Abstract
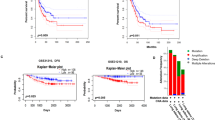
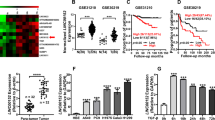
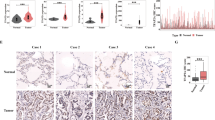
Metastasis remains the primary cause of mortality in lung adenocarcinoma (LUAD) patients. However, the molecular mechanisms underlying LUAD cell metastasis are only partially elucidated. Here, by performing integrated bioinformatic analysis of clinical data, RNA-binding protein (RBP) NOVA2 is identified as a pivotal LUAD metastasis-associated regulator. NOVA2 expression is elevated in metastatic LUAD tissues and correlates with poor prognosis of LUAD patients. Functionally, NOVA2 depletion suppresses epithelial-mesenchymal transition (EMT), migration, and invasion in vitro, and attenuates LUAD cell metastasis in vivo. Mechanistically, histone acetyltransferase p300 augments H3K27 acetylation level and facilitates the binding of STAT3 to the NOVA2 promoter, which in turn promotes NOVA2 transcription. Increased NOVA2 expression induces exon skipping (exons 6–7) in SMAD4 to generate a truncated splicing isoform (termed Δ-SMAD4). The resulting Δ-SMAD4 isoform evades E3 ubiquitin ligase β-TrCP-mediated ubiquitination, maintaining its ability to form complex with SMAD3 (R-SMAD) and sustain TGF-β/SMAD signaling. Moreover, in NOVA2-overexpressing LUAD cells, Δ-SMAD4 knockdown has stronger inhibitory effects on TGF-β-induced EMT and invasion than does SMAD4 knockdown. In summary, our findings identify a novel mechanism by which STAT3-mediated transcriptional upregulation of NOVA2 promotes SMAD4 splicing in metastatic LUAD, and suggest that the STAT3-NOVA2-Δ-SMAD4 axis drives EMT and LUAD metastasis, which may be a promising therapeutic target for treating LUAD.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data generated in Fig. 1A are retrieved from EuRBPDB [26], ATtRACT [27], CancerSEA [28], and prognostic gene signatures [29] online databases. The gene expression correlation analyzed in Fig. 4F, S3E and S3F are available from GEPIA database (http://gepia.cancer-pku.cn/). The RNA-seq data generated in Figs. 5A and 6A are included in supplementary materials, and all other raw data are available upon request from the corresponding author.
References
Chaffer CL, Weinberg RA. A perspective on cancer cell metastasis. Science. 2011;331:1559–64.
Brandt WS, Yan W, Zhou J, Tan KS, Montecalvo J, Park BJ. et al. Outcomes after neoadjuvant or adjuvant chemotherapy for cT2-4N0-1 non-small cell lung cancer: A propensity-matched analysis. J Thorac Cardiovasc Surg. 2019;157:743–753.e743.
Lou F, Huang J, Sima CS, Dycoco J, Rusch V, Bach PB. Patterns of recurrence and second primary lung cancer in early-stage lung cancer survivors followed with routine computed tomography surveillance. J Thorac Cardiovasc Surg. 2013;145:75–81.
Esposito M, Ganesan S, Kang Y. Emerging strategies for treating metastasis. Nat Cancer. 2021;2:258–70.
Watanabe K, Tsuboi M, Sakamaki K, Nishii T, Yamamoto T, Nagashima T, et al. Postoperative follow-up strategy based on recurrence dynamics for non-small-cell lung cancer. Eur J Cardiothorac Surg. 2016;49:1624–31.
Cancer Genome Atlas Research Network. Comprehensive molecular profi ling of lung adenocarcinoma. Nature. 2014;511:543–50.
Sterne-Weiler T, Sanford JR. Exon identity crisis: disease-causing mutations that disrupt the splicing code. Genome Biol. 2014;15:201.
Yang X, Coulombe-Huntington J, Kang S, Sheynkman GM, Hao T, Richardson A, et al. Widespread expansion of protein interaction capabilities by alternative splicing. Cell. 2016;164:805–17.
Chabot B, Shkreta L. Defective control of pre-messenger RNA splicing in human disease. J Cell Biol. 2016;212:13–27.
Lee SC, Abdel-Wahab O. Therapeutic targeting of splicing in cancer. Nat Med. 2016;22:976–86.
Lu ZX, Huang Q, Park JW, Shen S, Lin L, Tokheim CJ, et al. Transcriptome-wide landscape of pre-mRNA alternative splicing associated with metastatic colonization. Mol Cancer Res. 2015;13:305–18.
Trincado JL, Sebestyen E, Pages A, Eyras E. The prognostic potential of alternative transcript isoforms across human tumors. Genome Med. 2016;8:85.
Wang S, Sun Z, Lei Z, Zhang HT. RNA-binding proteins and cancer metastasis. Semin Cancer Biol. 2022;86:748–68.
Wang Y, Chen D, Qian H, Tsai YS, Shao S, Liu Q, et al. The splicing factor RBM4 controls apoptosis, proliferation, and migration to suppress tumor progression. Cancer Cell. 2014;26:374–89.
Oltean S, Bates DO. Hallmarks of alternative splicing in cancer. Oncogene. 2014;33:5311–8.
Buckanovich RJ, Yang YY, Darnell RB. The onconeural antigen Nova-1 is a neuron-specific RNA-binding protein, the activity of which is inhibited by paraneoplastic antibodies. J Neurosci. 1996;16:1114–22.
Meldolesi J. Alternative Splicing by NOVA Factors: From Gene Expression to Cell Physiology and Pathology. Int J Mol Sci. 2020;21:3941.
Campos-Melo D, Droppelmann CA, Volkening K, Strong MJ. RNA-binding proteins as molecular links between cancer and neurodegeneration. Biogerontology. 2014;15:587–610.
Pradella D, Deflorian G, Pezzotta A, Di Matteo A, Belloni E, Campolungo D, et al. A ligand-insensitive UNC5B splicing isoform regulates angiogenesis by promoting apoptosis. Nat Commun. 2021;12:4872.
Angiolini F, Belloni E, Giordano M, Campioni M, Forneris F, Paronetto MP, et al. A novel L1CAM isoform with angiogenic activity generated by NOVA2-mediated alternative splicing. Elife. 2019;8:e44305.
Cui W, Fowlis DJ, Bryson S, Duffie E, Ireland H, Balmain A, et al. TGFbeta1 inhibits the formation of benign skin tumors, but enhances progression to invasive spindle carcinomas in transgenic mice. Cell. 1996;86:531–42.
David CJ, Massague J. Contextual determinants of TGFbeta action in development, immunity and cancer. Nat Rev Mol Cell Biol. 2018;19:419–35.
Katsuno Y, Lamouille S, Derynck R. TGF-beta signaling and epithelial-mesenchymal transition in cancer progression. Curr Opin Oncol. 2013;25:76–84.
Massague J, Seoane J, Wotton D. Smad transcription factors. Genes Dev. 2005;19:2783–810.
Xu P, Lin X, Feng XH. Posttranslational Regulation of Smads. Cold Spring Harb Perspect Biol. 2016;8:a022087
Liao JY, Yang B, Zhang YC, Wang XJ, Ye Y, Peng JW, et al. EuRBPDB: a comprehensive resource for annotation, functional and oncological investigation of eukaryotic RNA binding proteins (RBPs). Nucleic Acids Res. 2020;48:D307–D313.
Giudice G, Sanchez-Cabo F, Torroja C, Lara-Pezzi E. ATtRACT-a database of RNA-binding proteins and associated motifs. Database (Oxford). 2016;2016:baw035.
Yuan H, Yan M, Zhang G, Liu W, Deng C, Liao G, et al. CancerSEA: a cancer single-cell state atlas. Nucleic Acids Res. 2019;47:D900–D908.
Zhang Y, Fu F, Zhang Q, Li L, Liu H, Deng C, et al. Evolutionary proteogenomic landscape from pre-invasive to invasive lung adenocarcinoma. Cell Rep Med. 2024;5:101358.
Jorgensen HF, Azuara V, Amoils S, Spivakov M, Terry A, Nesterova T, et al. The impact of chromatin modifiers on the timing of locus replication in mouse embryonic stem cells. Genome Biol. 2007;8:R169.
Song M, Yang X, Ren X, Maliskova L, Li B, Jones IR, et al. Mapping cis-regulatory chromatin contacts in neural cells links neuropsychiatric disorder risk variants to target genes. Nat Genet. 2019;51:1252–62.
Mortimer SA, Kidwell MA, Doudna JA. Insights into RNA structure and function from genome-wide studies. Nat Rev Genet. 2014;15:469–79.
Castel SE, Martienssen RA. RNA interference in the nucleus: roles for small RNAs in transcription, epigenetics and beyond. Nat Rev Genet. 2013;14:100–12.
Fiszbein A, Krick KS, Begg BE, Burge CB. Exon-mediated activation of transcription starts. Cell. 2019;179:1551–65.e1517.
Wang Z, Lei Z, Wang Y, Wang S, Wang JP, Jin E, et al. Bone-metastatic lung adenocarcinoma cells bearing CD74-ROS1 fusion interact with macrophages to promote their dissemination. Oncogene. 2024;43:2215–27.
Cai L, Sutter BM, Li B, Tu BP. Acetyl-CoA induces cell growth and proliferation by promoting the acetylation of histones at growth genes. Mol Cell. 2011;42:426–37.
Yang YY, Yin GL, Darnell RB. The neuronal RNA-binding protein Nova-2 is implicated as the autoantigen targeted in POMA patients with dementia. Proc Natl Acad Sci USA. 1998;95:13254–9.
Demagny H, Araki T, De Robertis EM. The tumor suppressor Smad4/DPC4 is regulated by phosphorylations that integrate FGF, Wnt, and TGF-beta signaling. Cell Rep. 2014;9:688–700.
Zhang Y, Liu K, Xu Z, Li B, Wu X, Fan R, et al. OncoSplicing 3.0: an updated database for identifying RBPs regulating alternative splicing events in cancers. Nucleic Acids Res. 2025;53:D1460–D1466.
Gerstberger S, Hafner M, Tuschl T. A census of human RNA-binding proteins. Nat Rev Genet. 2014;15:829–45.
Zhou MM, Cole PA. Targeting lysine acetylation readers and writers. Nat Rev Drug Discov. 2025;24:112–33.
Rice JC, Allis CD. Histone methylation versus histone acetylation: new insights into epigenetic regulation. Curr Opin Cell Biol. 2001;13:263–73.
Liu D, Zhang H, Cong J, Cui M, Ma M, Zhang F, et al. H3K27 acetylation-induced lncRNA EIF3J-AS1 improved proliferation and impeded apoptosis of colorectal cancer through miR-3163/YAP1 axis. J Cell Biochem. 2020;121:1923–33.
Zhang Y, Liu Z, Yang X, Lu W, Chen Y, Lin Y, et al. Erratum: H3K27 acetylation activated-COL6A1 promotes osteosarcoma lung metastasis by repressing STAT1 and activating pulmonary cancer-associated fibroblasts: Erratum. Theranostics. 2022;12:4604–5.
Ye P, Lv X, Aizemaiti R, Cheng J, Xia P, Di M. H3K27ac-activated LINC00519 promotes lung squamous cell carcinoma progression by targeting miR-450b-5p/miR-515-5p/YAP1 axis. Cell Prolif. 2020;53:e12797.
Han M, Qian X, Cao H, Wang F, Li X, Han N, et al. lncRNA ZNF649-AS1 Induces Trastuzumab Resistance by Promoting ATG5 Expression and Autophagy. Mol Ther. 2020;28:2488–502.
Wang YC, Wu YS, Hung CY, Wang SA, Young MJ, Hsu TI, et al. USP24 induces IL-6 in tumor-associated microenvironment by stabilizing p300 and beta-TrCP and promotes cancer malignancy. Nat Commun. 2018;9:3996.
Frank DA. Transcription factor STAT3 as a prognostic marker and therapeutic target in cancer. J Clin Oncol. 2013;31:4560–1.
Bai L, Zhou H, Xu R, Zhao Y, Chinnaswamy K, McEachern D, et al. A Potent and Selective Small-Molecule Degrader of STAT3 Achieves Complete Tumor Regression In Vivo. Cancer Cell. 2019;36:498–511.e417.
Liu J, Yu Q, Yang X. Circ_0102231 inactivates the PI3K/AKT signaling pathway by regulating the miR-635/NOVA2 pathway to promote the progression of non-small cell lung cancer. Thorac Cancer. 2023;14:3453–64.
Xia F, Xie M, He J, Cheng D. Circ_0004140 promotes lung adenocarcinoma progression by upregulating NOVA2 via sponging miR-330-5p. Thorac Cancer. 2023;14:3483–94.
Dhara C, Dhara A, Gantayat S. A Robust NSCLC Biomarker- Mir-7-5p: Its in-silico validation and potential SPR-based probe for detection. Microrna. 2025;14:112–23.
David CJ, Manley JL. Alternative pre-mRNA splicing regulation in cancer: pathways and programs unhinged. Genes Dev. 2010;24:2343–64.
Di Matteo A, Belloni E, Pradella D, Chiaravalli AM, Pini GM, Bugatti M. et al. Alternative splicing changes promoted by NOVA2 upregulation in endothelial cells and relevance for gastric cancer. Int J Mol Sci. 2023;24:8102.
Tress ML, Martelli PL, Frankish A, Reeves GA, Wesselink JJ, Yeats C, et al. The implications of alternative splicing in the ENCODE protein complement. Proc Natl Acad Sci USA. 2007;104:5495–5500.
Pierreux CE, Nicolas FJ, Hill CS. Transforming growth factor beta-independent shuttling of Smad4 between the cytoplasm and nucleus. Mol Cell Biol. 2000;20:9041–54.
Zhao C, Zhao JW, Zhang YH, Zhu YD, Yang ZY, Liu SL, et al. PTBP3 mediates IL-18 exon skipping to promote immune escape in gallbladder cancer. Adv Sci (Weinh). 2024;11:e2406633.
DaSilva JO, Yang K, Perez Bay AE, Andreev J, Ngoi P, Pyles E, et al. A biparatopic antibody that modulates MET trafficking exhibits enhanced efficacy compared with parental antibodies in MET-driven tumor models. Clin Cancer Res. 2020;26:1408–19.
Acknowledgements
We are grateful for the participation and cooperation from LUAD patients. This work was supported in part by National Natural Science Foundation of China (82273372, 82203773, 82303450), and the Natural Science Foundation of Jiangsu Province (BK20231233, BK20220250), and the “Qinglan Project” of Jiangsu Province universities, and the Talent Project of Kangda College of Nanjing Medical University (KD2024JXJH001), and Suzhou Key Laboratory for Molecular Cancer Genetics (SZS201209), and Collaborative Innovation Center of Molecular Medicine between Soochow University and Donghai County People’s Hospital (H230470), and A Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Contributions
SW, EJ, and H-TZ contributed to study concept and design. SW, XT, RS, XX, DC, and ZW performed all the experiments. HS, CW, XG, and DH provided technical support. SW, EJ, and H-TZ analyzed the data, wrote the manuscript, and contributed to the interpretation of data and critical revision of the manuscript. SW, XT, and H-TZ provided the funding and supervised the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics
Human LUAD tissues were obtained after informed consent was available from all patients. The study protocol was carried out in accordance with the ethical principles outlined in the Declaration of Helsinki and has been approved by the Ethics Committee of Soochow University.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, S., Tong, X., Sun, R. et al. STAT3-mediated transactivation of NOVA2 promotes lung adenocarcinoma metastasis by splicing SMAD4. Oncogene (2026). https://doi.org/10.1038/s41388-026-03752-6
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41388-026-03752-6