Abstract
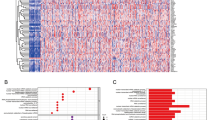
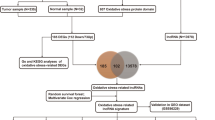
Although rRNA metabolism-related genes have been reported to be associated with human cancer, a systematic assessment of rRNA metabolism-related genes across human cancers is lacking. Thus, we performed a Pan-cancer analysis of rRNA metabolism-related genes across 20 human cancers. Here, we examined mRNA expression, mutation, DNA methylation, copy number variation (CNV) and clinical landscape of rRNA metabolism-related genes in more than 8600 patients across 20 human cancers from The Cancer Genome Atlas (TCGA) dataset. Besides, ten independent Gene Expression Omnibus (GEO) datasets, Cancer Cell Line Encyclopedia (CCLE) dataset and Project Achilles dataset were used to verify our study. A landscape of rRNA metabolism-related genes was established across 20 human cancers. The results suggest that rRNA metabolism-related genes are upregulated in multiple cancers, particularly in digestive and respiratory system cancers. Most of the upregulated genes were driven by CNV gain rather than mutation or DNA hypomethylation. We systematically identified CNV-driven rRNA metabolism-related genes with clinical relevance, including EXOSC8. Finally, functional experiments confirmed the oncogenic roles of EXOSC8 in colorectal carcinoma. Our study highlights the important roles of rRNA metabolism-related genes in tumorigenesis as prognostic biomarkers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Weinstein JN, Collisson EA, Mills GB, Shaw KR, Ozenberger BA, Ellrott K, et al. The cancer genome atlas pan-cancer analysis project. Nat Genet. 2013;45:1113–20.
Cancer Cell Line Encyclopedia Consortium GODS. Pharmacogenomic agreement between two cancer cell line data sets. Nature. 2015;528:84–7.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, et al. The cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483:603–7.
Beroukhim R, Mermel CH, Porter D, Wei G, Raychaudhuri S, Donovan J, et al. The landscape of somatic copy-number alteration across human cancers. Nature. 2010;463:899–905.
Ye Y, Hu Q, Chen H, Liang K, Yuan Y, Xiang Y, et al. Characterization of hypoxia-associated molecular features to aid hypoxia-targeted therapy. Nat Metab 2019;1:431–44.
Gong J, Li Y, Liu CJ, Xiang Y, Li C, Ye Y, et al. A pan-cancer analysis of the expression and clinical relevance of small nucleolar RNAs in human cancer. Cell Rep 2017;21:1968–81.
Schaub FX, Dhankani V, Berger AC, Trivedi M, Richardson AB, Shaw R, et al. Pan-cancer alterations of the MYC oncogene and its proximal network across the cancer genome atlas. Cell Syst 2018;6:282–300.e2.
Ge Z, Leighton JS, Wang Y, Peng X, Chen Z, Chen H, et al. Integrated genomic analysis of the ubiquitin pathway across cancer types. Cell Rep 2018;23:213–226.e3.
Wang Y, Xu X, Maglic D, Dill MT, Mojumdar K, Ng PK, et al. Comprehensive molecular characterization of the hippo signaling pathway in cancer. Cell Rep. 2018;25:1304–1317.e5.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–9.
Consortium TGO. Expansion of the gene ontology knowledgebase and resources. Nucleic Acids Res. 2017;45:D331–D338.
Dolezal JM, Dash AP, Prochownik EV. Diagnostic and prognostic implications of ribosomal protein transcript expression patterns in human cancers. BMC Cancer. 2018;18:275.
Srivastava L, Lapik YR, Wang M, Pestov DG. Mammalian DEAD box protein Ddx51 acts in 3’ end maturation of 28S rRNA by promoting the release of U8 snoRNA. Mol Cell Biol. 2010;30:2947–56.
Choudhury P, Hackert P, Memet I, Sloan KE, Bohnsack MT. The human RNA helicase DHX37 is required for release of the U3 snoRNP from pre-ribosomal particles. RNA Biol. 2019;16:54–68.
Allmang C, Kufel J, Chanfreau G, Mitchell P, Petfalski E, Tollervey D. Functions of the exosome in rRNA, snoRNA and snRNA synthesis. EMBO J. 1999;18:5399–410.
van Dijk EL, Schilders G, Pruijn GJ. Human cell growth requires a functional cytoplasmic exosome, which is involved in various mRNA decay pathways. RNA. 2007;13:1027–35.
Prieto JL, McStay B. Recruitment of factors linking transcription and processing of pre-rRNA to NOR chromatin is UBF-dependent and occurs independent of transcription in human cells. Genes Dev. 2007;21:2041–54.
Sloan KE, Bohnsack MT, Schneider C, Watkins NJ. The roles of SSU processome components and surveillance factors in the initial processing of human ribosomal RNA. RNA. 2014;20:540–50.
Chestkov IV, Jestkova EM, Ershova ES, Golimbet VE, Lezheiko TV, Kolesina NY et al. Abundance of ribosomal RNA gene copies in the genomes of schizophrenia patients. Schizophr Res. 2018;197 :305–14.
Ruggero D, Shimamura A. Marrow failure: a window into ribosome biology. Blood. 2014;124:2784–92.
Calo E, Gu B, Bowen ME, Aryan F, Zalc A, Liang J, et al. Tissue-selective effects of nucleolar stress and rDNA damage in developmental disorders. Nature. 2018;554:112–7.
Kulkarni S, Dolezal JM, Wang H, Jackson L, Lu J, Frodey BP, et al. Ribosomopathy-like properties of murine and human cancers. PLoS ONE. 2017;12:e0182705.
Bhattacharya C, Wang X, Becker D. The DEAD/DEAH box helicase, DDX11, is essential for the survival of advanced melanomas. Mol Cancer. 2012;11:82.
Cowling VH, Turner SA, Cole MD. Burkitt’s lymphoma-associated c-Myc mutations converge on a dramatically altered target gene response and implicate Nol5a/Nop56 in oncogenesis. Oncogene. 2014;33:3519–27.
Lin TC, Su CY, Wu PY, Lai TC, Pan WA, Jan YH. et al. The nucleolar protein NIFK promotes cancer progression via CK1alpha/beta-catenin in metastasis and Ki-67-dependent cell proliferation. Elife. 2016;5:pii: e11288.
Liu T, Fang Y, Zhang H, Deng M, Gao B, Niu N, et al. HEATR1 negatively regulates Akt to help sensitize pancreatic cancer cells to chemotherapy. Cancer Res. 2016;76:572–81.
Miluzio A, Beugnet A, Grosso S, Brina D, Mancino M, Campaner S, et al. Impairment of cytoplasmic eIF6 activity restricts lymphomagenesis and tumor progression without affecting normal growth. Cancer Cell. 2011;19:765–75.
Tang X, Shi L, Xie N, Liu Z, Qian M, Meng F, et al. SIRT7 antagonizes TGF-beta signaling and inhibits breast cancer metastasis. Nat Commun. 2017;8:318.
Tsoi H, Lam KC, Dong Y, Zhang X, Lee CK, Zhang J, et al. Pre-45s rRNA promotes colon cancer and is associated with poor survival of CRC patients. Oncogene. 2017;36:6109–18.
Liberzon A, Subramanian A, Pinchback R, Thorvaldsdottir H, Tamayo P, Mesirov JP. Molecular signatures database (MSigDB) 3.0. Bioinformatics 2011;27:1739–40.
Hanzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinforma. 2013;14:7.
Foroutan M, Bhuva DD, Lyu R, Horan K, Cursons J, Davis MJ. Single sample scoring of molecular phenotypes. BMC Bioinforma. 2018;19:404.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005;102:15545–50.
Helleday T, Eshtad S, Nik-Zainal S. Mechanisms underlying mutational signatures in human cancers. Nat Rev Genet. 2014;15:585–98.
Ehrlich M. DNA hypomethylation in cancer cells. Epigenomics. 2009;1:239–59.
Gamazon ER, Stranger BE. The impact of human copy number variation on gene expression. Brief Funct Genomics. 2015;14:352–7.
Henrichsen CN, Chaignat E, Reymond A. Copy number variants, diseases and gene expression. Hum Mol Genet. 2009;18:R1–8.
Maruvka YE, Tang M, Michor F. On the validity of using increases in 5-year survival rates to measure success in the fight against cancer. PLoS ONE. 2014;9:e83100.
Tai P, Yu E, Cserni G, Vlastos G, Royce M, Kunkler I, et al. Minimum follow-up time required for the estimation of statistical cure of cancer patients: verification using data from 42 cancer sites in the SEER database. BMC Cancer. 2005;5:48.
Driscoll JJ, Rixe O. Overall survival: still the gold standard: why overall survival remains the definitive end point in cancer clinical trials. Cancer J. 2009;15:401–5.
Hudis CA, Barlow WE, Costantino JP, Gray RJ, Pritchard KI, Chapman JA, et al. Proposal for standardized definitions for efficacy end points in adjuvant breast cancer trials: the STEEP system. J Clin Oncol. 2007;25:2127–32.
Punt CJ, Buyse M, Kohne CH, Hohenberger P, Labianca R, Schmoll HJ, et al. Endpoints in adjuvant treatment trials: a systematic review of the literature in colon cancer and proposed definitions for future trials. J Natl Cancer Inst. 2007;99:998–1003.
Meyers RM, Bryan JG, McFarland JM, Weir BA, Sizemore AE, Xu H, et al. Computational correction of copy number effect improves specificity of CRISPR-Cas9 essentiality screens in cancer cells. Nat Genet. 2017;49:1779–84.
Rohrmoser M, Holzel M, Grimm T, Malamoussi A, Harasim T, Orban M, et al. Interdependence of Pes1, Bop1, and WDR12 controls nucleolar localization and assembly of the PeBoW complex required for maturation of the 60S ribosomal subunit. Mol Cell Biol. 2007;27:3682–94.
Chung KY, Cheng IK, Ching AK, Chu JH, Lai PB, Wong N. Block of proliferation 1 (BOP1) plays an oncogenic role in hepatocellular carcinoma by promoting epithelial-to-mesenchymal transition. Hepatology. 2011;54:307–18.
Killian A, Sarafan-Vasseur N, Sesboue R, Le Pessot F, Blanchard F, Lamy A, et al. Contribution of the BOP1 gene, located on 8q24, to colorectal tumorigenesis. Genes Chromosomes Cancer. 2006;45:874–81.
Song J, Ma Z, Hua Y, Xu J, Li N, Ju C, et al. Functional role of RRS1 in breast cancer cell proliferation. J Cell Mol Med. 2018;22:6304–13.
Pan Y, Tong J, Kang W, Lung R, Chak WP, Chung LY, et al. EXOSC4 functions as a potential oncogene in development and progression of colorectal cancer. Mol Carcinog. 2018;57:1780–91.
Haag S, Kretschmer J, Bohnsack MT. WBSCR22/Merm1 is required for late nuclear pre-ribosomal RNA processing and mediates N7-methylation of G1639 in human 18S rRNA. RNA. 2015;21:180–7.
Yan D, Tu L, Yuan H, Fang J, Cheng L, Zheng X, et al. WBSCR22 confers oxaliplatin resistance in human colorectal cancer. Sci Rep. 2017;7:15443.
Gantenbein N, Bernhart E, Anders I, Golob-Schwarzl N, Krassnig S, Wodlej C, et al. Influence of eukaryotic translation initiation factor 6 on non-small cell lung cancer development and progression. Eur J Cancer. 2018;101:165–80.
McIver SC, Kang YA, DeVilbiss AW, O’Driscoll CA, Ouellette JN, Pope NJ, et al. The exosome complex establishes a barricade to erythroid maturation. Blood. 2014;124:2285–97.
Mayakonda A, Lin DC, Assenov Y, Plass C, Koeffler HP. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018;28:1747–56.
Shen H, Xing C, Cui K, Li Y, Zhang J, Du R, et al. MicroRNA-30a attenuates mutant KRAS-driven colorectal tumorigenesis via direct suppression of ME1. Cell Death Differ. 2017;24:1253–62.
Zhu Y, Gu L, Li Y, Lin X, Shen H, Cui K, et al. miR-148a inhibits colitis and colitis-associated tumorigenesis in mice. Cell Death Differ. 2017;24:2199–209.
Li W, Cui K, Prochownik EV, Li Y. The deubiquitinase USP21 stabilizes MEK2 to promote tumor growth. Cell Death Dis. 2018;9:482.
Acknowledgements
We thank the TCGA, GEO, CCLE and Project Achilles teams. We thank Professor Jinxiang Zhang (Wuhan Union Hospital, Wuhan) for providing the CRC patient samples. This work was supported by grants from the National Key R&D Program of China (2016YFC1302300), National Nature Science Foundation of China (81772609) and Medical Science Advancement Program (Basic Medical Sciences) of Wuhan University (TFJC2018005).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Cui, K., Liu, C., Li, X. et al. Comprehensive characterization of the rRNA metabolism-related genes in human cancer. Oncogene 39, 786–800 (2020). https://doi.org/10.1038/s41388-019-1026-9
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-019-1026-9
This article is cited by
-
The role of non-coding RNAs (ncRNAs) and their potential connection with cancer
Egyptian Journal of Medical Human Genetics (2025)
-
Atypical R-loops in cancer: decoding molecular chaos for therapeutic gain
Journal of Translational Medicine (2025)
-
Targeting ribosomes reprograms the tumour microenvironment and augments cancer immunotherapy
British Journal of Cancer (2025)
-
Identification of PFKFB3 as a key factor in the development of colorectal cancer and immunotherapy resistance
Clinical and Experimental Medicine (2024)
-
A pan-cancer analysis of Dyskeratosis congenita 1 (DKC1) as a prognostic biomarker
Hereditas (2023)