Abstract
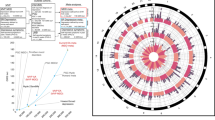
The neuropeptides oxytocin and vasotocin are predominantly produced in the supraoptic and paraventricular nuclei of the anterior-inferior, anterior-superior and tubular-superior hypothalamic subunits. Evidence suggests that oxytocin and vasotocin signaling play a role in both physiology and behavior, and that dysfunction of these signaling systems may contribute to the co-occurrence of metabolic and psychiatric conditions. The genetic pathways, however, that may underlie the connection between these physiological and behavioral traits are yet to be clearly delineated. We deployed bivariate mixture models and conjunctional FDR to estimate the global and local genetic overlap between three oxytocinergic-vasotocinergic hypothalamus subunits and ten psychiatric and metabolic traits related to oxytocin and vasotocin signaling. We show that these three subunits share moderate-to-extensive genetic overlap with the tested traits, therein stronger overlap with psychiatric than metabolic traits. We found most complete overlap between the anterior subunits and systolic blood pressure. Across all subunit and trait combinations, we pinpoint 95 novel, unique associated loci. The genes associated with these loci were enriched in gene sets linked to neuroimaging and neurodegeneration as well as metabolic markers, and were up-/down-regulated in tissues such as blood vessel and the liver. These findings help shed light on the genetic architecture of the hypothalamic subunits implicated in oxytocin and vasotocin and selected traits, and provide new avenues for future research.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The external GWAS summary statistics data is partially publicly available for download at the consortium’s official websites (PGC: https://pgc.unc.edu/, GIANT consortium: https://portals.broadinstitute.org/collaboration/giant/index.php/GIANT_consortium, DIAGRAM consortium: https://diagram-consortium.org/index.html). The internally generated GWAS summary statistics data for the three selected hypothalamus subunits are available at https://osf.io/k46gu/.
Code availability
Supplementary information, scripts used for analyses with information on specific parameter settings, and additional notes are available at https://osf.io/k46gu/.
References
Leng G, Leng RI. Oxytocin: A citation network analysis of 10 000 papers. J Neuroendocrinol. 2021;33:e13014.
Kenkel WM. Corpus colossal: A bibliometric analysis of neuroscience abstracts and impact factor. Front Integr Neurosci. 2019;13:18.
Howarth G, Botha DJ. Amniotomy plus intravenous oxytocin for induction of labour. Cochrane Database Syst Rev. 2001;2001:CD003250.
Ruis H, Rolland R, Doesburg W, Broeders G, Corbey R. Oxytocin enhances onset of lactation among mothers delivering prematurely. BMJ. 1981;283:340–2.
Bradley ER, Woolley JD. Oxytocin effects in schizophrenia: Reconciling mixed findings and moving forward. Neurosci Biobehav Rev. 2017;80:36–56.
Jurek B, Neumann ID. The oxytocin receptor: From intracellular signaling to behavior. Physiol Rev. 2018;98:1805–908.
Winterton A, Westlye LT, Steen NE, Andreassen OA, Quintana DS. Improving the precision of intranasal oxytocin research. Nat Hum Behav. 2021;5:9–18.
Vancampfort D, Stubbs B, Mitchell AJ, De Hert M, Wampers M, Ward PB, et al. Risk of metabolic syndrome and its components in people with schizophrenia and related psychotic disorders, bipolar disorder and major depressive disorder: a systematic review and meta-analysis. World Psychiatry. 2015;14:339–47.
Birkenaes AB, Opjordsmoen S, Brunborg C, Engh JA, Jonsdottir H, Ringen PA, et al. The level of cardiovascular risk factors in bipolar disorder equals that of schizophrenia: A comparative study. J Clin Psychiatry. 2007;68:10292.
Venkatasubramanian G, Chittiprol S, Neelakantachar N, Naveen MN, Thirthall J, Gangadhar BN, et al. Insulin and insulin-like growth factor-1 abnormalities in antipsychotic-naive schizophrenia. Am J Psychiatry. 2007;164:1557–60.
Winterton A, Bettella F, de Lange A-MG, Haram M, Steen NE, Westlye LT, et al. Oxytocin-pathway polygenic scores for severe mental disorders and metabolic phenotypes in the UK Biobank. Transl Psychiatry. 2021;11:1–9.
Theofanopoulou C, Gedman G, Cahill JA, Boeckx C, Jarvis ED. Universal nomenclature for oxytocin–vasotocin ligand and receptor families. Nature. 2021;592:747–55.
Bernstein H-G, Dobrowolny H, Bogerts B, Keilhoff G, Steiner J. The hypothalamus and neuropsychiatric disorders: Psychiatry meets microscopy. Cell Tissue Res. 2019;375:243–58.
Ivell R. Biosynthesis of oxytocin in the brain and peripheral organs. In: Ganten D, Pfaff D, editors. Neurobiology of Oxytocin, Berlin, Heidelberg: Springer; 1986. p. 1–18.
Leng G. The Heart of the Brain: The Hypothalamus and Its Hormones. MIT Press; 2018.
Buijs RM, Soto Tinoco EC, Hurtado Alvarado G, Escobar C. The circadian system: From clocks to physiology. In: Swaab DF, Kreier F, Lucassen PJ, Salehi A, Buijs RM, editors. Handbook of Clinical Neurology, Vol. 179, Elsevier; 2021. p. 233-47.
Althammer F, Grinevich V. Diversity of oxytocin neurones: Beyond magno- and parvocellular cell types? J Neuroendocrinol. 2018;30:e12549.
Kovács KJ. CRH: The link between hormonal-, metabolic- and behavioral responses to stress. J Chem Neuroanat. 2013;54:25–33.
Fekete C, Légrádi G, Mihály E, Huang Q-H, Tatro JB, Rand WM, et al. α-melanocyte-stimulating hormone is contained in nerve terminals innervating thyrotropin-releasing hormone-synthesizing neurons in the hypothalamic paraventricular nucleus and prevents fasting-induced suppression of prothyrotropin-releasing hormone gene expression. J Neurosci. 2000;20:1550–8.
Gwee P-C, Tay B-H, Brenner S, Venkatesh B. Characterization of the neurohypophysial hormone gene loci in elephant shark and the Japanese lamprey: Origin of the vertebrate neurohypophysial hormone genes. BMC Evol Biol. 2009;9:47.
Sartorius AM, Rokicki J, Birkeland S, Bettella F, Barth C, de Lange A-M G, et al. An evolutionary timeline of the oxytocin signaling pathway. Commun Biol. 2024;7:1–13.
Gruber CW. Physiology of invertebrate oxytocin and vasopressin neuropeptides. Exp Physiol. 2014;99:55–61.
Richter D. Oxytocin and vasopressin genes. In: Habener JF, editor. Molecular Cloning of Hormone Genes, Totowa, NJ: Humana Press; 1987. p. 173–206.
Bankir L, Bichet DG, Morgenthaler NG. Vasopressin: Physiology, assessment and osmosensation. J Intern Med Suppl. 2017;282:284–97.
Holt NF, Haspel KL. Vasopressin: A review of therapeutic applications. J Cardiothorac Vasc Anesth. 2010;24:330–47.
Pasquali R, Gagliardi L, Vicennati V, Gambineri A, Colitta D, Ceroni L, et al. ACTH and cortisol response to combined corticotropin releasing hormone-arginine vasopressin stimulation in obese males and its relationship to body weight, fat distribution and parameters of the metabolic syndrome. Int J Obes. 1999;23:419–24.
Heinrichs M, von Dawans B, Domes G. Oxytocin, vasopressin, and human social behavior. Front Neuroendocrinol. 2009;30:548–57.
Keverne EB, Curley JP. Vasopressin, oxytocin and social behaviour. Curr Opin Neurobiol. 2004;14:777–83.
Song Z, Albers HE. Cross-talk among oxytocin and arginine-vasopressin receptors: Relevance for basic and clinical studies of the brain and periphery. Front Neuroendocrinol. 2018;51:14–24.
Ramos L, Hicks C, Kevin R, Caminer A, Narlawar R, Kassiou M, et al. Acute prosocial effects of oxytocin and vasopressin when given alone or in combination with 3,4-methylenedioxymethamphetamine in rats: Involvement of the V1A receptor. Neuropsychopharmacol. 2013;38:2249–59.
Fujiwara Y, Hiroyama M, Sanbe A, Yamauchi J, Tsujimoto G, Tanoue A. Mutual regulation of vasopressin- and oxytocin-induced glucagon secretion in V1b vasopressin receptor knockout mice. J Endocrinol. 2007;192:361–9.
Young LJ, Flanagan-Cato LM. Editorial comment: Oxytocin, vasopressin and social behavior. Horm Behav. 2012;61:227–9.
Manning M, Misicka A, Olma A, Bankowski K, Stoev S, Chini B, et al. Oxytocin and vasopressin agonists and antagonists as research tools and potential therapeutics. J Neuroendocrinol. 2012;24:609–28.
Cid-Jofré V, Moreno M, Reyes-Parada M, Renard GM. Role of oxytocin and vasopressin in neuropsychiatric disorders: Therapeutic potential of agonists and antagonists. Int J Mol Sci. 2021;22:12077.
Kreier F, Swaab DF. History of hypothalamic research: “The spring of primitive existence”. In: Swaab DF, Kreier F, Lucassen PJ, Salehi A, Buijs RM, editors. Handbook of Clinical Neurology, vol. 179, Elsevier; 2021. p. 7–43.
Leibowitz SF, Alexander JT. Hypothalamic serotonin in control of eating behavior, meal size, and body weight. Biol Psychiatry. 1998;44:851–64.
Saper CB, Chou TC, Elmquist JK. The need to feed: Homeostatic and hedonic control of eating. Neuron. 2002;36:199–211.
Saper CB, Scammell TE, Lu J. Hypothalamic regulation of sleep and circadian rhythms. Nature. 2005;437:1257–63.
Le TM, Liao D-L, Ide J, Zhang S, Zhornitsky S, Wang W, et al. The interrelationship of body mass index with gray matter volume and resting-state functional connectivity of the hypothalamus. Int J Obes. 2020;44:1097–107.
Brown SSG, Westwater ML, Seidlitz J, Ziauddeen H, Fletcher PC. Hypothalamic volume is associated with body mass index. NeuroImage: Clinical. 2023;39:103478.
Rahmouni K. Cardiovascular regulation by the arcuate nucleus of the hypothalamus. Hypertension. 2016;67:1064–71.
Sapru HN. Role of the hypothalamic arcuate nucleus in cardiovascular regulation. Auton Neurosci: Basic and Clinical. 2013;175:38–50.
Guillemin R, Burgus R. The hormones of the hypothalamus. Sci Am. 1972;227:24–33.
Bernstein H-G, Müller S, Dobrowolny H, Wolke C, Lendeckel U, Bukowska A, et al. Insulin-regulated aminopeptidase immunoreactivity is abundantly present in human hypothalamus and posterior pituitary gland, with reduced expression in paraventricular and suprachiasmatic neurons in chronic schizophrenia. Eur Arch Psychiatry Clin Neurosci. 2017;267:427–43.
Guo L, Qi Y-J, Tan H, Dai D, Balesar R, Sluiter A, et al. Different oxytocin and corticotropin-releasing hormone system changes in bipolar disorder and major depressive disorder patients. EBioMedicine. 2022;84:104266.
Ross HE, Young LJ. Oxytocin and the neural mechanisms regulating social cognition and affiliative behavior. Front Neuroendocrinol. 2009;30:534–47.
Inada K, Tsujimoto K, Yoshida M, Nishimori K, Miyamichi K. Oxytocin signaling in the posterior hypothalamus prevents hyperphagic obesity in mice. eLife. 2022;11:e75718.
Chen S-D, You J, Zhang W, Wu B-S, Ge Y-J, Xiang S-T, et al. The genetic architecture of the human hypothalamus and its involvement in neuropsychiatric behaviours and disorders. Nat Hum Behav. 2024;8:1–15.
Satizabal CL, Adams HHH, Hibar DP, White CC, Knol MJ, Stein JL, et al. Genetic architecture of subcortical brain structures in 38,851 individuals. Nat Genet. 2019;51:1624–36.
Vattikuti S, Guo J, Chow CC. Heritability and genetic correlations explained by common SNPs for metabolic syndrome traits. PLoS Genet. 2012;8:e1002637.
DeForest N, Majithia AR. Genetics of type 2 diabetes: Implications from large-scale studies. Curr Diab Rep. 2022;22:227–35.
Hilker R, Helenius D, Fagerlund B, Skytthe A, Christensen K, Werge TM, et al. Heritability of schizophrenia and schizophrenia spectrum based on the nationwide Danish twin register. Biol Psychiatry. 2018;83:492–8.
Gordovez FJA, McMahon FJ. The genetics of bipolar disorder. Mol Psychiatry. 2020;25:544–59.
Hibar DP, Stein JL, Renteria ME, Arias-Vasquez A, Desrivières S, Jahanshad N, et al. Common genetic variants influence human subcortical brain structures. Nature. 2015;520:224–9.
Hibar DP, Adams HHH, Jahanshad N, Chauhan G, Stein JL, Hofer E, et al. Novel genetic loci associated with hippocampal volume. Nat Commun. 2017;8:13624.
Elvsåshagen T, Shadrin A, Frei O, van der Meer D, Bahrami S, Kumar VJ, et al. The genetic architecture of the human thalamus and its overlap with ten common brain disorders. Nat Commun. 2021;12:2909.
Elvsåshagen T, Bahrami S, van der Meer D, Agartz I, Alnæs D, Barch DM, et al. The genetic architecture of human brainstem structures and their involvement in common brain disorders. Nat Commun. 2020;11:4016.
Bahrami S, Nordengen K, Rokicki J, Shadrin AA, Rahman Z, Smeland OB, et al. The genetic landscape of basal ganglia and implications for common brain disorders. Nat Commun. 2024;15:8476.
Moberget T, van der Meer D, Bahrami S, Roelfs D, Frei O, Kaufmann T, et al. The genetic architecture of human cerebellar morphology supports a key role for the cerebellum in human evolution and psychopathology. 2024:2023.02.10.23285704.
Campbell ML, Shadrin A, van der Meer D, Tesfaye M, Smeland OB, O’Connell KS, et al. Regional genetic architecture of the corpus callosum and its overlap with psychiatric and substance use phenotypes. 2023:2023.12.03.23299351.
Frei O, Holland D, Smeland OB, Shadrin AA, Fan CC, Maeland S, et al. Bivariate causal mixture model quantifies polygenic overlap between complex traits beyond genetic correlation. Nat Commun. 2019;10:2417.
Andreassen OA, Thompson WK, Schork AJ, Ripke S, Mattingsdal M, Kelsoe JR, et al. Improved detection of common variants associated with schizophrenia and bipolar disorder using pleiotropy-informed conditional false discovery rate. PLoS Genet. 2013;9:e1003455.
Mountjoy E, Schmidt EM, Carmona M, Schwartzentruber J, Peat G, Miranda A, et al. An open approach to systematically prioritize causal variants and genes at all published human GWAS trait-associated loci. Nat Genet. 2021;53:1527–33.
Ghoussaini M, Mountjoy E, Carmona M, Peat G, Schmidt EM, Hercules A, et al. Open Targets Genetics: systematic identification of trait-associated genes using large-scale genetics and functional genomics. Nucleic Acids Res. 2021;49:D1311–D1320.
Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826.
Ishunina TA, Swaab DF. Vasopressin and oxytocin neurons of the human supraoptic and paraventricular nucleus; Size changes in relation to age and sex. J Clin Endocrinol Metab. 1999;84:4637–44.
Billot B, Bocchetta M, Todd E, Dalca AV, Rohrer JD, Iglesias JE. Automated segmentation of the hypothalamus and associated subunits in brain MRI. Neuroimage. 2020;223:117287.
Bocchetta M, Gordon E, Manning E, Barnes J, Cash DM, Espak M, et al. Detailed volumetric analysis of the hypothalamus in behavioral variant frontotemporal dementia. J Neurol. 2015;262:2635–42.
Winterton A, Bettella F, Beck D, Gurholt TP, Steen NE, Rødevand L, et al. The oxytocin signalling gene pathway contributes to the association between loneliness and cardiometabolic health. Psychoneuroendocrinology. 2022;144:105875.
O’Connor LJ, Schoech AP, Hormozdiari F, Gazal S, Patterson N, Price AL. Extreme polygenicity of complex traits is explained by negative selection. Am J Hum Genet. 2019;105:456–76.
Lappalainen T, Li YI, Ramachandran S, Gusev A. Genetic and molecular architecture of complex traits. Cell. 2024;187:1059–75.
Wendt FR, Pathak GA, Tylee DS, Goswami A, Polimanti R. Heterogeneity and polygenicity in psychiatric disorders: A genome-wide perspective. Chronic Stress. 2020;4:2470547020924844.
Doernberg E, Hollander E. Neurodevelopmental disorders (ASD and ADHD): DSM-5, ICD-10, and ICD-11. CNS Spectr. 2016;21:295–9.
Grzadzinski R, Huerta M, Lord C. DSM-5 and autism spectrum disorders (ASDs): an opportunity for identifying ASD subtypes. Mol Autism. 2013;4:12.
Greenwood TA, Shutes-David A, Tsuang DW. Endophenotypes in schizophrenia: Digging deeper to identify genetic mechanisms. J Psychiatr Brain Sci. 2019;4:e190005.
Donati FL, D’Agostino A, Ferrarelli F. Neurocognitive and neurophysiological endophenotypes in schizophrenia: An overview. Biomark Neuropsychiatry. 2020;3:100017.
Gazal S, Finucane HK, Furlotte NA, Loh P-R, Palamara PF, Liu X, et al. Linkage disequilibrium–dependent architecture of human complex traits shows action of negative selection. Nat Genet. 2017;49:1421–7.
Boyle EA, Li YI, Pritchard JK. An expanded view of complex traits: From polygenic to omnigenic. Cell. 2017;169:1177–86.
Hindley G, Frei O, Shadrin AA, Cheng W, O’Connell KS, Icick R, et al. Charting the landscape of genetic overlap between mental disorders and related traits beyond genetic correlation. Am J Psychiatry. 2022;179:833–43.
Nogueira FN, Carvalho RA. Metabolic remodeling triggered by salivation and diabetes in major salivary glands. NMR Biomed. 2017;30:e3683.
Rui L. Energy metabolism in the liver. Compr Physiol. 2014;4:177–97.
Lambert J-C, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet. 2013;45:1452.
Nho K, Kim S, Horgusluoglu E, Risacher SL, Shen L, Kim D, et al. Association analysis of rare variants near the APOE region with CSF and neuroimaging biomarkers of Alzheimer’s disease. BMC Med Genomics. 2017;10:29.
Kulminski AM, Philipp I, Shu L, Culminskaya I. Definitive roles of TOMM40-APOE-APOC1 variants in the alzheimer’s risk. Neurobiol Aging. 2022;110:122–31.
Jeong JH, Lee DK, Jo Y-H. Cholinergic neurons in the dorsomedial hypothalamus regulate food intake. Mol Metab. 2017;6:306–12.
Saper CB, Lowell BB. The hypothalamus. Curr Biol. 2014;24:R1111–R1116.
Border R, O’Rourke S, de Candia T, Goddard ME, Visscher PM, Yengo L, et al. Assortative mating biases marker-based heritability estimators. Nat Commun. 2022;13:660.
Border R, Athanasiadis G, Buil A, Schork AJ, Cai N, Young AI, et al. Cross-trait assortative mating is widespread and inflates genetic correlation estimates. Science. 2022;378:754–61.
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for StatisticalComputing, Vienna, Austria. https://www.R-project.org/ 2024.
Posit team. RStudio: Integrated Development Environment for R. Posit Software, PBC, Boston, MA. http://www.posit.co/ 2025.
Fischl B. FreeSurfer. Neuroimage. 2012;62:774–81.
Mbatchou J, Barnard L, Backman J, Marcketta A, Kosmicki JA, Ziyatdinov A, et al. Computationally efficient whole-genome regression for quantitative and binary traits. Nat Genet. 2021;53:1097–103.
Bulik-Sullivan BK, Loh P-R, Finucane HK, Ripke S, Yang J, Patterson N, et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet. 2015;47:291–5.
Finucane HK, Bulik-Sullivan B, Gusev A, Trynka G, Reshef Y, Loh P-R, et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat Genet. 2015;47:1228–35.
Bulik-Sullivan B, Finucane HK, Anttila V, Gusev A, Day FR, Loh P-R, et al. An atlas of genetic correlations across human diseases and traits. Nat Genet. 2015;47:1236–41.
Smeland OB, Frei O, Shadrin A, O’Connell K, Fan C-C, Bahrami S, et al. Discovery of shared genomic loci using the conditional false discovery rate approach. Hum Genet. 2020;139:85–94.
Wickham H, Averick M, Bryan J, Chang W, McGowan LD, François R, et al. Welcome to the tidyverse. J Open Source Softw. 2019;4:1686.
Audunsdottir K, Sartorius AM, Kang H, Glaser BD, Boen R, Nærland T, et al. The effects of oxytocin administration on social and routinized behaviors in autism: A preregistered systematic review and meta-analysis. Psychoneuroendocrinology. 2024;167:107067.
Chaves VE, Tilelli CQ, Brito NA, Brito MN. Role of oxytocin in energy metabolism. Peptides. 2013;45:9–14.
Gutkowska J, Jankowski M, Mukaddam-Daher S, McCann SM. Oxytocin is a cardiovascular hormone. Braz J Med Biol Res. 2000;33:625–33.
Li E, Wang L, Wang D, Chi J, Lin Z, Smith GI, et al. Control of lipolysis by a population of oxytocinergic sympathetic neurons. Nature. 2024;625:175–80.
Nakamura K, Velho G, Bouby N. Vasopressin and metabolic disorders: translation from experimental models to clinical use. J Intern Med Suppl. 2017;282:298–309.
Shapiro NL, Todd EG, Billot B, Cash DM, Iglesias JE, Warren JD, et al. In vivo hypothalamic regional volumetry across the frontotemporal dementia spectrum. NeuroImage: Clinical. 2022;35:103084.
Grove J, Ripke S, Als TD, Mattheisen M, Walters RK, Won H, et al. Identification of common genetic risk variants for autism spectrum disorder. Nat Genet. 2019;51:431–44.
O’Connell KS, Koromina M, van der Veen T, Boltz T, David FS, Yang JMK, et al. Genomics yields biological and phenotypic insights into bipolar disorder. Nature. 2025;639:968–75.
Trubetskoy V, Pardiñas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604:502–8.
Locke AE, Kahali B, Berndt SI, Justice AE, Pers TH, Day FR, et al. Genetic studies of body mass index yield new insights for obesity biology. Nature. 2015;518:197–206.
Shungin D, Winkler TW, Croteau-Chonka DC, Ferreira T, Locke AE, Mägi R, et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature. 2015;518:187–96.
Morris JA, Kemp JP, Youlten SE, Laurent L, Logan JG, Chai RC, et al. An atlas of genetic influences on osteoporosis in humans and mice. Nat Genet. 2019;51:258–66.
Klarin D, Damrauer SM, Cho K, Sun YV, Teslovich TM, Honerlaw J, et al. Genetics of blood lipids among ~300,000 multi-ethnic participants of the Million Veteran Program. Nat Genet. 2018;50:1514–23.
Giri A, Hellwege JN, Keaton JM, Park J, Qiu C, Warren HR, et al. Trans-ethnic association study of blood pressure determinants in over 750,000 individuals. Nat Genet. 2019;51:51–62.
Morris AP, Voight BF, Teslovich TM, Ferreira T, Segrè AV, Steinthorsdottir V, et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat Genet. 2012;44:981–90.
Gadin JR, Zetterberg R, Meijsen J, Schork AJ. Cleansumstats: Converting GWAS sumstats to a common format to facilitate downstream applications. Zenodo. https://doi.org/10.5281/zenodo.7441266 2022.
Sudlow C, Gallacher J, Allen N, Beral V, Burton P, Danesh J, et al. UK Biobank: An open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 2015;12:e1001779.
Rosen AFG, Roalf DR, Ruparel K, Blake J, Seelaus K, Villa LP, et al. Quantitative assessment of structural image quality. Neuroimage. 2018;169:407–18.
Bycroft C, Freeman C, Petkova D, Band G, Elliott LT, Sharp K, et al. The UK Biobank resource with deep phenotyping and genomic data. Nature. 2018;562:203–9.
van der Meer D, Hindley G, Shadrin AA, Smeland OB, Parker N, Dale AM, et al. Mapping the genetic landscape of psychiatric disorders with the MiXeR toolset. Biol Psychiatry. 2025;98:455–65.
Feizi A, Ray K. otargen: GraphQL-based R package for tidy data accessing and processing from Open Targets Genetics. Bioinformatics. 2023;39:btad441.
Open Targets Genetics. Assigning variants to genes (V2G): Open Targets Genetics documentation. https://genetics-docs.opentargets.org/our-approach/data-pipeline 2022. Accessed 19 February 2025.
Open Targets Genetics. FAQs: Open Targets Genetics documentation. https://genetics-docs.opentargets.org/faqs 2023. Accessed 19 February 2025.
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: Multitissue gene regulation in humans. Science. 2015;348:648–60.
Acknowledgements
This research was funded by Research Council of Norway (301767). Figure 1 was created with https://app.biorender.com/ and Adobe Illustrator 2024–2026, the grid for the Venn diagrams in Fig. 3 was created in Adobe Illustrator 2024–2026. This work was partly performed on the TSD (“Tjenester for Sensitive Data”) facilities, owned by the University of Oslo, operated and developed by the TSD service group at the University of Oslo, IT-Department (USIT) (tsd-drift@usit.uio.no). Computations were also performed on resources provided by UNINETT Sigma2 – the National Infrastructure for High-Performance Computing and Data Storage in Norway (NS9666S). All methods were performed in accordance with the relevant ethical guidelines and regulations.
Author information
Authors and Affiliations
Contributions
AIS, DSQ and DVDM conceived and planned the study. AIS analyzed the data, with contributions from JR for the hypothalamus subunits segmentation, AS for the GWAS, heritability, MiXeR and genetic correlation analyses, and DVDM for the conjFDR analyses. DSQ and DVDM supervised the study. DSQ provided funding for the study. AIS interpreted the results, with contributions from DSQ, DVDM, AS, JR, MC, AW, OAA, TN and LTW. AIS wrote the first and revised drafts of the manuscript, with DSQ, DVDM, AS, JR, MC, AW, OAA, ES, TN and LTW contributing to the first and revised drafts of the manuscript.
Corresponding author
Ethics declarations
Competing interests
ES received speaker fees from bfd buchholz-fachingormationsdienst GmbH and Lundbeckfonden as well as editorial fees from Lundbeckfonden and the Wellcome Trust. All other authors declare no potential competing interests with respect to the research, authorship and/or publication of this article.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sartorius, A.I., van der Meer, D., Shadrin, A. et al. Genetic pathways linking oxytocin-vasotocin hypothalamic subunit architecture with psychiatric and metabolic traits. Mol Psychiatry (2026). https://doi.org/10.1038/s41380-026-03508-4
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41380-026-03508-4