Abstract
Klinefelter syndrome (47,XXY; KS) impacts neurodevelopment. Furthermore, KS is associated with widespread alterations in the epigenome and transcriptome. Whether these epigenetic and transcriptomic alterations can be linked to the neurocognitive phenotype remains to be elucidated. We performed a comprehensive, integrative analysis of the neurocognitive profile and the methylome in blood from males with KS (n = 65) and male controls (n = 63) (Cohort 1). The results were validated in a second cohort of males with KS (n = 22) and male controls (n = 16) in which transcriptome data was also available (Cohort 2). The findings were further validated in neural precursor cells derived from human induced pluripotent stem cells from 47,XXY (n = 3) and 46,XY (n = 3) amniotic cells. In cohort 1, we identified five CG-sites within the TRAK1 gene which were hypomethylated in males with KS compared to male controls. TRAK1 hypomethylation was positively correlated with several neurocognitive variables among males with KS. In cohort 2, we identified a similar methylation pattern and demonstrated that the methylation levels at the five CG-sites were correlated with a high expression level of a specific short TRAK1 transcript (ENST00000341421.7). Neural precursor cells (NPCs) established from 47,XXY amniotic cells also exhibited hypomethylation at the five CG-sites and strong upregulation of ENST00000341421.7 compared to NPCs established from 46,XY amniotic cells. In conclusion, we demonstrate that the DNA methylation level at specific CG-sites within TRAK1, a gene highly expressed in the brain, is correlated with the neurocognitive phenotype of KS, implying a possible epigenetic underpinning for the observed neurocognitive impairments in KS.
Similar content being viewed by others
Introduction
Klinefelter syndrome (47,XXY; KS) is the most prevalent form of sex chromosome aneuploidy, affecting one in 660 males [1]. The phenotypic traits of KS include hypergonadotropic hypogonadism, small testes, infertility, altered body composition, and increased height (reviewed in [2]). KS also affects neurodevelopment and is associated with an increased risk of neurocognitive impairments, although the variability is large. Approximately 70–80% of males with KS have verbal deficits that affect both receptive and expressive language functions [3, 4]. Several studies [5,6,7,8] have also documented an increased frequency of impaired executive functioning in males with KS, including deficits in attention, working memory, cognitive flexibility, and inhibitory control. On average, full-scale IQ (FSIQ) of males with KS is about 10 points lower than that of the general population, with verbal IQ (VIQ) being more severely affected than performance IQ (PIQ) [5, 6, 8,9,10,11], which has been coined as an IQ split. This is also observed in Turner syndrome (45,X), but with a mirror-image of higher VIQ than PIQ [12].
Interestingly, a mirror-image of the DNA methylation pattern is also seen in KS and TS [13,14,15]. DNA methylation is crucial for normal brain development such as neuronal differentiation and maturation, and normal brain functions, including learning and memory [16,17,18]. Global cognitive ability and executive function have been reported to be associated with specific DNA methylation patterns [19]. We, and others, have previously shown that KS is associated with genome-wide alterations in DNA methylation across different tissues [13,14,15, 20,21,22]. Genes that are differentially methylated in KS are involved in a variety of biological processes related to neural development and neurotransmission, suggesting that alterations in DNA methylation could be implicated in the neurocognitive impairments seen in KS [14, 15].
Here, we present a comprehensive and integrative analysis of the neurocognitive profile and the methylome in KS, aiming to investigate the link between specific neurocognitive traits (processing speed, working memory, visual performance, verbal memory and learning, verbal fluency and comprehension, verbal performance, and executive functions) and DNA methylation patterns, and to identify potential epigenetic biomarkers of neurocognitive function in KS.
Materials and Methods
Participants
Cohort 1
Males with verified KS and male controls were included as described previously [6, 14]. Participants were enrolled between 2009 and 2012. Neurocognitive data and DNA methylation data were available for 65 males with KS (mean age 36.8 ± 10.5 (SD) years) and 63 controls matched for age and education (mean age 36.4 ± 10.3 (SD) years) (Fig. 1; Table 1).
DNA methylation data from peripheral blood samples and neurocognitive data were investigated for cohort 1 (left panel) that included 65 males with KS and 63 male controls. For cohort 2 (middle panel; 22 males with KS and 16 male controls), DNA methylation and RNA sequencing data from peripheral blood samples as well as neurocognitive data were examined. DNA methylation and RNA sequencing data were investigated in XXY (n = 3) and XY (n = 3) neural precursor cells (right panel) derived from human induced pluripotent stem cells (hiPSCs), obtained from amniocytes. This figure was made using BioRender.com.
Cohort 2
Cohort 2 consisted of 22 males with verified KS (mean age 38.3 ± 8.1 (SD) years) and 16 male controls matched for age and education (mean age 38.4 ± 12.0 (SD) years) [15]. Participants were enrolled between 2015 and 2019. Males with KS were recruited from fertility and endocrinology clinics across Denmark. Controls were included by public advertising. DNA methylation data, gene expression data and neurocognitive data were available for all participants (Fig. 1; Table 2).
Ethics
The study was approved by The Danish Data Protection Agency and the local ethics committee (Region Midtjylland, Denmark number M-20080238, Central Denmark Regional Committee on Health Research Ethics number 1-10-72-131-15) and conducted according to the principles expressed in the Declaration of Helsinki. Every participant provided informed consent. This research was registered at ClinicalTrials.gov (NCT00999310, NCT02526628).
Cognitive evaluation
As previously described [6], participants from cohort 1 underwent a comprehensive battery of standardized neuropsychological tests to assess their neurocognitive functions. In the present study, only domains and subtests in which males with KS and control males differed significantly were included. These tests included: processing speed (Trail Making A (TMT-A) and B (TMT-B) [23], and the Coding subtest (WAIS-C) of Wechsler Adult Intelligence Scale III (WAIS-III)) [24], working memory (Digit Span (WAIS-DS) and Letter-Number Sequencing (WAIS-LN) subtests of WAIS-III)), visual performance (Block Design (WAIS-BD) and Matrix Reasoning (WAIS-MR) subtests of WAIS-III), verbal memory and learning (total score of Rey Auditory Verbal Learning Test (RAVL-tot)), verbal fluency and comprehension (Sentence Repetition (SR)), verbal performance (Verbal fluency test (Fluency), Similarities (WAIS-S) and Vocabulary (WAIS-V) subtests of WAIS-III), executive function (cards administered of the Wisconsin Card Sorting Test (WCST-trai) [25], total move score (TOL-tms) and total correct score (TOL-tcs) of the Tower of London) [26], and FSIQ, VIQ and PIQ. FSIQ, VIQ and PIQ was determined by regression equations derived from summary statistics data (e.g. mean, standard deviation, correlations of the subtest) from the WAIS-III Danish reference material [24]:
Participants from cohort 2 underwent a three-hour battery of standardized cognitive tests, designed to assess major cognitive functions. The tests were administered and scored by trained research assistants under the supervision of a licensed psychologist. The test battery consisted of Wechsler Adult Intelligence Scale IV (WAIS-IV) [27], Wechsler Memory Scale III (WMS–III) [28] and the Wisconsin Card Sorting Test (WCST) [25]. Only data from the WCST and the subtest of the WAIS that were also available for cohort 1 were included.
DNA methylation and gene expression in blood
For cohort 1, all methylation data from peripheral blood were obtained from a previously published study [14]. For cohort 2, all methylation and RNA sequencing data from peripheral blood were obtained from a previously published study [15].
Raw intensity values for cohort 1 (Infinium HumanMethylation450k) and cohort 2 (Infinium MethylationEPIC) were imported and processed using the R package Minfi [29]. Cross-reactive probes and poorly performing probes, as indicated by a detection p-value < 0.01, were excluded from the analysis. The preprocess Funnorm normalization method [30] was applied to remove between-array variation inferred by control probes, followed by the conversion of methylation values to beta-values and M-values. For differential methylation analysis, M-values were analyzed using LIMMA [31], with differentially methylated positions defined as having an adjusted p-value < 0.05. Beta-values of the differentially methylated positions (DMPs) were used for downstream analysis.
For RNA expression, the fastq files underwent initial quality control using FastQC (Babraham Bioinformatics, Cambridge, UK). Adaptor removal and trimming of low-quality ends were performed using Trim Galore with default settings (Babraham Bioinformatics, Cambridge, UK). Transcript expression levels were quantified using Salmon [32], with a decoy-aware transcriptome index based on the hg38 transcriptome. Transcript abundances were summarized to the gene level using the R package Tximeta [33].
Neural precursor cells (NPCs)
Derivation of human induced pluripotent stem cells (hiPSCs)
KS (47,XXY, n = 3) and control male (46,XY, n = 3) human induced pluripotent stem cell (hiPSC) lines were derived from amniocytes using non-integrating episomal vectors (Fig. 1) [34]. KS amniotic fluid-derived cell lines X4, X5, and X6 were obtained from the Coriell Institute, Camden, NJ, USA, under the identifiers GM02269, GM03091, and GM03535, respectively, and karyotypically normal control male amniocytes M7, M8, and M9 were obtained from Dr. Daniel L. Van Dyke at the Mayo Clinic, Rochester, MN, USA. Briefly, 1×106 amniocytes from each line were reprogrammed via nucleofection with three non-integrating plasmids overexpressing human SOX2 and KL4 (pCXLE-hSK, plasmid #27078, Addgene, Watertown, MA, USA), human L-MYC and LIN28 (pCXLE-hUL, plasmid #27080, Addgene, Watertown, MA, USA), and human OCT3/4 and shRNA against p53 (pCXLE-OCT3/4-shp53, plasmid #27077, Addgene, Watertown, MA, USA) and plated onto gelatinized plates in DMEM media containing 10% FBS and 1% penicillin-streptomycin (Gibco, Grand Island, NY, USA). Following six-seven days in culture, cells were re-plated at single-cell density onto mouse embryo fibroblast (MEF) feeders in F12 media consisting of DMEM/F12, 20% Knockout Serum Replacer, 1x non-essential amino acids, 1x Glutamax-100, β-ME, and 1% penicillin-streptomycin (all from Gibco, Grand Island, NY, USA), supplemented with the HDAC inhibitors SAHA and Sodium Butyrate (Sigma, St. Louis, MO, USA). By day 20–28, following the emergence of iPSC colonies, cells were switched to F12 media supplemented with FGF-2 (Gibco, Grand Island, NY, USA) and without HDAC inhibitors. Colonies with typical iPSC morphology were individually picked up, expanded, and passaged with dispase on MEF feeders in F12 media supplemented with FGF. The derived iPSC clones were tested for the expression of the key endogenous pluripotency factors such as OCT4 and NANOG, and for episomal/viral clearance by testing for lack of the vector containing transcripts as described in Okita K. et al. [34]. Prior to differentiation, iPSCs were adapted to feeder-less conditions by passaging to matrigel- or laminin-coated plates using accutase.
hiPSC clones were karyotyped for commonly duplicated or deleted genomic regions using hPSC Genetic Analysis qPCR kit (STEMCELL Technologies, Vancouver, BC, Canada). X chromosome aneuploidy was confirmed by FISH, XIST genomic DNA-qPCR, and CNV analysis of 850 K DNA methylation array data. RT-qPCR for XIST expression was performed to screen for X-chromosome inactivation (XCI) erosion. Only the XXY cell lines with levels of XIST expression comparable to XX lines were used for differentiation and downstream molecular studies.
Differentiation of hiPSCs to neural precursor cells (NPCs)
hiPSCs were differentiated into neural precursor cells (NPCs) using a dual SMAD inhibition protocol adapted from previously published studies (Fig. 1) [35, 36]. Briefly, cells were densely plated into 24- or 12-well plates pre-coated with laminin-521 (BioLamina, Sundbyberg, Sweden). NPC differentiation was induced in BNMM media, consisting of a 1:1 mix of DMEM/F12 and neural basal media, supplemented with N2, B27, non-essential amino acids, Glutamax-100, Insulin-transferrin-selenium-sodium pyruvate (ITSA), β-ME, 1% penicillin-streptomycin (all from Gibco, Grand Island, NY, USA), and the SMAD inhibitors SB431542 and LDN193189 (Selleck, Houston, TX, USA). Following nine days of neural induction, cells were treated with Versene (Gibco, Grand Island, NY, USA) and collected using a cell scraper. After gentle trituration, cells were passaged into 12-well plates pre-coated with laminin-521 and fed with BNMM media without SMAD inhibitors for three days. Following the emergence of neural rosettes, cells were fed with BNMM media supplemented with FGF (Gibco, Grand Island, NY, USA) without SMAD inhibitors. Cell pellets were collected for downstream molecular analyses at day 19 of differentiation.
DNA methylation and gene expression in neuro precursor cells
DNA extraction and methylation analysis in NPCs
DNA lysates were prepared using the DNeasy blood and tissue kit (Qiagen, Hilden, Germany). DNA from NPCs was subjected to bisulfite conversion using the EZ DNA Methylation kit according to the manufacturer’s protocol (Zymo Research, Irvine, CA, USA). Methylation data were generated using Illumina 850 K methylation arrays (Illumina, San Diego, CA, USA). Raw Idat files were read into R using the Bioconductor Minfi package and processed using the adjustedFunnorm function from the Bioconductor wateRmelon package. Briefly, this method first corrects for background binding and dye bias using NOOB [37], and then normalizes using functional normalization [30, 38], with an additional interpolation of the probes on the X and Y chromosomes to avoid sex bias. Probes with low binding (not significantly different from background) were excluded from further analyses.
RNA extraction, cDNA library preparation, and sequencing
NPCs were collected and stored in Qiazol (Qiagen, Hilden, Germany) before processing. Total RNA was isolated using the Qiagen RNeasy mini kit (Qiagen, Hilden, Germany). Bulk RNA-seq indexed libraries were prepared using Illumina TruSeq RNA sample preparation kit V2 (Illumina, San Diego, CA, USA). All libraries were generated from 1 µg of total RNA starting material prior to mRNA isolation. cDNA libraries were constructed for analysis on a NextSeq sequencer and 75 bp single-end reads were generated.
RNAseq data analysis
The fastq files underwent initial quality control using FastQC (Babraham Bioinformatics, Cambridge, UK). Adaptor removal and trimming of low-quality ends were performed using Trim Galore with default settings (Babraham Bioinformatics, Cambridge, UK). Transcript expression levels were quantified using Salmon [32], with a decoy-aware transcriptome index based on the hg38 transcriptome. Transcript abundances were summarized to the gene level using the R package Tximeta [33].
Weighted correlation network analysis
A weighted gene correlation network analysis (WGCNA, v1.70.3) [39] was applied to methylation data from cohort 1. The analysis was restricted to males with KS, evaluating the potential link between methylation patterns and neurocognitive traits within the KS group.
The beta-values of the differentially methylated positions (DMPs) between KS and controls (nDMPs = 32 977) were used as input for the analysis. A signed co-expression network was constructed using a one-step approach, calculating adjacency by choosing an appropriate soft thresholding power with approximate scale-free topology. Clustering was performed on the signed Topology Overlap Matrix by hierarchical clustering, identifying modules via the blockwiseModules function. The module eigengenes were calculated via the moduleEigengenes function, and eigengene significance and corresponding p-value were obtained for each module-trait association.
Combined methylation score of TRAK1 and multiple linear regression including ENST00000341421.7
To create a combined methylation score, principal component analysis (PCA) was performed on the methylation levels of the CG-sites that were highly correlated to FSIQ in cohort 2. The first principal component (PC1) was used as the combined methylation score, capturing the largest variance in the methylation data. Multiple linear regression analysis was then conducted using the combined methylation score and ENST00000341421.7 expression as predictors, and with FSIQ as the dependent variable.
Protein sequence alignment analysis
We obtained the amino acid sequence of full-length TRAK1 (Transcript ID: ENST00000327628.10) and TRAK1 isoform 686 (Transcript ID: ENST00000341421.7) from Ensembl [40]. The sequence alignment analysis was conducted using Clustal Omega (version 1.2.4, European Bioinformatics Institute, Hinxton, UK) with default parameters [41].
Statistical analyses
Two-sample comparisons were performed using Welch T-tests for parametric and Mann-Whitney U-tests for non-parametric data. Linear regressions were performed using Pearson correlations for parametric and Spearman correlations for non-parametric data. Normality was evaluated using Quantile-Quantile plots. Equality of variance between groups was tested with Levene’s test. P < 0.05 was used as the significant threshold. Data was presented as boxplots showing the median and spread (IQR). Graphs were drawn in R (R Foundation for Statistical Computing, Vienna, Austria) and BioRender (BioRender, Toronto, ON, Canada).
Results
Weighted gene correlation network analysis (WGCNA) revealed potential links between DNA methylation patterns and neurocognitive traits in KS
To investigate potential links between DNA methylation patterns and neurocognitive traits in KS, we performed WGCNA on methylation data from peripheral blood samples. DMPs between KS and male controls from cohort 1 were used as input. This initial analysis was restricted to males with KS from cohort 1 to evaluate the potential link between methylation patterns and neurocognitive traits within the KS group (Fig. 2A: Supplementary fig. 1: Table 1). Here, we examined correlations between the collective methylation status of co-methylated positions, termed modules, and neurocognitive traits. One module correlated with several neurocognitive traits (“white”), indicating that this set of CG-sites could be related to general neurocognitive function in KS. The strongest correlation was found to PIQ (r = 0.44, p = 3e–04). To further pinpoint which CG-sites within the “white” module had the strongest significance for PIQ, we calculated at the module-membership of the CG-sites within the “white” module in relation to significance for PIQ (Fig. 2B). Interestingly, five CG-sites within the TRAK1 gene had both high module-membership and high significance for PIQ. The five identified CG-sites within the TRAK1 gene from the “white” module were identified as cg01855070, cg03168947, cg08508763, cg08804892, and cg23715029. To further explore these five CG-sites from the “white” module, we analyzed the methylation levels at these sites in peripheral blood samples from cohort 1. Our analysis revealed that the five CG-sites were hypomethylated in males with KS compared to male controls (Fig. 3A).
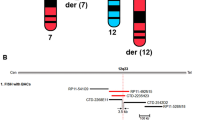
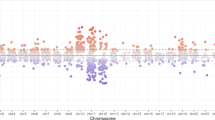
A WGCNA was performed on DNA methylation data from peripheral blood samples of individuals with KS (47,XXY) from cohort 1, to construct modules of CG-sites with similar methylation patterns and correlate these with neurocognitive traits. A Module-trait correlation plot. Color indicates correlation value, with negative correlations marked in green and positive correlations in orange. Numbers in each square indicate correlation values (top) and p-values (bottom, in parentheses). Only module-trait associations containing significant correlations are shown. The complete plot can be found in Supplementary fig. 1. B Module membership and trait significance. Scatterplot displaying the membership within the “white” module and significance for performance IQ (PIQ) for each CG-site within the “white” module. Dark points indicate X-chromosomal CG-sites whereas light points indicate autosomal CG-sites. Labels indicate UCSC gene name(s) for each CG-site, if available. Five CG-sites within the TRAK1 gene with high significance for PIQ are highlighted.
A ß-values for the five TRAK1 CG-sites from the “white” module, comparing methylation levels in peripheral blood samples from males with KS (XXY) and male controls (XY). B Correlation values between the five TRAK1 CG-sites from the “white” module and neurocognitive traits. Color indicates correlation value, with negative correlations marked in green and positive correlations in orange. C-G Correlations between the five TRAK1 CG-sites from the “white” module within peripheral blood samples of males with KS (XXY) and male controls (XY), and full scale IQ (FSIQ,C), performance IQ (PIQ, D), verbal IQ (VIQ, E), Trail Making B (TMT-B, F) and Wisconsin Card Sorting Test (WCST-trai, G).
Our WGCNA analysis did identify methylation patterns of CG-sites within other modules, exhibiting high module-membership and significance for neurocognitive traits (Supplementary Table 1). However, the correlation between other modules and neurocognitive traits were not as strong as for the “white” module. Therefore, we focused on the five CG-sites within the TRAK1 gene throughout the remainder of this article.
The methylation level at five CG-sites within TRAK1 correlated specifically with neurocognitive performance in KS, and not in male controls
To examine the relationship between the methylation level at the five identified CG-sites within the TRAK1 gene and the neurocognitive traits measured in males with KS, we performed additional correlation analyses (Fig. 3B-G: Supplementary fig. 2). The methylation levels at the five CG-sites were strongly correlated with the majority of neurocognitive traits in KS, with the strongest correlation observed for FSIQ (cg01855070: r = 0.45, p = 0.000160; cg03168947: r = 0.50, p = 2.1e–05; cg08508763: r = 0.63, p = 2e–08; cg08804892: r = 0.51, p = 1.7e–05; cg23715029: r = 0.54, p = 3.7e–06) and PIQ (cg01855070: r = 0.52, p = 8e–06; cg03168947: r = 0.49, p = 3.2e–05; cg08508763: r = 0.58, p = 4.6e–07; cg08804892: r = 0.53, p = 4.8e–06; cg23715029: r = 0.51, p = 1.73e–05). Positive correlations were observed for most neurocognitive traits, and negative correlations for TMT-A, TMT-B, TOL-tms, and WCST-trai. This was expected, since a lower score indicates better performance for these latter four variables. To determine whether the correlation between the methylation level at the five TRAK1 CG-sites and the neurocognitive traits was specific to KS, we also investigated the correlations between the CG-sites and the neurocognitive traits in male controls from cohort 1 (Fig. 3C-G: Supplementary fig. 2). No significant correlations were found in male controls, indicating that the correlation between the methylation levels at the five CG-sites within TRAK1 and neurocognitive performance is specific to KS.
The methylation pattern of the five CG-sites within TRAK1 in KS are consistent across cohorts
We next validated our findings from cohort 1 by assessing the methylation level at the five identified CG-sites within TRAK1 in peripheral blood samples from a second cohort (cohort 2). The analysis of peripheral blood samples from cohort 2 showed the same methylation pattern at the five CG-sites with hypomethylation in KS compared to male controls (Fig. 4A).
A ß-values for the five TRAK1 CG-sites from the “white” module, comparing methylation levels in peripheral blood samples from males with KS (XXY) and controls (XY) from cohort 2. B Expression of TRAK1 transcripts, comparing expression levels in peripheral blood samples from males with KS (XXY) and controls (XY) from cohort 2. C Scatter plot of ß-values for the five TRAK1 CG-sites from the “white” module and expression of ENST00000341421.7 (st-TRAK1) in peripheral blood samples from males with KS (XXY) and controls (XY) from cohort 2.
The expression level of a short TRAK1 transcript correlated negatively with the methylation level at the five CG-sites within TRAK1 in KS
To investigate the transcriptomic effect of the altered methylation signature at the five identified CG-sites within TRAK1, we analyzed TRAK1 expression in peripheral blood samples from cohort 2. No overall difference in TRAK1 expression was detected between KS and controls (Supplementary fig. 3). However, analysis of the different TRAK1 transcripts revealed that transcript ENST00000341421.7 (hereafter referred to as (short transcript TRAK1 (st-TRAK1)) was upregulated in KS compared to controls (Fig. 4B). No difference in the expression of other TRAK1 transcripts was observed between the two groups in cohort 2. We then examined the relationship between the expression of st-TRAK1 and the methylation level at the five identified CG-sites within TRAK1. Interestingly, the methylation level at these CG-sites was negatively correlated with the expression level of st-TRAK1 in KS, but not in controls (Fig. 4C). This agreed with the location of the five CG-sites, which are positioned upstream of the transcription start site (TSS) of st-TRAK1 (Supplementary fig. 4). Specifically, cg01855070, cg08508763, and cg08804892 were within 1500 bp upstream of the TSS (TSS1500) while cg03168947 and cg23715029 were within 200 bp upstream of TSS (TSS200). Thus, the five CG-sites may influence the promoter activity of st-TRAK1.
To further validate our findings from cohort 1, we investigated the correlation between the five CG-sites within TRAK1 and the neurocognitive traits in cohort 2 (Table 2). Despite the smaller number of individuals in cohort 2 compared to cohort 1, we found that these CG-sites correlated with several neurocognitive traits in KS (Supplementary fig. 5). The strongest correlations were again observed for FSIQ (cg01855070: r = 0.2, p = 0.37; cg03168947: r = 0.54, p = 0.0093; cg08508763: r = 0.4, p = 0.068; cg08804892: r = 0.48, p = 0.025; cg23715029: r = 0.52, p = 0.014) and PIQ (cg01855070: r = 0.27, p = 0.22; cg03168947: r = 0.48, p = 0.024; cg08508763: r = 0.4, p = 0.064; cg08804892: r = 0.57, p = 0.0055; cg23715029: r = 0.53 p = 0.012).
As described above, four of the five CG-sites within TRAK1 were highly correlated with FSIQ in cohort 2 (cg08508763, cg23715029, cg03168947, cg08804892). As gene expression data (RNAseq) was also present for all KS individuals in cohort 2, we created a combined methylation score for these four CG-sites to capture the overall methylation pattern. This was done using principal component analysis (PCA). Component 1 (PC1) was used as the combined methylation score and was then included in a linear multiple regression model along with st-TRAK1 expression to assess their joint effect on FSIQ. The model was significant (p = 0.014), indicating that the combined predictors significantly contribute to explaining the variation in FSIQ. Similarly, when PIQ and VIQ were tested as outcome variables in our multiple linear regression analysis, significant associations were observed for both PIQ (p = 0.048) and VIQ (p = 0.045). Furthermore, the combined model indicates that while st-TRAK1 alone is not a strong predictor of FSIQ, its interaction with the combined methylation pattern might be relevant for cognitive outcomes.
DNA methylation and neurocognitive functions were stable over time
Eleven of the males with KS were included in both cohort 1 and cohort 2 and therefore had blood samples taken for DNA methylation analysis and underwent cognitive testing at two different time points. This enabled us to investigate whether the methylation levels measured by two different arrays (Infinium HumanMethylation450k and Infinium MethylationEPIC) were consistent and stable over time. Our data revealed a high degree of concordance across the five CG-sites (Supplementary fig. 6A). Furthermore, neurocognitive performance also revealed a high level of concordance over time (Supplementary fig. 6B).
Hypomethylation at the five CG-sites within TRAK1 and upregulation of the short TRAK1 transcript observed in 47,XXY neural precursor cells
In the neurocognitive phenotype of KS, the brain is the primary target organ of interest. Due to the inaccessibility of brain samples from males with KS, we used neural precursor cells differentiated from human induced pluripotent stem cells derived from amniotic fluid cells with either a 47,XXY or a 46,XY karyotype. This confirmed that the five CG-sites within TRAK1 were hypomethylated in neural cells with a 47,XXY karyotype compared to neural cells with a 46,XY karyotype (Fig. 5A). Additionally, we investigated the expression level of the st-TRAK1 in the neural precursor cells. In 47,XXY cells, st-TRAK1 was highly upregulated compared to 46,XY cells (Fig. 5B). No difference in the expression of other TRAK1 transcripts was observed (Supplementary fig. 7). Next, we analyzed the relation between the methylation level at the five CG-sites within TRAK1 and the expression level of st-TRAK1. For all CG-sites, we found a clear pattern between hypomethylation and higher expression levels (Fig. 5C). Thus, the differences observed in blood cells were also observed in neuronal cells.
A ß-values for the five TRAK1 CG-sites from the “white” module, comparing methylation levels in XXY and XY neural precursors. B Expression of ENST00000341421.7 (st-TRAK1) in XXY and XY neural precursors. C Scatter plot of ß-values for the five TRAK1 CG-sites from the “white” module and expression of ENST00000341421.7 (st-TRAK1) in XXY and XY neural precursors. Solid and dotted lines indicate the range of methylation and expression, respectively, within the karyotype groups.
The short TRAK1 transcript encodes a 686-amino acid isoform of TRAK1 that lacks segments from both the N-terminal and C-terminal regions compared to full-length TRAK1
The TRAK1 gene encodes trafficking kinesin protein 1 (TRAK1), a 953-amino acid long protein [42]. St-TRAK1 encodes a 686-amino acid isoform of TRAK1 (hereafter referred to as TRAK1 isoform 686) [40]. To compare TRAK1 isoform 686 with full-length TRAK1, we performed a protein sequence alignment analysis. The analysis showed that most of the protein sequence was identical except that TRAK1 isoform 686 is missing segments from both the N-terminal and C-terminal (Supplementary fig. 8), suggesting that the full-length TRAK1 and TRAK1 isoform 686 may have functional differences.
Discussion
Here, we present an integrated analysis of the neurocognitive profile, the methylome, and the transcriptome in peripheral blood samples and neuronal precursor cells from males with KS and male controls. We identified correlations, exclusively in KS, between the methylation level at five specific CG-sites (cg01855070, cg03168947, cg08508763, cg08804892, cg23715029) within TRAK1 and several neurocognitive traits, including FSIQ, VIQ, and PIQ. Furthermore, we demonstrated a significant correlation between the expression level of st-TRAK1 and the methylation level at the five CG-sites within TRAK1 in KS. The five CG-sites within TRAK1 are located upstream of the TSS of the st-TRAK1, suggesting that they control its expression. In neural precursor cells derived from 47,XXY amniotic cells, we similarly observed hypomethylation at these CG-sites and highly increased expression of st-TRAK1 compared to control cells. To our knowledge, this is the first study linking a specific methylation signature to the neurocognitive phenotype in KS.
The TRAK1 gene, mapping to 3p22.1, is widely expressed, including in the fetal and adult brain [43]. It encodes trafficking kinesin protein 1 (TRAK1), that is a 953-amino acid long protein with a molecular mass of 106 kDa (Fig. 6A) [42]. TRAK1 variants and single nucleotide variants have been associated with a wide range of neurological and neuropsychological disorders/conditions including schizophrenia [44], autism spectrum disorders [45,46,47], trigeminal neuralgia [48], encephalopathy [49], seizures, epilepsy, developmental delay [47, 49,50,51,52,53,54], as well as anatomical brain alterations, including brain atrophy [49,50,51,52]. The broad spectrum of neurological and neuropsychological disorders/conditions linked to variants in TRAK1 suggests that TRAK1 is crucial for normal neuronal cell function. Recently, an association between the methylation level at one CG-site near the TRAK1 gene (not one of the five CG-sites found in this study) and the cognitive function in monozygotic twins with normal karyotype was demonstrated [55], further supporting that altered methylation at specific CG-site in relation to TRAK1 might be related to neurocognitive performance.
A Diagram of the full-length TRAK1 protein with important regions for mitochondrial transport. The kinesin-1 binding region (residues within 1–360) is shown in green, the dynein/dynactin binding region (118–392 residues) is depicted with stripes, the MIRO1 binding region (394–431 residues) is shown in light blue, and the microtubules binding region (636–953 residues) is shown in dark blue. B Diagram of the TRAK1 isoform 686 protein. Sequence alignment analysis has shown that TRAK1 isoform 686 contains the dynein/dynactin binding region (60–334 residues; striped), the MIRO1 binding region (336–373 residues; light blue), and only part of the binding region for kinesin-1 (38–302 residues; green) and microtubules (578–596 residues; dark blue). Dotted lines demonstrate where the sequence of the full-length TRAK1 and the TRAK1 isoform 686 is identical. Parts of the C-terminal and N-terminal region of TRAK isoform 686 (shown in orange) differ from the full-length. This figure was made using BioRender.com.
Several studies have demonstrated that TRAK1 plays an essential role in mitochondrial mobility. In neurons, the TRAK1 protein is abundantly localized in axons to facilitate the axonal transport of mitochondria along microtubules by acting as an adaptor protein that links mitochondria to kinesin-1 (for anterograde transport) and dynein/dynactin (for retrograde transport) [56]. Knockdown of TRAK1 in neurons significantly reduces the mitochondrial motility in the axons, highlighting its importance for proper axonal transport of mitochondria [56, 57]. Moreover, the mitochondrial distribution and motility are disrupted in cells carrying a pathogenic TRAK1 variant [49]. In neurons, mitochondria are crucial for growth, survival, and function, including initiation of action potentials and synaptic transmission, by producing ATP and buffering intracellular Ca2+ [58]. Mitochondrial transport within neurons is not only responsible for distributing mitochondria to distal regions like synapses to meet local energy demands but also for removing and replacing aged and dysfunctional mitochondria with healthy ones [58]. Defects in mitochondrial transport are linked to several neurodegenerative diseases including Alzheimer’s disease [59].
The N-terminal domain of TRAK1 is essential for both the kinesin-1 and dynein/dynactin-dependent transport (Fig. 6A) [56, 60]. Based on comparisons with the cryo-EM structures of other adaptors that interact with dynein/dynactin, it is predicted that the region of TRAK1 implicated in binding dynein/dynactin spans amino acids 118 to 392 [61]. This region of TRAK1 contains the CC1 box and the Spindly motif that bind dynein light intermediate chain and the pointed-end complex of dynactin, respectively [60, 62, 63]. The exact binding site of TRAK1 for kinesin-1 is unknown but appears to involve residues within 1–360 [56, 60, 64]. Co-immunoprecipitation experiments further revealed that residues 100–360 of TRAK1 could efficiently bind to kinesin-1 [56], while a more recent study using crosslinking mass spectrometry demonstrated that residues 23–138 were essential for the interaction of TRAK1 with kinesin-1 [64]. TRAK1 interacts with the mitochondria by binding MIRO proteins anchored to the mitochondrial outer membrane [65]. Just like TRAK1, MIRO proteins are crucial for proper mitochondrial transport, with MIRO1 identified as the main isoform involved in mitochondria trafficking in neurons [66]. Baltrusaitis et al. demonstrated that TRAK1 residues 394–431 are sufficient for binding MIRO1 [61]. The C-terminal region of TRAK1 (residues 636–953) also plays a role in mitochondrial trafficking by interacting with microtubules [67], which increases the processivity of the TRAK1-kinesin-1 complex, as shown by increased run length and microtubule interaction time of the complex for full-length TRAK1 compared to a TRAK1 mutant lacking its C-terminal [67].
In the present study, we demonstrated that most of the protein sequence of full-length TRAK1 and TRAK1 isoform 686 is identical. However, TRAK1 isoform 686 lacks segments from both the N-terminal and C-terminal (Fig. 6). Based on our protein sequence alignment analysis, TRAK1 isoform 686 contains binding sites for dynein/dynactin and MIRO1, but only part of the binding site for kinesin-1, suggesting that its affinity for kinesin-1 may be reduced compared to full-length TRAK1 (Fig. 6B). Additionally, its truncated C-terminal may prevent or diminish its microtubule binding, leading to reduced processivity of the TRAK1-kinesin-1 complex, according to Henrichs et al [67]. Consequently, we hypothesize that TRAK1 isoform 686 may be less effective at transporting mitochondria compared to full-length TRAK1. The observed upregulation of st-TRAK1 in KS indicates a disruption in the balance between full-length TRAK1 and TRAK1 isoform 686, potentially leading to impaired mitochondrial distribution and motility in neurons in KS. This disruption may contribute to the neurocognitive phenotype observed in KS, although experimental validation will be necessary. Future experiments should investigate the impact of st-TRAK1 upregulation on mitochondria trafficking in KS and examine its effect on cognitive functions including memory in animal models.
Besides being involved in mitochondria trafficking, TRAK1 has been implicated in other biological processes, including regulation of endosomal trafficking [68] and the endocytic trafficking of the gamma-aminobutyric acid (GABA)-A receptor [69]. However, it is unclear whether these processes are affected by hypomethylation at the five CG-sites within TRAK1 and upregulation of st-TRAK1 in KS. TRAK1 isoform 686 could also be involved in additional functions in neurons as well as in other cell types. Thus, the link between TRAK1 and the neurocognitive profile in KS may be influenced by factors beyond its role in mitochondrial transport.
Our findings demonstrate a strong correlation between the “white” module and general cognitive measures such as FSIQ and PIQ. While these measures are broad and not the most specific neurocognitive traits associated with KS, prior research has established that individuals with KS consistently exhibit FSIQ scores approximately 10 points lower than controls [5, 6, 8,9,10,11], underscoring its relevance as a marker of cognitive differences. Notably, we also observed significant correlations between the “white” module and neurocognitive traits more characteristic of KS, such as verbal performance, and verbal memory and learning. These results suggest that the methylation pattern of CG-sites in the “white” module is implicated not only in general cognitive performance, but also in cognitive domains more directly associated with the KS phenotype. Overall, these findings highlight the potential role of TRAK1 in shaping both KS-specific and general cognitive outcomes, offering deeper insights into the genotype-phenotype correlations in this condition.
The relationship between hypomethylation at the TRAK1 sites and the presence of an additional X chromosome in XXY remains unclear. However, our findings are consistent with our prior studies and the broader literature in that while hypermethylation is the predominant epigenetic signature observed in XXY, instances of hypomethylation are also evident [15]. Hypomethylation at specific loci, including TRAK1, may represent a nuanced and locus-specific response to the additional X chromosome. Such responses could arise from factors such as the disruption of chromatin remodeling, altered transcription factor binding, or differential regulatory influences associated with having an additional X chromosome. Moreover, while TRAK1 hypomethylation may appear contradictory to the global hypermethylation trend, it may highlight a unique regulatory mechanism at this locus. The coexistence of hyper- and hypomethylated sites reflects the complexity of epigenetic regulation in XXY. Further research, including transcriptomic and chromatin accessibility studies, is warranted to elucidate the X-linked biological drivers underlying these specific methylation changes and their functional implications.
Our findings highlight methylation at an autosomal gene, TRAK1, as the primary correlate of neurocognitive outcomes in KS, despite the presence of five X-chromosomal CG-sites within the “white” module, three of which localize within known genes (CXorf23/BCLAF3, SH2D1A, KLHL13). The relative paucity of module-specific methylation on X linked genes underscores the unexpected prominence of an autosomal in this context. This raises the possibility that TRAK1 may be subject to indirect regulation by X chromosome-encoded transcription factors or that epigenetic modifiers could be at play. Prior studies, including our own, have demonstrated that X chromosome dosage effects induce widespread transcriptomic and epigenomic alterations across the genome, suggesting that X linked regulatory elements may exert transacting effects on autosomal genes [13,14,15, 20,21,22]. However, whether such mechanisms contribute to TRAK1 regulation remains unclear. Additionally, given the well-documented endocrine alterations in KS (reviewed in [2]), it is plausible that TRAK1 expression and methylation may be influenced by perturbed androgen signaling, potentially through androgen receptor binding sites or other hormone-responsive elements. While no direct link between testosterone-related pathways and the TRAK1 promoter region has been established, investigating such interactions may provide novel insights into the molecular underpinnings of KS-associated neurocognitive phenotypes. These findings collectively reinforce the broader relevance of our work within the framework of sex chromosome aneuploidy and highlight critical avenues for future research into the interplay between X-linked regulatory factors, hormonal signaling, and autosomal gene expression.
We demonstrate that in KS, DNA hypomethylation at specific CG-sites within TRAK1 and increased expression of st-TRAK1 are associated with specific neurocognitive phenotypes, including lower FSIQ, VIQ and PIQ, indicating a potential epigenetic basis for the neurocognitive impairments observed in KS. These specific CG-sites may be useful as epigenetic biomarkers for determining the level of neurocognitive function in KS. However, further validation using functional assays will be necessary to clarify their impact on the neurocognitive traits in KS.
Data access
Human data used in this study has previously been published and is available from the European Genome-phenome Archive (EGA) under the accession number EGAS00001002797 (cohort 1), EGAS00001006996 and EGAS00001007020 (cohort 2). Data from neural precursor cells are available upon request.
References
Bojesen A, Juul S, Gravholt CH. Prenatal and postnatal prevalence of Klinefelter syndrome: a national registry study. J Clin Endocrinol Metab. 2003;88:622–6.
Gravholt CH, Chang S, Wallentin M, Fedder J, Moore P, Skakkebæk A. Klinefelter syndrome: integrating genetics, neuropsychology, and endocrinology. Endocr Rev. 2018;39:389–423.
Leonard MF, Schowalter JE, Landy G, Ruddle FH, Lubs HA. Chromosomal abnormalities in the New Haven newborn study: a prospective study of development of children with sex chromosome anomalies. Birth Defects Orig Artic Ser. 1979;15:115–59.
Ratcliffe SG. Speech and learning disorders in children with sex chromosome abnormalities. Dev Med Child Neurol. 1982;24:80–4.
Fjermestad KW, Huster R, Thunberg C, Stokke S, Gravholt CH, Solbakk AK. Neuropsychological functions, sleep, and mental health in adults with Klinefelter syndrome. Am J Med Genet C Semin Med Genet. 2020;184:482–92.
Skakkebæk A, Gravholt CH, Rasmussen PM, Bojesen A, Jensen JS, Fedder J, et al. Neuroanatomical correlates of Klinefelter syndrome studied in relation to the neuropsychological profile. Neuroimage Clin. 2014;4:1–9.
Kompus K, Westerhausen R, Nilsson LG, Hugdahl K, Jongstra S, Berglund A, et al. Deficits in inhibitory executive functions in Klinefelter (47, XXY) syndrome. Psychiatry Res. 2011;189:135–40.
van Rijn S, Swaab H. Executive dysfunction and the relation with behavioral problems in children with 47,XXY and 47,XXX. Genes Brain Behav. 2015;14:200–8.
Bender BG, Puck MH, Salbenblatt JA, Robinson A. Dyslexia in 47,XXXY boys identified at birth. Behav Genet. 1986;16:343–54.
Ratcliffe SG, Masera N, Pan H, McKie M. Head circumference and IQ of children with sex chromosome abnormalities. Dev Med Child Neurol. 1994;36:533–44.
Netley C, Rovet J. Verbal deficits in children with 47,XXY and 47,XXX karyotypes: a descriptive and experimental study. Brain Lang. 1982;17:58–72.
Björlin Avdic H, Kleberg JL, van der Poll M, Frisén L, Hutley M, Sarjanen M, et al. Cognitive profile in adult women with turner syndrome: IQ split and associations with ADHD and ASD. Cogn Neuropsychiatry. 2023;28:207–25.
Sharma A, Jamil MA, Nuesgen N, Schreiner F, Priebe L, Hoffmann P, et al. DNA methylation signature in peripheral blood reveals distinct characteristics of human X chromosome numerical aberrations. Clin Epigenetics. 2015;7:76.
Skakkebæk A, Nielsen MM, Trolle C, Vang S, Hornshøj H, Hedegaard J, et al. DNA hypermethylation and differential gene expression associated with Klinefelter syndrome. Sci Rep. 2018;8:13740.
Viuff M, Skakkebæk A, Johannsen EB, Chang S, Pedersen SB, Lauritsen KM, et al. X chromosome dosage and the genetic impact across human tissues. Genome Med. 2023;15:21.
Halder R, Hennion M, Vidal RO, Shomroni O, Rahman RU, Rajput A, et al. DNA methylation changes in plasticity genes accompany the formation and maintenance of memory. Nat Neurosci. 2016;19:102–10.
Zocher S, Overall RW, Berdugo-Vega G, Rund N, Karasinsky A, Adusumilli VS, et al. De novo DNA methylation controls neuronal maturation during adult hippocampal neurogenesis. Embo j. 2021;40:e107100.
Wu H, Coskun V, Tao J, Xie W, Ge W, Yoshikawa K, et al. Dnmt3a-dependent nonpromoter DNA methylation facilitates transcription of neurogenic genes. Science. 2010;329:444–8.
Marioni RE, McRae AF, Bressler J, Colicino E, Hannon E, Li S, et al. Meta-analysis of epigenome-wide association studies of cognitive abilities. Mol Psychiatry. 2018;23:2133–44.
Viana J, Pidsley R, Troakes C, Spiers H, Wong CC, Al-Sarraj S, et al. Epigenomic and transcriptomic signatures of a Klinefelter syndrome (47,XXY) karyotype in the brain. Epigenetics. 2014;9:587–99.
Wan ES, Qiu W, Morrow J, Beaty TH, Hetmanski J, Make BJ, et al. Genome-wide site-specific differential methylation in the blood of individuals with Klinefelter syndrome. Mol Reprod Dev. 2015;82:377–86.
Zhang X, Hong D, Ma S, Ward T, Ho M, Pattni R, et al. Integrated functional genomic analyses of Klinefelter and turner syndromes reveal global network effects of altered X chromosome dosage. Proc Natl Acad Sci USA. 2020;117:4864–73.
Reitan RM. Validity of the trail making test as an indicator of organic brain damage. Percept Mot Skills. 1958;8:271–6.
Wechsler D Wechsler Adult Intelligence Scale, Third edition. San Antonio, TX:The Psychological Corporation;1997.
Heaton RK, Chelune GJ, Talley JL, Kay GG, Curtiss G Wisconsin Card Sorting Test Manual, Revised and Expanded. Odessa, FL; Psychological Assessment Resources, Inc, p 1993.
Culbertson WC, Zillmer EA. Tower of London — Drexel University. 2nd Edition. NY: Technical Manual Multi-Health Systems Inc; 2005.
Wechsler D Wechsler adult intelligence scale – Fourth Edition (4th ed.). Psychological Corporation. 2008.
Wechsler D Wechsler memory scale – Third edition manual. Psychological Corporation. 1997.
Fortin JP, Triche TJ Jr., Hansen KD. Preprocessing, normalization and integration of the Illumina HumanMethylationEPIC array with minfi. Bioinformatics. 2017;33:558–60.
Fortin JP, Labbe A, Lemire M, Zanke BW, Hudson TJ, Fertig EJ, et al. Functional normalization of 450k methylation array data improves replication in large cancer studies. Genome Biol. 2014;15:503.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47.
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C. Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods. 2017;14:417–9.
Love MI, Soneson C, Hickey PF, Johnson LK, Pierce NT, Shepherd L, et al. Tximeta: Reference sequence checksums for provenance identification in RNA-seq. PLoS Comput Biol. 2020;16:e1007664.
Okita K, Matsumura Y, Sato Y, Okada A, Morizane A, Okamoto S, et al. A more efficient method to generate integration-free human iPS cells. Nat Methods. 2011;8:409–12.
Knupp A, Mishra S, Martinez R, Braggin JE, Szabo M, Kinoshita C, et al. Depletion of the AD risk gene SORL1 selectively impairs neuronal endosomal traffic independent of amyloidogenic APP processing. Cell Rep. 2020;31:107719.
Mishra S, Knupp A, Szabo MP, Williams CA, Kinoshita C, Hailey DW, et al. The Alzheimer’s gene SORL1 is a regulator of endosomal traffic and recycling in human neurons. Cell Mol Life Sci. 2022;79:162.
Triche TJ Jr., Weisenberger DJ, Van Den Berg D, Laird PW, Siegmund KD. Low-level processing of Illumina Infinium DNA methylation beadarrays. Nucleic Acids Res. 2013;41:e90.
Wang Y, Gorrie-Stone TJ, Grant OA, Andrayas AD, Zhai X, McDonald-Maier KD, et al. InterpolatedXY: a two-step strategy to normalize DNA methylation microarray data avoiding sex bias. Bioinformatics. 2022;38:3950–7.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinforma. 2008;9:559.
Harrison PW, Amode MR, Austine-Orimoloye O, Azov AG, Barba M, Barnes I, et al. Ensembl 2024. Nucleic Acids Res. 2024;52:D891–d9.
Madeira F, Pearce M, Tivey ARN, Basutkar P, Lee J, Edbali O, et al. Search and sequence analysis tools services from EMBL-EBI in 2022. Nucleic Acids Res. 2022;50:W276–w9.
Iyer SP, Akimoto Y, Hart GW. Identification and cloning of a novel family of coiled-coil domain proteins that interact with O-GlcNAc transferase. J Biol Chem. 2003;278:5399–409.
Kikuno R, Nagase T, Ishikawa K, Hirosawa M, Miyajima N, Tanaka A, et al. Prediction of the coding sequences of unidentified human genes. XIV. The complete sequences of 100 new cDNA clones from brain which code for large proteins in vitro. DNA Res. 1999;6:197–205.
Xu B, Roos JL, Dexheimer P, Boone B, Plummer B, Levy S, et al. Exome sequencing supports a de novo mutational paradigm for schizophrenia. Nat Genet. 2011;43:864–8.
Iossifov I, O’Roak BJ, Sanders SJ, Ronemus M, Krumm N, Levy D, et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature. 2014;515:216–21.
Bacchelli E, Cameli C, Viggiano M, Igliozzi R, Mancini A, Tancredi R, et al. An integrated analysis of rare CNV and exome variation in autism spectrum disorder using the infinium psycharray. Sci Rep. 2020;10:3198.
Turner TN, Wilfert AB, Bakken TE, Bernier RA, Pepper MR, Zhang Z, et al. Sex-based analysis of de novo variants in neurodevelopmental disorders. Am J Hum Genet. 2019;105:1274–85.
Dong W, Jin SC, Allocco A, Zeng X, Sheth AH, Panchagnula S, et al. Exome sequencing implicates impaired GABA signaling and neuronal ion transport in trigeminal neuralgia. iScience. 2020;23:101552.
Barel O, Malicdan MCV, Ben-Zeev B, Kandel J, Pri-Chen H, Stephen J, et al. Deleterious variants in TRAK1 disrupt mitochondrial movement and cause fatal encephalopathy. Brain. 2017;140:568–81.
Li RK, Xiong YR, Pan SJ, Lei WT, Shu XM, Shi XQ, et al. Role of TRAK1 variants in epilepsy: genotype-phenotype analysis in a pediatric case of epilepsy with developmental disorder. Front Mol Neurosci. 2024;17:1342371.
Anazi S, Maddirevula S, Salpietro V, Asi YT, Alsahli S, Alhashem A, et al. Expanding the genetic heterogeneity of intellectual disability. Hum Genet. 2017;136:1419–29.
Sagie S, Lerman-Sagie T, Maljevic S, Yosovich K, Detert K, Chung SK, et al. Expanding the phenotype of TRAK1 mutations: hyperekplexia and refractory status epilepticus. Brain. 2018;141:e55.
Mitani T, Isikay S, Gezdirici A, Gulec EY, Punetha J, Fatih JM, et al. High prevalence of multilocus pathogenic variation in neurodevelopmental disorders in the Turkish population. Am J Hum Genet. 2021;108:1981–2005.
Deciphering Developmental Disorders Study. Prevalence and architecture of de novo mutations in developmental disorders. Nature. 2017;542:433-8.
Liu J, Wang W, Luo J, Duan H, Xu C, Tian X, et al. Mediation role of DNA methylation in association between handgrip strength and cognitive function in monozygotic twins. J Hum Genet. 2024;69:357–63.
van Spronsen M, Mikhaylova M, Lipka J, Schlager MA, van den Heuvel DJ, Kuijpers M, et al. TRAK/Milton motor-adaptor proteins steer mitochondrial trafficking to axons and dendrites. Neuron. 2013;77:485–502.
Brickley K, Stephenson FA. Trafficking kinesin protein (TRAK)-mediated transport of mitochondria in axons of hippocampal neurons. J Biol Chem. 2011;286:18079–92.
Sheng ZH. Mitochondrial trafficking and anchoring in neurons: new insight and implications. J Cell Biol. 2014;204:1087–98.
Sheng ZH, Cai Q. Mitochondrial transport in neurons: impact on synaptic homeostasis and neurodegeneration. Nat Rev Neurosci. 2012;13:77–93.
Canty JT, Hensley A, Aslan M, Jack A, Yildiz A. TRAK adaptors regulate the recruitment and activation of dynein and kinesin in mitochondrial transport. Nat Commun. 2023;14:1376.
Baltrusaitis EE, Ravitch EE, Fenton AR, Perez TA, Holzbaur ELF, Dominguez R. Interaction between the mitochondrial adaptor MIRO and the motor adaptor TRAK. J Biol Chem. 2023;299:105441.
Gama JB, Pereira C, Simões PA, Celestino R, Reis RM, Barbosa DJ, et al. Molecular mechanism of dynein recruitment to kinetochores by the Rod-Zw10-Zwilch complex and Spindly. J Cell Biol. 2017;216:943–60.
Celestino R, Henen MA, Gama JB, Carvalho C, McCabe M, Barbosa DJ, et al. A transient helix in the disordered region of dynein light intermediate chain links the motor to structurally diverse adaptors for cargo transport. PLoS Biol. 2019;17:e3000100.
Tan Z, Yue Y, Leprevost F, Haynes S, Basrur V, Nesvizhskii AI, et al. Autoinhibited kinesin-1 adopts a hierarchical folding pattern. Elife. 2023;12:RP86776 https://doi.org/10.7554/eLife.86776
Fransson S, Ruusala A, Aspenström P. The atypical Rho GTPases Miro-1 and Miro-2 have essential roles in mitochondrial trafficking. Biochem Biophys Res Commun. 2006;344:500–10.
López-Doménech G, Higgs NF, Vaccaro V, Roš H, Arancibia-Cárcamo IL, MacAskill AF, et al. Loss of dendritic complexity precedes neurodegeneration in a mouse model with disrupted mitochondrial distribution in mature dendrites. Cell Rep. 2016;17:317–27.
Henrichs V, Grycova L, Barinka C, Nahacka Z, Neuzil J, Diez S, et al. Mitochondria-adaptor TRAK1 promotes kinesin-1 driven transport in crowded environments. Nat Commun. 2020;11:3123.
Webber E, Li L, Chin LS. Hypertonia-associated protein trak1 is a novel regulator of endosome-to-lysosome trafficking. J Mol Biol. 2008;382:638–51.
Gilbert SL, Zhang L, Forster ML, Anderson JR, Iwase T, Soliven B, et al. Trak1 mutation disrupts GABA(A) receptor homeostasis in hypertonic mice. Nat Genet. 2006;38:245–50.
Acknowledgements
We thank Lone Kvist and Pamela Celis for their technical assistance. Some of the computing for this project was performed on the GenomeDK cluster. We would like to thank GenomeDK and Aarhus University for providing computational resources and support that contributed to these research results.
Funding
HBLT and CHG were supported by Novo Nordisk Foundation (NNF15OC0016474, NNF20OC0060610), Independent Research Fund Denmark (2096-00165 A), “Fonden til lægevidenskabens fremme” and “sygesikringen danmark” (2022-0189). EBH was supported by Aarhus University and the Independent Research Fund Denmark (2096-00165 A). SC was supported by University of Southern Denmark and The Region of Southern Denmark. CDRB was supported by Danish Cancer Society (Grants #R296-A16987 and #R312-A18151) and Independent Research Fund in Denmark (#0132-00059B). CD, XD, GF, JB were supported by NIH grants (AG073918; GM131745). AS was supported by the Independent Research Fund Denmark (0134-00130B). Data analysis was done in part at the University of Washington Interdisciplinary Center for Exposures, Diseases, Genomics and Environment (Grant: P30ES007033). Open access funding provided by Aarhus University Hospital.
Author information
Authors and Affiliations
Contributions
XD, JBB, GNF, CHG, CMD, JJ and AS designed research. GNF, JBB, XD, SC and AS performed research. HBLT, EBH, JWM, TKB, CDRB, JJ and AS analyzed data. HBLT, EBH, JBB, GNF, JWM, TKB, CHG, CMD, JJ and AS wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tallaksen, H.B.L., Hasselholm, E.B., Berletch, J.B. et al. DNA hypomethylation at specific CG-sites within TRAK1 is linked to the neurocognitive profile in Klinefelter syndrome. Mol Psychiatry 31, 1409–1419 (2026). https://doi.org/10.1038/s41380-025-03254-z
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41380-025-03254-z