Abstract
Cancer cells exhibit abnormally altered proteome to satisfy the metabolic demands that arise from heightened proliferation. Trypsin-like serine proteases are a class of proteolytic enzymes whose expression is often dysregulated in cancer. Serine protease 22 (PRSS22) has been associated with the tumorigenesis of several types of cancers. In this study, we identified PRSS22 as a key driver of inflammation-cancer transition (ICT) in colorectal cancer (CRC). PRSS22 expression was positively correlated with the pathological progression of colitis-associated ICT. Genetic disabling of PRSS22 inhibited the growth and migration of and caused redox stress in CRC cells. Mechanistically, knocking down PRSS22 promoted the expression of HMOX1, which fine-tuned the inflammatory response and led to ferroptosis. Loss of PRSS22 also prevented the cleavage of osteopontin and reduced the migratory capacity of CRC cells. We also observed a reduction of M0-to-M2 macrophage polarization in a co-culture system of CRC cells and THP-1-derived macrophages. Altogether, this study reveals a tumor-promoting function of PRSS22 and positions it as a key driver of CRC progression. Targeting PRSS22 represents a promising therapeutic strategy against CRC.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) ranks among the most prevalent malignant tumors globally, claiming nearly one million lives annually due to its high mortality rate. Although extensive research has identified multiple associated driver genes that promote inflammation-cancer transition (ICT) in the gut [1, 2], therapeutic benefits of existing interventions targeting those ICT drivers remain significantly limited. Therefore, fully elucidating the underlying mechanisms implicated in ICT to CRC and developing new targeted therapeutic strategies are urgent imperatives.
Proteases are a large group of enzymes that catalyze the hydrolysis of peptide bonds, they can be classified based on the amino acid residue involved in the peptide bond cleavage, as serine, cysteine and aspartic proteases. Proteases play a crucial role in the onset and progression of cancer, therefore, there is a huge interest in therapeutically exploiting the reliance of cancer cells on proteases to develop new therapies [3]. Serine protease 22 (PRSS22, also known as tryptase ε, BSSP-4) is a member of the serine protease family [4]. Previous studies have demonstrated that PRSS22 is overexpressed in several types of cancers, including ovarian, breast, lung and colon [5]. Mechanistically, PRSS22 enhances lung cancer cell migration through evoking the ERK1/2-C/EBPβ-VEGF cascade [6]; it also promotes the cleavage of ANXA1 and thus activates the FPR2/ERK signaling axis to enhance breast cancer invasiveness [7]. PRSS22 has recently been found to influence the proliferation, migration and invasion of CRC cells, and is associated with the risk of recurrence following surgery to remove primary colorectal tumours [8, 9]. However, prior to this study, how PRSS22 drives the tumorigenesis of CRC remains largely unknown.
Materials and methods
Animal studies
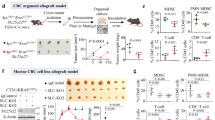
All animal experiments were conducted in accordance with protocols approved by Shanghai University of Traditional Chinese Medicine’s Animal Ethics Committee (Approval Number: PZSHUTCM2406110018). For xenograft studies, 8-week-old BALB/c nude mice were randomly divided into two groups. shPRSS22- or control shRNA (shCtrl)-expressing cells were injected subcutaneously into the right hind paw of mice. Animal body weight and tumor size (V = 0.52 × L × W2) were measured every three days.
Bioinformatic analyses
scRNA-seq and spatial transcriptomic data were retrieved from two public datasets: Xiaojing Chu et al. [10] and Michelli F. Oliveira et al. [11], respectively, and were used to analyze the cell-type-specific expression pattern and spatial localization of PRSS22 within tumor tissues. Data processing and statistical analyses were performed using Seurat (version 5.2.1) [12] and Loupe Browser (version 8.1.2). M1/M2 score-related cell markers were selected based on Sijin Cheng et al. [13].
Clinical CRC transcriptomic data were downloaded from the TCGA database [14], GSE41568 [15], GSE95132 [16], GSE35982 [17], GSE41258 [18], GSE68468 [19] and GSE100179 [20], respectively. Inflammatory bowel disease transcriptomic data was from TaMMa [21]. Colitis transcriptomic data were from GSE131032 [22] and GSE178145 [23]. Patient survival curves were analyzed using KM Plotter [12] and TCGAplot (version 7.0.0) [14].
Cell culture
All cell lines were obtained from the American Type Culture Collection. HCT116 cells were cultured in RPMI-1640 supplemented with 10% (v/v) fetal bovine serum (FBS), Caco-2 and HEK293 cells were cultured in Dulbecco’s Modified Eagle’s medium supplemented with 10% (v/v) FBS, THP-1 cells were cultured in RPMI-1640 medium supplemented with 10% (v/v) FBS and 0.05 mM β-mercaptoethanol.
Western blotting
Western blotting was performed as previously described [24]. Proteins were extracted from cell lysates using Triton lysis buffer (50 mM HEPES pH 7.5, 1% (v/v) Triton, 150 mM NaCl, 10 mM CaCl2 and 1× protease inhibitors (Epizyme Biomedical Technology, China)). Lysates were spun at 16 000 × g for 10 min, the supernatants were kept and total protein concentration was measured using the BCA Protein Assay Kit (Epizyme Biomedical Technology). Samples were mixed with Laemmli sample buffer and boiled at 100 °C for 3 min before loading onto SDS-PAGE gels (Epizyme Biomedical Technology), and transferred to PVDF membranes (Epizyme Biomedical Technology). The membranes were blocked in InterceptTM TBS Blocking Buffer (927-60001, LI-COR Biosciences, NE, USA) for 1 h, and then incubated with the following primary antibodies: anti-PRSS22 (ab197158, Abcam, UK), anti-OPN (22952-1-AP, Proteintech, China), anti-HO-1 (43966, Cell Signaling, MA, USA), anti-beta-actin (66009, Proteintech), anti-V5 (14440-1-AP, Proteintech), anti-Flag (20543-1-AP, Proteintech); overnight at 4 °C. Membranes were then washed thrice in phosphate buffered saline (PBS) supplemented with 0.1% (v/v) Tween-20 (PBST) and incubated with fluorescently tagged secondary antibody for 1 h, washed again thrice in PBST. Fluorescent signals were visualized using an Odyssey CLx imaging system.
Real-time quantitative RT-PCR (qPCR) amplification analysis
Total RNA was extracted using the Trizol Reagent (B610409, Sangon Biotech, China). RNA quality was assessed by measuring the A260/A280 ratio. cDNA was produced using a Reverse Transcription Kit (XKL0511, Guangzhou Xinkailai Biotechnology, China). qPCR was performed using the Fluorescent Quantitative Detection Kit (XKL0412, Guangzhou Xinkailai Biotechnology) on a Real-time PCR instrument (AE31E, Tuojie, China). Table S1 includes a list of primer sequences used in this study.
Plasmids
Human PRSS22 was amplified by qPCR from HCT116 cell DNA extracts, and then cloned into a pcDNA3.1 vector using HindIII and EcoRI restriction sites. The resulting pcDNA3.1-PRSS22 plasmid was validated by Sanger sequencing. Flag-OPN and HMOX1-V5 plasmids were synthesized by Gentlegen (China).
Lentiviral transduction
Recombinant lentiviral vectors expressing shRNA against PRSS22 (pLKO-shPRSS22) or PRSS22 (pLVX-PRSS22) were constructed via PCR and double restriction enzyme digestion (AgeI and XhoI for pLKO; AgeI and SalI for pLVX). PRSS22 siRNA/shRNA sequences are: 5’- CCCACTGTTTCAAGGACAA-3’ and 5’- GGTGGAGAAGATCGTGCAA-3’, respectively. siRNAs were synthesized by Sangon Biotech. Lentiviruses were packaged in HEK293T cells by co-transfecting recombinant lentiviral vectors with packaging plasmids psPAX2 and pMD2.G using Lipofectamine 3000 (Thermofisher, MA, USA). Viral supernatants were collected both at 48 and 72 h post-transfection. For infection, target cells were incubated with viral particles in the presence of 8 μg/mL polybrene (C0351, Beyotime, China). Transfected cells were selected with either 2 μg/mL (HCT116) or 12 μg/mL (Caco-2) puromycin (E607054, Sangon Biotech).
Cell line transfection
When cells reached 40-60% confluence in culture, transfection was performed using TransIntro® EL Transfection Reagent (FT201, Transgen, China).
Cell growth and apoptosis assays
Cell proliferation was measured using the Cell Counting Kit 8 (CCK-8, Beyotime). % Growth (confluence) and death was measured by Incucyte S1 live cell imaging and analysis platform using SYTOXTM Green dye. In addition, cell death was also determined by the V-FITC/PI Apoptosis Detection Kit (C1062S, Beyotime, China) using flow cytometry (BD FACSCelesta, NJ, USA). Data were quantified using the FlowJo software (version 10.9.0).
Colony formation assay
HCT116 (700 cells per well) cells were seeded in 6-well plates for 3 weeks. Cells were fixed with 4% (v/v) paraformaldehyde for 10 min, and then stained with crystal violet solution for 30 min. Stained cells were gently washed with water. Images were captured (AE2000, Panthera, China), and the number of colonies (>50 micrometers) was counted.
Cell migration assay
Transwell and cell scratch wound-healing assays were used to measure cell migration. For transwell assays, cells were resuspended in growth medium without FBS at a concentration of 2×10⁵ cells/mL, and added to the upper chamber of the Transwell insert (Jetbiofil, China), while full growth medium was added to the lower chamber. After 24 h of incubation at 37 °C in a CO₂ incubator, cells were fixed with 0.1% (m/v) paraformaldehyde and stained with crystal violet. Migrating cells were then counted by ImageJ (version 1.54) software (https://imagej.nih.gov/ij/). For scratch wound-healing assays, cells were seeded into 6-well plates at 5×105 cells/well and cultured to a 90% confluence. Cell surface was scratched using a P-200 pipette tip forming a “wound” across its diameter. Detached cells were removed by washing twice with PBS, fresh serum-free medium was then readded. Images of the scratched area were captured by a inverted microscope (AE2000, Panthera) at indicated time points. Images were quantified by ImageJ software.
RNA sequencing
Total RNA was extracted from HCT116 cells using the TRIzol reagent. For each treatment condition, three biological replicates were generated. mRNA was enriched from total RNA, processed to prepare cDNA libraries, and sequenced after quality control. Samples were sequenced using the DNBSEQ platform at Beijing Genomics Institute with PE150 (read length). Raw sequencing reads were trimmed and filtered for short sequences using SOAPnuke [25] (version 1.56) with parameters -l 20 -q 0.4 -n 0.011 -G. Trimmed FASTQ files were analyzed and quality-assessed using FastQC (version 0.12.1). Read alignment to the human reference genome (Gencode GRCh38) was performed using HISAT [26] (version 2.2.1) with default parameters. Bowtie2 [27] (version 2.3.4.3) was employed to map clean data to the reference gene set. Gene expression quantification was performed using RSEM [28] (version 1.3.1), and differential expression analysis was conducted with DESeq2 [29] (version 1.4.5), Limma [30] (version 3.58.1) and EdgeR [31] (version 4.0.16) in R (version 4.3.1). Gene set enrichment analysis (GSEA) [32] was performed using ClusterProfiler [33] (version 4.10.1).
Structural analysis and in silico modelling
Amino acid sequences of human PRSS22 (Q9GZN4), thrombin (P00734) and osteopontin (OPN, P10451) were obtained from Uniprot. Protein-protein interaction models between each serine protease (PRSS22, activated PRSS22 and thrombin) and osteopontin were generated using the AlphaFold3’s multimer prediction module.
Interaction interfaces were visualized and structurally aligned using PyMOL (version 3.1.4.1). PROtein binDIng enerGY prediction (PRODIGY) was employed to predict the binding affinity and dissociation constant of protein-protein complexes. PDBePISA was employed to quantify interface properties, including the solvation free energy gain upon formation of the interface, Accessible Surface Area, Buried Surface Area, Solvation energy effect and Buried area percentage were applied to assess the relative binding strengths between different serine proteases and osteopontin.
Molecular dynamics simulations were conducted using GROMACS (v2024.2) with the CHARMM36m force field (via CHARMM-GUI), where the activated PRSS22-osteopontin complex was embedded in a TIP3P water box containing 0.15 M NaCl. Following energy minimization and equilibration (NVT and NPT ensembles), an 80 ns production run was performed, with trajectories saved every 100 ps. Post-simulation analyses included: root-mean-square deviation (RMSD) of backbone atoms, root-mean-square fluctuation of active site residues, and hydrogen bond persistence. Free energy landscape (FEL) calculations based on principal component analysis were performed to characterize interactions between the PRSS22 catalytic triad and osteopontin cleavage motif, identifying dominant conformational states and dynamic interplays within the enzyme-substrate complex.
Statistical analysis
All experiments were performed independently for at least three times. Statistical method details are included in the figure legends. Statistical analyses were performed using GraphPad Prism 10.2.0 and R (version 4.3.1).
Results
PRSS22 is highly expressed in CRC
Given that inflammation is a major driver of CRC tumorigenesis, we sought to identify core genes that connect inflammatory processes with CRC progression and thereby serve as key regulators of CRC development. To this end, we analysed differential gene expression profiles in both CRC (TCGA) and colitis-associated cancer (CAC) datasets [23] (Fig. 1A). Intersecting the upregulated genes from these two datasets yielded 43 shared candidates (Fig. 1B, Fig. S1A, B). Among them, members of the MMP and CXCL family genes, as well as SPP1 and PRSS22, were prominently enriched. Notably, while the functions of several identified genes have been well studied, the role of PRSS22 in CRC remains poorly understood, prompting us to focus on its mechanistic and functional characterization in subsequent analyses.
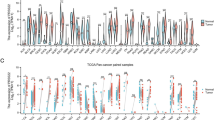
A Workflow illustrating the strategy of identifying core differentially expressed genes (DEGs) between tumour and normal tissues in the TCGA CRC cohort and in a murine CAC model [14, 23]. B Bar plot showing the top overlapping upregulated genes identified in the two models from (A). C mRNA expression levels of PDIAs in primary colon (n = 473) and rectal (n = 167) tumours compared with their respective benign tissues (n = 41 for colon; and n = 10 for rectal). Groups were compared using two-way ANOVA and Sidak’s multiple comparison tests. D Expression levels of PRSS22 in metastatic (MET) (n = 94) versus primary CRC (PT) (n = 39) from GSE41568 [15]. Groups were compared using unpaired t tests. E Kaplan–Meier curves for overall survival (OS) with high (n = 443) and low PRSS22 expression (n = 618) patient cohorts. F Kaplan–Meier curves for recurrence-free survival (RFS) with high (n = 883) and low PRSS22 expression (n = 453) patient cohorts. G, H ROC curve analyses of PRSS22 in colon (AUC = 0.983) and rectal (AUC = 0.919) cancers. I Analysis of a Spatially-resolved CRC primary tumour samples transcriptomic profile dataset illustrating spatial distribution of PRSS22 and the CRC marker CEACAM6 [11]. In all panels: *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
Based on the TCGA clinical transcriptomic dataset [34] and several independent cohorts [16,17,18,19,20], the mRNA level of PRSS22 was upregulated in primary CRC compared to non-malignant tissues (Fig. 1C, S1C–G). Analysis of a metastatic CRC dataset [15] also revealed that PRSS22 transcript levels were elevated in metastatic CRC compared to primary tumours (Fig. 1D). Survival analysis (Fig. 1E, F) revealed that high PRSS22 expression was correlated with poor overall survival (OS) and recurrence-free survival (RFS). Receiver Operating Characteristic (ROC) Curve analysis [14] (Fig. 1G, H) also demonstrated that high PRSS22 expression is positively associated with CRC positivity. High AUC values (0.983 for colon cancer and 0.919 for rectal cancer, respectively) on the ROC curves strongly indicated that PRSS22 can serve as a robust marker for the diagnosis of CRC. Finally, a single-cell spatial transcriptomic dataset [11] (Fig. 1I) revealed that PRSS22 was colocalized with the CRC marker CEACAM6.
PRSS22 enhances CRC growth and survival
The elevated expression of PRSS22 in CRC suggested that it may be important for CRC pathogenesis. Since PRSS22 expression appeared to be associated with Wnt/β-catenin signaling, to investigate the role of PRSS22 in CRC, we selected HCT116 cells, which possess an intact and constitutively active Wnt/β-catenin pathway and exhibit moderate endogenous PRSS22 expression, as the primary model for our functional experiments. To verify that the observed effects were not cell line–specific, Caco-2 cells, a differentiated, microsatellite-stable CRC line, were also included for comparison. HCT116 and Caco-2 cells were genetically modified with lentiviruses to express shRNAs targeting PRSS22 (shPRSS22). Knockdown of PRSS22 was confirmed by Western blotting (Fig. 2A) and qPCR (Fig. 2B). Silencing PRSS22 modestly suppressed the proliferation of (Fig. 2C) and evoked death (Fig. 2D) to the HCT116 cells. Representative live-cell imaging micrographs further illustrate these dynamic changes, showing slower cell confluence increase and stronger green fluorescence signals corresponding to apoptosis in shPRSS22 cells compared with shCtrl (Fig. 2E). Because our data (Fig. 2D, E) indicated that knocking down PRSS22 caused cytotoxicity, we also measured cell viability using Annexin V-propidium iodide (PI) assays. Annexin V binds to phosphatidylserine on the surface of cells that are undergoing programmed cell death, while PI stains the DNA base pairs of cells at mid/late programmed death stages where cell membrane integrity is impaired. Targeting PRSS22 caused modestly increase in cell death (Fig. 2F, G). We also found that the loss of PRSS22 indeed elicited profound tumor suppressive effects according to colony formation assays (Fig. 2H, I). Silencing of PRSS22 also inhibited growth of and evoked death to HCT116 and Caco-2 cells (Fig. S2B, D). Conversely, PRSS22 overexpressing cells exhibited a slightly elevated growth rate and improved cell viability (Fig. S2A, C). Next, we used the three-dimensional transwell migration system to assess the ability of shCtrl- or shPRSS22-expressing HCT116 cells to migrate through an extracellular matrix (Fig. 2J, K). Knocking down the PRSS22 gene impaired the migratory capacity of these cells. Scratch assays (Fig. 2L, M) also corroborated with our findings in the transwell assays, as loss of PRSS22 slowed down cell migration into the “wounded” area.
HCT116 cells stably expressing shCtrl or 2 different shRNAs against PRSS22 were lysed and subjected to: A immunoblotting. PRSS22:actin intensity ratio quantified; B qPCR (n = 3); C CCK-8 assay (n = 10); D, E Live-cell imaging using IncuCyte S1 (n = 20) to monitor cell growth and cell death; representative time-lapse images are shown in (E); F, G Annexin V-propidium iodide (PI) staining followed by flow cytometry analysis in (G) and quantification in (F) (n = 3); Colony formation assay, with representative images in (H) and quantification in (I) (n = 3); J Transwell migration assay, quantified in (K) (n = 4); L Scratch wound-healing assay, quantified in (M) (n = 12). N Subcutaneous xenograft assay, tumor size quantified in (O). In all panels: Data are means ± SD. Groups were compared using two-way ANOVA and Dunnett’s multiple comparison tests. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
Having demonstrated the importance of PRSS22 for CRC cell growth and survival in vitro, we next undertook xenograft studies to evaluate its relevance to tumour growth in vivo. When grown as subcutaneous xenografts, knocking down PRSS22 substantially reduced tumour weight and volume (Fig. 2N, O). Altogether, these in vitro and in vivo experiments demonstrate that PRSS22 plays an important role in CRC cell growth and survival.
PRSS22 improves CRC cell survival via inhibition of HMOX1-mediated ferroptosis
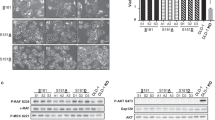
Having established its important role in regulating CRC cell growth and viability, we focused on exploring specific mechanisms by which PRSS22 influences cell fate. Transcriptomic analysis identified 179 differentially regulated genes between shCtrl- and shPRSS22-expressing cells, 131 of these genes were upregulated whereas 48 were downregulated in shPRSS22 cells (Fig. 3A, B). ‘Over-representation analysis’ (ORA) of these genes revealed that knocking down PRSS22 enriched the expression of genes implicated in ferroptosis and glutathione metabolism, whereas the expression of genes involved in oncogenic signalling pathways, such as MAPK, Hippo and Wnt (Fig. 3C), were downregulated. Findings from GSEA were generally consistent with the ORA outputs and further revealed that PRSS22 also plays a role in the maintenance of cell REDOX homeostasis, as evidenced by an increase in expression of genes implicated in the reactive oxygen species (ROS) and the heme metabolism pathways (Fig. 3D–F). To confirm these predictions experimentally, we treated shPRSS22 and shCtrl HCT116 cells with a panel of cell-death inhibitors, including ferrostatin-1 (Fer-1), Z-VAD-FMK, Necrostatin-1, belnacasan and 3-methyladenine (3-MA), alongside hemin (heme analog) as a positive control. CCK-8 assays showed that only Fer-1 rescued the reduction in cell viability caused by PRSS22 knockdown or hemin treatment, whereas combining shPRSS22 with hemin further exacerbated cell death, consistent with the hypothesis that PRSS22 loss sensitizes cells to ferroptosis driven by excessive heme metabolism (Fig. 3G). In a complementary assay using a ferroptosis detection kit, we visualized intracellular Fe²⁺ and ROS levels by fluorescence microscopy and quantitative fluorimetry. shPRSS22 cells exhibited marked increases in both Fe²⁺ accumulation and ROS intensity relative to controls. Fer-1 treatment effectively reduced ROS fluorescence but did not significantly alter Fe²⁺ levels, confirming that PRSS22 depletion induces ferroptotic stress through excessive iron loading and lipid peroxidation (Fig. 3H, I). The enzyme heme oxygenase-1 (HO-1), encoded by HMOX1 whose expression was induced by shPRSS22 (Fig. 3A), releases Fe2+ from heme, and may in turn elevated ROS levels and triggered ferroptosis. Notably, the expression of HMOX1 (Fig. S3A) and genes involved in heme metabolism (Fig. S3B) was markedly reduced in primary CRC tumours from the TCGA cohort. Immunoblotting analysis also confirmed that HMOX1 protein level was indeed increased in shPRSS22 cells (Fig. 3J). Exogenously overexpressing HMOX1 significantly inhibited HCT116 cell growth (Fig. 3K, L). Treatment of shPRSS22-expressing cells with the HMOX1 inhibitor Heme Oxygenase-1-IN-1 partially rescued the cell growth inhibitory effect of PRSS22 knockdown (Fig. 3M). Altogether, these findings elucidated that the loss of PRSS22 impaired CRC cell growth and viability by evoking HMOX1-mediated ferroptosis.
A–F RNA lysates from HCT116 cells stably expressing shCtrl or shPRSS22 were subjected to transcriptomic analysis. The following data analyses were performed: A Volcano plot displaying differentially expressed genes. Values with p < 0.05 and log2(fold change) >1 were considered as significant and were labelled in either blue (upregulated in shCtrl) or red (upregulated in shPRSS22). B Heatmap illustration of differentially regulated genes. C ORA of the differentially abundant genes. D–F GSEA. NES normalised enrichment score. G HCT116 cells stably expressing shCtrl or shPRSS22 were treated with hemin alone or in combination with ferrostatin-1 (Fer-1) (2 μM), Z-VAD-FMK (20 μM), Necrostatin-1 (20 μM), belnacasan (20 μM) and 3-methyladenine (3-MA) (60 μM), respectively. After 48 h of treatment cell viability was assayed. Data are means ± SD (n = 3). Groups were compared using two-way ANOVA and Tukey’s multiple comparison tests. H, I Detection of intracellular Fe²⁺ accumulation and ROS generation in HCT116 cells using a ferroptosis detection kit. Fluorescence microscopy was used to visualize Fe²⁺ (RhoNox-6) and ROS (CM-H2DCFDA) signals. Representative images in (I) and quantification of fluorescence intensity in (H) are shown. Data are means ± SD (n = 3). Groups were compared using two-way ANOVA and Tukey’s multiple comparison tests. J, K Western blotting analysis of HCT116 cells stably expressing shCtrl or shPRSS22. Cells were also transfected with an empty vector or a C-terminal V5-tagged HMOX1 (HMOX1-V5) for 48 h in (K). L CCK-8 assay of cells in (K). M CCK-8 assay of shPRSS22 cells treated with either vehicle or HO1 inhibitor (Heme Oxygenase-1-IN-1). Data are means ± SD (n = 10). Groups were compared using two-way ANOVA and Dunnett’s multiple comparison tests. In all panels: *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
PRSS22 positively regulates cell migration through the cleavage of osteopontin (OPN)
OPN is a multifunctional glycoprotein encoded by the SPP1 gene [35]. Splicing variants of OPN have been observed in multiple human cancers [36,37,38]. For example, full length OPN can be cleaved by the serine protease thrombin at a conserved site (R159GDSVVYGLRSK170), yielding two active fragments: an N-terminal fragment (M1-R168), which is subsequently processed into another active variant (M1-L167), and a C-terminal fragment [39]. Thrombin-cleaved OPN fragments were shown to promote cell adhesion, migration, motility and immune responses in a rat glioma model [40]. Given that SPP1, the gene encoding OPN, was also identified as a core upregulated gene in our initial datamining analysis (Fig. 1A, B), which was significantly elevated in CRC tissues, and was strongly associated with poor prognosis (Fig. 1B, Fig. S4A, B), we investigated the potential interaction between OPN and PRSS22, where the latter is also a serine protease like thrombin. As both thrombin and PRSS22 belong to the trypsin family of serine proteases, PRSS22 may also catalyze OPN cleavage to accelerate CRC cell migration. To test this hypothesis, we first used AlphaFold3 to predict the catalytically active sites of thrombin and PRSS22 (Fig. 4A, B). Trombin’s active site consists of 3 key amino acid residues: His-406, Asp-462 and Ser-568. Similarly, PRSS22’s catalytic triad likely consists of His-90, Asp-141 and Ser-242. Furthermore, PRSS22 undergoes self-activation cleavage at Arg-49 to generate an active fragment (PRSS22*). Based on this information, AlphaFold3 multimeric modelling and PRODIGY affinity prediction suggested a similar binding model between thrombin-OPN, PRSS22-OPN and PRSS22*-OPN at their corresponding active sites (Fig. 4A, B). Active PRSS22 exhibits a strong binding affinity for OPN (-15.9 kcal/mol; Kd = 2.2 × 10⁻¹² M), similar to the case of thrombin (-19.5 kcal/mol; Kd = 4.8 × 10⁻¹⁵ M). PDBePISA analysis further elucidated that the predicted active PRSS22*-OPN complex has a high solvation free energy gain and a similar buried surface area at the binding interface as thrombin-OPN. In contrast, the inactivated full-length PRSS22-OPN complex is weaker in solvation free energy gain than the PRSS22*-OPN complex (Fig. 4C, D). Immunoblotting analysis validated our in silico observations as shPRSS22-expressing cells were unable to evoke OPN cleavage (Fig. 4E).
A, B Protein–protein interaction models of OPN binding to active PRSS22 or thrombin, as predicted by AlphaFold3. Binding energy magnitudes were predicted by PRODIGY. C, D Quantitative analysis of the binding affinity and interface properties using PDBePISA. E Western blotting analysis of OPN cleavage in HCT116 cells stably expressing shCtrl or shPRSS22. F, G Molecular dynamics simulations of the interaction between active PRSS22 and OPN. RMSD (F): root-mean-square deviation; FEL (G): free energy landscape. H qPCR analysis of HCT116 cells stably overexpressing PRSS22 (n = 3). Groups were compared using unpaired t tests. I Western blotting analysis of HCT116 cells transfected with a plasmid expressing C-terminal V5-tagged OPN. J, K Scratch wound-healing assay of HCT116 cells stably overexpressing PRSS22 and/or transiently overexpressing OPN for 48 h. Data represent means ± SD (n = 3). Groups were compared using two-way ANOVA and Tukey’s multiple comparison tests. In all panels: *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
To further substantiate our findings that PRSS22 catalyzes OPN cleavage, we also performed molecular dynamics simulations with RMSD and FEL, which showed that OPN cleavage motif stably binds to the active PRSS22 catalytic triad pocket (Fig. 4F, G, Fig. S4C–F). To further delineate how OPN contributes to the oncogenic properties of PRSS22 in CRC, we created DNA vectors expressing OPN and HCT116 cell lines stably overexpressing PRSS22 (Fig. 4H, I). Exogenously expressing OPN in PRSS22-overexpressing cells exhibited the strongest pro-migratory capacity compared to when overexpressing either OPN or PRSS22 alone (Fig. 4J, K). Collectively, these results strongly suggest that increased levels of PRSS22 may aid CRC cell migration by cleaving OPN.
Increased PRSS22 was associated with ICT by inducing macrophage polarization
ICT is a key tumorigenesis process during CRC development [41]. To study the potential role of PRSS22 during colonic ICT, we first examined PRSS22 mRNA expression level in clinical samples; these samples comprised a meta-analysis of publicly available RNA-seq datasets from 26 independent studies, encompassing a variety of tissues [21]. We noted that PRSS22 expression was low in healthy individuals and patients without inflammatory bowel disease (noneIBD), yet elevated in patients suffering ulcerative colitis (UC) or Crohn’s disease (CD), and was peaked in colorectal tumour samples (Fig. 5A), implying a strong correlation between PRSS22 expression levels and tissue inflammation. Similarly, in a CAC mouse model, PRSS22 mRNA expression levels were low in the control group, gradually increased during the inflammatory phase (at weeks 2 and 4), and peaked when tumor was developed (at weeks 7 and 10) [23] (Fig. 5B). In a short-term, dextran sulphate sodium (DSS)-induced, acute colitis mouse model [22], PRSS22 mRNA expression level was peaked on day 6 during the inflammation induction phase, and was gradually declined during the recovery phase (after day 7) (Fig. 5C). Single sample GSEA (ssGSEA) of shPRSS22-expressing CRC cells (Fig. 3A) revealed that the loss of PRSS22 significantly reduced the infiltration of pro-oncogenic M2 macrophages (Fig. 5D). Analysis of a CRC single-cell transcriptomic dataset [10] also showed that patients with high PRSS22 mRNA expression levels had higher M2 scores (Fig. 5E, F) and proportion of SPP1⁺ macrophages (Fig. 5G). SPP1⁺ subtype of macrophages are known to predominate in autologous samples from CRC patients with liver metastasis, are highly proangiogenic and thus closely associated with CRC metastasis [42]. In support of this notion, co-culture of HCT116 cells and THP-1-derived macrophages unraveled that knocking down PRSS22 significantly reduced HCT116 cells’ ability to induce M0-to-M2 polarization in macrophages (Fig. 5H, I). To further explore the tumor-derived factors responsible for this immunomodulatory effect, we revisited our transcriptomic data and identified transforming growth factor beta (TGFβ) as one of the most significantly downregulated cytokines following PRSS22 depletion (Fig. 5J). Consistent with this, intracellular ELISA quantification confirmed a marked decrease of TGFβ protein levels in shPRSS22 HCT116 cells compared with controls (Fig. 5K). Given that TGFβ functions both as an upstream inducer and a downstream marker of M2 macrophage polarization, these findings suggest that PRSS22 promotes an immunosuppressive, M2-dominant tumor microenvironment in CRC at least in part through regulation of tumor-cell–derived TGFβ.
A PRSS22 mRNA expression in intestinal tissue samples from normal individuals (n = 41), non-IBD (n = 50), UC (n = 74), CD (n = 127) and CRC groups (n = 473) in the TCGA and the TaMMa patient cohorts [14, 21]. B Time-course analysis of PRSS22 expression in a CAC mouse model. Data are means ± SD (n = 3). Groups were compared using two-way ANOVA and Dunnett’s multiple comparison tests. C Time-course analysis of PRSS22 expression in an acute DSS-induced colitis mouse model [22]. Data are means ± SD (n = 3). D ssGSEA of transcriptomic data presented in Fig. 3A–F. Data are means ± SD (n = 3). Groups were compared using two-way ANOVA and Sidak’s multiple comparison tests. E–G Analysis of a CRC patient tumour sample single-cell transcriptomic dataset [10]. E UMAP clustering showing differential PRSS22 expression levels at a single-cell level. F M2 macrophage scores in PRSS22-high (n = 88) and low groups (n = 104). G Proportion of SPP1⁺ and other subtypes of macrophages in PRSS22-high (n = 88) and low (n = 104) groups. H, I qPCR analysis of M1 and M2 macrophage markers following the co-culture of THP-1–derived macrophages and HCT116 cells stably expressing shCtrl or shPRSS22. Data represent means ± SD (n = 3). Groups were compared using two-way ANOVA and Sidak’s multiple comparison tests. J Bar plot showing TGFβ expression from transcriptomic data presented in Fig. 3A–F. Data are means ± SD (n = 3). Groups were compared using unpaired t tests. K Intracellular enzyme-linked immunosorbent assay (ELISA) quantification of TGFβ protein levels in HCT116 cells stably expressing shCtrl or shPRSS22. Data are means ± SD (n = 3). Groups were compared using unpaired t tests. In all panels: *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
In conclusion, these data demonstrated that elevated expression of PRSS22 was associated with intestinal inflammation and M2 macrophage polarization, which may aid colorectal ICT.
Discussion
Increasing evidence supports the notion that serine proteases play a crucial role in reshaping cancer proteome to aid tumorigenesis [43] and thus represent a therapeutic vulnerability. However, progress in this field has been hampered because the diversity and scope of protease substrates means that they have pleiotropic effects, which in turn has made it difficult to precisely pinpoint the cancer-promoting mechanisms of these enzymes. Recent reports suggest that the serine protease PRSS22 plays a role in the development of CRC, influencing the proliferation, migration, and invasive capabilities of CRC cells and correlating with recurrence risk following therapeutic surgery [8, 9], yet prior to this study, the underlying mechanism remains to be elucidated. Ours is the first study to undertake integrative functional characterization and to provide mechanistic insights of how PRSS22 promotes CRC tumorigenesis. Findings presented herein highlight the therapeutic potential of targeting PRSS22 in CRC treatment, and will have broad impact in terms of influencing research into serine protease inhibitors as a therapeutic strategy in other cancers/diseases. PRSS22 was initially discovered in human airway epithelial cells and expressed in a developmentally regulated manner [4]. It is highly expressed in liver and breast cancers, aids cancer metastasis without affecting cancer cell growth [6, 7]. In this study, we demonstrated that PRSS22 expression is significantly upregulated in IBD and CRC patients, plays an important role in colorectal ICT and is critical for CRC cell growth, survival and migration.
An unbiased transcriptomic approach revealed that PRSS22 deficiency led to a significant increase in HMOX1, activated ROS- and ferroptosis-associated pathways. HMOX1-mediated heme degradation can elevate intracellular Fe2+ levels and induce lipid peroxidation, which are hallmarks to ferroptosis [44]. Conversely, elevated PRSS22 levels may aid CRC cell survival by preventing HMOX1-driven ferroptosis during tumorigenesis. In agreement to our findings, small molecule compounds such as tagitinin C and polysaccharides from maggot extracts are shown to increase HMOX1 expression to induce ferroptosis in CRC [45, 46]. It is also noteworthy that HMOX1 levels are decreased in CRC patients and may thus confer resistance to cancer cells against chemotherapies [47]. It is tempting to speculate that HMOX1 inducers can be a promising anti-CRC strategy, which certainly warrants further investigations.
OPN cleavage, catalyzed by serine proteases such as thrombin, is known to trigger a range of pro-oncogenic cell properties including growth, pro-survival, migration and immune escape [40, 48, 49]. We noted that the thrombin gene is scarcely expressed in CRC cells, whereas PRSS22 serves as a crucial catalytic factor for OPN cleavage in these cells. Our in silico analysis indicated that the PRSS22-OPN interaction model and the corresponding OPN cleaved site resembles that of thrombin-OPN. Future structural analyses of the PRSS22-OPN complex will be important to further delineate subtle differences in the mode of action between PRSS22 and thrombin against OPN and other serine protease substrates.
The immune microenvironment surrounding cancer cells is under gradual transformation during ICT, this is a process also known to be regulated by the serine proteases, and thus prompted us to investigate the role of PRSS22 in modulating immune response in a CRC context. Macrophage polarization is a key immune response process during ICT, which can be influenced by a plethora of factors. Polarized and thus active macrophages are typically categorized into two major types: M1 macrophages are primarily pro-inflammatory, whereas M2 macrophages are anti-inflammatory in response to cytokines [50]. Macrophages associated with the tumor tissues do not strictly adhere to an M1 or an M2 subtype, but rather resembles M2 macrophages to facilitate immune suppression [51]. Beyond the suppressive effects against ROS production and ferroptosis, and acting as a pro-cell migratory factor, we found that PRSS22 also encouraged macrophage polarization toward an M2-like phenotype. Mechanistically, our data revealed that PRSS22 positively regulates tumor-intrinsic TGFβ expression, a cytokine that exerts pleiotropic immunoregulatory functions within the tumor microenvironment. TGFβ serves as both an upstream inducer of M2 macrophage polarization and a downstream marker of fully polarized M2 macrophages, forming an autocrine–paracrine feedback loop that amplifies immunosuppressive signaling. This dual role of TGFβ may explain how PRSS22-driven TGFβ upregulation reinforces an M2-dominant microenvironment, favoring immune evasion and tumor growth. SPP1, which encodes OPN, is specifically linked with a subtype of tumour-associated macrophages (TAMs) [52]. It has been suggested that TAMs would be better defined according to the expression levels of CXCL9 and SPP1 instead of the traditional M1/M2 classification system [52]. High levels of SPP1+ macrophages confer resistance to immune therapies and predicts poor prognosis, such as head and neck squamous cell carcinoma, hepatocellular carcinoma and metastatic castration-resistant prostate cancer [53,54,55]; they are also associated with liver metastasis in CRC [42, 56]. Mining of public single cell and spatial transcriptomic datasets unraveled that high levels of PRSS22 positively correlates with increased proportions of SPP1⁺ macrophages. Collectively, our findings established PRSS22 as a key pro-oncogenic factor that aids CRC cell against host immune response. Although high levels of PRSS22 favors an immune microenvironment shifting toward an “anti-inflammatory” state, which is cytoprotective during IBD, yet such “benefits” can be exploited by tumor stem cells to escape immune attack under ICT.
In summary, this study revealed PRSS22 as a key driver of the onset and progression of CRC in the context of chronic inflammation (Fig. 6). PRSS22 mRNA levels are barely detectable in normal colorectal tissues, implying that it might be dispensable for the growth of normal/healthy cells, inhibition of PRSS22 thus represents a therapeutic potential against CRC. However, currently there is no PRSS22-specific inhibitor available and therefore should be a priority for future research for a prompt clinical translation.
Within the context of chronic inflammation, PRSS22 expression is progressively upregulated in colorectal epithelial cells. Elevated PRSS22 suppresses ferroptosis by inhibiting HMOX1-mediated iron release and ROS accumulation, thereby supporting tumor cell survival. Concurrently, PRSS22 proteolytically cleaves OPN into active fragments that enhance the migratory and metastatic potential of CRC cells. In the tumor microenvironment, PRSS22 promotes the polarization of macrophages toward an immunosuppressive M2-like phenotype, particularly increasing the proportion of SPP1⁺ macrophages, which further contributes to immune evasion and poor prognosis. The schematic illustration was created using Adobe Illustrator (version 27.5).
Data availability
RNA-seq data are available at the Gene Expression Omnibus database (GSE308833).
References
Schmitt M, Greten FR. The inflammatory pathogenesis of colorectal cancer. Nat Rev Immunol. 2021;21:653–67.
Dekker E, Tanis PJ, Vleugels JLA, Kasi PM, Wallace MB. Colorectal cancer. Lancet. 2019;394:1467–80.
Eatemadi A, Aiyelabegan HT, Negahdari B, Mazlomi MA, Daraee H, Daraee N, et al. Role of protease and protease inhibitors in cancer pathogenesis and treatment. Biomed Pharmacother. 2017;86:221–31.
Wong GW, Yasuda S, Madhusudhan MS, Li L, Yang Y, Krilis SA, et al. Human tryptase epsilon (PRSS22), a new member of the chromosome 16p13.3 family of human serine proteases expressed in airway epithelial cells. J Biol Chem. 2001;276:49169–82.
Mitsui S, Okui A, Kominami K, Konishi E, Uemura H, Yamaguchi N. A novel serine protease highly expressed in the pancreas is expressed in various kinds of cancer cells. FEBS J. 2005;272:4911–23.
Chen CY, Chung IH, Tsai MM, Tseng YH, Chi HC, Tsai CY, et al. Thyroid hormone enhanced human hepatoma cell motility involves brain-specific serine protease 4 activation via ERK signaling. Mol Cancer. 2014;13:162.
Song L, Li H, Ma RR, Liu S, Zhang GH, Guo XY, et al. E2F1-initiated transcription of PRSS22 promotes breast cancer metastasis by cleaving ANXA1 and activating FPR2/ERK signaling pathway. Cell Death Dis. 2022;13:982.
Xu C, Zhou Z, Zhu D, Zhang Q, Zhong S, Li Z. Single-cell transcriptomics in colorectal cancer uncover the potential of metastasis and immune dysregulation of a cell cluster overexpressed PRSS22. Front Immunol. 2025;16:1586428.
Mohamed A, AbdelMageed M, Zahran F, Zein N, Olsson L, Lindmark G, et al. Combined serine protease PRSS22 and CEA mRNA analysis identifies the majority of colon cancer patients that recur within 12 years. Front Oncol. 2025;15:1628069.
Chu X, Li X, Zhang Y, Dang G, Miao Y, Xu W, et al. Integrative single-cell analysis of human colorectal cancer reveals patient stratification with distinct immune evasion mechanisms. Nat Cancer. 2024;5:1409–26.
Oliveira MF, Romero JP, Chung M, Williams SR, Gottscho AD, Gupta A, et al. High-definition spatial transcriptomic profiling of immune cell populations in colorectal cancer. Nat Genet. 2025;57:1512–23.
Gyorffy B. Integrated analysis of public datasets for the discovery and validation of survival-associated genes in solid tumors. Innovation. 2024;5:100625.
Cheng S, Li Z, Gao R, Xing B, Gao Y, Yang Y, et al. A pan-cancer single-cell transcriptional atlas of tumor infiltrating myeloid cells. Cell. 2021;184:792–809.e723.
Liao C, Wang X. TCGAplot: an R package for integrative pan-cancer analysis and visualization of TCGA multi-omics data. BMC Bioinformatics. 2023;24:483.
Lu M, Zessin AS, Glover W, Hsu DS. Activation of the mTOR pathway by oxaliplatin in the treatment of colorectal cancer liver metastasis. PLoS ONE. 2017;12:e0169439.
Hanley MP, Hahn MA, Li AX, Wu X, Lin J, Wang J, et al. Genome-wide DNA methylation profiling reveals cancer-associated changes within early colonic neoplasia. Oncogene. 2017;36:5035–44.
Fu J, Tang W, Du P, Wang G, Chen W, Li J, et al. Identifying microRNA-mRNA regulatory network in colorectal cancer by a combination of expression profile and bioinformatics analysis. BMC Syst Biol. 2012;6:68.
Martin ML, Zeng Z, Adileh M, Jacobo A, Li C, Vakiani E, et al. Logarithmic expansion of LGR5(+) cells in human colorectal cancer. Cell Signal. 2018;42:97–105.
Sheffer M, Bacolod MD, Zuk O, Giardina SF, Pincas H, Barany F, et al. Association of survival and disease progression with chromosomal instability: a genomic exploration of colorectal cancer. Proc Natl Acad Sci USA 2009;106:7131–6.
Szigeti KA, Kalmar A, Galamb O, Valcz G, Bartak BK, Nagy ZB, et al. Global DNA hypomethylation of colorectal tumours detected in tissue and liquid biopsies may be related to decreased methyl-donor content. BMC Cancer. 2022;22:605.
Massimino L, Lamparelli LA, Houshyar Y, D’Alessio S, Peyrin-Biroulet L, Vetrano S, et al. The inflammatory bowel disease transcriptome and metatranscriptome meta-analysis (IBD TaMMA) framework. Nat Comput Sci. 2021;1:511–5.
Czarnewski P, Parigi SM, Sorini C, Diaz OE, Das S, Gagliani N, et al. Conserved transcriptomic profile between mouse and human colitis allows unsupervised patient stratification. Nat Commun. 2019;10:2892.
Chen L, Luo Z, Zhao C, Li Q, Geng Y, Xiao Y, et al. Dynamic chromatin states coupling with key transcription factors in colitis-associated colorectal cancer. Adv Sci. 2022;9:e2200536.
Xie J, Shen K, Liang W, Kuang Z, Shrestha RK, Hanson AR, et al. Protein disulfide isomerases regulate androgen receptor stability and promote prostate cancer cell growth and survival. Proc Natl Acad Sci USA 2025;122:e2509222122.
Li R, Li Y, Kristiansen K, Wang J. SOAP: short oligonucleotide alignment program. Bioinformatics. 2008;24:713–4.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9.
Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics. 2011;12:323.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005;102:15545–50.
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation. 2021;2:100141.
Cancer Genome Atlas N. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–7.
Gimba ER, Tilli TM. Human osteopontin splicing isoforms: known roles, potential clinical applications and activated signaling pathways. Cancer Lett. 2013;331:11–7.
Shi Z, Mirza M, Wang B, Kennedy MA, Weber GF. Osteopontin-a alters glucose homeostasis in anchorage-independent breast cancer cells. Cancer Lett. 2014;344:47–53.
Bastos A, Blunck CB, Emerenciano M, Gimba ERP. Osteopontin and their roles in hematological malignancies: Splice variants on the new avenues. Cancer Lett. 2017;408:138–43.
Tang X, Li J, Yu B, Su L, Yu Y, Yan M, et al. Osteopontin splice variants differentially exert clinicopathological features and biological functions in gastric cancer. Int J Biol Sci. 2013;9:55–66.
Jung S, Ha J, Park JH, Yoo KH. Decoding SPP1 regulation: genetic and nongenetic insights into its role in disease progression. Mol Cells. 2025;48:100215.
Leung LL, Myles T, Morser J. Thrombin cleavage of osteopontin and the host anti-tumor immune response. Cancers. 2023;15:3480.
Keum N, Giovannucci E. Global burden of colorectal cancer: emerging trends, risk factors and prevention strategies. Nat Rev Gastroenterol Hepatol. 2019;16:713–32.
Liu Y, Zhang Q, Xing B, Luo N, Gao R, Yu K, et al. Immune phenotypic linkage between colorectal cancer and liver metastasis. Cancer Cell. 2022;40:424–37.e425.
Song R, Qiao W, He J, Huang J, Luo Y, Yang T. Proteases and their modulators in cancer therapy: challenges and opportunities. J Med Chem. 2021;64:2851–77.
Hassannia B, Vandenabeele P, Vanden Berghe T. Targeting ferroptosis to iron out cancer. Cancer Cell. 2019;35:830–49.
Wei R, Zhao Y, Wang J, Yang X, Li S, Wang Y, et al. Tagitinin C induces ferroptosis through PERK-Nrf2-HO-1 signaling pathway in colorectal cancer cells. Int J Biol Sci. 2021;17:2703–17.
Tang X, Mo D, Jiang N, Kou Y, Zhang Z, Peng R, et al. Polysaccharides from maggot extracts suppressed colorectal cancer progression by inducing ferroptosis via HMOX1/GPX4 signaling pathway. Int J Biol Macromol. 2025;296:139734.
Rah B, Shafarin J, Karim A, Bajbouj K, Hamad M, Muhammad JS. Iron overloading potentiates the antitumor activity of 5-fluorouracil by promoting apoptosis and ferroptosis in colorectal cancer cells. Cell Biochem Biophys. 2024;82:3763–80.
Liu C, Xia S, Wang B, Li J, Wang X, Ren Y, et al. Osteopontin promotes tumor microenvironment remodeling and therapy resistance. Cancer Lett. 2025;617:217618.
Xue YH, Zhang XF, Dong QZ, Sun J, Dai C, Zhou HJ, et al. Thrombin is a therapeutic target for metastatic osteopontin-positive hepatocellular carcinoma. Hepatology. 2010;52:2012–22.
Yunna C, Mengru H, Lei W, Weidong C. Macrophage M1/M2 polarization. Eur J Pharmacol. 2020;877:173090.
Mehla K, Singh PK. Metabolic regulation of macrophage polarization in cancer. Trends Cancer. 2019;5:822–34.
Bill R, Wirapati P, Messemaker M, Roh W, Zitti B, Duval F, et al. CXCL9:SPP1 macrophage polarity identifies a network of cellular programs that control human cancers. Science. 2023;381:515–24.
Su X, Liang C, Chen R, Duan S. Deciphering tumor microenvironment: CXCL9 and SPP1 as crucial determinants of tumor-associated macrophage polarity and prognostic indicators. Mol Cancer. 2024;23:13.
Liu Y, Xun Z, Ma K, Liang S, Li X, Zhou S, et al. Identification of a tumour immune barrier in the HCC microenvironment that determines the efficacy of immunotherapy. J Hepatol. 2023;78:770–82.
Lyu A, Fan Z, Clark M, Lea A, Luong D, Setayesh A, et al. Evolution of myeloid-mediated immunotherapy resistance in prostate cancer. Nature. 2025;637:1207–17.
Sathe A, Mason K, Grimes SM, Zhou Z, Lau BT, Bai X, et al. Colorectal cancer metastases in the liver establish immunosuppressive spatial networking between tumor-associated SPP1+ macrophages and fibroblasts. Clin Cancer Res. 2023;29:244–60.
Acknowledgements
The authors would like to thank Mr. Peiwen Zhou (South China University of Technology) for assistance with lentiviral experiments. This work was supported by: Australian NHMRC Ideas Grant (2047980 to JX); South China University of Technology’s Double First Class Initiative Fund (to JX); the Program for Guangdong Introducing Innovative and Entrepreneurial Teams (2019ZT08Y318 to JX); the Flinders Foundation (to JX); Masonic Charities Trust via the Freemasons Centre for Male Health & Wellbeing (to JX); Shanghai Youth Top Talent Program of Eastern Talent Plan (to KS).
Author information
Authors and Affiliations
Contributions
ZK, KS and JX conceived the idea, analysed the data, and wrote the paper with input from all authors. ZK performed most of the experiments. QS and KS designed and performed the in vivo xenograft experiments. WL designed and performed some lentiviral and macrophage experiments. KS and JX provided supervision. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All methods were performed in accordance with the relevant guidelines and regulations. All animal procedures were approved by Shanghai University of Traditional Chinese Medicine’s Animal Ethics Committee (Approval Number: PZSHUTCM2406110018). No human subjects were involved in this study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kuang, Z., Su, Q., Liang, W. et al. PRSS22 inhibits HMOX1-mediated ferroptosis and induces osteopontin cleavage to promote M2 macrophage polarization and colitis-associated carcinogenesis. Oncogene 45, 1275–1286 (2026). https://doi.org/10.1038/s41388-026-03729-5
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-026-03729-5