Abstract
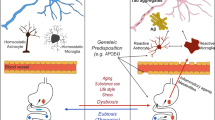
Alzheimer’s disease (AD) is a progressive neurodegenerative disorder with a decade-long preclinical pathological period that can be divided into several stages. Emerging evidence has revealed that the microbiota-gut-brain axis plays an important role in AD pathology. However, the role of gut microbiota in different AD stages has not been well characterized. In this study, we performed fecal shotgun metagenomic analysis on a Chinese cohort with 476 participants across five stages of AD pathology to characterize stage-specific alterations in gut microbiota and evaluate their diagnostic potential. We discovered extensive gut dysbiosis that is associated with neuroinflammation and neurotransmitter dysregulation, with over 10% of microbial species and gene families showing significant alterations during AD progression. Furthermore, we demonstrated that microbial gene families exhibited strong diagnostic capabilities, evidenced by an average AUC of 0.80 in cross-validation and 0.75 in independent external validation. In the optimal model, the most discriminant gene families are primarily involved in the metabolism of carbohydrates, amino acids, energy, glycan and vitamins. We found that stage-specific microbial gene families in AD pathology could be validated by an in vitro gut simulator and were associated with specific genera. We also observed that the gut microbiota could affect the progression of cognitive decline in 5xFAD mice through fecal microbiota transplantation, which could be used for early intervention of AD. Our multi-stage large cohort metagenomic analysis demonstrates that alterations in gut microbiota occur from the very early stages of AD pathology, offering important etiological and diagnostic insights.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Shotgun metagenomic sequencing reads of our AD cohort is available in the China National Center for Bioinformation (CNCB) under accession number PRJCA022804. The publicly available Japan cohort can be accessed in the National Center for Biotechnology Information BioProject PRJNA679346. The five 16S rRNA amplicon sequencing datasets can be downloaded from the BioProject of NCBI: PRJNA489760, PRJNA554111, PRJNA633959, PRJNA734525, and PRJNA811324. The publicly available cohorts for other brain disorders can also be accessed in the BioProject of NCBI: PRJNA798058 for preclinical AD, PRJNA834801 for PD, PRJEB29127 for SCZ, and PRJNA516054, PRJEB23052 and PRJNA451479 for ASD. The publicly available BD cohort can be accessed in China National GeneBank DataBase (CNGBdb) with accession number CNP0002003.
Code availability
All custom code is available at the following GitHub repository: https://github.com/ZhaoXM-Lab/metaAD_analysis.
References
Scheltens P, De Strooper B, Kivipelto M, Holstege H, Chetelat G, Teunissen CE, et al. Alzheimer’s disease. Lancet. 2021;397:1577–90.
McKhann GM, Knopman DS, Chertkow H, Hyman BT, Jack CR Jr, Kawas CH, et al. The diagnosis of dementia due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimer’s & dementia. 2011;7:263–9.
Long S, Benoist C, Weidner W. World Alzheimer report 2023: reducing dementia risk: never too early, never too late. London, England: Alzheimer’s Disease International; 2023.
Long JM, Holtzman DM. Alzheimer disease: an update on pathobiology and treatment strategies. Cell. 2019;179:312–39.
Self WK, Holtzman DM. Emerging diagnostics and therapeutics for Alzheimer disease. Nature Medicine. 2023;29:2187–99.
Huang L, Chen K, Liu Z, Guo Q. A conceptual framework for research on cognitive impairment with no dementia in memory clinic. Current Alzheimer Research. 2020;17:517–25.
Edmonds EC, Delano-Wood L, Galasko DR, Salmon DP, Bondi MW, Brandt J. Subtle cognitive decline and biomarker staging in preclinical Alzheimer’s disease. Journal of Alzheimer’s Disease. 2015;47:231–42.
Bondi MW, Edmonds EC, Jak AJ, Clark LR, Delano-Wood L, McDonald CR, et al. Neuropsychological criteria for mild cognitive impairment improves diagnostic precision, biomarker associations, and progression rates. Journal of Alzheimer’s Disease. 2014;42:275–89.
Thomas KR, Bangen KJ, Weigand AJ, Edmonds EC, Wong CG, Cooper S, et al. Objective subtle cognitive difficulties predict future amyloid accumulation and neurodegeneration. Neurology. 2020;94:e397–e406.
Papp KV, Buckley R, Mormino E, Maruff P, Villemagne VL, Masters CL, et al. Clinical meaningfulness of subtle cognitive decline on longitudinal testing in preclinical. AD. Alzheimer’s & Dementia. 2020;16:552–60.
Van Dyck CH, Swanson CJ, Aisen P, Bateman RJ, Chen C, Gee M, et al. Lecanemab in early Alzheimer’s disease. New England Journal of Medicine. 2023;388:9–21.
Jack CR Jr, Bennett DA, Blennow K, Carrillo MC, Dunn B, Haeberlein SB, et al. NIA-AA research framework: toward a biological definition of Alzheimer’s disease. Alzheimer’s & Dementia. 2018;14:535–62.
Khedr EM, Omeran N, Karam-Allah Ramadan H, Ahmed GK, Abdelwarith AM. Alteration of gut microbiota in Alzheimer’s disease and their relation to the cognitive impairment. Journal of Alzheimer’s Disease. 2022;88:1103–14.
Dubois B, Villain N, Frisoni GB, Rabinovici GD, Sabbagh M, Cappa S, et al. Clinical diagnosis of Alzheimer’s disease: recommendations of the International Working Group. The Lancet Neurology. 2021;20:484–96.
Cryan JF, O’Riordan KJ, Cowan CSM, Sandhu KV, Bastiaanssen TFS, Boehme M, et al. The Microbiota-Gut-Brain Axis. Physiological Reviews. 2019;99:1877–2013.
Morais LH, Schreiber IVHL, Mazmanian SK. The gut microbiota–brain axis in behaviour and brain disorders. Nature Reviews Microbiology. 2021;19:241–55.
Loh JS, Mak WQ, Tan LKS, Ng CX, Chan HH, Yeow SH, et al. Microbiota–gut–brain axis and its therapeutic applications in neurodegenerative diseases. Signal Transduction and Targeted Therapy. 2024;9:37.
Varesi A, Pierella E, Romeo M, Piccini GB, Alfano C, Bjorklund G, et al. The potential role of Gut Microbiota in Alzheimer’s disease: from diagnosis to treatment. Nutrients. 2022;14:668.
Li B, He Y, Ma J, Huang P, Du J, Cao L, et al. Mild cognitive impairment has similar alterations as Alzheimer’s disease in gut microbiota. Alzheimer’s & Dementia. 2019;15:1357–66.
Haran JP, Bhattarai SK, Foley SE, Dutta P, Ward DV, Bucci V, et al. Alzheimer’s disease microbiome is associated with dysregulation of the anti-inflammatory P-Glycoprotein pathway. mBio. 2019;10:e00632–19.
Ferreiro AL, Choi J, Ryou J, Newcomer EP, Thompson R, Bollinger RM, et al. Gut microbiome composition may be an indicator of preclinical Alzheimer’s disease. Science Translational Medicine. 2023;15:eabo2984.
Liang Y, Liu C, Cheng M, Geng L, Li J, Du W, et al. The link between gut microbiome and Alzheimer’s disease: from the perspective of new revised criteria for diagnosis and staging of Alzheimer’s disease. Alzheimer’s & Dementia. 2024;20:5771–88.
Chandra S, Sisodia SS, Vassar RJ. The gut microbiome in Alzheimer’s disease: what we know and what remains to be explored. Molecular Neurodegeneration. 2023;18:9.
Li M, Liu J, Zhu J, Wang H, Sun C, Gao NL, et al. Performance of gut microbiome as an independent diagnostic tool for 20 diseases: cross-cohort validation of machine-learning classifiers. Gut Microbes. 2023;15:2205386.
Xiao L, Zhang F, Zhao F. Large-scale microbiome data integration enables robust biomarker identification. Nature Computational Science. 2022;2:307–16.
Ranjan R, Rani A, Metwally A, McGee HS, Perkins DL. Analysis of the microbiome: Advantages of whole genome shotgun versus 16S amplicon sequencing. Biochemical and Biophysical Research Communications. 2016;469:967–77.
Liu P, Wu L, Peng G, Han Y, Tang R, Ge J, et al. Altered microbiomes distinguish Alzheimer’s disease from amnestic mild cognitive impairment and health in a Chinese cohort. Brain, Behavior, and Immunity. 2019;80:633–43.
Ueda A, Shinkai S, Shiroma H, Taniguchi Y, Tsuchida S, Kariya T, et al. Identification of Faecalibacterium prausnitzii strains for gut microbiome-based intervention in Alzheimer’s-type dementia. Cell Reports Medicine. 2021;2:100398.
Wu H, Esteve E, Tremaroli V, Khan MT, Caesar R, Mannerås-Holm L, et al. Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nature Medicine. 2017;23:850–8.
Koh A, Molinaro A, Ståhlman M, Khan MT, Schmidt C, Mannerås-Holm L, et al. Microbially produced imidazole propionate impairs insulin signaling through mTORC1. Cell. 2018;175:947–61.e917.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet Journal. 2011;17:3.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
Wood DE, Lu J, Langmead B. Improved metagenomic analysis with Kraken 2. Genome Biology. 2019;20:1–13.
Lu J, Rincon N, Wood DE, Breitwieser FP, Pockrandt C, Langmead B, et al. Metagenome analysis using the Kraken software suite. Nature Protocols. 2022;17:2815–39.
Li D, Liu CM, Luo R, Sadakane K, Lam TW. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics. 2015;31:1674–6.
Hyatt D, Chen G-L, LoCascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics. 2010;11:119.
Fu L, Niu B, Zhu Z, Wu S, Li W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics. 2012;28:3150–2.
Cantalapiedra CP, Hernández-Plaza A, Letunic I, Bork P, Huerta-Cepas J, Tamura K. eggNOG-mapper v2: functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Molecular Biology and Evolution. 2021;38:5825–9.
Dixon P. VEGAN, a package of R functions for community ecology. Journal of Vegetation Science. 2003;14:927–30.
Ma S, Shungin D, Mallick H, Schirmer M, Nguyen LH, Kolde R, et al. Population structure discovery in meta-analyzed microbial communities and inflammatory bowel disease using MMUPHin. Genome Biology. 2022;23:208.
Nearing JT, Douglas GM, Hayes MG, MacDonald J, Desai DK, Allward N, et al. Microbiome differential abundance methods produce different results across 38 datasets. Nature Communications. 2022;13:342.
Mallick H, Rahnavard A, McIver LJ, Ma S, Zhang Y, Nguyen LH, et al. Multivariable association discovery in population-scale meta-omics studies. PLOS Computational Biology. 2021;17:e1009442.
Wang DD, Nguyen LH, Li Y, Yan Y, Ma W, Rinott E, et al. The gut microbiome modulates the protective association between a Mediterranean diet and cardiometabolic disease risk. Nature Medicine. 2021;27:333–43.
Beck LC, Masi AC, Young GR, Vatanen T, Lamb CA, Smith R, et al. Strain-specific impacts of probiotics are a significant driver of gut microbiome development in very preterm infants. Nature Microbiology. 2022;7:1525–35.
Huang B, Chau SWH, Liu Y, Chan JWY, Wang J, Ma SL, et al. Gut microbiome dysbiosis across early Parkinson’s disease, REM sleep behavior disorder and their first-degree relatives. Nature Communications. 2023;14:2501.
Wallen ZD, Demirkan A, Twa G, Cohen G, Dean MN, Standaert DG, et al. Metagenomics of Parkinson’s disease implicates the gut microbiome in multiple disease mechanisms. Nature Communications. 2022;13:6958.
Zhu F, Ju Y, Wang W, Wang Q, Guo R, Ma Q, et al. Metagenome-wide association of gut microbiome features for schizophrenia. Nature Communications. 2020;11:1612.
Li Z, Lai J, Zhang P, Ding J, Jiang J, Liu C, et al. Multi-omics analyses of serum metabolome, gut microbiome and brain function reveal dysregulated microbiota-gut-brain axis in bipolar depression. Molecular Psychiatry. 2022;27:4123–35.
Morton JT, Jin D-M, Mills RH, Shao Y, Rahman G, McDonald D, et al. Multi-level analysis of the gut–brain axis shows autism spectrum disorder-associated molecular and microbial profiles. Nature Neuroscience. 2023;26:1208–17.
Breiman L. Random forests. Machine learning. 2001;45:5–32.
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, et al. Scikit-learn: machine learning in python. Journal of Machine Learning Research. 2011;12:2825–30.
Ling Z, Zhu M, Yan X, Cheng Y, Shao L, Liu X, et al. Structural and functional dysbiosis of fecal microbiota in Chinese patients with Alzheimer’s disease. Frontiers in Cell and Developmental Biology. 2021;8:634069.
Zhuang Z-Q, Shen L- L, Li W-W, Fu X, Zeng F, Gui L, et al. Gut microbiota is altered in patients with Alzheimer’s disease. Journal of Alzheimer’s Disease. 2018;63:1337–46.
Kaiyrlykyzy A, Kozhakhmetov S, Babenko D, Zholdasbekova G, Alzhanova D, Olzhayev F, et al. Study of gut microbiota alterations in Alzheimer’s dementia patients from Kazakhstan. Scientific Reports. 2022;12:15115.
Yıldırım S, Nalbantoğlu ÖU, Bayraktar A, Ercan FB, Gündoğdu A, Velioğlu HA, et al. Stratification of the gut microbiota composition landscape across the Alzheimer’s disease continuum in a Turkish cohort. Msystems. 2022;7:e00004–00022.
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nature Biotechnology. 2019;37:852–7.
Douglas GM, Maffei VJ, Zaneveld JR, Yurgel SN, Brown JR, Taylor CM, et al. PICRUSt2 for prediction of metagenome functions. Nature Biotechnology. 2020;38:685–8.
Ling W, Lu J, Zhao N, Lulla A, Plantinga AM, Fu W, et al. Batch effects removal for microbiome data via conditional quantile regression. Nature Communications. 2022;13:5418.
Grabrucker S, Marizzoni M, Silajdžić E, Lopizzo N, Mombelli E, Nicolas S, et al. Microbiota from Alzheimer’s patients induce deficits in cognition and hippocampal neurogenesis. Brain. 2023;146:4916–34.
Farias ST, Mungas D, Reed BR, Cahn-Weiner D, Jagust W, Baynes K, et al. The measurement of everyday cognition (ECog): scale development and psychometric properties. Neuropsychology. 2008;22:531–44.
Katzman R, Zhang M, Ouang Ya Q, Wang Z, Liu WT, Yu E, et al. A Chinese version of the mini-mental state examination; Impact of illiteracy in a Shanghai dementia survey. Journal of Clinical Epidemiology. 1988;41:971–8.
Chen KL, Xu Y, Chu AQ, Ding D, Liang XN, Nasreddine ZS, et al. Validation of the Chinese version of Montreal cognitive assessment basic for screening mild cognitive impairment. Journal of the American Geriatrics Society. 2016;64:e285–e290.
Pan F-F, Wang Y, Huang L, Huang Y, Guo Q-H. Validation of the Chinese version of Addenbrooke’s cognitive examination III for detecting mild cognitive impairment. Aging & Mental Health. 2022;26:384–91.
Seo DO, O’Donnell D, Jain N, Ulrich JD, Herz J, Li Y, et al. ApoE isoform- and microbiota-dependent progression of neurodegeneration in a mouse model of tauopathy. Science. 2023;379:eadd1236.
Vogt NM, Kerby RL, Dill-McFarland KA, Harding SJ, Merluzzi AP, Johnson SC, et al. Gut microbiome alterations in Alzheimer’s disease. Scientific Reports. 2017;7:13537.
Nagpal R, Neth BJ, Wang S, Craft S, Yadav H. Modified Mediterranean-ketogenic diet modulates gut microbiome and short-chain fatty acids in association with Alzheimer’s disease markers in subjects with mild cognitive impairment. EBioMedicine. 2019;47:529–42.
Zhang D, Li N, Wang Y, Lu W, Zhang Y, Chen Y, et al. Methane ameliorates post-operative cognitive dysfunction by inhibiting microglia NF-κB/MAPKs pathway and promoting IL-10 expression in aged mice. International Immunopharmacology. 2019;71:52–60.
Robinson N, Grabowski P, Rehman I. Alzheimer’s disease pathogenesis: is there a role for folate? Mechanisms of Ageing and Development. 2018;174:86–94.
Qian X, Hai W, Chen S, Zhang M, Jiang X, Tang H. Multi-omics data reveals aberrant gut microbiota-host glycerophospholipid metabolism in association with neuroinflammation in APP/PS1 mice. Gut Microbes. 2023;15:2282790.
Xi Y, Zhang Y, Zhou Y, Liu Q, Chen X, Liu X, et al. Effects of methionine intake on cognitive function in mild cognitive impairment patients and APP/PS1 Alzheimer’s disease model mice: role of the cystathionine-β-synthase/H2S pathway. Redox Biology. 2023;59:102595.
Lin C-H, Chen P-K, Chang Y-C, Chuo L-J, Chen Y-S, Tsai GE, et al. Benzoate, a D-amino acid oxidase inhibitor, for the treatment of early-phase Alzheimer disease: a randomized, double-blind, placebo-controlled trial. Biological Psychiatry. 2014;75:678–85.
Paley EL. Discovery of Gut Bacteria specific to Alzheimer’s associated diseases is a clue to understanding disease etiology: meta-analysis of population-based data on human gut metagenomics and metabolomics. Journal of Alzheimer’s Disease. 2019;72:319–55.
Connell E, Le Gall G, Pontifex MG, Sami S, Cryan JF, Clarke G, et al. Microbial-derived metabolites as a risk factor of age-related cognitive decline and dementia. Molecular Neurodegeneration. 2022;17:43.
Fang P, Kazmi SA, Jameson KG, Hsiao EY. The microbiome as a modifier of neurodegenerative disease risk. Cell Host Microbe. 2020;28:201–22.
Thu Thuy Nguyen V, Endres K. Targeting gut microbiota to alleviate neuroinflammation in Alzheimer’s disease. Advanced Drug Delivery Reviews. 2022;188:114418.
Jones RB, Alderete TL, Kim JS, Millstein J, Gilliland FD, Goran MI. High intake of dietary fructose in overweight/obese teenagers associated with depletion of Eubacterium and Streptococcus in gut microbiome. Gut Microbes. 2019;10:712–9.
Shkoporov AN, Hill C. Bacteriophages of the human gut: the “known unknown” of the microbiome. Cell Host Microbe. 2019;25:195–209.
Almeida A, Nayfach S, Boland M, Strozzi F, Beracochea M, Shi ZJ, et al. A unified catalog of 204,938 reference genomes from the human gut microbiome. Nature Biotechnology. 2021;39:105–14.
Heintz-Buschart A, Wilmes P. Human gut microbiome: function matters. Trends in Microbiology. 2018;26:563–74.
Liu N-N, Jiao N, Tan J-C, Wang Z, Wu D, Wang A-J, et al. Multi-kingdom microbiota analyses identifybacterial–fungal interactions and biomarkers of colorectal cancer acrosscohorts. Nature Microbiology. 2022;7:238–50.
Kalynych S, Morona R, Cygler M. Progress in understanding the assembly process of bacterial O-antigen. FEMS Microbiology Reviews. 2014;38:1048–65.
Laman JD, ’t Hart BA, Power C, Dziarski R. Bacterial peptidoglycan as a driver of chronic brain inflammation. Trends in Molecular Medicine. 2020;26:670–82.
Feng M, Hou T, Zhou M, Cen Q, Yi T, Bai J, et al. Gut microbiota may be involved in Alzheimer’s disease pathology by dysregulating pyrimidine metabolism in APP/PS1 mice. Frontiers in Aging Neuroscience. 2022;14:967747.
Chen C, Liao J, Xia Y, Liu X, Jones R, Haran J, et al. Gut microbiota regulate Alzheimer’s disease pathologies and cognitive disorders via PUFA-associated neuroinflammation. Gut. 2022;71:2233–52.
Chaudhuri J, Bains Y, Guha S, Kahn A, Hall D, Bose N, et al. The role of advanced glycation end products in aging and metabolic diseases: bridging association and causality. Cell Metabolism. 2018;28:337–52.
Liu Q, Xi Y, Wang Q, Liu J, Li P, Meng X, et al. Mannan oligosaccharide attenuates cognitive and behavioral disorders in the 5xFAD Alzheimer’s disease mouse model via regulating the gut microbiota-brain axis. Brain, Behavior, and Immunity. 2021;95:330–43.
Ansoleaga B, Jové M, Schlüter A, Garcia-Esparcia P, Moreno J, Pujol A, et al. Deregulation of purine metabolism in Alzheimer’s disease. Neurobiology of Aging. 2015;36:68–80.
Liu P, Fleete MS, Jing Y, Collie ND, Curtis MA, Waldvogel HJ, et al. Altered arginine metabolism in Alzheimer’s disease brains. Neurobiology of Aging. 2014;35:1992–2003.
Guo M, Peng J, Huang X, Xiao L, Huang F, Zuo Z. Gut microbiome features of Chinese patients newly diagnosed with Alzheimer’s disease or mild cognitive impairment. Journal of Alzheimer’s Disease. 2021;80:299–310.
Rossi M, Amaretti A, Raimondi S. Folate production by probiotic bacteria. Nutrients. 2011;3:118–34.
Avalos M, Garbeva P, Vader L, van Wezel GP, Dickschat JS, Ulanova D. Biosynthesis, evolution and ecology of microbial terpenoids. Natural Product Reports. 2022;39:249–72.
Schoeler M, Caesar R. Dietary lipids, gut microbiota and lipid metabolism. Reviews in Endocrine and Metabolic Disorders. 2019;20:461–72.
Song L, Chen J, Lo C-YZ, Guo Q, Feng J, Zhao X-M Impaired type I interferon signaling activity implicated in the peripheral blood transcriptome of preclinical Alzheimer’s disease. EBioMedicine 2022; 82.
Song L, Yang YT, Guo Q, Zhao X-M, the ZIBC. Cellular transcriptional alterations of peripheral blood in Alzheimer’s disease. BMC Medicine. 2022;20:266.
Acknowledgements
The authors thank the patients and healthy participants who made this study possible and provided their fecal samples. This work was partly supported by National Natural Science Foundation of China (T2225015, 61932008, 62433008), Shanghai Science and Technology Commission Program (23JS1410100), Key Science and Technology Project of Hainan Province (ZDYF2024SHFZ058), National Key R&D Program of China (2023YFF1204800, 2020YFA0712403), Lingang Laboratory & National Key Laboratory of Human Factors Engineering Joint Grant (LG-TKN-202203-01) and Lingang Laboratory (LG-GG-202401-ADA010100 and LG-GG-202401-ADA050100). The computations in this research were performed using the CFFF platform of Fudan University.
Author information
Authors and Affiliations
Consortia
Contributions
XMZ and JD conceived the study and supervised the project. LHJ performed metagenomic analysis and wrote the first draft of the manuscript. QHG managed the sampling of fecal samples. YZK and HW conducted experimental validation by an in vitro intestinal simulator and fecal microbiota transplantation. SW, YJL, FC and JW performed behavioral monitoring based on novel object recognition and Y-maze tests. WHC and JXL provided useful suggestions. XHL and JXC helped curate metadata. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jia, L., Ke, Y., Zhao, S. et al. Metagenomic analysis characterizes stage-specific gut microbiota in Alzheimer’s disease. Mol Psychiatry 30, 3951–3962 (2025). https://doi.org/10.1038/s41380-025-02973-7
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41380-025-02973-7
This article is cited by
-
Cross-domain metabolic interactions link Methanobrevibacter smithii to colorectal cancer microbial ecosystems
Nature Communications (2026)
-
Compositional and functional gut microbiota alterations in mild cognitive impairment: links to Alzheimer’s disease pathology
Alzheimer's Research & Therapy (2025)