution of Borassus flabellifer and Corypha umbraculifera across different Asian countries, as classified by PLNet.
Corypha umbraculifera prevailed across mainland Southeast Asia, exceeding 95% prevalence in China (99.96%, n = 21,083), Laos (99.93%, n = 3004), Thailand (99.67%, n = 612), Vietnam (99.43%, n = 2947), and Nepal (100%, n = 40). Its near-exclusive dominance suggests established cultivation networks and cultural preference for its workability in humid tropical regions. Borassus flabellifer exhibited concentrated adoption in archipelagic and arid zones. Indonesia has the highest percentage of Borassus flabellifer at 64.66% (n = 2497). India also has a substantial proportion of Borassus flabellifer at 25.54% (n = 28,204). The Indonesian anomaly (Borassus flabellifer > Corypha umbraculifera) reflects deep-rooted ecological adaptation. Unlike mainland regions reliant on monsoon-driven Corypha umbraculifera cultivation, eastern Indonesian communities historically leveraged Borassus flabellifer—a species uniquely adapted to semi-arid climates41. Its fibrous leaves provided superior durability for maritime trade manuscripts, while its economic value in local crafts (thatching, sugar production) established pre-existing processing techniques42. This predates Indian cultural influence, as evidenced by 9th-century Javanese inscriptions on Borassus flabellifer folios. In India, Borassus flabellifer usage (25.54%) correlates with manuscripts from arid western regions (Rajasthan, Gujarat), where its drought tolerance offered reliable material sourcing—contrasting with Corypha umbraculifera’s dominance in the humid Ganges basin (74.46%, n = 82,241).
PLNet enabled this first biogeographical analysis of pan-Asian PLMs by overcoming traditional identification bottlenecks. It processed 142,679 manuscripts in <1 day (vs. >16 years via destructive sampling), and achieved species identification across diverse degradation states. Further research could explore the reasons behind these distribution patterns, including ecological factors, historical trade routes, and cultural practices in PLM production across different regions.
Model sensitivity to image resolution
The practical application of PLNet across diverse cultural institutions necessitates addressing variability in image acquisition standards. Our analysis reveals that spatial resolution significantly impacts classification confidence, as quantified by logit values in Fig. S6. The findings demonstrate that the model achieves its highest confidence for Corypha umbraculifera at 33.96 pixels/mm, while the peak for Borassus flabellifer occurs earlier, at 17.91 pixels/mm. Excessively low resolutions fail to capture the fine, discriminative morphological details, while excessively high resolutions may introduce noise or artifacts that confuse the classifier. To ensure the most reliable classification results, we suggest capturing images within a range of 10 to 40 pixels/mm. This range encompasses the peak performance for both species studied and is expected to provide a balance between capturing sufficient detail and avoiding potential high-resolution artifacts.
Discussion
In this study, we proposed PLNet, a deep learning framework for non-destructive and rapid identification of plant species of PLMs. By integrating Lion-optimized EfficientNetB2, transfer learning, and task-specific data augmentation strategies, PLNet achieved 99.07% accuracy on a diverse test set, outperforming state-of-the-art models like ResNet50, DenseNet169, and YOLO11x-cls, while operating at 0.31 s/folio with only 7.7 M parameters. A critical component of PLNet, the Lion optimizer, demonstrated superior performance by achieving lower loss on the validation set compared to the widely used Adam optimizer. Beyond its loss gains, Lion exhibited enhanced robustness and greater insensitivity to hyperparameter variations, making it more reliable and easier to deploy across diverse experimental setups.
The interpretability analysis using Guided GradCAM revealed that PLNet focuses on key features such as surface patterns, vein structures, and edge characteristics for species identification, providing transparency to the decision-making process of the model. To facilitate practical use, we developed PLNet-GUI, an easy-to-use software that enables heritage researchers to perform species identification without requiring technical expertise. Our analysis also highlighted the sensitivity of the model to image resolution, emphasizing the importance of digitization quality for optimal performance.
The real-world application of PLNet on 142,679 PLMs from 8 countries demonstrated its ability to analyze geographical distributions of palm species, offering valuable insights into historical trade routes and regional preferences in manuscript production. These findings bridge AI efficiency with ethical conservation, eliminating destructive sampling (5–10 cm2 loss/folio). PLNet establishes a new method in cultural heritage informatics, offering museums and archives a scalable, non-destructive tool to safeguard humanity’s fragile written legacy. In the future, PLNet can be extended to classify additional palm species and improve recognition in low-quality or degraded images. With further enrichment of training data from diverse regions, PLNet has the potential to become a universal tool for the preservation and study of palm-leaf manuscripts worldwide.
Data availability
Raw data for the palm-leaf manuscripts dataset was acquired from the British Library and the Potala Palace. The authors do not have permission to share the data.
Code availability
We provide the Python code and software for the method. It is available at https://github.com/yxcsu/PLNet.
References
Lawson, P. Palm leaf books and their conservation. Libr. Conserv. N. 16, 5–7 (1987).
Suryawanshi, D., Sinha, P. & Agrawal, O. Basic studies on the properties of palm leaf. Restaurator 15, 65–78 (1994).
Cai, M. Research on the cataloging items of the Beiyejing in the main collection databases of the UK. Libr. J. Shandong. 6, 75–83 (2020).
Xu, Z. & Liu, H. Palm leaves Buddhism Sutra culture of Xishuangbanna Dai and plant diversity. Biodiv. Sci. 3, 174–179 (1995).
Nishanthi, M., Kumara, H. & Konpola, K. Research conception of palm leaf manuscript conservation: Bibliometric Analysis of Scopus database. Int. J. Multidiscip. Stud. 10, 13–28 (2023).
Cabral, U. & Rathnabahu, R. N. Report on the best practices for conservation of Palm-Leaf Manuscripts in Sri Lankan Libraries. Sri Lankan J. Librariansh. Inf. Manag. 1, 1–4 (2021).
Zhang, M., Song, X., Wang, J. & Lyu, X. Preservation characteristics and restoration core technology of palm leaf manuscripts in Potala Palace. Arch. Sci. 22, 501–519 (2022).
Nichols, K. An alternative approach to loss compensation in palm leaf manuscripts. Pap. Conserv. 28, 105–109 (2004).
Crowley, A. S. Repair and conservation of palm-leaf manuscripts. Restaurator 1, 105–114 (1970).
Van Dyke, Y. Sacred leaves: the conservation and exhibition of early Buddhist manuscripts on palm leaves. Book Pap. Group Annu. 28, 83–97 (2009).
Zysk, K. G. Conjugal love in India: Ratiâsåastra and ratiramaòna: text, translation, and notes. Brill, Leiden (2002).
de Bernon, O., Sopheap, K. & An, L. K. Provisional inventory of Cambodian manuscripts, Part Two. EFEO, Bangkok (2018).
Aid, M. B. The documentary heritage of Myanmar: selected case studies. UNESCO, Paris (2018).
Tomlinson, P. B. The uniqueness of palms. Bot. J. Linn. Soc. 151, 5–14 (2006).
ICOMOS. Principles for the preservation of historic timber structures. ICOMOS, Paris (1999).
Alvisi, C. Un’anima per il diritto: andare più in alto. Mucchi Editore, Modena (2023).
Glassman, S. Systematic studies in the leaf anatomy of palm genus Syagrus. Am. J. Bot. 59, 775–788 (1972).
Sharma, D., Krist, G. & Velayudhan, N. M. Structural characterisation of 18th century Indian palm leaf manuscripts of India. Int. J. Conserv. Sci. 9, 257–264 (2018).
Hundius, H. & Wharton, D. The digital library of Lao manuscripts: Making the literary heritage of Laos available via the Internet. Microform Digit. Rev. 39, 142–144 (2010).
Mehta, R. V. K. & Challa, N. P. Facilitating enhanced user access through Palm-leaf manuscript digitization—Challenges and solutions. 2017 Second International Conference on Electrical, Computer and Communication Technologies (ICECCT). 1-4, (2017).
Wu, Q., Zhou, C. & Wang, C. Feature extraction and automatic recognition of plant leaf using artificial neural network. Adv. Artif. Intell. 3, 5–12 (2006).
Wu, S. G. et al. A leaf recognition algorithm for plant classification using probabilistic neural network. 2007 IEEE international symposium on signal processing and information technology. 11-16, (2007).
Prasad, S., Kudiri, K. M. & Tripathi, R. Relative sub-image based features for leaf recognition using support vector machine. Proceedings of the 2011 International Conference on Communication, Computing & Security. 343-346, (ICCCS, 2011).
Priya, C. A., Balasaravanan, T. & Thanamani, A. S. An efficient leaf recognition algorithm for plant classification using support vector machine. International conference on pattern recognition, informatics and medical engineering. 428-432, (PRIME, 2012).
Caglayan, A., Guclu, O. & Can, A. B. A plant recognition approach using shape and color features in leaf images. Int. Conf. Image Anal. Process. 8157, 161–170 (2013).
Munisami, T., Ramsurn, M., Kishnah, S. & Pudaruth, S. Plant leaf recognition using shape features and colour histogram with K-nearest neighbour classifiers. Procedia Comput. Sci. 58, 740–747 (2015).
Jeon, W. S. & Rhee, S. Y. Plant leaf recognition using a convolution neural network. Int. J. Fuzzy Log. Intell. Syst. 17, 26-34 (2017).
Shah, M. P., Singha, S. & Awate, S. P. Leaf classification using marginalized shape context and shape+ texture dual-path deep convolutional neural network. 2017 IEEE International Conference on Image Processing, 860-864, (ICIP, 2017).
Beikmohammadi, A., Faez, K. & Motallebi, A. SWP-LeafNET: a novel multistage approach for plant leaf identification based on deep CNN. Expert Syst. Appl. 202, 117470 (2022).
Tan, M. & Le, Q. Efficientnet: rethinking model scaling for convolutional neural networks. Int. Conf. Mach. Learn. 97, 6105–6114 (2019).
Buslaev, A. et al. Albumentations: fast and flexible image augmentations. Information 11, 125 (2020).
Müller, R., Kornblith, S. & Hinton, G. E. When does label smoothing help?. Adv. Neural Inf. Process. Syst. 32, 4694–4703 (2019).
Szegedy, C. et al. Rethinking the inception architecture for computer vision. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 2818-2826 (CVPR, 2016).
Chen, X. et al. Symbolic discovery of optimization algorithms. Adv. Neural Inf. Process. Syst. 36, 2931–2950 (2024).
Shannon, C. E. A mathematical theory of communication. Bell Syst. Tech. J. 27, 379–423 (1948).
Selvaraju, R. R. et al. Grad-cam: Visual explanations from deep networks via gradient-based localization. Proceedings of the IEEE international conference on computer vision (ICCV), 618-626, (2017).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. 2016 IEEE Conference on Computer Vision and Pattern Recognition, 770-778, (CVPR, 2016).
Huang, G., Liu, Z., Van Der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. 2017 IEEE Conference on Computer Vision and Pattern Recognition, 2261-2269, (CVPR, 2017).
Jocher, G., Qiu, J. & Chaurasia, A. YOLOv11 by Ultralytics. GitHub repository https://github.com/ultralytics/ultralytics (2023).
Snoek, J., Larochelle, H. & Adams, R. P. Practical Bayesian optimization of machine learning algorithms. Adv. Neural Inf. Process. Syst. 25, 2960–2968 (2012).
Eagleton, G. E. Persistent pioneers; Borassus L. and Corypha L. in Malesia. Biodiversitas 17, 716–732 (2016).
Gabriel, A. A. & Mardhiyyah, Y. S. Utilization of Siwalan (Borassus Flabellifer L.) plantation waste for kraft paper production. J. Teknol. Ind. Pertan. Indones. 11, 1–5 (2019).
Acknowledgements
This work is financially supported by the National Key Research and Development Program of China (grant No. 2023YFF0906702) and the National Natural Science Foundation of China (grant No. 22273120). We are grateful for resources from the High-Performance Computing Center of Central South University.
Author information
Authors and Affiliations
Contributions
Lu and Zhang conceived the idea and supervised the project. Yang was the primary executor of the experiments, conducted the majority of the analyses, and was a major contributor to writing the manuscript. Zhou, Gao and Yang obtained the rights to collect the PLM image data and annotate the species of the PLMs. Chen, Tan, and Wang contributed to the experimental design and assisted with data processing. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, X., Chen, S., Tan, L. et al. A lion-optimized efficientnet framework for non-destructive and rapid plant species identification in palm-leaf manuscripts. npj Herit. Sci. 13, 618 (2025). https://doi.org/10.1038/s40494-025-02149-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s40494-025-02149-0