Abstract
Microalgae, including Cyanobacteria, have been shown to be pioneers among microbial colonizers at many archaeological sites, particularly on outdoor stone monuments. Being predominant pollutants, microalgae cause significant deterioration of historic buildings and materials, together with other colonizers. This perspective comprehensively explores the resources of algicidal bacteria, revisits their algicidal mechanisms as an eco-friendly mitigation against harmful microalgae, and proposes a paradigm shift in heritage conservation to address microalgal impacts.
Similar content being viewed by others
Introduction
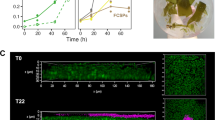
Microalgae, in the strictest definition, are eukaryotic, unicellular microorganisms that are photosynthetic and typically have an aquatic lifestyle1. Although cyanobacteria are prokaryotic and therefore not true algae, they are usually included because they share a similar physiology and ecology with eukaryotic microalgae. Moreover, photosynthesis originated around 3.5 billion years ago in cyanobacteria, with eukaryotes emerging much later, following 1–1.5 billion years of prokaryotic evolution2. This photoautotrophic ability enables microalgae to be widely distributed in aquatic environments with low organic nutrient levels, such as epilithic and endolithic niches3,4. For example, microalgae have been widely identified as pioneer colonizers on outdoor stone monuments and historical buildings5,6,7, particularly when water is available8,9 (Fig. 1a–e). Subsequently, their growth provides sufficient organic nutrients for heterotrophic colonizers, which are usually harmful microorganisms (e.g., archaea, bacteria, and fungi) to stone heritage10,11,12. However, case studies on microalgal colonization of heritage sites have been reported frequently, especially in stone heritage (Fig. 1f). Therefore, developing an efficient mitigation strategy to control microalgal growth is essential for the conservation of stone heritage.
a Microscopy of microalgal colonization of stone materials. b Microalgal blooming at the Jinsha earthen site, Chengdu, China. c Microalgal colonization at the gate tower of Ming Xiaoling Mausoleum, Nanjing, China. d Microalgal growth on the brick wall of ancient tombs, Luoyang, China. e Cyanobacterial colonization of the Longmen Grottoes, Luoyang, China. f Statistics of microalgae-related cases at the archeological sites. Based on Scopus, bibliographic databases were curated using the following search algorithms: TITLE-ABS-KEY (cyanobacteria) OR TITLE-ABS-KEY (alga) AND TITLE-ABS-KEY (heritage) OR TITLE-ABS-KEY (monument) OR TITLE-ABS-KEY (Archaeological). Images courtesy of X.L., J.Q., F.W., and R.Y.
Over the past two decades, numerous bacterial species capable of lysing algal cells or suppressing algal growth—particularly those associated with harmful algal blooms—have been isolated and identified13,14. These bacteria are collectively referred to as algicidal bacteria and can inhibit or kill algae directly or indirectly15,16. Algicidal bacteria are highly diverse in their ecological roles, including shaping phytoplankton species composition in pelagic environments17. Their algicidal modes have been primarily categorized into two types17,18: direct attack, which requires bacteria to actively seek out and attach to microalgal cells19; and indirect contact, which depends on the secretion of algicidal substances20. The variety of algicidal mechanisms and the diversity of algicidal metabolites open up the possibility of developing eco-friendly mitigations for the biocontrol of harmful microalgae that colonize stone heritage.
Despite the achievements of applying algicidal bacteria to control harmful algal blooms, several issues and challenges remain, including the development of algicidal-specific predation strategies and molecular mechanisms, the mechanisms of action of algicidal substances, and the development of effective algicides17,18. Moreover, there are few studies on the application of algicidal bacteria or their metabolites in the conservation of stone heritage. These issues have largely impeded the application of algicidal bacteria in controlling harmful microalgae on stone heritage.
Here, we provide a comprehensive overview of the diversity and distribution of algicidal bacteria, their mechanisms of action, and their potential applications as eco-friendly biocontrol agents, aiming to offer a reference framework for future research on sustainable protection of stone heritage from harmful microalgae. In this review, cyanobacteria are treated operationally as microalgae for ecological and heritage-colonization purposes, while remaining taxonomically distinct.
Diversity and distribution of algicidal bacteria
Algicidal bacteria are mostly Gram-negative bacteria that belong to the three phyla Proteobacteria, Bacilli, and Bacteroidetes18. Also, some Gram-positive bacteria from the Actinobacteria and Firmicutes phyla have been reported to be algicidal. Currently, algicidal bacteria are found to belong to the genera Bacillus, Vibrio, Alteromonas, and Pseudoalteromonas (Fig. 2).
The tree topology was inferred using maximum-likelihood methodology and constructed from 16S rRNA sequences of the representative species. Bootstrap values (expressed as percentages of 1000 replications) are shown at the branch nodes. The species that kill microalgae by direct attack are highlighted in blue, while the others kill them indirectly.
Vibrio
Vibrio alginolyticus is one of the most common algicidal bacteria in marine environments21. It has been found to kill or inhibit a wide range of marine microalgae, such as Chaetoceros22, Akashiwo23, Alexandrium24, and Heterosigma25. Polysaccharides constitute a significant portion of both dissolved and particulate organic matter in the ocean, and the degradation of polysaccharides plays a central role in the marine carbon cycle26. Many studies suggest that marine Vibrio species can utilize extracellular enzymes to degrade algal cell polysaccharides, such as glucosidases and chitinases26,27. Moreover, algicidal Vibrio species, such as Vibrio sp. W13, Vibrio sp. NJ-04 and V. splendidus OU02 possess genes encoding alginate lyase, an enzyme that can degrade alginate into unsaturated oligosaccharides by cleaving β-glycosidic bonds, thereby breaking down the algal cell wall27,28,29.
In addition to secreting extracellular enzymes to lyse algal cell walls, the Vibrio species can use other algicidal mechanisms. Wang et al.30 co-cultured two algicidal Vibrio species, V. brasiliensis and V. tubiashii, and found that their metabolites increased reactive oxygen species (ROS) levels of Akashiwo sanguinea. This ultimately led to cell membrane damage and further inhibited the antioxidant system, resulting in a sharp decrease in catalase (CAT) activity and ultimately the death of algal cells.
Alteromonas and Pseudoalteromonas
In Proteobacteria, Alteromonas and Pseudoalteromonas are primarily algicidal bacterial genera distributed in marine environments. Pseudoalteromonas is one of the most widely distributed and extensively studied algicidal genera31. Initially, many species of Pseudoalteromonas were members of the genus Alteromonas, but in 1995, a taxonomic reorganization of the genus Alteromonas was carried out based on phylogenetic analysis32. They are often the algicidal bacteria of diatoms and dinoflagellates.
The algicidal activity of species from the genera Alteromonas and Pseudoalteromonas primarily occurs through indirect action involving extracellular exudates. This algicidal characteristic may help bacterial cells compete for nutrients, regulate algal blooms, and defend against surface predators31,33,34. Sun et al.35 isolated a novel alginate lyase from Pseudoalteromonas sp. Alg6B, which showed a hydrolysis rate of up to 97% for Laminaria japonica within 24 h. Wang et al.36 reported that Pseudoalteromonas sp. LD-B6 showed intense algicidal activity against Noctiluca scintillans, achieving an algicidal rate of 90.5% within 12 h. Also, Alteromonas FDHY-03 and A. abrolhosensis JY-JZ1 have been shown to produce an algicidal substance that disrupts algal cell structures by digesting cell wall polysaccharides, leading to intracellular leakage37,38.
Bacillus
Bacillus spp. are Gram-positive, rod-shaped bacteria widely distributed in soil and water environments. Importantly, they are simple nutritional requirements and can form spores under unfavorable environmental conditions. Bacillus can secrete various antimicrobial substances and extracellular hydrolases, which have been reported to inhibit the growth of microalgae, especially cyanobacteria.
Most Bacillus species inhibit algal growth by suppressing photosynthesis and stimulating oxidative stress in host cells. For example, Bacillus altitudinis G3 exhibits high algicidal activity against Microcystis aeruginosa by secreting algicidal substances that can partially block the photosynthetic electron transfer from primary (QA) and secondary (QB) quinone electron acceptors39. Furthermore, the interruption of photosynthesis, especially the inhibition of electron transfer downstream of QA−, generates excessive ROS, leading to a significant increase in malondialdehyde (MDA) content, which is responsible for severe lipid oxidation and, ultimately, oxidative stress in M. aeruginosa. Additionally, Bacillus pumilus SU8S0818 has also been reported to kill cyanobacteria through algicidal substances that produce excessive ROS, leading to their oxidative stress death40.
Apart from the above-mentioned algicidal bacteria, some species of the genera Pseudomonas, Bdellovibrio, Spirillum, Flavobacterium, and Lysozymimonas, which belong to the phyla Bacteroidetes and Proteobacteria, can also act as microalgal predators through direct and/or indirect interactions to kill algae (Table 1). Unfortunately, the algicidal potential of these bacteria is not well explored, indicating that further work is required to unravel their novel algicidal processes that might be applied to biocontrol.
Mechanisms of algicidal action
Generally, algicidal modes of action in bacteria are categorized into indirect and direct mechanisms. Indirect action mainly relies on the secretion of extracellular active substances to lyse algae or competition with algae for limited nutrients, as exemplified by Vibrio spp. and Bacillus spp. However, direct action requires physical contact, in which algicidal bacteria adhere to algal cells and may even penetrate the intracellular space to exert their algicidal effects, a strategy predominantly employed by Myxobacteria spp. and Bdellovibrio spp.
Indirect action
Indirect algicidal contact is the primary mode of algicidal action reported for most bacteria. Algicidal bacteria release specific or non-specific extracellular substances that act as algicides, indirectly killing algae or inhibiting their growth and reproduction. These substances include proteins, peptides, amino acids, fatty acids and their derivatives, alkaloids, and antibiotics. Common bacteria that exhibit this activity include Vibrio41, Pseudomonas42, Flavobacterium43, and Alteromonas44.
Algicides derived from different natural product categories have been shown to primarily exhibit three main algicidal actions: 1) disruption of the cell membrane system, 2) oxidative stress, and 3) inhibition of photosynthetic respiration17.
Disruption of the cell membrane system
For indirect algicidal contact, algicidal bacteria release substances that disrupt the cell wall, membrane, and thylakoids of algal cells. This leads to loss of cell integrity and inhibits algal growth, resulting in lysis and cell death.
In Vibrio, some extracellular hydrolases could dissolve algal cells by digesting the cell wall and cell membrane. Due to its amphipathic properties, the algal plasma membrane can interact with cationic peptides via electrostatic interactions, thereby facilitating membrane permeation. The chloroplast membrane of algal cells can also be similarly disrupted by cationic peptides (Fig. 3a)45. Park et al.45 determined the cationic and hydrophobic content by altering amino acid arrangements, ultimately synthesizing four hybrid peptides with algicidal activity. Among them, glycine, proline-derived cyclic dipeptides, and diketopiperazines are significant classes of antimicrobial compounds that inhibit the growth of Microcystis aeruginosa by compromising cell wall integrity and intracellular structures46.
a Degradation of the cell membrane system by extracellular enzymes. b Inhibition of photosynthesis through disruption of the chloroplast membrane. c Interference with membrane protein synthesis through the introduction of exogenous amino acids.
Exogenous polyunsaturated fatty acids can be incorporated into bacterial phospholipids, altering the plasma membrane’s ultrastructure46. This induces changes in membrane permeability and the dissociation of phycobilin from the thylakoid membrane, leading to potassium ion leakage through the cell wall. The leakage of ions from the plasma membrane through enzymatic hydrolysis or pore formation is considered irreparable damage16.
Wu et al.47. found that indole alkaloids can disrupt the cell membrane and chloroplast membrane, deteriorating the internal materials of the thylakoid membrane, interrupting the electron transfer in photosystem II, reducing the effective quantum yield, and ultimately leading to the failure of photosynthesis, thereby inhibiting the growth of cyanobacteria (Fig. 3b). Kodani et al.42 isolated a tricyclic β-carboline, an indole alkaloid, from an algicidal freshwater Pseudomonas species, inhibiting various cyanobacteria growth. Moreover, 2-n-pentyl-4-quinoline, a quinoline alkaloid produced by marine Pseudoalteromonas and marine Pseudomonas species, can inhibit the growth of cyanobacteria and diatoms32.
L-valine has been demonstrated to affect the morphology and integrity of Microcystis, by which exogenous L-valine can substitute for other amino acids and be incorporated into proteins, thereby producing defective proteins. This ultimately leads to the collapse or perforation of Microcystis cells (Fig. 3c)48. Additionally, the bacteriolysin of Bacillus amyloliquefaciens FZB42 can disrupt the plasma membrane of Microcystis aeruginosa49. An et al.50 found that (2-isobutoxyphenyl)amine from Brevibacterium sp. BS01 can alter the cell structure of Alexandrium tamarense, leading to cytoplasmic degradation and loss of organelle integrity.
Oxidative stress
The accumulation of ROS in algae is another common mode of algicidal action, which leads to oxidative stress in algal cells (Fig. 4). ROS include superoxide anion radicals (O2−·), hydrogen peroxide (H2O2), and hydroxyl radicals (OH·), which are well known to be associated with cell aging and death. In most eukaryotes, the primary sources of ROS are the mitochondrial electron transport chain and peroxisomes. However, plant cells contain chloroplasts with intense electron flow, leading to a high rate of ROS production. Due to the strong oxidizing nature of ROS, many essential biomolecules, including DNA, proteins, and lipids, cannot withstand oxidative damage, resulting in loss of cellular function14,51.
a Some substances can enter the interior of algal cells through transport proteins, causing the chloroplasts to produce large amounts of ROS. b High concentrations of metals generate ROS through the Fenton and Haber-Weiss reactions. c With the onset of oxidative stress in algal cells, the antioxidant system is crucial, including antioxidant enzymes and antioxidants. The main antioxidant enzymes include superoxide dismutase (SOD), catalase (CAT), and peroxidase (POD). Due to the dysfunction of the antioxidant system in dying cells, their activity usually increases in the early stages to remove accumulated ROS, then sharply declines in the later stages of the algicidal process. d ROS can lead to lipid peroxidation, which could inhibit photosynthesis by damaging the thylakoid membrane and/or oxidizing photosynthetic pigments (d1), damage the cell membrane and cause leakage of cellular contents (d2), and produce excessive malondialdehyde (MDA) that damages the membrane (d3). e ROS could damage essential biomolecules, such as DNA, RNA, and functional proteins. Red arrows indicate the damage effects to the corresponding cellular structures, whereas black arrows represent the inductive or promotive effects to the corresponding processes.
Chen et al.52 isolated pyocyanin from Pseudomonas aeruginosa as an algicidal compound, which exhibits high toxicity to eukaryotic microalgae and other aquatic organisms. In our previous studies, we found that Enterobacter cloacae NP23 and Gibberella moniliformis AM1 exhibited a high lysis rate for four algal species (i.e., Chlorella vulgari, Scenedesmus sp., Microcystis wesenbergii, and Chlorella pyrenoidosa) by inhibiting their host antioxidase activities13,14,53. These two algicidal bacteria would provide a new member for the biocontrol of microalgal populations in eco-technology54. An algicidal compound secreted by Bacillus sp. strain B1, N-acetylsalicylic acid, can induce excessive ROS production in Heterosigma akashiwo, affecting the antioxidant system, photosynthetic pigment contents, and protein levels, leading to the death of H. akashiwo51. L-lysine can induce oxidative stress in Microcystis aeruginosa, reducing chlorophyll content and disrupting the cell membrane in cyanobacteria55,56. Bacillus licheniformis strain sp34 can induce oxidative stress, lipid peroxidation, morphological damage in algal cells, DNA damage, and dysfunction of DNA repair mechanisms, weakening the photosynthetic system and inhibiting Microcystis aeruginosa54.
Inhibition of photosynthetic respiration
Photosynthesis depends on the absorption of sunlight by chlorophyll molecules (e.g., chlorophyll a) in photosystem I (PSI) and photosystem II (PSII). Microalgae can synthesize specific pigments (e.g., carotenoids) that help prevent photo-oxidative damage caused by highly reactive by-products of photosynthesis. However, algicidal bacteria could disrupt photosynthetic processes and activities in algal cells, including electron transfer in photosynthetic pigments, photochemical efficiency, and synthesis and maintenance (Fig. 5), thereby interfering with energy transformation and eventually leading to cell death55.
Tryptophan could bind to the enzyme FtsH, which degrades the protein D1, thereby inhibiting its degradation. Also, some molecules, such as 2-hydroxychalcone, arginine A, and β-diketone, could inhibit microalgal photosynthesis by impeding electron transfer.
Recent studies demonstrated that a novel algicidal bacterium, Pseudomonas fragi YB2, could significantly decrease chlorophyll content by 4.74 times in 120 h and the activity of succinate dehydrogenase by 103 times in 36 h56. Su et al.57 found that Acinetobacter sp. J25 could reduce chlorophyll a in algae by up to 90% after a 3-day treatment. Also, arginine A, which is a potent anti-cyanobacterial compound produced by Sphingomonas sp. M-17 could act as a photosynthetic inhibitor, interrupting the electron transport chain before PSII58. Additionally, tryptophan hinders photosynthesis by inhibiting the degradation of the D1 protein by the FtsH protease during PSII quality control in chloroplasts and kills algae by significantly reducing photosynthetic efficiency and carbon assimilation (Fig. 5), inhibiting photochemical electron transfer, and increasing the number of closed reaction centers and energy loss59.
Based on the type of electron acceptors on the reduced (acceptor) side of the photosystems, the reaction centers (RC) of the PSI and PSII core complexes are divided into FeS type and quinone type. In photosynthetic reaction centers of PSII, the photoinduced charge separation is terminated by an electron transfer between the primary (QA) and secondary (QB) quinones. Therefore, Zhang et al.60 found that 2-hydroxychalcone can replace HCO3– as a ligand for non-heme iron, thereby disrupting the electron transfer from QA to QB (Fig. 5). Based on this, Yilimulati et al.61 further inferred that β-diketones can bind to the Fe-S cluster of Fd, inhibiting photosynthesis.
However, these three modes of action are not mutually exclusive. ROS attacks the polyunsaturated fatty acids in biological membranes, leading to lipid peroxidation and the production of lipid peroxides in algal cells. At the same time, the accumulation of ROS disrupts pigment synthesis and membrane integrity, inhibiting or ultimately killing algal cells62. Observations showed that the supernatant of Brevibacterium sp. BS01 inhibits psbA gene expression, blocks the electron transport chain, and significantly increases ROS levels and the excessive activity of the antioxidant system. The accumulation of ROS disrupts pigment synthesis and membrane integrity, inhibiting or killing algal cells63. On the other hand, impaired photosynthesis also promotes ROS production17. Dysfunction of the PSII system transfers excitation energy to singlet oxygen, which then forms ROS. Decreased chlorophyll a and carotenoid levels reduce the capacity to inhibit ROS generation64. The pyoluteorin secreted by Hahella sp. KA22 completely inhibits photochemical reactions and the carbon-fixation system in PSII, thereby suppressing photosynthesis, promoting excessive ROS production, inducing severe oxidative damage to cells and organelles, and ultimately killing algal cells64,65.
Direct action
Some algicidal bacteria can directly contact algal cells and digest algal cell walls by releasing hydrolases17. Enzyme activity depends on site specificity, protein concentration, and bacterial cell density. They mainly correspond to two types of enzyme activities: glycoside hydrolases (carbohydrate hydrolases) and proteases, based on their functions16.
Carbohydrate enzymes target the degradation of polysaccharides that constitute the algal cell wall and are involved in cell adhesion. It has been reported that extracellular algicidal glycoside hydrolases produced by bacteria include α-glucanases, β-glucanases, or cellulases16. β-glucanases primarily act on the digestion of algal cell walls, while α-glucanases mainly act on the degradation of intracellular starch38,66. The activities of amylase, cellulase, and xylanase are also associated with the degradation of the algal cell wall67.
Proteases may have dual functions of algicidal activity and promoting the utilization of host polymers through extracellular degradation. For example, a marine Pseudoalteromonas species that is algicidal to diatoms can produce two proteases (probably belonging to the peptidase families S8 and M11, respectively) responsible for cell wall digestion during the cell life cycle68. In addition to their lytic activity, bacterial proteases can also reduce microalgae activity by affecting cell motility69,70.
Importantly, algicidal bacteria that employ direct action are usually predatory, and their modes of action can be divided into endobiotic and epibiotic predation19. Endobiotic predators can enter the periplasm of the prey cell, utilizing its biomass to reproduce and lyse the prey cell from within71. In contrast, epibiotic predators induce lysis of the prey from the outside and feed on the released biomass72.
Epibiotic predation
Epibiotic predation is a mode by which bacteria attach to the surface of host cells and consume the cytoplasm from the outside19. In the 1980s, Guerrero et al. described the attack by Vampirococcus on various Chromatium species as a typical epibiotic predatory strategy73. In direct contact with algicidal activity, epibiotic predation is more common. For example, some studies found that Tenacibaculum sp. GD3 could aggregate and bind to the cell membrane of Karenia mikimotoi, then kill the host through lipid peroxidation and membrane protein lysis (Fig. 6a)15. As the culture time increased, CAT activity and MDA levels decreased, indicating the collapse of the algal defense system (Fig. 4). Furthermore, all chlorophyll fluorescence parameters declined significantly, indicating damage to the photosynthetic system. The expression of genes encoding glutathione S-transferase (GST), pheophorbide a oxygenase (PAO), pabA1, cellulase, and cyclin B was significantly inhibited, leading to irreversible rupture of cells and chloroplasts after 8 h. Moreover, Shi et al.74 showed that Bacillus cereus DC22 can secrete mucus that binds algicidal bacterial cells to cyanobacterial cells and lyses the latter. Additionally, Saprospira sp. PdY3 lyses cyanobacteria through direct contact: it exhibits distinct group behavior on solid media, forming orderly bundle-like group structures; however, when co-cultured with Anabaena in a liquid medium, PdY3 also shows group behavior, forming a three-dimensional mesh structure and lysing Anabaena75.
a Epibiotic predation through lipid peroxidation and membrane protein lysis. b Endobiotic predation through cytolysis and/or periplasmic invasion. c Endobiotic predation through cytolysis in Cyanoraptor togatus LGM-1.
Endobiotic predation
Endobiotic predation is a mode by which algicidal bacteria penetrate host cells and consume them from the inside73. In 1984, Caiola76 isolated a bacterium that could enter Microcystis aeruginosa, causing the host cell to lyse and die. In 1985, Moulder referred to the process in which predatory cells directly enter the host cell cytoplasm as cytolysis77, while the process in which predatory cells invade and grow in specific compartments of Gram-negative cells was called periplasmic invasion (Fig. 6b)73.
One of the most famous predatory bacteria is the endobiotic Bdellovibrio bacteriovorus19,71. During the attack phase, Bdellovibrio signals the bacterial population to release deacetylase, which alters the host cell wall. Once prey is located, it releases enzymes such as glycanases and peptidases to form a “reinforced circular pore” through which the predator enters the host cell. Inside, B. bacteriovorus degrades periplasmic components and consumes them, preventing the host cell from self-replicating. After an extended phase, Bdellovibrio separates into single cells and exits the prey by producing lysozyme. This specifically destroys the previously deacetylated host cell wall, thereby preventing the predator’s self-destruction. The progeny released by Bdellovibrio then begins the cycle anew.
In addition, Julie et al.78 isolated and identified a strain of endobiotic predatory algicidal bacterium, Cyanoraptor togatus LGM-1. Under the microscope, LGM-1 docks with the sheath of microalgae, enters the interior of the microalgae, and undergoes morphological changes. After multiple divisions, a fibrous cocoon forms around the bacteria, while the peptidoglycan transverse walls of the microalgae gradually degrade. In the mid-to-late stage of predation, extracellular propagules of LGM-1 appear outside the fibrous cocoon and, finally, the microalgae lyse (Fig. 6c).
Application paradigm shift to heritage conservation and future perspectives
Currently, the application of algicidal bacteria as a biocontrol agent has been successfully demonstrated to mitigate harmful algal bloom18. This strategy is an eco-friendly treatment for environmental biological contaminants, urgently required for the sustainable development of our society. However, a comprehensive understanding of algicidal mechanisms will help develop additional resources for algicidal bacteria, thereby advancing the field of biocontrol agents of microorganisms15,54,55,58,65.
Moving forward, this mitigation strategy could serve as a successful paradigm for heritage conservation when the risk of on-site application is fully evaluated. In the future, multidisciplinary incorporation from microbiologists, ecologists, and heritage conservators would achieve this great goal through the following endeavors:
First, the prerequisite for effective control of harmful microalgae is that microbiologists provide accurate identification of the species present at the archaeological site. With this information, we can identify the corresponding algicidal bacteria that kill or inhibit the harmful microalgae. Here, we propose to curate a database or bio-archive of microalgae and their algicidal bacteria to support the development of an efficient biocontrol policy for sustainable heritage conservation79. Based on the algicidal mechanisms underlying the interaction between microalgae and algicidal bacteria, we can produce algicidal bacteria or their metabolites as algicides using current biotechnology approaches. However, more work is required to ensure that these insights from algal bloom control can be non-trivially translated to stone heritage systems. In contrast to aquatic bloom systems, subaerial biofilms on stone heritage markedly differ in hydration regimes, nutrient fluxes, surface chemistry, and ecological stability. Given these differences, we have to explicitly identify which algicidal mechanisms are likely transferable and which are not. For example, severe microenvironments (e.g., oligotrophy, limited water availability, and large temperature fluctuations) on stone surfaces could threaten the persistence and survival of algicidal bacteria, thereby reducing their algicidal effectiveness. To address these issues, microbiologists can focus on biosynthetic engineering of algicidal metabolites, in situ isolation of algicidal bacteria from stone heritage systems, or screening for algicidal bacteria that adapt to microenvironments. Nevertheless, it is more controllable in practice on stone surfaces than in the aquatic environment, as interactions between bacteria and phytoplankton in aqueous ecosystems are more complex and dynamic80, and water also reduces the density of algicidal bacteria or their metabolites17. Thus, whether direct or indirect mechanisms are more efficient depends on whether algicidal bacteria can survive on stone surfaces.
Next, our ecologists could track the timelines and patterns of microalgal blooms at the archaeological site. This provides us with accurate information on when and what algicidal bacteria should be used to address the microalgal bloom on site. Furthermore, a comprehensive assessment of the ecological effects of algicidal bacteria or their metabolites should be conducted to confirm that there is no environmental risk. For example, whether potential interactions with existing stone-associated microbial consortia (e.g., fungi and lichens) would exacerbate the biodeterioration of stone heritage must be considered. These potential side effects will constitute the pertinent regulatory and ethical barriers to on-site release of microorganisms, as one cannot expect to freely apply living organisms such as these to surfaces without some form of regulation17. As our treatment is based on microbial control, this ecological risk could be lower than that of chemical algicides, which are strictly forbidden for heritage conservation due to their proven ecological risks.
Lastly, but most importantly, heritage conservators are responsible for a comprehensive on-site assessment of the long-term effects on heritage materials themselves, for at least years. For one, microalgal biofilms can, in some cases, protect covered heritage materials, especially when natural weathering processes outweigh biodeterioration caused by microalgae81. In this case, killing or inhibiting microalgae could not be a positive strategy. For another, we should ensure that either algicidal bacteria used for microalgal biocontrol or algicides produced by algicidal bacteria do not damage heritage materials. Thus, the risks of unintended biodeterioration caused by metabolites (e.g., organic acids, enzymes, and extracellular polymeric substances) produced by algicidal bacteria must be evaluated. If these assessments are successful, we can deliver algicidal bacteria as an eco-friendly approach to mitigate microalgal blooms and support sustainable heritage conservation. Based on these endeavors, we will develop a comprehensive mitigation strategy to address the harmful microalgal bloom at the archaeological site.
Data availability
All data supporting the conclusions of this study are available from the author, Xiaobo Liu, upon request.
References
Thoré, E. S. J., Muylaert, K., Bertram, M. G. & Brodin, T. Microalgae. Curr. Biol. 33, R91–R95 (2023).
Olson, J. M. & Pierson, B. K. Photosynthesis 3.5 thousand million years ago. Photosynth. Res. 9, 251–259 (1986).
Roush, D., Couradeau, E., Guida, B., Neuer, S. & Garcia-Pichel, F. A new niche for anoxygenic phototrophs as endoliths. Appl. Environ. Microbiol. 84, e02055–02017 (2018).
Walker, J. J. & Pace, N. R. Endolithic microbial ecosystems. Annu. Rev. Microbiol. 61, 331–347 (2007).
Gaylarde, C. C. Influence of environment on microbial colonization of historic stone buildings with emphasis on cyanobacteria. Heritage 3, 1469–1482 (2020).
Zhang, Y. et al. Dominance by cyanobacteria in the newly formed biofilms on stone monuments under a protective shade at the Beishiku Temple in China. Environ. Res. 251, 118576 (2024).
Wu, F. et al. Community assembly, potential functions and interactions between fungi and microalgae associated with biodeterioration of sandstone at the Beishiku Temple in Northwest China. Sci. Total Environ. 835, 155372 (2022).
Wang, L. et al. Water determines geomicrobiological impact on stone heritage. Nat. Geosci. 18, 108–111 (2025).
Liu, X., Meng, H., Wang, Y., Katayama, Y. & Gu, J.-D. Water is a critical factor in evaluating and assessing microbial colonization and destruction of Angkor sandstone monuments. Int. Biodeterior. Biodegrad. 133, 9–16 (2018).
Wu, F. et al. Community structures of bacteria and archaea associated with the biodeterioration of sandstone sculptures at the Beishiku Temple. Int. Biodeterior. Biodegrad. 164, 105290 (2021).
Liu, X., Koestler, R. J., Warscheid, T., Katayama, Y. & Gu, J.-D. Microbial deterioration and sustainable conservation of stone monuments and buildings. Nat. Sustain. 3, 991–1004 (2020).
Gadd, G. M., Fomina, M. & Pinzari, F. Fungal biodeterioration and preservation of cultural heritage, artwork, and historical artifacts: extremophily and adaptation. Microbiol. Mol. Biol. Rev. 88, e00200–e00222 (2024).
Liao, C., Liu, X., Liu, R. & Shan, L. Two novel algicidal isolates kill Chlorella pyrenoidosa by inhibiting their host antioxidase activities. Appl. Biochem. Biotechnol. 177, 567–576 (2015).
Liao, C., Liu, X. & Shan, L. Optimization of liquid media and biosafety assessment for algae-lysing bacterium NP23. Can. J. Microbiol. 60, 593–597 (2014).
Shi, J. et al. Efficient inactivation of harmful algae K. mikimotoi by a novel algicidal bacterium via a rare direct contact pathway: Performances and mechanisms. Sci. Total Environ. 892, 164401 (2023).
Demuez, M., González-Fernández, C. & Ballesteros, M. Algicidal microorganisms and secreted algicides: New tools to induce microalgal cell disruption. Biotechnol. Adv. 33, 1615–1625 (2015).
Meyer, N., Bigalke, A., Kaulfuss, A. & Pohnert, G. Strategies and ecological roles of algicidal bacteria. FEMS Microbiol. Rev. 41, 880–899 (2017).
Morón-López, J., Serwecińska, L., Balcerzak, Ł, Glińska, S. & Mankiewicz-Boczek, J. Algicidal bacteria against cyanobacteria: Practical knowledge from laboratory to application. Crit. Rev. Environ. Sci. Technol. 54, 239–266 (2024).
Bauer, A. & Forchhammer, K. Bacterial predation on cyanobacteria. Microb. Physiol. 31, 99–108 (2021).
Van Le, V. et al. Algicide capacity of Paucibacter aquatile DH15 on Microcystis aeruginosa by attachment and non-attachment effects. Environ. Pollut. 302, 119079 (2022).
Smith, D. S. et al. Polyunsaturated fatty acids cause physiological and behavioral changes in Vibrio alginolyticus and Vibrio fischeri. MicrobiologyOpen 10, e1237 (2021).
Imai, I. & Kimura, S. Resistance of the fish-killing dinoflagellate Cochlodinium polykrikoides against algicidal bacteria isolated from the coastal sea of Japan. Harmful Algae 7, 360–367 (2007).
Hu, J. M. et al. Differentiation of microbial communities in coastal seawater before and during an Akashiwo sanguinea (Dinophyceae) bloom in the urban area of Antofagasta city (northern Chile). Harmful Algae 142, 102782 (2025).
Li, D. et al. A novel algicide: evidence of the effect of a fatty acid compound from the marine bacterium, Vibrio sp. BS02 on the harmful dinoflagellate, Alexandrium tamarense. PLoS One 9, e91201 (2014).
Cho, J. Y. Algicidal activity of marine Alteromonas sp. KNS-16 and isolation of active compounds. Biosci. Biotechnol. Biochem. 76, 1452–1458 (2012).
Matthias, W. et al. Bacterial community dynamics during polysaccharide degradation at contrasting sites in the Southern and Atlantic Oceans. Environ. Microbiol. 17, 3822–3831 (2015).
Tang, L. et al. Effects of module truncation of a new alginate lyase VxAly7C from marine Vibrio xiamenensis QY104 on biochemical characteristics and product distribution. Int. J. Mol. Sci. 23, 4795 (2022).
Zhu, B., Ni, F., Ning, L., Sun, Y. & Yao, Z. Cloning and characterization of a new pH-stable alginate lyase with high salt tolerance from marine Vibrio sp. NJ-04. Int. J. Biol. Macromolecules 115, 1063–1070 (2018).
Yu, Z., Zhu, B., Wang, W., Tan, H. & Yin, H. Characterization of a new oligoalginate lyase from marine bacterium Vibrio sp. Int. J. Biol. Macromolecules 112, 937–942 (2018).
Wang, Y. et al. Continuous production of algicidal compounds against Akashiwo sanguinea via a Vibrio sp. co-culture. Bioresour. Technol. 295, 122246 (2020).
Wang, M., Chen, S., Zhou, W., Yuan, W. & Wang, D. Algal cell lysis by bacteria: A review and comparison to conventional methods. Algal Res. 46, 101794 (2020).
Bowman, J. P. Bioactive compound synthetic capacity and ecological significance of marine bacterial genus Pseudoalteromonas. Mar. Drugs 5, 220–241 (2007).
Cai, G., Yu, X., Wang, H., Zheng, T. & Azam, F. Nutrient-dependent interactions between a marine copiotroph Alteromonas and a diatom Thalassiosira pseudonana. mBio 14, e0094023 (2023).
Xu, F., Cha, Q.-Q., Zhang, Y.-Z. & Chen, X.-L. Degradation and utilization of alginate by marine Pseudoalteromonas: a review. Appl. Environ. Microbiol. 87, e00368–00321 (2021).
Sun, C., Zhou, J., Duan, G. & Yu, X. Hydrolyzing Laminaria japonica with a combination of microbial alginate lyase and cellulase. Bioresour. Technol. 311, 123548 (2020).
Wang, J. et al. Isolation and characterization of a high-efficiency algicidal bacterium Pseudoalteromonas sp. LD-B6 against the harmful dinoflagellate Noctiluca scintillans. Front. Microbiol. 13, 1091561 (2022).
Yang, J. et al. Algicidal bacteria in phycosphere regulate free-living Symbiodinium fate via triggering oxidative stress and photosynthetic system damage. Ecotoxicol. Environ. Saf. 263, 115369 (2023).
Shi, X. et al. Isolation of an algicidal bacterium and its effects against the harmful-algal- bloom dinoflagellate Prorocentrum donghaiense (Dinophyceae). Harmful Algae 80, 72–79 (2018).
Hou, X. et al. An insight into algicidal characteristics of Bacillus altitudinis G3 from dysfunctional photosystem and overproduction of reactive oxygen species. Chemosphere 310, 136767 (2022).
Nájera, A. F. et al. Algicidal bacteria induce a molecular stress response in Microcystis aeruginosa and Aphanizomenon gracile leading to physiological alterations and cell death. Int. Biodeterior. Biodegrad. 189, 105763 (2024).
Matsumura, Y., Sato, K., Al-saari, N., Nakagawa, S. & Sawabe, T. Enhanced hydrogen production by a newly described heterotrophic marine bacterium, Vibrio tritonius strain AM2, using seaweed as the feedstock. Int. J. Hydrog. Energy 39, 7270–7277 (2014).
Kodani, S., Imoto, A., Mitsutani, A. & Murakami, M. Isolation and identification of the antialgal compound, harmane (1-methyl-β-carboline), produced by the algicidal bacterium, Pseudomonas sp. K44-1. J. Appl. Phycol. 14, 109–114 (2002).
Inoue, A., Nishiyama, R., Mochizuki, S. & Ojima, T. Identification of a 4-Deoxy-l-erythro-5-hexoseulose uronic acid reductase, flred, in an alginolytic bacterium Flavobacterium sp. strain UMI-01. Mar. Drugs 13, 493–508 (2015).
Wigglesworth-Cooksey, B., Cooksey, K. E. & Long, R. Antibiotic from the marine environment with antimicrobial fouling activity. Environ. Toxicol. 22, 275–280 (2007).
Park, S.-C., Lee, J.-K., Kim, S. W. & Park, Y. Selective algicidal action of peptides against harmful algal bloom species. PLoS One 6, e26733 (2018).
Li, Z. et al. A freshwater bacterial strain, Shewanella sp. Lzh-2, isolated from Lake Taihu and its two algicidal active substances, hexahydropyrrolo[1,2-a]pyrazine-1,4-dione and 2, 3-indolinedione. Appl. Microbiol. Biotechnol. 98, 4737–4748 (2014).
Wu, Y. et al. Allelopathic control of cyanobacterial blooms by periphyton biofilms. Environ. Microbiol. 13, 604–615 (2011).
Zhang, B.-H. et al. L-valine, an antialgal amino acid from Streptomyces jiujiangensis JXJ 0074T. Appl. Microbiol. Biotechnol. 100, 4627–4636 (2016).
Wu, L. et al. Bacilysin from Bacillus amyloliquefaciens FZB42 has specific bactericidal activity against harmful algal bloom species. Appl. Environ. Microbiol. 80, 7512–7520 (2014).
An, X. et al. Discovery of an algicidal compound from Brevibacterium sp. BS01 and its effect on a harmful algal bloom-causing species, Alexandrium tamarense. Front. Microbiol. 6, 1235 (2015).
Zhu, Q., Wu, B. & Zhao, L. Effect of algicidal compound Nω-acetylhistamine on physiological response and algal toxins in Heterosigma akashiwo. Ecotoxicol. Environ. Saf. 208, 111423 (2021).
Chen, S., Haga, M., Imai, I., Sakai, R. & Fujita, M. J. Function of the algicidal bacterium Pseudomonas sp. Go58 isolated from the biofilm on a water plant, and its active compounds, pyoluteorins. Sci. Total Environ. 872, 162088 (2023).
Liao, C., Liu, X., Liu, R. & Shan, L. Characterization and effects of two algicidal isolates on antioxidase activities of Chlorella pyrenoidosa. Environ. Prog. Sustain. Energy 34, 1647–1651 (2015).
Liu, J. et al. Algicidal characterization and mechanism of Bacillus licheniformis Sp34 against Microcystis aeruginosa in Dianchi Lake. J. Basic Microbiol. 59, 1112–1124 (2019).
Lee, H. W. et al. Cyanobacteria-specific algicidal mechanism of bioinspired naphthoquinone derivative, NQ 2-0. Sci. Rep. 8, 1–12 (2018).
Zhang, Y., Wang, X. & Sun, Y. A newly identified algicidal bacterium of Pseudomonas fragi YB2: Algicidal compounds and effects. J. Hazard. Mater. 478, 135490 (2024).
Su, J. F. et al. Algicidal and denitrification characterization of Acinetobacter sp. J25 against Microcystis aeruginosa and microbial community in eutrophic landscape water. Mar. Pollut. Bull. 107, 233–239 (2016).
Hibayashi, R. & Imamura, N. Action mechanism of a selective anti-cyanobacterial compound, argimicin A. J. Antibiotics 56, 154–159 (2003).
Kato, Y. et al. Characterization of tryptophan oxidation affecting D1 degradation by FtsH in the photosystem II quality control of chloroplasts. eLife 12, RP88822 (2023).
Zhang, X. et al. 2-Hydroxychalcone as a novel natural photosynthesis inhibitor against bloom-forming cyanobacteria. J. Agric. Food Chem. 70, 15069–15079 (2022).
Yilimulati, M. et al. Regulation of photosynthesis in bloom-forming cyanobacteria with the simplest β-diketone. Environ. Sci. Technol. 55, 14173–14184 (2021).
Li, Y. et al. The first evidence of deinoxanthin from Deinococcus sp. Y35 with strong algicidal effect on the toxic dinoflagellate Alexandrium tamarense. J. Hazard. Mater. 290, 87–95 (2015).
Zhang, H. et al. Effect of oxidative stress induced by Brevibacterium sp. BS01 on a HAB causing species-Alexandrium tamarense. PLOS One 8, e63018 (2017).
Zhang, H. et al. Toxic effects of prodigiosin secreted by Hahella sp KA22 on harmful alga Phaeocystis globosa. Front. Microbiol. 8, 999 (2017).
Ke, Y. et al. The algicidal mechanism of prodigiosin from Hahella sp. KA22 against Microcystis aeruginosa. Sci. Rep. 7, 7750 (2017).
Kim, J.-D., Kim, J.-Y., Park, J.-K. & Lee, C.-G. Selective control of the Prorocentrum minimum harmful algal blooms by a novel algal-lytic bacterium Pseudoalteromonas haloplanktis AFMB-008041. Mar. Biotechnol. 11, 463–472 (2009).
Chen, C.-Y., Bai, M.-D. & Chang, J.-S. Improving microalgal oil collecting efficiency by pretreating the microalgal cell wall with destructive bacteria. Biochem. Eng. J. 81, 170–176 (2013).
Lee, S.-O. et al. Cloning and characterization of extracellular metal protease gene of the algicidal marine bacterium Pseudoalteromonas sp. strain A28. Biosci. Biotechnol. Biochem. 66, 1366–1369 (2002).
Demuez, M., Mahdy, A., Tomás-Pejó, E., González-Fernández, C. & Ballesteros, M. Enzymatic cell disruption of microalgae biomass in biorefinery processes. Biotechnol. Bioeng. 112, 1955–1966 (2015).
Mayali, X., Franks, P. J. S., Tanaka, Y. & Azam, F. Bacteria-induced motility reduction in Lingulodinium polyedrum (Dinophyceae). J. Phycol. 44, 923–928 (2008).
Sockett, R. E. Predatory lifestyle of Bdellovibrio bacteriovorus. Annu. Rev. Microbiol. 63, 523–539 (2009).
Pasternak, Z. et al. In and out: an analysis of epibiotic vs periplasmic bacterial predators. ISME J. 8, 625–635 (2014).
Martin, M. O. Predatory prokaryotes: An emerging research opportunity. J. Mol. Microbiol. Biotechnol. 4, 467–477 (2002).
Shi, S., Liu, Y., Shen, Y., Li, G. & Li, D. Lysis of Aphanizomenon flos-aquae (Cyanobacterium) by a bacterium Bacillus cereus. Biol. Control 39, 345–351 (2006).
Shi, M. et al. A novel bacterium Saprospira sp. strain PdY3 forms bundles and lyses cyanobacteria. Front. Biosci. 11, 1916–1923 (2006).
Caiola, M. G. & Pellegrini, S. Lysis of Microcystis aeruginosa (Kütz.) by Bdellovibrio-like bacteria. J. Phycol. 20, 471–475 (1984).
Moulder, J. W. Comparative biology of intracellular parasitism. Microbiol. Mol. Biol. Rev. 49, 298–337 (1985).
Julie, B., Lynn, J. S. & Ferran, G. P. High impact of bacterial predation on cyanobacteria in soil biocrusts. Nat. Commun. 13, 4835 (2022).
Fu, X., Wu, F. & Liu, X. Bio-archive of cultural heritage microbiomes for sustainable conservation in the multi-omics era. Adv. Genet. 1, e00046 (2025).
Coyne, K. J., Wang, Y. & Johnson, G. Algicidal bacteria: A review of current knowledge and applications to control harmful algal blooms. Front. Microbiol. 13, 871177 (2022).
Liu, X. et al. Biofilms on stone monuments: biodeterioration or bioprotection? Trends Microbiol. 30, 816–819 (2022).
Lu, L. et al. The algicidal efficacy and the mechanism of Enterobacter sp. EA-1 on Oscillatoria dominating in aquaculture system. Environ. Res. 197, 111105 (2021).
Syhapanha, K. S. et al. Transcriptomics-guided identification of an algicidal protease of the marine bacterium Kordia algicida OT-1. MicrobiologyOpen 12, e1387 (2023).
Lu, Q. et al. Impacts of a bacterial algicide on metabolic pathways in Chlorella vulgaris. Ecotoxicol. Environ. Saf. 249, 114451 (2023).
Li, D. et al. Algicidal mechanism of Raoultella ornithinolytica against Microcystis aeruginosa: Antioxidant response, photosynthetic system damage and microcystin degradation. Environ. Pollut. 287, 117644 (2021).
Lu, X. et al. A marine algicidal Thalassospira and its active substance against the harmful algal bloom species Karenia mikimotoi. Appl. Microbiol. Biotechnol. 100, 5131–5139 (2016).
Li, Y. et al. Increasing the autotrophic growth of Chlorella USTB-01 via the control of bacterial contamination by Bdellovibrio USTB-06. J. Appl. Microbiol. 124, 1131–1138 (2018).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (grant nos. 32370105 and 32570139) and the Basic Research Program of Jiangsu (grant no. BK20250086).
Author information
Authors and Affiliations
Contributions
Conceptualization, X.L. Writing—original draft preparation, R.L. and J.Q. Writing—review and editing, R.Y., C.M., and F.W. Supervision, X.L. All authors have read and agreed to the submitted version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial or non-financial interests. Author Xiaobo Liu is an editorial board member of npj Heritage Science. Xiaobo Liu was not involved in the journal’s review of, or decisions related to, this manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liang, R., Qu, J., Yang, R. et al. Algicidal bacteria as an eco-friendly mitigation of microalgae: an application paradigm shift to heritage conservation. npj Herit. Sci. 14, 196 (2026). https://doi.org/10.1038/s40494-026-02474-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s40494-026-02474-y