Abstract
Global cancer incidence continues to rise, emphasizing the urgent need for improved diagnostics and management strategies. DNA methylation biomarkers in liquid biopsies offer a promising, minimally invasive solution. Despite their potential, only a few tests have successfully transitioned from research to clinical practice. This review addresses key aspects influencing successful biomarker development and clinical implementation—including liquid biopsy source selection, biomarker discovery workflow and targeted validation in clinical sample series—and provide strategies to improve accuracy, reproducibility and clinical utility. Altogether, these considerations could aid in bridging the translational gap from research to clinical application, and to increase the number of clinically implemented liquid biopsy tests.
Similar content being viewed by others
Background
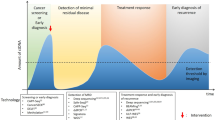
The International Agency for Research on Cancer (IARC) recently predicted a significant increase in cancer incidence world-wide, expecting over 35 million new diagnoses in 2050Footnote 1,Footnote 2. The anticipated rise in cancer cases places great demands on the healthcare systems and underscores the need for enhanced strategies in cancer management. To meet this challenge, liquid biopsies offer a minimally invasive source for detection of a broad range of cancer biomarkers, including markers for screening, diagnosis, assessment of prognosis, detection of residual disease, early identification of recurrence, and prediction of treatment response [1] (Fig. 1). Tumor material, in the form of circulating tumor DNA (ctDNA), circulating tumor cells (CTCs) and exosomes, are shed by the tumors into various body fluids, including blood, urine, saliva, bile and seminal fluid [2], and show several advantages for biomarker discovery compared to tissue biopsies. While tissue biopsies offer a limited view of only one section of the tumor, liquid biopsies reflect both the entire tumor burden of a patient and the molecular cancer heterogeneity [1, 3]. Liquid biopsies are also typically less invasive, and enables repeated sampling, e.g. for monitoring treatment response and cancer progression [1, 4].
Liquid biopsies offer a minimally invasive method with diverse applications in cancer care, including early screening, precise diagnosis, prognostic assessment, real-time disease monitoring, personalized treatment selection, and evaluation of treatment response.
DNA methylation refers to the addition of a methyl group to the 5’ position of cytosine, typically at CpG dinucleotides, resulting in 5-methylcytosine. This epigenetic modification regulates gene expression and chromatin structure without altering the underlying DNA sequence. DNA methylation is essential for processes such as genomic imprinting, X chromosome inactivation, transposon silencing, and ultimately cellular development and differentiation [5]. In cancer, DNA methylation patterns are frequently altered, with tumors typically displaying both genome-wide hypomethylation and hypermethylation of CpG rich gene promoters [6,7,8]. Promoter hypermethylation of key tumor suppressor genes is commonly associated with gene silencing, while global hypomethylation can induce chromosomal instability. Together, these events can disrupt normal growth pathways and drive malignant transformation.
DNA methylation alterations often emerge early in tumorigenesis and remain stable throughout tumor evolution [7, 9, 10], making cancer-specific DNA methylation patterns highly relevant as biomarkers. The inherent stability of the DNA double helix – arising from complementary base pairing and its helical conformation – provides additional protection compared to single-stranded nucleic acid-based biomarkers. However, the rapid clearance of circulating cell-free DNA (cfDNA), with estimated half-lives ranging from minutes up to a few hours [11, 12], represents a challenge for blood-based biomarker analyses. In this setting, DNA methylation biomarkers offer an advantage, as DNA methylation seems to impact cfDNA fragmentation [13]. Nucleosome interactions help to protect methylated DNA from nuclease degradation [14, 15], resulting in a relative enrichment of methylated DNA fragments within the cfDNA pool [16]. Collectively, these features contribute to the high potential of DNA methylation-based biomarkers, including their enhanced resistance to degradation during sample collection, storage, and processing, especially compared to more labile molecules such as RNA [17].
Various methods exist for analysis of DNA methylation and have been comprehensively reviewed [18,19,20]. Whole-genome bisulfite sequencing (WGBS) and reduced representation bisulfite sequencing (RRBS) are widely used for biomarker discovery, providing broad methylome coverage through bisulfite-based chemical conversion. Enzymatic methyl-sequencing (EM-seq), along with emerging third generation sequencing technologies such as nanopore and single-molecule real-time sequencing, offers comprehensive methylation profiling without chemical conversion, thereby better preserving DNA integrity. This is particularly promising for liquid biopsy analyses, where DNA quantity is often limited. Microarrays and enrichment-based techniques, including methylated DNA immunoprecipitation sequencing (MeDIP-seq), support both biomarker discovery and clinical validation by balancing profiling breadth with cost and throughput. However, these methods provide less detailed methylation information compared to whole-genome sequencing methods. Targeted methods, such as quantitative real-time PCR (qPCR) and digital PCR (dPCR), offer highly sensitive, locus-specific analysis, making them particularly suited for clinical validation.
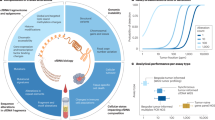
Despite their potential, DNA methylation-based cancer biomarkers have seen limited adoption in routine clinical use. Searching PubMed with the MeSH terms “Biomarkers” and “DNA Methylation” together with the term “cancer” returns 6,191 publications since 1996. Yet, this vast amount of studies is not reflected in the number of DNA methylation biomarkers in clinical use, underscoring the challenges that remain in developing robust, high performance DNA methylation-based biomarkers. Several factors affect the successful clinical implementation of DNA methylation biomarkers (Fig. 2). Low levels of tumor material in liquid biopsies require advanced technologies for biomarker discovery and validation, including optimized discovery workflows and standardized targeted analyses. In addition, the choice of liquid biopsy source, selection of appropriate control groups, in both discovery and validation phases, sufficient independent validation, and large-scale clinical studies to demonstrate clinical utility, are all important factors that impact the chance of succeeding with clinical translation. These aspects are addressed in the following review, mainly focusing on diagnostic DNA methylation biomarkers. Other challenges, such as regulatory and economic barriers, are discussed elsewhere [21].
Successful clinical implementation requires careful selection of an appropriate liquid biopsy source, paired with an optimized biomarker discovery approach and standardized targeted analyses. Emphasizing standardization, optimization and quality control across all stages increase the chances of developing robust biomarker tests that can be validated in clinical trials, demonstrating clinical utility and enabling eventual clinical implementation.
Liquid biopsy source
Blood
Blood is one of the most frequently used sources for liquid biopsy analyses [22]. The circulatory system reaches virtually every tissue in the body, allowing blood to serve as a reservoir for cancer-specific material shed from tumors, regardless of their anatomic location. Both ctDNA, CTCs and extracellular vesicles can be released into the blood stream as tumors grow and evolve [23]. Blood is also easily accessible through minimally invasive procedures, making it attractive from a patient perspective [24]. However, detection of circulating cancer biomarkers is challenging due to the high degree of dilution of tumor-derived signals within the total blood volume of an average adult (4–5 L). As a result, these signals are often present at extremely low concentrations, making them inherently challenging to detect. ctDNA is also highly unstable and degrades rapidly [25]. In addition, blood contains a variety of other components, including white blood cells and cfDNA fragments originating from various healthy tissues. This complex background not only dilutes the tumor-derived signal but may also contribute to false positive biomarker signals, thereby challenging the specificity of targeted analyses.
For blood-based analyses of cancer biomarkers, plasma is frequently used. While serum has a higher concentration of total cfDNA (including both normal cfDNA and ctDNA), plasma is usually enriched for ctDNA, and has less contamination of genomic DNA from lysed cells [3]. The stability of ctDNA is also higher in plasma [26]. Interestingly, while plasma ctDNA is mostly used for analyzing cancer-specific molecular features, such as mutations and DNA methylation alterations, the quantitative level of cfDNA in plasma has in numerous studies been proposed as a promising biomarker per se for various aspects of cancer management [27]. In breast cancer, cfDNA concentrations were found to be significantly higher in cancer patients compared to both healthy controls and individuals with benign tumors, indicating a diagnostic potential for cfDNA [28]. However, clinical utility requires more than a statistically significant difference between cancer and control groups. For a biomarker to be clinically useful, there should be minimal overlap in concentration ranges between groups—a criterion that often presents a challenge for using cfDNA concentration as a stand-alone biomarker [29]. Several reports have further explored how cfDNA and ctDNA yields vary across tumor types and how this variability influences the implementation of DNA methylation-based diagnostic biomarkers. The variability in overall cfDNA among patients with the same cancer type is often larger than the variability seen between different cancer types [30, 31]. In contrast, the fraction of ctDNA tends to differ more consistently between cancer types and stages [32]. This is highly relevant for DNA methylation-based diagnostics, since biomarker sensitivity will be limited not by total cfDNA abundance, but by the proportion of ctDNA present in the sample. At low ctDNA fractions, which are commonly seen in early-stage disease and e.g. cancers of the central nervous system [32], the robustness of DNA methylation-based detection will decrease.
Technological advances, including the development of highly sensitive methods, have facilitated the use of blood as a liquid biopsy source. Accordingly, numerous blood-based DNA methylation tests for cancer management have been proposed [33, 34]. Still, only a few tests have achieved FDA-approval (including Epi proColon and Shield for detection of colorectal cancer) or the designation “FDA Breakthrough Device” (e.g., the two multi-cancer tests Grail’s Galleri and OverC MCDBT; Table 1). Thus, the use of blood as a liquid biopsy source demonstrates promising potential but also presents clear challenges—where a focus on quality and optimization at each step could greatly improve the prospects for successful clinical implementation of blood-based DNA methylation biomarkers.
Local liquid biopsy sources
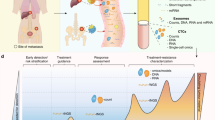
Depending on their anatomical location, various cancer types are likely to shed material into nearby body fluids other than blood, including urine, saliva, cerebrospinal fluid, bile, stool, pleural effusions, peritoneal fluid and seminal fluid [2, 35]. In contrast to the systemic nature of blood, local body fluids often offer distinct advantages, including a higher biomarker concentration and reduced background noise from other tissues (Fig. 3). Thus, for certain cancer types, using a local liquid biopsy source could increase biomarker precision and reliability.
Blood as a systemic liquid biopsy source, circulates throughout the body, collecting tumor materials irrespective of anatomical location. In contrast, local liquid biopsy sources often provide distinct advantages, including higher target concentrations and reduced background signals. For urological cancers, urine serves as an ideal local liquid biopsy source, offering non-invasive collection and potentially higher biomarker sensitivity compared to blood. The illustrated examples demonstrate these comparative advantages. *Reference to example 1 [36] and example 2 [40].
For urological cancers such as bladder-, prostate- and kidney cancer, urine is a natural liquid biopsy source due to its proximity to the urinary tract. Urine samples can be obtained non-invasively, minimizing patient discomfort and supporting better compliance with testing protocols. Urine is particularly effective for bladder cancer management, as most tumors are in direct contact with the urine, resulting in higher concentrations of tumor-derived biomarkers compared with blood. This leads to greater accuracy for urine-based tests (Fig. 3). A relevant example is detection of TERT mutations in matched urine and blood samples from bladder cancer patients, where the sensitivity was 87% in urine versus 7% in plasma [36]. Table 1 highlights the strong representation of urine-based tests for bladder cancer among commercially available and FDA-designated breakthrough devices. In comparison, fewer urine-based DNA methylation biomarker tests exist for prostate and renal cancer [37], likely due to lower amount of CTCs and ctDNA shed into the urine from these cancers [38, 39]. Nevertheless, a significantly higher biomarker sensitivity in urine compared to plasma has also been reported in prostate cancer [40].
For biliary tract cancers including cholangiocarcinoma, bile has emerged as a promising liquid biopsy source [41]. Studies indicate that bile samples often outperform plasma in detecting tumor-related somatic mutations [42], providing more sensitive and specific clinical information. Similar findings have been reported for stool and cerebrospinal fluid, which offer superior performance compared to plasma in detecting cancer-specific DNA methylation biomarkers in early-stage colorectal cancer [43] and brain tumors [44]. While some local liquid biopsy sources, such as urine and saliva, can be collected non-invasively, others - including bile, cerebrospinal fluid, pleural fluid and peritoneal fluid - typically require more invasive procedures. This could in principle limit their utility. However, many of these fluids are routinely obtained during diagnostic or therapeutic procedures, so their use for biomarker testing does not involve additional invasiveness or risk. For instance, bile can be obtained during endoscopic retrograde cholangiopancreatography (ERCP), a procedure frequently used in the diagnostic evaluation of suspected cholangiocarcinoma [45].
Conclusion
Blood plasma is the most widely used liquid biopsy source and shows considerable clinical promise across cancer types and applications. However, for some cancer types, local body fluids may offer advantages such as higher concentration of tumor-derived signals and a reduced background noise, which can significantly improve the accuracy and clinical utility of biomarker tests.
Discovery of DNA methylation biomarkers
The STARD (Standards for Reporting of Diagnostic Accuracy Studies) guidelines were developed to improve the completeness and transparency of diagnostic accuracy study reports [46], including those reporting new cancer biomarkers. These guidelines provide comprehensive recommendations covering essential aspects such as study design, results presentation, and the intended clinical application. An effective discovery strategy is necessary for identifying biomarkers with high diagnostic accuracy. This strategy should involve careful selection of appropriate liquid biopsy sources, optimized pre-analytical procedures, inclusion of relevant control groups and state-of-the-art high-throughput technologies. Additionally, it should incorporate an appropriate bioinformatics workflow to ensure robust data generation, accurate analysis and reliable interpretation, thereby increasing the likelihood of developing clinically relevant biomarkers.
Clinical material
To increase the chances of succeeding with a clinical liquid biopsy test, an appropriate sample type should be selected already in the biomarker discovery phase. To date, tissue samples have been most frequently used for biomarker discovery and have resulted in highly accurate cancer biomarkers that have subsequently been detected in liquid biopsies (Table 2). Incorporating liquid biopsy samples directly into the discovery phase may offer distinct advantages, allowing identification of biomarkers that are inherently detectable in biofluids, potentially reducing the number of steps from discovery to clinical application. An increasing number of methylation-based high-throughput methods are using cfDNA directly as starting material and suggest that biomarkers identified using this approach have robust performance (Table 2). As a proof of concept, Hlady et al. have demonstrated that biomarker candidates identified directly from cfDNA performed better than biomarkers primarily identified from tissue samples [47]. Efforts such as the Circulating Cell-free Genome Atlas (CCGA) study have also successfully contributed to the discovery of biomarker panels for detection and prediction of tissue of origin of several cancer types at high specificity [48,49,50].
Pre-analytical steps
The performance and reliability of a biomarker test ultimately depend on the quantity and quality of DNA used in the analysis, both during the discovery phase and in subsequent targeted analyses. Thus, optimizing pre-analytical steps is critical - in particular for liquid biopsy material, where target DNA is often present in low quantities, prone to degradation and surrounded by a complex background of other cells and DNA fragments [23]. Indeed, cfDNA degradation and contamination with normal leukocyte DNA are major pre-analytical variables that can compromise the analytical performance of a blood-based test [3]. Factors such as collection tube type, processing time, storage temperature, centrifugation protocol and DNA purification method all significantly influence the quality and quantity of cfDNA available for downstream analyses [3]. For urine samples, time to processing and storage temperature are especially important. While sedimentation protocols have traditionally been the gold standard for urine processing, alternative solutions – such as stabilizing buffers [51] and urine filtration systems [52] – have been developed to simplify sample handling and preservation during transportation and storage. For bile, phenol-chloroform extraction has been shown to yield higher levels of amplifiable DNA compared to other isolation kits [41]. Efforts to standardize pre-analytical variables are ongoing, with several consortia - including the CANCER-ID project [53], the European Liquid Biopsy Society (ELBS; www.elbs.eu), and BLOODPAC (https://www.bloodpac.org/) [54, 55] - making important contributions to improve the clinical translation of liquid biopsy biomarkers.
To detect DNA methylation in downstream analyses, a DNA pre-treatment step is frequently required. Bisulfite conversion is considered the gold standard for this purpose, and various commercial kits are available, each with differences in DNA recovery and conversion efficiency [56]. Studies have also reported variability in DNA yield and recovery when combining different bisulfite conversion- and cfDNA isolation kits for plasma DNA methylation analysis [57]. Despite its status as the gold standard, bisulfite conversion is a harsh treatment, resulting in considerable DNA degradation, which is particularly challenging for liquid biopsy material where DNA is already limited. Recently, enzymatic conversion using TET2 and APOBEC enzymes (EM-seq; New England BioLabs) has been presented as more gentle and promising alternative to bisulfite conversion [58]. This approach yields longer DNA fragments compared to bisulfite conversion, but results in lower average cfDNA recovery (38% vs. 68%) [59]. As a result, bisulfite conversion seems to remain the preferred method for PCR-based analyses of DNA methylation biomarkers in liquid biopsies, while enzymatic conversion may be better suited for sequencing-based approaches that require longer DNA fragments.
Methylomic-based biomarker discovery
Discovery of novel biomarker candidates has benefitted from advances in high-throughput technologies and streamlined bioinformatic workflows. Microarray-based profiling and genome-wide methylation sequencing are widely used for this purpose (Table 2). DNA methylation arrays have an affordable price and streamlined bioinformatic workflows. Differentially methylated CpG sites between tumors and controls have been identified from arrays by numerous studies across malignancies [47, 60,61,62,63,64,65,66,67,68] and may constitute promising biomarker candidates for cancer detection. In particular, the commercially available EarlyTect BCD test for bladder cancer detection resulted from an array-based biomarker discovery [60] (Tables 1 and 2). While microarrays cover a set of selected CpG sites, methylome sequencing offers broader genome coverage [69], which facilitates the design and development of more precise and potentially more accurate biomarker assays for subsequent targeted validation. The broader coverage also allows the identification of sequences with relevant methylation patterns in previously unexplored regions, independent of their genomic location or potential function. A commonly used strategy to identify new DNA methylation biomarkers has focused on functionally relevant locations, such as CpG islands close to transcription start sites, where DNA methylation status will likely affect gene expression. However, recent studies have shown that highly performing DNA methylation biomarkers are not restricted to genomic regions with known biological function [70, 71]. This suggests that expanding the search beyond traditionally explored regions can reveal novel high-performing biomarkers, potentially overlooked by conventional strategies.
Several promising discovery studies have used WGBS to analyze plasma cfDNA [49, 50, 71]. Among these initiatives, the Grail’s Galleri test has emerged as a commercially available tool for early detection of multiple cancers (Tables 1 and 2). This blood-based test uses next-generation sequencing (NGS) data combined with a machine learning classifier to identify DNA methylation patterns in cfDNA, potentially detecting over 50 cancer types before symptoms appear. In parallel, enrichment-based techniques have proven to be cost-effective methods for genome-wide methylation profiling. One example is MeDIP combined with fragment sequencing, which has been successfully used for detecting cfDNA methylation in multiple types of cancers [72]. With RRBS, the amount of sequencing required is lowered compared to WGBS, while it still captures most genomic regions affected by DNA methylation [73, 74]. There is also considerable optimism tied to third generation sequencing technology, such as nanopore sequencing [75], which enables advanced, real-time detection of DNA methylation without the need for bisulfite conversion. This technology is expected to overcome several challenges associated with analyses of cfDNA in liquid biopsies, including issues related to DNA fragment interruption, GC bias, and degradation caused by bisulfite treatment. As a proof-of-principle, Lau and colleagues successfully used nanopore sequencing to monitor longitudinal cfDNA methylation dynamics during treatment of cancer patients [76].
Optimized bioinformatic workflows for analysis of sequencing-based data may increase the likelihood of discovering suitable biomarkers. Differentially methylated regions (DMRs) between tumors and controls are considered promising biomarker candidates [70, 72, 77,78,79,80,81]. Compared to single CpG sites, DMRs increase the robustness of subsequent targeted validation analyses, as demonstrated by e.g. the BladMetrix biomarker panel, detecting bladder cancer in urine with a sensitivity of 92% and a specificity of 93% [70]. Although sequencing-based approaches offer numerous advantages, their high cost remains a barrier for using large sample cohorts in the biomarker discovery phase. This may partly explain why a lower number of liquid biopsy-based DNA methylation biomarker panels are so far developed from sequencing-based discoveries compared to the more cost-effective array-based methods (Table 2). However, continuous decrease in price and improved sequencing technology, combined with enhanced bioinformatics capability, are likely to encourage increased use of sequencing-based approaches for liquid biopsy-based biomarker discovery and their direct application in a clinical setting.
Control groups
Biomarker discovery approaches are commonly designed to detect candidates with DNA methylation levels significantly above a threshold established from control cases, thereby robustly separating cancer cases from controls. This distinction is particularly important in plasma or serum samples, where the signal-to-noise ratio can be high. Consequently, the selection of control groups directly affects the specificity of a biomarker test and should be guided by the intended clinical application of the test. Non-malignant tissue adjacent to cancers or samples from healthy individuals are frequently used as controls in the discovery phase (Table 2). However, samples from individuals presenting with symptoms similar to those of cancer patients may provide more clinically relevant controls. Such controls help ensure that the biomarker test can accurately distinguish cancer from other conditions with overlapping clinical presentations, thereby improving reliability. For example, Hlady et al. identified candidate biomarkers for hepatocellular carcinoma using plasma samples from patients with cirrhosis, a patient group at increased risk for this cancer [47]. Similarly, samples from patients with haematuria - a symptom of bladder cancer but also of other genitourinary conditions - are frequently used as controls in bladder cancer detection [37]. For some cancers, including control samples from patients with tumors in anatomically adjacent organs can be beneficial. This approach enables the test to discriminate not only between cancer and normal tissue (cancer specificity), but also between different types of malignancy (cancer type specificity). For instance, prostate and renal cancer patients serve as useful controls in studies aiming to detect bladder cancer from tumor DNA in urine [70]. Importantly, for cancers preceded by inflammatory lesions that increase the risk of tumor development, it is crucial to include individuals with such inflammation in the control group. This is exemplified by the use of samples from primary sclerosing cholangitis [82] or Barrett’s esophagus [83] for the discovery of biomarkers for cholangiocarcinoma and esophageal adenocarcinoma, respectively. Finally, as DNA methylation generally increases with age [84], age-matched controls are particularly important in DNA methylation studies.
An instructive example of the importance of proper control group selection is provided by the CxBladder test [85]. Although Cxbladder initially showed promise for non-invasive detection of bladder cancer and advanced to clinical implementation, an elevated rate of false positives ultimately lowered its positive predictive value [86], leading to the loss of Medicare reimbursement in the USFootnote 3. Retrospective analyses revealed that validation studies did not include patient groups with non-urothelial malignancies or inflammatory urinary tract conditions [85, 87]. This case illustrates how the absence of important controls can undermine clinical validity and hinder successful translation of otherwise promising biomarker assays.
Zero background biomarkers
The discovery of cancer biomarkers with zero background signals in control samples has emerged as a highly accurate approach for blood-based testing and has been successfully applied in several studies [88,89,90,91]. This strategy could be particularly advantageous for early-stage cancers, where the amount of ctDNA released into the bloodstream is minimal. Feber et al. identified biomarker candidates for bladder cancer showing no or very low methylation signals in normal urothelium, blood and non-cancer urine samples [91], while Draškovič et al. identified candidate DMRs for several cancer types that were unmethylated in cfDNA of healthy individuals [90]. Moreover, the commercially available ColveraTM test for detection of colorectal cancer recurrence also utilizes DNA methylation biomarkers with no background signal in plasma from healthy individuals [92]. A zero-background approach differs from the conventional strategy, which establishes a threshold based on background signals in the control group to dichotomize samples. Although commonly used across studies, the threshold-based approach may risk filtering out true but weak (low-level) cancer-derived signals, potentially leading to false negatives. Adapting discovery workflows to prioritize biomarker candidates with zero or minimal background signals in controls could mitigate interference during validation, thereby potentially enhancing sensitivity and accuracy for cancer detection.
Conclusion
Successful discovery of liquid biopsy DNA methylation biomarkers requires optimized study designs, careful sample type selection, rigorous pre-analytical procedures, and the use of advanced molecular and bioinformatic technologies. Integrating liquid biopsy material in the discovery phase, applying sensitive genome-wide approaches, and selecting clinically informed control groups all contribute to improved diagnostic accuracy and potential utility. Approaches that prioritize biomarkers with zero background signals may also improve the translation of promising candidates into robust clinical tests.
Targeted biomarker analysis
To successfully translate biomarker candidates from large-scale discoveries to suitable tests for clinical routine, targeted methods and assays that are sufficiently sensitive, robust, fast, and cost efficient are required [93].
Methods for targeted analysis
Among existing FDA-approved or commercially available DNA methylation liquid biopsy tests for cancer management, the majority rely on quantitative methylation-specific PCR (qMSP; Table 1). PCR-based methods have long been considered the gold standard for targeted analyses in DNA methylation biomarker research, due to their reliability and precision. Additionally, PCR platforms are widely available and well-established in diagnostic laboratories, making them a practical choice for clinical applications. Moreover, qMSP allows high-throughput analysis of large clinical sample series. However, the detection of extremely rare alterations within a complex background may require a more sensitive method, such as digital PCR (dPCR). The dPCR technology has been reported to have a significantly lower limit of detection (LOD); approximately 0.001% compared to around 0.1% for qPCR [94]. This increased sensitivity is achieved by the division of the sample into thousands of partitions, with a separate PCR reaction running inside each partition [95]. The partitioning significantly reduces competition from background DNA and increases the detection of rare events [95]. Moreover, recent advancements in dPCR technology allow for multiplexing, i.e. analyzing multiple targets simultaneously in one reaction. This capability is particularly valuable for liquid biopsy analysis, where the amount of target DNA often is limited. Although none of the FDA-approved DNA methylation tests currently used for cancer management are dPCR-based, this method shows promising potential for future clinical use [96].
While PCR-based targeted methods are restricted to analyzing a small number of biomarkers, NGS allows for the detection of larger biomarker panels, potentially increasing test accuracy [97]. Some of the most sensitive NGS-based methods, such as whole genome sequencing, have an estimated LOD of ~0.001%. This sensitivity outcompetes qPCR and is comparable to dPCR in detecting rare alterations [94]. NGS is widely used in biomarker discovery, but its application in routine clinical practice has so far been limited by high costs and complex data analysis. However, advancement in sequencing technology and bioinformatics has opened new possibilities for more targeted liquid biopsy NGS tests to receive FDA-approved and/or become commercially available for cancer management (Table 1). For example, the blood-based Shield test (Guardant Health), approved by the FDA in 2024 for early detection and screening of colorectal cancer, uses NGS to detect a multi-marker panel consisting of DNA methylation, mutations and fragmentation patterns (Table 1).
Importantly, the choice of methodology for biomarker discovery may influence the compatibility of downstream targeted analyses. Robust qMSP and dPCR assays typically require high CpG density in the target sequence to adequately differentiate between methylated and unmethylated sites, rendering them well-suited for biomarkers identified via methylome sequencing platforms. In contrast, translating microarray-derived biomarkers to qMSP or dPCR assays can be more challenging, as microarray probes often target regions with lower CpG content. Validation of microarray data could instead be conducted on single CpG sites using e.g. pyrosequencing. However, pyrosequencing offers lower resolution and sensitivity compared to dPCR and is thus less suited for liquid biopsy applications, which typically require high analytical performance.
Standardization of targeted analyses
Regardless of the targeted method, ensuring high quality and standardization at every step will improve the likelihood for a biomarker test to achieve clinical utility. First, the precise selection of CpG sites targeted by a biomarker assay is important, as it can significantly influence the observed DNA methylation frequency [98]. For example, studies of the MAL gene promoter in colorectal cancer have reported widely diverging methylation frequencies, ranging from 6% [99] to 80% [100]. MAL has a variable DNA methylation pattern in its promoter region and the inclusion of different CpG sites in various targeted PCR assays will lead to discrepant results [101]. Such discrepancies in methylation frequencies for the same gene in the same cancer type can reduce the clinical value of a biomarker test [3]. Second, a variety of parameters in the experimental pipeline for PCR-based DNA methylation analyses can impact the consistency of methylation values. Among these, the choice of reference for normalization, or internal control, is particularly critical [102]. Since cancer often is characterized by chromosomal aberrations, using single-copy genes as references for normalization may introduce bias in the end-results. Internal controls targeting multiple loci, like ALU sequences in qPCR [102, 103] or the 4Plex control in dPCR [104], have been shown to increase the consistency of methylation measurements. For dPCR specifically, another step that directly affects final methylation concentrations is partition classification, i.e. classifying droplets into positive and negative clusters. Various partition classification algorithms have been developed and compared [105] and should be carefully selected for each study. Of note, the PoDCall algorithm has been specifically designed for DNA methylation analyses on the dPCR platform, and has been shown to increase the robustness of such analyses [106].
Although various recommendations are found in the literature, there is still no consensus in the scientific community on a standardized pipeline for targeted methylation analyses. Nevertheless, an important step towards improving transparency and quality in PCR-based studies has been the development of the MIQE (The Minimum Information for Publication of Quantitative Real-Time PCR Experiments) guidelines [107,108,109]. These guidelines have been widely adopted, with several scientific journals requiring authors to submit a standardized MIQE table. The implementation of MIQE guidelines has significantly increased awareness of the importance of maintaining quality at every step of the experimental pipeline.
Control groups
Selecting appropriate control groups is crucial when developing a biomarker test for clinical use. As discussed in the biomarker discovery phase, the choice of controls depends on both the cancer type under investigation and the intended clinical application of the test. While the most essential controls are typically included during discovery, additional relevant control groups should also be incorporated in targeted analyses. These may comprise patients with benign or inflammatory conditions, cancer-free individuals at high cancer risk, or patients with other cancer types. Achieving high biomarker specificity often requires inclusion of multiple control groups in the study design, particularly for liquid biopsies, where target biomarkers often coexist with DNA fragments and cells from other tissues. Blood-based cancer biomarkers often perform well when compared to healthy controls but may struggle to distinguish cancer from non-cancerous diseases, highlighting the challenge of identifying reliable biomarkers for cancer detection [110]. For urine-based biomarkers, specificity against other urological cancers is natural. However, evaluating their performance against non-urological malignancies (such as lung, breast and colorectal cancer), is also important, as cfDNA from these cancer types can be present in urine [111].
Conclusion
The choice of targeted method may influence a biomarker test’s potential to achieve clinical utility. Various methods differ in sensitivity and suitability for routine use, each with advantages and limitations. Regardless of method, rigorous quality control, optimization and standardization throughout the pipeline are essential to generate robust and consistent methylation results that can be validated in clinical trials.
Demonstrating analytical validity, clinical validity and clinical utility
The National Academy of Medicine advises that tumor biomarkers should not be employed in general practice until they have demonstrated both analytical validity and clinical utility [112]. Analytical validity is defined as “the ability of an assay to detect and measure, with statistical significance, the presence of a biomarker of interest accurately, reproducibly, and reliably” [3]. Key measures for evaluating analytical validity include sensitivity, specificity, accuracy, and robustness [3], and these should be assessed using samples representative of real clinical settings. The required analytic performance depends on intended clinical use, such as general population screening, high-risk monitoring, prediction of treatment response, or detection of relapse. Higher sensitivity is typically prioritized for biomarkers used in high-risk populations, whereas higher specificity is essential for screening in the general population [113].
Clinical validity refers to the “ability of an assay to divide, with statistical significance, one population into two or more groups on the basis of outcome, such as presence of cancer and treatment response” [3]. While most biomarkers are initially measured as continuous variables, clinical decisions frequently rely on establishing thresholds or cut-off values [112]. The validation of these thresholds requires assessment of parameters such as sensitivity, specificity, negative predictive value (NPV), positive predictive value (PPV), and positive and negative likelihood ratios (LR+, LR−) [114]. Both retrospective and prospective patient sample series may be used for clinical validation, preferably using samples series that are blinded and independent of the discovery series. The sample size needed for robust clinical validation depends on several factors, including the biomarker’s intended use and the required test performance (e.g. accuracy, sensitivity, specificity) [114]. Also, included patients should preferentially have characteristics that mirrors the target population to ensure applicability. Many studies that demonstrate clinical validity of a biomarker, yet fail to establish its clinical utility, have relied on convenience samples collected without specific consideration for the biomarker’s intended use [112].
To establish clinical utility, a biomarker must possess high analytical and clinical validity, and demonstrate improved patient outcomes when used to guide clinical strategy [112]. If a new biomarker fails to improve patient outcomes and to prove value beyond existing tests and biomarkers already in clinical use, it may never reach clinical implementation [115]. The most reliable way to demonstrate clinical utility is through well-powered prospective clinical trials, conducted either as observational studies, where the test does not direct patient management, or as intervention studies, where the test directs patient management [112]. In rare cancers, validation through well-powered clinical trials may be challenging. In such cases, smaller or adaptive trials, potentially using surrogate endpoints may be appropriate [116, 117]. Additionally, clinical utility can be assessed through prospective-retrospective studies using archived specimens collected during previously conducted prospective trials. In such cases, it is recommended to include at least two data sets from independent clinical trials to strengthen the validity of the findings [118]. Irrespective of design, selecting a patient population that accurately reflects those intended for the clinical application of the liquid biopsy assay will likely increases the chance that the observed biomarker accuracy also applies to real-world scenarios.
Summary
Several aspects affect the chance of succeeding with moving biomarkers from bench to bedside, some of which have been discussed in this review. The type of liquid biopsy source could significantly affect the accuracy of a biomarker test. This decision should be guided by the principle that the sensitivity of an accurate biomarker is generally proportional to the quantity of ctDNA copies analyzed: the more copies analyzed, the higher the sensitivity [10]. In many cases, the concentration of ctDNA in nearby body fluids is higher than in blood. Consequently, for certain cancers, using local body fluids as a liquid biopsy source may offer superior performance over blood-based assays. It is also important to consider whether to use tissue samples or liquid biopsies directly in the discovery phase, as this could impact the accuracy of the final test. Biomarkers with high sensitivity in tissue samples do not necessarily show the same accuracy in liquid biopsies. Although successful liquid biopsy biomarkers have been identified from tissue, using liquid biopsies already in the discovery phase may increase the accuracy of the final biomarkers and maximize the chances of developing clinically relevant and impactful biomarkers.
Both sequencing- and array-based approaches have their limitations and advantages. The choice of method in the discovery phase depends on several factors, including available resources, cost and intended downstream analysis or validation methods. Compared to sequencing-based approaches, arrays allow for more samples to be analyzed at a lower cost and have traditionally been the preferred method. However, one should keep in mind that transferring array-based biomarkers to targeted analyses may be cumbersome, while biomarker candidates identified from sequencing-based approaches could be more easily developed into PCR-based assays for validation. Such PCR-based assays can further be quite straightforwardly implemented into clinical use. Additionally, the cost of sequencing has decreased in recent years and is likely to continue doing so in the coming years, facilitating the use of NGS approaches, or even third generation sequencing (e.g. nanopore sequencing), for both discovery and as targeted panels for direct clinical use. Regardless of the chosen method, selecting appropriate clinical cases and controls that accurately represent the target population for the intended liquid biomarker test, is of importance both in the discovery and validation phase. Failing to include relevant controls may lead to difficulties in replicating the original findings, resulting in reduced accuracy when the biomarker is tested or evaluated in the actual study population.
Finally, clinical validation in independent sample series and performing clinical studies to prove clinical utility is essential. These steps ensure that the biomarkers provide actionable information to improve patient management and outcomes. Ultimately, the goal is to develop biomarkers that not only show statistical significance in controlled studies but also have meaningful impact on clinical decision making and patient care. Altogether, this comprehensive approach, from careful method selection and sample preparation to rigorous validation and clinical utility demonstration, is key to successful transitioning of biomarkers from bench to bedside.
Data availability
No datasets were generated or analyzed during the current study.
Notes
https://gco.iarc.who.int/tomorrow/en, accessed 18.03.2025
Medicare Coverage of Cxbladder Expected to Cease » Pacific Edge
References
Crowley E, Di Nicolantonio F, Loupakis F, Bardelli A. Liquid biopsy: monitoring cancer-genetics in the blood. Nat Rev Clin Oncol. 2013;10:472–84.
Tivey A, Church M, Rothwell D, Dive C, Cook N. Circulating tumour DNA—looking beyond the blood. Nat Rev Clin Oncol. 2022;19:600–12.
Merker JD, Oxnard GR, Compton C, Diehn M, Hurley P, Lazar AJ, et al. Circulating tumor DNA analysis in patients with cancer: American Society of Clinical Oncology and College of American Pathologists joint review. JCO. 2018;36:1631–41.
Lone SN, Nisar S, Masoodi T, Singh M, Rizwan A, Hashem S, et al. Liquid biopsy: a step closer to transform diagnosis, prognosis and future of cancer treatments. Mol Cancer. 2022;21:79.
Berdasco M, Esteller M. Aberrant epigenetic landscape in cancer: how cellular identity goes awry. Dev Cell. 2010;19:698–711.
Herman JG, Baylin SB. Gene silencing in cancer in association with promoter hypermethylation. N Engl J Med. 2003;349:2042–54.
Esteller M. Epigenetics in cancer. N Engl J Med. 2008;358:1148–59.
Ehrlich M. DNA methylation in cancer: too much, but also too little. Oncogene. 2002;21:5400–13.
Kim M, Costello J. DNA methylation: an epigenetic mark of cellular memory. Exp Mol Med. 2017;49:e322.
Worm Ørntoft M-B. Review of blood-based colorectal cancer screening: how far are circulating cell-Free DNA methylation markers from clinical implementation?. Clinical Colorectal Cancer. 2018;17:e415–33.
Kustanovich A, Schwartz R, Peretz T, Grinshpun A. Life and death of circulating cell-free DNA. Cancer Biol Ther. 2019;20:1057–67.
Diehl F, Schmidt K, Choti MA, Romans K, Goodman S, Li M, et al. Circulating mutant DNA to assess tumor dynamics. Nat Med. 2008;14:985–90.
An Y, Zhao X, Zhang Z, Xia Z, Yang M, Ma L, et al. DNA methylation analysis explores the molecular basis of plasma cell-free DNA fragmentation. Nat Commun. 2023;14:287.
Choy JS, Wei S, Lee JY, Tan S, Chu S, Lee T-H. DNA methylation increases nucleosome compaction and rigidity. J Am Chem Soc. 2010;132:1782–3.
Collings CK, Waddell PJ, Anderson JN. Effects of DNA methylation on nucleosome stability. Nucleic Acids Res. 2013;41:2918–31.
Noë M, Mathios D, Annapragada AV, Koul S, Foda ZH, Medina JE, et al. DNA methylation and gene expression as determinants of genome-wide cell-free DNA fragmentation. Nat Commun. 2024;15:6690.
Huang L-H, Lin P-H, Tsai K-W, Wang L-J, Huang Y-H, Kuo H-C, et al. The effects of storage temperature and duration of blood samples on DNA and RNA qualities. PLoS One. 2017;12:e0184692.
Neefs I, Ibrahim J, Peeters M, Van Camp G, Op De Beeck K. The technology landscape for detection of DNA methylation in cancer liquid biopsies. Epigenetics. 2025;20:2453273.
Yang Y, Wen X, Wang L. Advancements in DNA methylation technologies and their application in cancer diagnosis. Epigenetics. 2025;20:2539995.
Hum M, Lee ASG. DNA methylation in breast cancer: early detection and biomarker discovery through current and emerging approaches. J Transl Med. 2025;23:465.
Sheriff S, Saba M, Patel R, Fisher G, Schroeder T, Arnolda G, et al. A scoping review of factors influencing the implementation of liquid biopsy for cancer care. J Exp Clin Cancer Res. 2025;44:50.
Luo H, Wei W, Ye Z, Zheng J, Xu R-H. Liquid biopsy of methylation biomarkers in cell-Free DNA. Trends Mol Med. 2021;27:482–500.
Heitzer E, Haque IS, Roberts CES, Speicher MR. Current and future perspectives of liquid biopsies in genomics-driven oncology. Nat Rev Genet. 2019;20:71–88.
Lee MJ, Hueniken K, Kuehne N, Lu L, Jiang SX, Id Said B, et al. Cancer patient-reported preferences and knowledge for liquid biopsies and blood biomarkers at a comprehensive cancer center. Cancer Manag Res. 2020;12:1163–73.
Dang DK, Park BH. Circulating tumor DNA: current challenges for clinical utility. J Clin Investig. 2022;132:e154941.
Lee J-S, Kim M, Seong M-W, Kim H-S, Lee YK, Kang HJ. Plasma vs. serum in circulating tumor DNA measurement: characterization by DNA fragment sizing and digital droplet polymerase chain reaction. Clin Chem Lab Med. 2020;58:527–32.
Pessoa LS, Heringer M, Ferrer VP. ctDNA as a cancer biomarker: a broad overview. Crit Rev Oncol Hematol. 2020;155:103109.
Huang ZH, Li LH, Hua D. Quantitative analysis of plasma circulating DNA at diagnosis and during follow-up of breast cancer patients. Cancer Lett. 2006;243:64–70.
Hench IB, Hench J, Tolnay M. Liquid biopsy in clinical management of breast, lung, and colorectal cancer. Front Med. 2018;5:9.
LaFramboise WA, Petrosko P, Gallo PH, Gil L, Lam TL, Barr RM, et al. Cell-free DNA, tumor molecular concordance, and clinical correlates of patients with cancer treated in a large community health care network. J Mol Diagn. 2025;27:882–98.
Bryzgunova OE, Konoshenko MY, Laktionov PP. Concentration of cell-free DNA in different tumor types. Expert Rev Mol Diagn. 2021;21:63–75.
Sánchez-Herrero E, Serna-Blasco R, Robado de Lope L, González-Rumayor V, Romero A, Provencio M. Circulating tumor DNA as a cancer biomarker: an overview of biological features and factors that may impact on ctDNA analysis. Front Oncol. 2022;12:943253.
Warton K, Mahon KL, Samimi G. Methylated circulating tumor DNA in blood: power in cancer prognosis and response. Endocr Relat Cancer. 2016;23:R157–171.
Wang B, Wang M, Lin Y, Zhao J, Gu H, Li X. Circulating tumor DNA methylation: a promising clinical tool for cancer diagnosis and management. Clin Chem Lab Med. 2024;62:2111–27.
Leygo C, Williams M, Jin HC, Chan MWY, Chu WK, Grusch M, et al. DNA methylation as a noninvasive epigenetic biomarker for the detection of cancer. Disease Markers. 2017;2017:1–13.
Avogbe PH, Manel A, Vian E, Durand G, Forey N, Voegele C, et al. Urinary TERT promoter mutations as non-invasive biomarkers for the comprehensive detection of urothelial cancer. eBioMedicine. 2019;44:431–8.
Larsen LK, Lind GE, Guldberg P, Dahl C. DNA-methylation-based detection of urological cancer in urine: overview of biomarkers and considerations on biomarker design, source of DNA, and detection technologies. IJMS. 2019;20:2657.
Fujita K, Pavlovich CP, Netto GJ, Konishi Y, Isaacs WB, Ali S, et al. Specific detection of prostate cancer cells in urine by multiplex immunofluorescence cytology. Hum Pathol. 2009;40:924–33.
Zill OA, Banks KC, Fairclough SR, Mortimer SA, Vowles JV, Mokhtari R, et al. The Landscape of actionable genomic alterations in cell-free circulating tumor DNA from 21,807 advanced cancer patients. Clin Cancer Res. 2018;24:3528–38.
Payne SR, Serth J, Schostak M, Kamradt J, Strauss A, Thelen P, et al. DNA methylation biomarkers of prostate cancer: Confirmation of candidates and evidence urine is the most sensitive body fluid for non-invasive detection. Prostate. 2009;69:1257–69.
Vedeld HM, Grimsrud MM, Andresen K, Pharo HD, von Seth E, Karlsen TH, et al. Early and accurate detection of cholangiocarcinoma in patients with primary sclerosing cholangitis by methylation markers in bile. Hepatology. 2022;75:59–73.
Gou Q, Zhang CZ, Sun ZH, Wu LG, Chen Y, Mo ZQ, et al. Cell-free DNA from bile outperformed plasma as a potential alternative to tissue biopsy in biliary tract cancer. ESMO Open. 2021;6:100275.
Liu Y, Zhao G, Miao J, Li H, Ma Y, Liu X, et al. Performance comparison between plasma and stool methylated SEPT9 tests for detecting colorectal cancer. Front Genet. 2020;11:324.
Zuccato JA, Patil V, Mansouri S, Voisin M, Chakravarthy A, Shen SY, et al. Cerebrospinal fluid methylome-based liquid biopsies for accurate malignant brain neoplasm classification. Neuro Oncol. 2023;25:1452–60.
Lazaridis KN, LaRusso NF. Primary sclerosing cholangitis. N Engl J Med. 2016;375:1161–70.
Bossuyt PM, Reitsma JB, Bruns DE, Gatsonis CA, Glasziou PP, Irwig L, et al. STARD 2015: an updated list of essential items for reporting diagnostic accuracy studies. BMJ. 2015;351:h5527.
Hlady RA, Zhao X, Pan X, Yang JD, Ahmed F, Antwi SO, et al. Genome-wide discovery and validation of diagnostic DNA methylation-based biomarkers for hepatocellular cancer detection in circulating cell free DNA. Theranostics. 2019;9:7239–50.
Liu MC, Klein E, Hubbell E, Maddala T, Aravanis AM, Beausan JF, et al. Plasma cell-free DNA (cfDNA) assays for early multi-cancer detection: the circulating cell-free genome atlas (CCGA) study. Ann Oncol. 2018;29:viii14–viii57.
Liu MC, Oxnard GR, Klein EA, Swanton C, Seiden MV, Liu MC, et al. Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA. Annals Oncol. 2020;31:745–59.
Klein EA, Richards D, Cohn A, Tummala M, Lapham R, Cosgrove D, et al. Clinical validation of a targeted methylation-based multi-cancer early detection test using an independent validation set. Annals Oncol. 2021;32:1167–77.
Nel I, Münch C, Shamkeeva S, Heinemann ML, Isermann B, Aktas B. The Challenge to Stabilize, Extract and Analyze Urinary Cell-Free DNA (ucfDNA) during Clinical Routine. Diagnostics. 2023;13:3670.
Andersson E, Dahmcke CM, Steven K, Larsen LK, Guldberg P. Filtration device for on-site collection, storage and shipment of cells from urine and its application to DNA-based detection of bladder cancer. PLoS ONE. 2015;10:e0131889.
Lampignano R, Neumann MHD, Weber S, Kloten V, Herdean A, Voss T, et al. Multicenter evaluation of circulating cell-free DNA extraction and downstream analyses for the development of standardized (Pre)analytical work flows. Clin Chem. 2020;66:149–60.
Grossman RL, Abel B, Angiuoli S, Barrett JC, Bassett D, Bramlett K, et al. Collaborating to compete: blood profiling atlas in cancer (BloodPAC) consortium. Clin Pharm Ther. 2017;101:589–92.
Lockwood CM, Merker JD, Bain E, Compton C, Grossman RL, Johann D, et al. Towards preanalytical best practices for liquid biopsy studies: a BLOODPAC landscape analysis. Clin Pharm Ther. 2025;117:28–33.
Worm Ørntoft M-B, Jensen SØ, Hansen TB, Bramsen JB, Andersen CL. Comparative analysis of 12 different kits for bisulfite conversion of circulating cell-free DNA. Epigenetics. 2017;12:626–36.
Kresse SH, Brandt-Winge S, Pharo H, Flatin BTB, Jeanmougin M, Vedeld HM, et al. Evaluation of commercial kits for isolation and bisulfite conversion of circulating cell-free tumor DNA from blood. Clin Epigenet. 2023;15:151.
Vaisvila R, Ponnaluri VKC, Sun Z, Langhorst BW, Saleh L, Guan S, et al. Enzymatic methyl sequencing detects DNA methylation at single-base resolution from picograms of DNA. Genome Res. 2021;31:1280–9.
Kresse SH, Thorkildsen EM, Brandt-Winge S, Pharo H, Vedeld HM, Lind GE. Comparison of enzymatic and bisulfite conversion of circulating cell-free tumor DNA for DNA methylation analyses. Clin Epigenet. 2025;17:93.
Oh TJ, Lim E, Bang B-R, Lee JJ, Na YG, Shin JH, et al. Identification and validation of methylated PENK gene for early detection of bladder cancer using urine DNA. BMC Cancer. 2022;22:1195.
Lin P-C, Lin J-K, Lin C-H, Lin H-H, Yang S-H, Jiang J-K, et al. Clinical relevance of plasma DNA methylation in colorectal cancer patients identified by using a genome-wide high-resolution array. Ann Surg Oncol. 2015;22:1419–27.
Xu R, Wei W, Krawczyk M, Wang W, Luo H, Flagg K, et al. Circulating tumour DNA methylation markers for diagnosis and prognosis of hepatocellular carcinoma. Nature Mater. 2017;16:1155–61.
Liu L, Toung JM, Jassowicz AF, Vijayaraghavan R, Kang H, Zhang R, et al. Targeted methylation sequencing of plasma cell-free DNA for cancer detection and classification. Ann Oncol. 2018;29:1445–53.
Liang N, Li B, Jia Z, Wang C, Wu P, Zheng T, et al. Ultrasensitive detection of circulating tumour DNA via deep methylation sequencing aided by machine learning. Nat Biomed Eng. 2021;5:586–99.
Lange CPE, Campan M, Hinoue T, Schmitz RF, Van Der Meulen-de Jong AE, Slingerland H, et al. Genome-scale discovery of dna-methylation biomarkers for blood-based detection of colorectal cancer. Marsit CJ, editor. PLoS ONE. 2012;7:e50266.
Gouda MA, Duose DY, Lapin M, Zalles S, Huang HJ, Xi Y, et al. Mutation-agnostic detection of colorectal cancer using liquid biopsy-based methylation-specific signatures. Oncologist. 2023;28:368–72.
Sabedot TS, Malta TM, Snyder J, Nelson K, Wells M, deCarvalho AC, et al. A serum-based DNA methylation assay provides accurate detection of glioma. Neuro-Oncol. 2021;23:1494–508.
Gallardo-Gómez M, Rodríguez-Girondo M, Planell N, Moran S, Bujanda L, Etxart A, et al. Serum methylation of GALNT9, UPF3A, WARS, and LDB2 as noninvasive biomarkers for the early detection of colorectal cancer and advanced adenomas. Clin Epigenet. 2023;15:157.
Beck D, Ben Maamar M, Skinner MK. Genome-wide CpG density and DNA methylation analysis method (MeDIP, RRBS, and WGBS) comparisons. Epigenetics. 2022;17:518–30.
Pharo HD, Jeanmougin M, Ager-Wick E, Vedeld HM, Sørbø AK, Dahl C, et al. BladMetrix: a novel urine DNA methylation test with high accuracy for detection of bladder cancer in hematuria patients. Clin Epigenet. 2022;14:115.
Chan KCA, Jiang P, Chan CWM, Sun K, Wong J, Hui EP, et al. Noninvasive detection of cancer-associated genome-wide hypomethylation and copy number aberrations by plasma DNA bisulfite sequencing. Proc Natl Acad Sci USA. 2013;110:18761–8.
Shen SY, Singhania R, Fehringer G, Chakravarthy A, Roehrl MHA, Chadwick D, et al. Sensitive tumour detection and classification using plasma cell-free DNA methylomes. Nature. 2018;563:579–83.
Meissner A. Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis. Nucleic Acids Res. 2005;33:5868–77.
De Koker A, Raman L, Van Paemel R, Van der Linden M, Van de Velde S, van der Leest P, et al. Development of circulating cell-free DNA reduced representation bisulfite sequencing for clinical methylomics diagnostics. bioRxiv. 2025;663195.
Si H-Q, Wang P, Long F, Zhong W, Meng Y-D, Rong Y, et al. Cancer liquid biopsies by Oxford Nanopore Technologies sequencing of cell-free DNA: from basic research to clinical applications. Mol Cancer. 2024;23:265.
Lau BT, Almeda A, Schauer M, McNamara M, Bai X, Meng Q, et al. Single-molecule methylation profiles of cell-free DNA in cancer with nanopore sequencing. Genome Med. 2023;15:33.
Ooki A, Maleki Z, Tsay J-CJ, Goparaju C, Brait M, Turaga N, et al. A panel of novel detection and prognostic methylated DNA markers in primary non–small cell lung cancer and serum DNA. Clin Cancer Res. 2017;23:7141–52.
Kandimalla R, Xu J, Link A, Matsuyama T, Yamamura K, Parker MI, et al. EpiPanGI Dx: a cell-free DNA methylation fingerprint for the early detection of gastrointestinal cancers. Clin Cancer Res. 2021;27:6135–44.
Nuzzo PV, Berchuck JE, Korthauer K, Spisak S, Nassar AH, Abou Alaiwi S, et al. Detection of renal cell carcinoma using plasma and urine cell-free DNA methylomes. Nat Med. 2020;26:1041–3.
Li W, Li Q, Kang S, Same M, Zhou Y, Sun C, et al. CancerDetector: ultrasensitive and non-invasive cancer detection at the resolution of individual reads using cell-free DNA methylation sequencing data. Nucleic Acids Res. 2018;46:e89–e89.
Cristall K, Bidard F-C, Pierga J-Y, Rauh MJ, Popova T, Sebbag C, et al. A DNA methylation-based liquid biopsy for triple-negative breast cancer. npj Precis Onc. 2021;5:53.
Yang JD, Ghoz H, Aboelsoud MM, Taylor WR, Yab TC, Berger CK, et al. DNA methylation markers for detection of cholangiocarcinoma: discovery, validation, and clinical testing in biliary brushings and plasma. Hepatol Commun. 2021;5:1448–59.
Stone TC, Ward V, Hogan A, Alexander Ho KM, Wilson A, McBain H, et al. Using saliva epigenetic data to develop and validate a multivariable predictor of esophageal cancer status. Epigenomics. 2024;16:109–25.
Horvath S. DNA methylation age of human tissues and cell types. Genome Biol. 2013;14:3156.
O’Sullivan P, Sharples K, Dalphin M, Davidson P, Gilling P, Cambridge L, et al. A multigene urine test for the detection and stratification of bladder cancer in patients presenting with hematuria. J Urol. 2012;188:741–7.
Chou R, Gore JL, Buckley D, Fu R, Gustafson K, Griffin JC, et al. Urinary biomarkers for diagnosis of bladder cancer: a systematic review and meta-analysis. Ann Intern Med. 2015;163:922–31.
Holyoake A, O’Sullivan P, Pollock R, Best T, Watanabe J, Kajita Y, et al. Development of a multiplex RNA urine test for the detection and stratification of transitional cell carcinoma of the bladder. Clin Cancer Res. 2008;14:742–9.
Vrba L, Futscher BW. A suite of DNA methylation markers that can detect most common human cancers. Epigenetics. 2018;13:61–72.
Vrba L, Oshiro MM, Kim SS, Garland LL, Placencia C, Mahadevan D, et al. DNA methylation biomarkers discovered in silico detect cancer in liquid biopsies from non-small cell lung cancer patients. Epigenetics. 2020;15:419–30.
Draškovič T, Hauptman N. Discovery of novel DNA methylation biomarker panels for the diagnosis and differentiation between common adenocarcinomas and their liver metastases. Sci Rep. 2024;14:3095.
Feber A, Dhami P, Dong L, De Winter P, Tan WS, Martínez-Fernández M, et al. UroMark—a urinary biomarker assay for the detection of bladder cancer. Clin Epigenet. 2017;9:8.
Mitchell SM, Ross JP, Drew HR, Ho T, Brown GS, Saunders NF, et al. A panel of genes methylated with high frequency in colorectal cancer. BMC Cancer. 2014;14:54.
BLUEPRINT consortium. Quantitative comparison of DNA methylation assays for biomarker development and clinical applications. Nat Biotechnol. 2016;34:726–37.
Siravegna G, Marsoni S, Siena S, Bardelli A. Integrating liquid biopsies into the management of cancer. Nat Rev Clin Oncol. 2017;14:531–48.
Hindson BJ, Ness KD, Masquelier DA, Belgrader P, Heredia NJ, Makarewicz AJ, et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem. 2011;83:8604–10.
Batool SM, Muralidharan K, Hsia T, Falotico S, Gamblin AS, Rosenfeld YB, et al. Highly sensitive EGFRvIII detection in circulating extracellular vesicle RNA of glioma patients. Clin Cancer Res. 2022;28:4070–82.
Reese K-L, Pantel K, Smit DJ. Multibiomarker panels in liquid biopsy for early detection of pancreatic cancer—a comprehensive review. J Exp Clin Cancer Res. 2024;43:250.
Lind GE, Van Engeland M. Details matter: the role of genomic location and assay standardization in DNA methylation analyses. Epigenomics. 2017;9:933–5.
Mori Y, Cai K, Cheng Y, Wang S, Paun B, Hamilton JP, et al. A genome-wide search identifies epigenetic silencing of somatostatin, tachykinin-1, and 5 other genes in colon cancer. Gastroenterology. 2006;131:797–808.
Lind GE, Ahlquist T, Kolberg M, Berg M, Eknaes M, Alonso MA, et al. Hypermethylated MAL gene - a silent marker of early colon tumorigenesis. J Transl Med. 2008;6:13.
Lind GE, Ahlquist T, Lothe RA. DNA hypermethylation of MAL: a promising diagnostic biomarker for colorectal tumors. Gastroenterology. 2007;132:1631–2.
Pharo HD, Honne H, Vedeld HM, Dahl C, Andresen K, Liestøl K, et al. Experimental factors affecting the robustness of DNA methylation analysis. Sci Rep. 2016;6:33936.
Weisenberger DJ, Trinh BN, Campan M, Sharma S, Long TI, Ananthnarayan S, et al. DNA methylation analysis by digital bisulfite genomic sequencing and digital MethyLight. Nucleic Acids Res. 2008;36:4689–98.
Pharo HD, Andresen K, Berg KCG, Lothe RA, Jeanmougin M, Lind GE. A robust internal control for high-precision DNA methylation analyses by droplet digital PCR. Clin Epigenet. 2018;10:24.
Vynck M, Chen Y, Gleerup D, Vandesompele J, Trypsteen W, Lievens A, et al. Digital PCR partition classification. Clin Chem. 2023;69:976–90.
Jeanmougin M, Brodal HP, Dietrichson Pharo H, Vedeld HM, Lind GE. PoDCall: positive droplet calling and normalization of droplet digital PCR DNA methylation data. Bioinformatics. 2023;39:btac766.
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem. 2009;55:611–22.
Huggett JF, Foy CA, Benes V, Emslie K, Garson JA, Haynes R, et al. The digital MIQE guidelines: minimum information for publication of quantitative digital PCR experiments. Clin Chem. 2013;59:892–902.
The dMIQE Group, Whale AS, De Spiegelaere W, Trypsteen W, Nour AA, Bae Y-K, et al. The digital MIQE guidelines update: minimum information for publication of quantitative digital PCR experiments for 2020. Clin Chem. 2020;66:1012–29.
Smelik M, Zhao Y, Mansour Aly D, Mahmud AF, Sysoev O, Li X, et al. Multiomics biomarkers were not superior to clinical variables for pan-cancer screening. Commun Med. 2024;4:234.
Salfer B, Li F, Wong DTW, Zhang L. Urinary cell-free DNA in liquid biopsy and cancer management. Clin Chem. 2022;68:1493–501.
Hayes DF. Defining clinical utility of tumor biomarker tests: a clinician’s viewpoint. JCO. 2021;39:238–48.
Passaro A, Al Bakir M, Hamilton EG, Diehn M, André F, Roy-Chowdhuri S, et al. Cancer biomarkers: emerging trends and clinical implications for personalized treatment. Cell. 2024;187:1617–35.
Jennings L, Van Deerlin VM, Gulley ML. College of American Pathologists Molecular Pathology Resource Committee. Recommended principles and practices for validating clinical molecular pathology tests. Arch Pathol Lab Med. 2009;133:743–55.
Parikh NI, Vasan RS. Assessing the clinical utility of biomarkers in medicine. Biomark Med. 2007;1:419–36.
Krendyukov A, Singhvi S, Zabransky M. Value of adaptive trials and surrogate endpoints for clinical decision-making in rare cancers. Front Oncol. 2021;11:636561.
Hilgers R. Design and analysis of clinical trials for small rare disease populations. J Rare Dis Res Treat. 2016;1:53–60.
Simon RM, Paik S, Hayes DF. Use of archived specimens in evaluation of prognostic and predictive biomarkers. JNCI J Natl Cancer Inst. 2009;101:1446–52.
Lofton-Day C, Model F, DeVos T, Tetzner R, Distler J, Schuster M, et al. DNA methylation biomarkers for blood-based colorectal cancer screening. Clin Chem. 2008;54:414–23.
deVos T, Tetzner R, Model F, Weiss G, Schuster M, Distler J, et al. Circulating methylated SEPT9 DNA in plasma is a biomarker for colorectal cancer. Clin Chem. 2009;55:1337–46.
Potter NT, Hurban P, White MN, Whitlock KD, Lofton-Day CE, Tetzner R, et al. Validation of a real-time PCR–based qualitative assay for the detection of methylated SEPT9 DNA in human plasma. Clin Chem. 2014;60:1183–91.
Shirley M. Epi proColon® for colorectal cancer screening: a profile of its use in the USA. Mol Diagn Ther. 2020;24:497–503.
Chung DC, Gray DM, Singh H, Issaka RB, Raymond VM, Eagle C, et al. A cell-free DNA blood-based test for colorectal cancer screening. N Engl J Med. 2024;390:973–83.
Imperiale TF, Ransohoff DF, Itzkowitz SH, Levin TR, Lavin P, Lidgard GP, et al. Multitarget stool DNA testing for colorectal-cancer screening. N Engl J Med. 2014;370:1287–97.
Imperiale TF, Porter K, Zella J, Gagrat ZD, Olson MC, Statz S, et al. Next-generation multitarget stool DNA test for colorectal cancer screening. N Engl J Med. 2024;390:984–93.
Fiorentino V, Pizzimenti C, Franchina M, Rossi ED, Tralongo P, Carlino A, et al. Bladder Epicheck test: a novel tool to support urothelial carcinoma diagnosis in urine samples. IJMS. 2023;24:12489.
Witjes JA, Morote J, Cornel EB, Gakis G, van Valenberg FJP, Lozano F, et al. Performance of the bladder EpiCheckTM methylation test for patients under surveillance for non-muscle-invasive bladder cancer: results of a multicenter, prospective, blinded clinical trial. Eur Urol Oncol. 2018;1:307–13.
Pierconti F, Martini M, Fiorentino V, Cenci T, Racioppi M, Foschi N, et al. Upper urothelial tract high-grade carcinoma: comparison of urine cytology and DNA methylation analysis in urinary samples. Hum Pathol. 2021;118:42–8.
Pedersen SK, Baker RT, McEvoy A, Murray DH, Thomas M, Molloy PL, et al. A two-gene blood test for methylated DNA sensitive for colorectal cancer. PLoS ONE. 2015;10:e0125041.
Murray DH, Baker RT, Gaur S, Young GP, Pedersen SK. Validation of a circulating tumor-derived DNA blood test for detection of methylated BCAT1 and IKZF1 DNA. J Appl Lab Med. 2017;2:165–75.
Lin N, Lin Y, Xu J, Liu D, Li D, Meng H, et al. A multi-analyte cell-free DNA–based blood test for early detection of hepatocellular carcinoma. Hepatol Commun. 2022;6:1753–63.
Cai G, Cai M, Feng Z, Liu R, Liang L, Zhou P, et al. A multilocus blood-based assay targeting circulating tumor DNA methylation enables early detection and early relapse prediction of colorectal cancer. Gastroenterology. 2021;161:2053–2056.e2.
Chen X, Gole J, Gore A, He Q, Lu M, Min J, et al. Non-invasive early detection of cancer four years before conventional diagnosis using a blood test. Nat Commun. 2020;11:3475.
Piatti P, Chew YC, Suwoto M, Yamada T, Jara B, Jia X-Y, et al. Clinical evaluation of Bladder CARE, a new epigenetic test for bladder cancer detection in urine samples. Clin Epigenet. 2021. Available from: https://clinicalepigeneticsjournal.biomedcentral.com/articles/10.1186/s13148-021-01029-1
Bang B-R, Zhong J, Oh TJ, Lee JY, Seo Y, Woo MA, et al. EarlyTect BCD, a streamlined PENK methylation test in urine DNA, effectively detects bladder cancer in patients with hematuria. J Mol Diagn. 2024;26:613–23.
Gao Q, Lin YP, Li BS, Wang GQ, Dong LQ, Shen BY, et al. Unintrusive multi-cancer detection by circulating cell-free DNA methylation sequencing (THUNDER): development and independent validation studies. Ann Oncol. 2023;34:486–95.
Chalasani NP, Ramasubramanian TS, Bhattacharya A, Olson MC, Edwards V DK, Roberts LR, et al. A Novel blood-based panel of methylated dna and protein markers for detection of early-stage hepatocellular carcinoma. Clin Gastroenterol Hepatol. 2021;19:2597–2605.e4.
Chalasani NP, Porter K, Bhattacharya A, Book AJ, Neis BM, Xiong KM, et al. Validation of a novel multitarget blood test shows high sensitivity to detect early stage hepatocellular carcinoma. Clin Gastroenterol Hepatol. 2022;20:173–182.e7.
Weiss G, Schlegel A, Kottwitz D, König T, Tetzner R. Validation of the SHOX2/PTGER4 DNA methylation marker panel for plasma-based discrimination between patients with malignant and nonmalignant lung disease. J Thorac Oncol. 2017;12:77–84.
Roperch J-P, Grandchamp B, Desgrandchamps F, Mongiat-Artus P, Ravery V, Ouzaid I, et al. Promoter hypermethylation of HS3ST2, SEPTIN9 and SLIT2 combined with FGFR3 mutations as a sensitive/specific urinary assay for diagnosis and surveillance in patients with low or high-risk non-muscle-invasive bladder cancer. BMC Cancer. 2016;16:704.
Uehiro N, Sato F, Pu F, Tanaka S, Kawashima M, Kawaguchi K, et al. Circulating cell-free DNA-based epigenetic assay can detect early breast cancer. Breast Cancer Res. 2016;18:129.
Kang S, Li Q, Chen Q, Zhou Y, Park S, Lee G, et al. CancerLocator: non-invasive cancer diagnosis and tissue-of-origin prediction using methylation profiles of cell-free DNA. Genome Biol. 2017;18:53.
Jensen SØ, Øgaard N, Ørntoft M-BW, Rasmussen MH, Bramsen JB, Kristensen H, et al. Novel DNA methylation biomarkers show high sensitivity and specificity for blood-based detection of colorectal cancer—a clinical biomarker discovery and validation study. Clin Epigenet. 2019;11:158.
Manoochehri M, Borhani N, Gerhäuser C, Assenov Y, Schönung M, Hielscher T, et al. DNA methylation biomarkers for noninvasive detection of triple-negative breast cancer using liquid biopsy. Intl J Cancer. 2023;152:1025–35.
Van Paemel R, De Koker A, Vandeputte C, Van Zogchel L, Lammens T, Laureys G, et al. Minimally invasive classification of paediatric solid tumours using reduced representation bisulphite sequencing of cell-free DNA: a proof-of-principle study. Epigenetics. 2021;16:196–208.
Funding
This work was supported by the Norwegian Cancer Society (project number 216129 and 223073), the Norwegian Esophageal Cancer Consortium (NORECa, project number 220115) and the South-Eastern Norway Regional Health Authority (project number 2024032).
Author information
Authors and Affiliations
Contributions
HP: Investigation, Writing: original draft, Writing: review and editing, HMV: Investigation, Writing: original draft, Writing: review and editing, IVS: Investigation, Writing: review and editing, Visualization RP: Investigation, Writing: original draft, Writing: review and editing, Visualization. GEL: Conceptualization, Writing: review and editing, Supervision, Funding acquisition. All authors have agreed to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
GEL has been granted a patent (EP2630261B1/US09797016B2) for biomarkers related to bladder cancer detection. Additionally, GEL and HP are named inventors on a pending PCT patent application (WO/2020/099938) covering biomarkers utilized in the BladMetrix test for bladder cancer detection. GEL and HP are named inventors on a granted patent (US11555222B2) for the 4Plex control, which is mentioned in the present study for standardization of DNA methylation ddPCR results. The 4Plex control has been made freely available to the scientific community. Remaining authors declare that they have no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pharo, H., Vedeld, H.M., Sjurgard, I.V. et al. From concept to clinic: a roadmap for DNA methylation biomarkers in liquid biopsies. Oncogene 44, 4814–4831 (2025). https://doi.org/10.1038/s41388-025-03624-5
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-025-03624-5