Abstract
Background:
White matter injury (WMI) is a major cause of neurodevelopmental impairment in preterm infants (PTIs), yet early molecular biomarkers remain elusive. Circulating microRNAs (miRNAs) hold promise, but neonatal biosample limitations challenge the use of high-throughput methods like next-generation sequencing (NGS). We evaluated the feasibility of quantitative PCR (qPCR) as a primary discovery tool for circulating miRNA biomarkers of WMI, supported by systems biology modeling and selective NGS validation.
Methods:
Plasma-derived miRNAs from PTIs with and without WMI were profiled using qPCR after stringent hemolysis screening. Candidate miRNAs were curated from brain-specific regulatory networks. ROC bootstrapping and gene ontology/pathway enrichment were used to assess diagnostic and functional relevance. Boolean logic modeling simulated miRNA-mediated regulation of oligodendrocyte maturation.
Results:
miR-23a-3p and miR-17-5p were differentially expressed across WMI states and showed moderate discriminative power (AUCs: 0.71 and 0.68), while oligodendrocyte miRNAs (miR-219a-2-3p, miR-338-5p) were consistently low. Boolean simulations confirmed miR-23a and miR-17 modulate myelination via PTEN repression and PI3K/Akt pathway activation. qPCR and model predictions aligned strongly; NGS showed discordant trends likely due to detection biases.
Conclusion:
This pilot study demonstrates that qPCR, when combined with systems modeling, proposes a viable and sensitive discovery tool for miRNA biomarkers in clinically constrained populations.
Impact
-
This study demonstrates that qPCR combined with systems biology modeling, is a viable and biologically coherent approach for identifying circulating miRNA biomarkers of white matter injury in preterm infants.
-
It challenges the conventional reliance on next-generation sequencing as the default discovery platform, and repositions qPCR as a powerful primary discovery method for diagnostic miRNA biomarkers when coupled with systems biology modeling, showing that qPCR can yield translational insights in ethically and clinically constrained neonatal populations.
-
By integrating Boolean logic simulations, the study links key miRNAs (miR-23a-3p and miR-17-5p) to mechanistic pathways of oligodendrocyte maturation, offering a functional layer to biomarker discovery.
Similar content being viewed by others
Introduction
Several forms of brain injury have been identified in preterm infants (PTIs), including intraventricular hemorrhage and white matter injury (WMI) which are best characterized using standardized scoring systems like the Kidokoro classification. WMI has become the primary neurological issue affecting PTIs leading to long-term neurodevelopmental disorders including, cerebral palsy, cognitive deficits, and behavioral defects1,2,3,4,5. Despite advances in neonatal care, early detection of WMI remains elusive. Conventional imaging techniques often fail to identify the subtle, diffuse lesions characteristic of WMI, which account for up to 80% of injuries in very preterm infants (vPTIs)6,7. This gap highlights the urgent need for reliable and economical biomarkers that can facilitate early diagnosis and intervention.
Circulating microRNAs (miRNAs) are promising candidates for this purpose due to their stability in body fluids, including blood, and their ability to reflect underlying pathological processes in tissues, including the brain8,9,10,11. Specific miRNA profiles have been associated with neuroinflammatory responses and glial cell damage12,13, both of which are pivotal in the pathogenesis of WMI. However, the discovery of RNA biomarker candidates is often limited within practical constraints in neonatal research. To enable clinical implementation, biomarkers must first undergo validation through extensive prospective trials, incorporating comparisons with standard care practices at specific predetermined intervals14,15. Besides this, small sample volumes, low RNA abundance in plasma, sample quality, and the ethical challenges of sampling from preterm infants limit the feasibility of high-throughput approaches like next-generation sequencing (NGS), which remains the valuable but pricey gold standard for transcriptomic discovery16,17,18,19,20.
Prior to the use of sequencing technologies, methods such as qPCR, northern blotting, and high-content imaging played a crucial role in the early identification and confirmation of disease biomarkers21,22. qPCR remains an economical and sensitive technique for detecting miRNA but it doesn’t offer the scalability and discovery breadth of sequencing.
Given the constraints of studying fragile populations such as preterm infants, Boolean modeling offers a powerful, qualitative systems biology framework that overcomes these limitations by enabling simulation of gene-regulatory interactions using logic-based rules rather than precise kinetic parameters. This approach has been successfully applied to model signal transduction, differentiation programs, and disease mechanisms in systems where quantitative data is scarce but topological network structure is known23,24,25. In the context of WMI, where our qPCR-derived miRNA expressions point to specific regulatory miRNAs, Boolean logic enables us to test mechanistic hypotheses about how these miRNAs influence downstream targets involved in myelination failure –thereby bridging molecular expression data with developmental outcomes in silico.
To this end, we designed a pilot study that integrates targeted qPCR profiling, systems-level pathway enrichment, and Boolean modeling to simulate miRNA-regulated networks underlying WMI. This hybrid strategy allowed us to explore not only differential expression patterns but also the mechanistic plausibility of observed miRNA–gene interactions and their potential therapeutic modulation. This approach allowed us to focus on a narrowed list of candidate miRNAs, using NGS as a validation tool in a select subset of 5 samples while miRNA–target interactions were annotated through miRNet 2.0 and miRPathDB26,27,28,29. This integrative approach aims to present the feasibility of using qPCR as a frontline discovery tool in preterm populations, reinforced by in silico modeling to bridge molecular findings with disease mechanisms.
Methods
This pilot study employed a hybrid approach, integrating experimental qPCR data with computational systems biology analyses to identify and validate miRNA biomarkers associated with WMI in preterm infants. The workflow involved three primary components as shown in Fig. 1: (1) qPCR-based expression analysis in plasma samples, (2) in silico pathway and network modeling to broaden the discovery scope, and (3) targeted validation of select miRNAs using NGS of 5 samples.
Candidate microRNAs associated with preterm birth-related insults were identified through publicly available datasets and refined using a systems-level prioritization strategy. Selected miRNAs were quantified in preterm infant plasma using qPCR, followed by computational functional enrichment analysis and Boolean network modeling to predict regulatory roles. In parallel, a subset of samples was analyzed using NGS to validate qPCR findings and assess congruency between methods.
Patients
Preterm newborns who were admitted at the Neonatal Intensive Care Unit (NICU) of University Hospital Puerta del Mar, in Cádiz, between the years 2021 and 2022 and were less than 32 weeks of gestation and/or 1500 grams at birth were included for this study. The exclusion criteria were defined as the presence of congenital or chromosomal anomalies, metabolic disorders, central nervous system infections or death before 40 weeks of corrected age. This study was approved by the Biomedical Research Ethics Committee of Cádiz, and all parents or guardians of the participants provided informed consent.
The patients were grouped into: PTIs without visible signs of WMI on the MRI scan at term corrected age were classed as the control group and, PTIs with WMI were the pathological group. Blood samples were collected at two time points: (a) <7 days of life and (b) >28 days of life. The early time point was chosen to capture miRNA expression profiles in response to the acute insults associated with preterm birth, including hypoxia and inflammation, which are known to contribute to WMI5,30. The late time point was selected to assess miRNA dynamics during the recovery phase, as well as to confirm the presence or absence of WMI through neuroimaging at term-equivalent age7. This dual-time-point design allowed us to track the progression of the candidate miRNAs and evaluate their potential as early biomarkers. No extremely preterm infants developed WMI; hence, all WMI samples were from very preterm infants. Extremely and very preterm infants with no WMI (NWMI) were merged into a single control group as their miRNA profiles (NWMI.a) did not differ significantly (Supplementary Fig. S4).
The clinical characteristics of the included preterm infants are summarized in Table 1. The cohort consisted of 6 infants NWMI and 8 infants with WMI, with detailed metrics provided for gestational age (GA), and postmenstrual age (PMA) at both early (a) and late (b) sampling timepoints.
Materials
In this study, the materials used were: from Norgen Biotek Canada - Plasma/Serum RNA Purification kit (product #55000) and, Cel-miR-39 Spike-in kit (product #59000); from Takara Bio USA - Mir-X miRNA First-Strand Synthesis and TB Green qRT-PCR and, TB Green Advantage qPCR Premix.
Blood sample collection and plasma extraction
Blood was drawn via venous sampling concomitantly with clinically indicated routine blood tests, and plasma was separated from EDTA-treated whole blood. Blood was drawn within the first 7 days of life and 28 days after birth. The blood was processed within 4 hours of collection. After blood centrifugation (1500 × g, 4 °C, and 10 min), the plasma was divided into aliquots of 200 µL and stored at −80 °C until RNA extraction.
RNA extraction
Total plasma RNA was extracted from 200 µL of frozen plasma samples using the total plasma/serum purification Kit (NorGen Biotek) according to the manufacturer’s instructions. In brief, 200 µl of plasma combined with 600 µl lysis buffer was briefly vortexed, followed by an addition of 99fmol of Cel-miR-39 spike-in, and finally 800 µl of 96% ethanol. After a quick vortex, 650 µl of the mixture was loaded onto a spin column for centrifugation until the mixture was exhausted and RNA was isolated. Total RNA including the spike-in was eluted to a 20 µL elution solution and transferred again to the column for centrifugation to achieve maximum RNA recovery. RNAs were stored at −80 °C for further analysis.
Hemolysis
Hemolysis is a common bias found in circulating RNA or DNA analysis. Addressing miRNA markers of hemolysis is particularly crucial in miRNA qPCR analysis of liquid biopsies, as erythrocytes, one of the main contributors of miRNAs in blood samples, release substantial amounts of exogenous miRNAs during hemolysis. This improves the reliability of biomarker discovery and validation.
Accordingly, samples were filtered following an optimized protocol of determining hemolysis within the sample population. We designed and conducted a pre-study to classify samples through (i) colourimetric tests, (ii) Thermo Fisher’s nanodrop ‘oxyhemoglobin’ package at 414 nm absorbance and, (iii) delta CT of miR-23a – miR-451a (Fig. S1). If the samples passed the colourimetric check, they were selected for downstream experiments. If the colourimetric test came undetermined, we checked for hemolysis using nanodrop. Our pre-study determined that an absorbance at 414 nm of <1.2 and deltaCT of <7 was deemed unhemolysed, so samples passing at least one criterion were retained for downstream qPCR profiling.
MicroRNA expression and analytical workflow
MiRNA candidate selection
To select microRNAsfor expression analysis of differentially expression in the brain of PTIs we conducted a systematic literature search on GEO (from inception to December 2023) using the following query search: (ischemic hypoxic encephalopathy) AND “Homo sapiens”[porgn:__txid9606]), (preterm infant microrna) AND “Homo sapiens”[porgn:__txid9606]), (cerebral palsy blood microrna (cerebral palsy brain) AND ““Homo sapiens”“[porgn:__txid9606]), ((autism asd rna) AND blood infant) AND “Homo sapiens”[porgn:__txid9606]).
Regulatory networks of miRNA-disease interactions were constructed and visualized using miRNet 2.026,27. This tool integrates data from multiple databases, including KEGG31,32,33, Reactome34, and Gene Ontology (GO)35,36, to allow for the identification of pathways and biological processes that are significantly enriched among the target genes of specific miRNAs. To optimize our search we explored any potential networks of miRNAs in commonly known perinatal and or WMI related disorders and oligodendrocyte-related disorders. The following query search was used: autism spectrum disorder, multiple sclerosis, bronchopulmonary dysplasia, neuroinflammation, hypoxic-ischemic encephalopathy, preterm labor, preeclampsia. The results were further narrowed using the following features: Degree filter and Betweenness of 2.0 and microRNAs only; Betweenness and Shortest paths were microRNAs only (Fig. S2).
RT-qPCR
Reverse transcription (RT) reactions were performed using the Mir-X™ miRNA First Strand Synthesis Kit (Takara) in a total reaction volume of 10 uL. RT reactions were run for 60 min at 42 °C, 5 min at 95 °C. cDNA was stored at −20 °C. qPCR was performed in a total volume of 10ul using the Mir-X™ miRNA qRT-PCR TB Green® Kit (Takara) at 95 °C for 2 min, followed by 40 cycles of 95 °C for 10 s, and 56 °C for 60 s. For primer sequences see Supplementary Data File, Table 1. A common reverse primer provided by the company was used for all reactions.
The cycle threshold (Ct) of target transcripts across all samples were normalized by standard delta-delta Ct to determine relative fold quantification with Cel-miR-39 CTs as the external control. Any Ct value of higher than 35 cycles was deemed undetected and excluded from analysis.
RNA sequencing
After RNA extraction, samples were sent to Genomics Core, Germany for sequencing where Dr. Vladimir Benes generously offered free services for this pilot study. They prepared the RNA sequencing library using the Nextflex kit by Bioscientific/PerkinElmer, with an adapter dilution of 1:10 and 25 PCR cycles. Finally, sequencing was performed on an Illumina Miseq sequencer with 70 bp single-end reads. Sequence reads were mapped against the reference genome GRCh38 and gene annotation was performed with Bowtie2 on Galaxy.
Statistical analyses
Normalization of qPCR Data
To ensure accurate quantification of miRNA expression levels, a spike-in control – cel-miR-39, was used to normalize the Ct values of the target miRNAs. 33fmol of cel-miR-39 was added to each sample prior to RNA extraction. miRNA Cts were normalized by subtracting the Ct value of the spike-in control from the target miRNA Ct value, resulting in a ΔCt value for each sample. This normalization process corrects for technical variations, such as differences in RNA input. Normalization was performed with Microsoft Excel (online).
Assessment of normal distribution
The distribution of the ΔCt values was assessed to determine if they followed a normal distribution. Q-Q plots were constructed to compare the observed ΔCt values of each sample against their expected Z-score. The Shapiro-Wilk test was performed to statistically evaluate the normality of the data. A p-value greater than 0.05 in the Shapiro-Wilk test was considered indicative of a normal distribution. The plots were performed using Microsoft Excel and Python Package 3.9.6 was used to generate the Shapiro-Wilk score. (Fig/ S3, Table S2).
Differential expression analysis
All statistical analyses were conducted to evaluate the differential expression of miRNAs among different sample groups and over time. The relative abundance of miRNAs was compared between the PTIs who had no white matter injury (NWMI.a) and/or, and late NWMI (NWMI.b), early White Matter Injury (WMI.a) and late WMI (WMI.b) using a t-test. A temporal analysis was also performed to assess changes in miRNA expression between early and late sampling times within each condition (WMI and NWMI). For this, fold changes were calculated, and a t-test was applied to detect significant temporal variations in miRNA expression. Relative differential analysis was performed with Microsoft Excel (online). P-values were two-tailed, with a significance threshold set at p < 0.05. Bootstrapping (n = 1000 iterations) was performed to estimate AUC confidence intervals. Despite some low absolute scores, bootstrapping confirmed stability (narrow 95% CIs: ±0.07).
Functional pathway analysis
To investigate the biological implication of the differentially expressed miRNAs, we conducted a functional pathway enrichment analysis utilizing MiRPathDB29, DAVID37,38, Reactome34, and KEGG tools31,32,33. Pathways having an enrichment p-value under 0.05 were deemed significant. The enrichment p-value was determined through a hypergeometric test, which evaluates the excess of miRNA target genes in particular pathways compared to the overall number of genes in that pathway. The Benjamini-Hochberg adjustment was utilized to correct for multiple comparisons, aiming to manage the false discovery rate.
In silico systems modeling
Boolean modeling
We developed a Boolean regulatory network incorporating experimentally validated human miRNA-mRNA interactions. Target genes for miR-23a-3p and miR-17-5p were retrieved from miRPathDB and cross-validated with published literature on neurodevelopment. The network included core regulators such as PTEN, LMNB1, and EphA4, which influence myelination through the PI3K/Akt/mTOR pathway. Boolean logic gates were used to simulate node activity: miRNA inputs were modeled as binary (ON/OFF), while target gene inhibition and downstream activation cascades were simulated accordingly. The model was constructed using Python’s networkx library and visualized to demonstrate logical propagation from miRNA expression to final myelination state.
Boolean Logic of the Model:
Inputs: miR-23a-3p and miR-17-5p levels (ON/OFF states)
Outputs: Oligodendrocyte Differentiation (Promoted/Inhibited)
Myelination (Enhanced/Impaired)
Logic Gates:
NOT Gate: miR-23a-3p and miR-17-5p inhibit their target genes (PTEN, LMNB1, EphA4)39,40.
AND Gate: Concurrent suppression of PTEN by both miRNAs leads to robust activation of the PI3K/Akt/mTOR pathway.
Model Dynamics:
Scenario 1: High miR-23a-3p and miR-17-5p Levels (Both ON)
↓ PTEN and LMNB1 expression
↑ PI3K/Akt/mTOR pathway activation
↑ Oligodendrocyte differentiation
↑ Myelination
Scenario 2: Low miR-23a-3p and miR-17-5p Levels (Both OFF)
↑ PTEN and LMNB1 expression
↓ PI3K/Akt/mTOR pathway activation
↓ Oligodendrocyte differentiation
↓ Myelination
Scenario 3: High miR-23a-3p and Low miR-17-5p Levels
↓ PTEN (due to miR-23a-3p activity)
Partial activation of PI3K/Akt/mTOR pathway
Moderate promotion of oligodendrocyte differentiation and myelination
Ethical approval
Informed consent was obtained from the parents or legal guardians of all participants included in the study. The study was conducted in accordance with the Declaration of Helsinki and was approved by the Research and Ethics Committee (Protocol number: PIGE-0026-2020; code: 117.21).
Results
Oligodendrocyte miRNAs miR-219a and miR-338 show low plasma expression and poor diagnostic utility in preterm white matter injury.
To contextualize our findings within established neurobiology, we first analyzed two oligodendrocyte miRNAs, miR-219a and miR-338, which are well-defined regulators of OL differentiation in model systems41,42,43. In transgenic models miR-219a, and miR-338 were found to be highly expressed during OL differentiation, with miR-219 emerging as the most differentially expressed44. Similarly, miR-338 promoted OL differentiation and was found to be upregulated in a perinatal hypoxia-ischemia mouse model. Strikingly, and in contrast to predictions from animal studies, we found their levels were significantly lower in all preterm infants plasma samples compared to other candidate miRNAs.
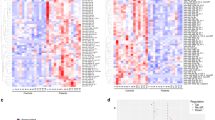
NGS analysis revealed no detectable expression of miR-219a-2-3p and miR-338-5p in all samples, reinforcing the qPCR data that demonstrated very low levels of these miRNAs (Fig. 2a). The absence of detectable reads suggests that these miRNAs may be present at concentrations below the threshold of detection for NGS or may not be actively circulating in plasma. Our data suggest that miR-219a-2-3p and miR-338-5p signatures in PTIs might not reflect the same regulatory mechanisms described in animal models, potentially due to the incomplete maturation of oligodendrocytes at this gestational stage.
a Box-and-whisker plots showing log10-transformed relative expression of miR-219a-2-3p, miR-338-5p, miR-23a, and miR-17 across WMI and non-WMI preterm infant plasma samples, as assessed by qPCR. MiR-219a and miR-338 were consistently the lowest expressed. b ROC-derived AUC scores for miR-219a and miR-338, representing their ability to discriminate WMI from non-WMI states. The vertical lines on each bar indicate 95% confidence intervals derived from bootstrap resampling, revealing high uncertainty for miR-219a (AUC ≈ 0.54) and moderate predictive strength for miR-338 (AUC ≈ 0.76). c Temporal expression patterns of miR-219a-2-3p and miR-338-5p across early (a) and late (b) postnatal time points in WMI and non-WMI groups. Both miRNAs show a non-specific decline over time, independent of WMI status. d Venn diagram showing overlap of predicted gene targets between miR-219a-2-3p and miR-338-5p, identifying 58 co-regulated genes. e Tissue enrichment analysis of co-targeted genes reveals significant associations with brain-specific regions and immune cell types. f Functional enrichment of co-targeted genes highlights roles in synaptic plasticity, organelle organization, immune signaling, and apoptosis-related pathways.
To estimate the diagnostic confidence of AUC scores despite small sample sizes, we employed nonparametric bootstrapping (1000 resamples, with replacement) to generate 95% confidence intervals. This method allows estimation of variability without assuming normality, mitigating the inflation of performance metrics due to limited data. Bootstrapping analysis indicated that miR-219a-2-3p showed poor diagnostic reliability, achieving an AUC score of 0.54, which suggests a performance nearly equivalent to random classification. In comparison, miR-338-5p showed a somewhat higher predictive confidence with an AUC of 0.76, indicating a somewhat more reliable connection with WMI (Fig. 2b). These results indicate that, although both miRNAs are found at low concentrations in preterm plasma, their effectiveness as biomarkers for WMI is uncertain, especially for miR-219a-2-3p, whose expression trends may better represent general low levels rather than being regulated by disease. This suggests that though qPCR and NGS produce similar results, the reliability of their classification potential can not be assumed for these low‑abundance miRNAs in this pilot cohort.
Moreover, our analysis revealed that miR-219a-2-3p and miR-338-5p exhibit a similar temporal expression pattern, independent of WMI status (Fig. 2c). This observation aligns with previous studies suggesting that these miRNAs play a more fundamental role in developmental processes rather than serving as direct indicators of pathological white matter injury. Their expression significantly declines over time, which may reflect the natural waning of developmental regulatory signals or a reduction in injury-induced reparative responses during later stages of preterm brain maturation. The markedly low levels of miR-219a-2-3p and miR-338-5p in our dataset (Fig. 2a) support the hypothesis that their regulatory activation may be developmentally delayed or dormant at the gestational age of very preterm infants, potentially limiting their role in acute injury response.
We identified through computational analysis, 58 co-targeted genes highly enriched in the fetal brain (Supplementary List SL1) which play regulatory roles in dendritic spine development and synaptic plasticity amongst other biological functions in adult tissues (Fig. 2d, e). Additionally, we found that miR-219a-2-3p and miR-338-5p are significantly involved in apoptotic and inflammatory pathways (Fig. 2f), suggesting a potential role in cellular stress responses following preterm birth-related insults29,38,45,46,47.
Taken together, these findings demonstrate that model-derived oligodendrocyte regulatory networks do not entirely recapitulate the circulating miRNA profile in human preterm infants. The low levels of miR‑219 and miR‑338 in preterm infant plasma suggest that NGS‑derived oligodendrocyte miRNA signatures from experimental models may not directly translate to circulating biomarkers in human neonates, supporting our decision to use a systems biology framework to prioritize other, more reliably detected miRNAs for biomarker exploration. Future investigations should explore whether other developmental stage-dependent miRNAs compensate for their function at this early gestational stage or whether alternative regulatory mechanisms are at play in very preterm infants.
Regulators of adipogenesis and myelination miR-23a and miR-17 are differentially expressed in qPCR but not in RNA-seq
MiR-23a plays a crucial role in synaptic plasticity, a process closely tied to myelination, through pathways such as fatty acid elongation, glutamatergic signaling, and GABAergic signaling48,49. In our study, we found that miR-23a was highly expressed in the samples with a distinct differential expression pattern in relation to disease state (WMI or NWMI) (Fig. 3a). MiR-23a appeared to be lower in the diseased state samples compared to the healthy sample. Notably, miR-23a exhibited a subtle recovery in abundance in the WMI.b samples after 28 days, suggesting its potential role in later stages of white matter injury. Bootstrapping analysis further validated the reliability of our findings, revealing an AUC score of 0.71 with a 95% confidence interval (CI) of 0.481–0.909 (Fig. 3a, purple bar). It is important to note that while miR-23a demonstrates some discriminatory power between WMI and NWMI groups, the wide CI indicates variability, highlighting the need for further investigation in larger cohorts.
a Fold expression of miR-23a-3p across WMI and NWMI samples at early (a) and late (b) postnatal stages. Expression was reduced in WMI.a but showed partial recovery in WMI.b. The purple bar indicates the AUC score from bootstrapping analysis (AUC = 0.71, 95% CI: 0.481–0.909). b Gene ontology enrichment of miR-23a target genes, categorized by biological process, cellular component, and molecular function. Key pathways include fatty acid elongation, synaptic signaling, and membrane organization –processes linked to myelination and neuronal plasticity. Full list in Supplementary Table S3. c Fold expression of miR-17-5p shows a marked decline in WMI.b, with significant differences from NWMI groups. The purple bar represents bootstrapped AUC score (AUC = 0.68, 95% CI: 0.498-0.892). d KEGG pathway enrichment of miR-17 target genes, showing significant involvement in cancer-related, autophagic, and axon guidance pathways. Bubble size represents the number of enriched genes; color indicates false discovery rate (FDR). e Venn diagram displaying 233 co-targeted genes shared between miR-23a and miR-17. f Comparison of log2 fold-change of miR-23a and miR-17 from qPCR and NGS data across the same early-life WMI and NWMI samples. NGS findings showed no statistically significant changes, underscoring potential detection limitations for these miRNAs in ultra-low-volume samples.
To elucidate the regulatory functions of miR-23a for the design of our boolean model, we conducted a GO analysis selecting the top 10 enriched pathways (Fig. 3b). We saw that miR-23a plays a crucial role in synaptic plasticity, a process closely tied to myelination, through pathways such as fatty acid elongation, glutamatergic signaling, and GABAergic signaling. Recent studies have shown that axon guidance cues such as ephrins and semaphorins also regulate oligodendrocyte migration, positioning, and maturation –key processes disrupted in WMI, as well as neuroinflammation50,51. This suggests that miRNA-mediated modulation of axon guidance may influence white matter development and remyelination following injury. Furthermore, our qPCR analysis revealed that miR-17 was the most abundant of our candidate miRNAs (Fig. 2a) and showed the steepest decline in abundance in our WMI samples after 28 days (Fig. 3c). Computational exploration of miR-17 target genes showed regulatory roles in various biological and disease pathways including autophagy, cancer, as well as axon guidance (Fig. 3d). The decline in miR-17 in WMI could suggest a disruption in pathways critical to brain development, including cell proliferation, apoptosis regulation, autophagy and inflammation, which are essential for oligodendrocyte proliferation and myelin sheath formation. Through a comprehensive interactome analysis, we identified several overlapping target genes for miR-17 and miR-23a, including JAK1, KPNA4, and FAS. These genes are integral to pathways such as sustained proliferative signaling, adipogenesis, and leptin signaling (Fig. S5). Leptin, in particular, has been shown to influence myelination processes, as evidenced by studies where leptin replenishment improved myelination in leptin-deficient mice without affecting OPC differentiation between postnatal days 7 to 2849. This underscores leptin’s role in creating a conducive environment for white matter repair and further aligns with the regulatory effects of miR-17 and miR-23a on neural tissue plasticity. Finally, we found that the co-targeted genes (Fig. 3e, Supplementary List SL2) of miR-23a and miR-17 play key regulatory roles in presynaptic and postsynaptic activity.
Given that the aim of this study was to assess the feasibility of performing a robust experimental investigation using qPCR and computational tools, with NGS reserved for the validation step, we limited our dataset to two WMI samples as an investigative set and three NWMI samples as our control. We hypothesized that the most notable discrepancies in expression would occur shortly after birth, so all five samples were collected within the first seven days of life. Interestingly, our NGS analysis showed a minor and statistically insignificant upregulation of hsa-miR-23a and hsa-miR-17 (Fig. 3f) in WMI patients.
Logic-based modeling supports miR-23a/miR-17 as regulators of myelination in WMI
To simulate how changes in miR-23a-3p and miR-17-5p expression might propagate through a regulatory network affecting myelination, we constructed a Boolean logic-based model integrating experimentally validated miRNA–target interactions. The network incorporated target genes of both miRNAs such as PTEN, a negative regulator of oligodendrocyte differentiation52, and downstream mediators including Akt, Olig2, and myelin basic protein (MBP) – a marker of myelin production53,54. In this framework, both miR-23a and miR-17 were modeled as upstream repressors of PTEN and additional targets such as LMNB1 and EphA4, enabling activation of differentiation and myelin gene expression under physiologic conditions (Fig. 4a).
a Boolean regulatory network model depicting the regulatory role of miR-23a-3p and miR-17-5p on human oligodendrocyte differentiation and myelination. miR-23a-3p and miR-17-5p downregulate PTEN, a key repressor of the PI3K/Akt/mTOR signaling pathway. miR-23a-3p also targets LMNB1, a gene whose overexpression disrupts myelin structure, while miR-17-5p inhibits EphA4, a regulator of axonal guidance. Activation of PI3K/Akt/mTOR promotes oligodendrocyte differentiation, which drives myelination. Arrows denote activation; blunt lines indicate repression. Node colors represent functional classes: red (miRNAs), blue (targets), green (signaling pathway), yellow (cellular outcomes). b Comparison of Boolean model predictions with experimental qPCR expression data for miR-23a-3p and miR-17-5p. Bars represent relative fold change across clinical states: NWMI.a, NWMI.b, WMI.a, and WMI.b. qPCR data (lighter bars) are plotted against model-predicted activity states (darker bars). The model correctly predicts the downregulation of both miRNAs in WMI.a and partial restoration in WMI.b, mirroring experimental observations. c Simulated Boolean network outputs for key regulators of myelination in relation to miR-23a-3p and miR-17-5p expression. Heatmap shows simulated activation states (0 = OFF, 1 = ON) for each node across clinical conditions. Inputs (miR-23a and miR-17) regulate PTEN, which in turn controls activation of the PI3K/Akt pathway. This pathway promotes oligodendrocyte differentiation and MBP expression. High miRNA expression in NWMI states leads to full downstream activation, while their loss in WMI results in suppression of myelination.
We first compared the predicted input logic of the model to our experimental expression profiles. As shown in Fig. 4b, miR-23a and miR-17 were consistently high in NWMI samples and markedly reduced in WMI, with WMI.b samples showing partial recovery –a pattern recapitulated by the model proving that the Boolean framework accurately captured miRNA-driven regulatory shifts across the disease spectrum.
We next simulated downstream network activity across the four clinical conditions. In the NWMI states, high miR-23a/miR-17 expression repressed PTEN, thereby activating PI3K/Akt signaling and enabling Olig2 and MBP expression. In contrast, in WMI.a where both miRNAs were absent, the model predicted sustained PTEN activation, resulting in complete silencing of the myelination axis (Fig. 4c). Notably, WMI.b presented a more complex case: although qPCR data showed moderate recovery of both miRNAs, the model still predicted insufficient activity to reactivate downstream targets -highlighting a potential threshold-dependent behavior not fully resolved by binary logic55.
Collectively, our modeling supports the hypothesis that miR-23a and miR-17 act as upstream regulators of a PTEN-Akt-Olig2-MBP axis in human white matter development. Their concerted downregulation in WMI may contribute to impaired oligodendrocyte maturation and myelination failure observed in preterm brain injury, with partial recovery in WMI.b potentially representing a delayed compensatory response.
Discussion
This study evaluated the feasibility of using qPCR as a primary method for miRNA biomarker discovery in preterm WMI, with NGS serving as a secondary validation step. Traditionally, NGS is favored for its unbiased genome-wide capacity, but it poses challenges in vulnerable populations due to cost, technical demands, and biosample limitations. Our data reveal that when coupled with robust normalization strategies (e.g., spike-in controls) and systems biology tools, qPCR yields biologically relevant findings even in low-input settings. The congruence between qPCR and NGS for key miRNAs like miR-23a and miR-17 supports the reliability of qPCR in early-phase discovery.
Biological relevance of miR-23a and miR-17 in WMI
We identified miR-23a and miR-17 as potential regulators of oligodendrocyte maturation and myelination, based on differential expression across WMI states and enrichment in neurodevelopmental pathways. MiR-23a, known to influence OPC differentiation and fatty acid metabolism, showed reduced expression in early WMI (WMI.a) with partial recovery at later time points (WMI.b), suggesting a role in remyelination (Fig. 3a, c). The subtle recovery of miR-23a abundance in WMI.b samples after 28 days may indicate its involvement in the later stages of WMI repair, particularly through remyelination and the regulation of OPCs. Studies have shown that miR-23a promotes the differentiation of OPCs, which are crucial for remyelination. For instance, miR-23a, especially when delivered via extracellular vesicles, has been found to directly target genes like Olig3, which is essential for OPC differentiation and white matter repair56. In experimental models of brain injury, such as cerebral ischemia, the presence of miR-23a-5p led to improved white matter recovery and remyelination by promoting OPC survival and differentiation. This suggests that the recovery of miR-23a in WMI.b may contribute to ongoing repair mechanisms in the brain by supporting the formation of new myelin sheaths, which are crucial for restoring neural connectivity after injury.
We decided to explore the role of miR-17, the most abundant miRNA in our dataset (Fig. 2a), in biological processes by performing a KEGG analysis. MiR-17 was found to play a central role in regulating various biological and disease pathways (Fig. 3d) including autophagy, cancer, as well as axon guidance. The high abundance of miR-17 might be explained by its critical role in other physiological states. In conditions such as preterm brain injury, where WMI is prevalent, the downregulation of miR-17 could contribute to a failure in the proliferation and maturation of OPCs, ultimately resulting in defective myelination. Though not in the top 20 enriched pathways, miR-17 has been shown to target key genes involved in neuroinflammatory responses and cell cycle regulation through its miR-17-92 cluster53, both of which are crucial for the expansion and differentiation of the oligodendrocyte pool.
Low circulating abundance of miR-219 and miR-338 in preterm infants contrasts with model systems
Despite their established roles in oligodendrocyte differentiation through animal and cellular models, miR-219 and miR-338 exhibited low expression in qPCR and were undetectable in NGS. Methodologically, the convergence of both qPCR and NGS on this finding underscores its biological validity and demonstrates the utility of a multi-platform approach for validating low-abundance signals. Our findings suggest that the regulatory signatures described in in vitro oligodendrocyte models of miR-219a-2-3p and miR-338-5p in PTIs may not recapitulate their circulatory signatures in human PTIs. Unlike in transgenic models or cell culture systems where blood volumes can be collected repeatedly, and environmental variables tightly controlled, human neonatal research is constrained by ethical and biological limitations, including minimal allowable sample volumes and high inter-individual variability57. Their low AUCs further question their diagnostic utility in plasma-derived assays at this gestational stage. Moreover, the human in vivo biological environment exhibits systemic complexity and developmental variability that is rarely captured in reductionist experimental models, complicating direct translational inference58.
Of note, these miRNAs served as benchmark candidates to bridge our findings with established biology. Their absence became a critical result, revealing a fundamental disconnect between the circulating milieu of the human preterm infant and the regulatory mechanisms defined in reductionist models. This paradox justified our need to move beyond a conventional candidate approach and instead use systems biology to interpret the functional role of other, more abundant miRNAs like miR-23a and miR-17 that were mechanistically linked to WMI in our cohort. This raises a broader question for the field: should biomarker discovery rely solely on ‘modern’ technologies like NGS, which may lack sensitivity in ultra-low input clinical samples? Or does the robustness, sensitivity, and contextual versatility of established platforms like qPCR warrant a re-evaluation? In fragile populations such as PTIs, perhaps ‘old’ is not obsolete, but essential.
Systems biology enables deeper insights
Integrating qPCR findings with Boolean logic modeling enabled us to simulate how miR-23a and miR-17 regulate key targets like PTEN, a master inhibitor of PI3K/Akt signaling. The model is grounded in validated miRNA-target interactions, notably the repression of PTEN, a master inhibitor of the PI3K/Akt/mTOR pathway59,60,61. Boolean simulation revealed that the concurrent expression of both miRNAs is necessary to relieve PTEN-mediated inhibition, activate downstream signaling, and promote MBP expression (Fig. 4c). The Boolean network recapitulated experimental trends: miRNA expression was high in NWMI conditions and low in WMI.a, consistent with a repressive PTEN state and diminished myelination. Likewise, miR-17 is implicated in neuronal differentiation and axon pathfinding through its modulation of EphA4, a guidance cue whose dysregulation impairs oligodendrocyte-neuron coordination39.
Interestingly, in WMI.b, qPCR data showed partial miRNA recovery, yet the Boolean model retained an OFF state across the downstream pathway. This highlights a known limitation of binary logic: its inability to represent graded biological effects or threshold-dependent transitions55. Future models may benefit from fuzzy Boolean networks or probabilistic extensions to account for intermediate regulatory states, particularly in recovery phases of injury62.
Limitations and critical perspectives
While our study highlighted qPCR as a highly sensitive tool with impressive alignment with modeled biology, it is fundamentally a targeted method, which restricts its ability to discover novel transcripts. Therefore, our findings support a complementary role for NGS, particularly for discovery breadth. Interestingly, NGS analysis showed a weak and statistically insignificant upregulation of miR-23a and miR-17, in contrast to the robust downregulation observed in qPCR. This discrepancy may reflect several technical and biological confounders such as low RNA input or PCR amplification bias during library prep63,64. These considerations underscore the need to interpret platform-specific data cautiously and support the utility of dual-platform validation such as with a Boolean network.
Furthermore, the pilot nature of this study with limited sample sizes, although reduced by different normalization methods, still present a threat to statistical power and reliability65. Underpowered studies can produce inflated effect sizes and misinterpretations, particularly when evaluating subtle differences in miRNA expression. This concern is magnified in the context of preterm infants, where biological variability is high, and the consequences of misdiagnosis can be severe. Moreover, while the study suggests that qPCR is more sensitive for low-abundance miRNAs, this does not inherently validate its use as a standalone tool. We acknowledge that the small sample size, especially for NGS (n = 5), limits statistical power and increases risk of model overfitting. While bootstrapping provided confidence intervals that partially address this limitation, these findings remain exploratory and require validation in larger, independent cohorts. The combination of both methodologies could yield a more robust understanding of miRNA dynamics in preterm brain injury, rather than relying solely on the strengths of qPCR.
Future directions and clinical implications
Historical precedent reinforces the idea that qPCR, when strategically applied and interpreted within a systems framework, retains utility not only for validation but for targeted discovery, particularly in clinically constrained settings such as neonatal biomarker research. Therefore, future work should re-evaluate the historical performance of qPCR in other discovery pipelines to develop best practices for qPCR –when guided by mechanistic modeling and supported by select NGS biomarker studies. Such approaches might strengthen our understanding of miRNA-mediated regulation for initial discovery in constrained clinical settings.
Conclusion
Our results demonstrate that qPCR, when integrated with systems biology, could serve as a credible and cost-effective tool for pilot phase in miRNA discovery in neonatal WMI. A strategically designed hybrid qPCR-NGS-systems biology pipeline may offer the best balance between sensitivity, specificity, and translational feasibility for biomarker research in fragile populations such as preterm infants.
Data availability
Data from the current study will be made available at the request of qualified investigators if approved by our Research Ethics Board at the end of the overall study period.
References
Murray, A. L. et al. ‘White matter abnormalities and impaired attention abilities in children born very preterm’. NeuroImage 124, 75–84 (2016).
Cainelli, E., Arrigoni, F. & Vedovelli, L. ‘White matter injury and neurodevelopmental disabilities: a cross-disease (dis)connection’. Prog. Neurobiol. 193, 101845 (2020).
Bax, M., Tydeman, C. & Flodmark, O. ‘Clinical and MRI correlates of cerebral palsy: the European Cerebral Palsy Study’. JAMA: J. Am. Med. Assoc. 296, 1602–1608 (2006).
Shah, D. K. et al. Adverse neurodevelopment in preterm infants with postnatal sepsis or necrotizing enterocolitis is mediated by white matter abnormalities on magnetic resonance imaging at term’. J. Pediatr. 153, 170–5–175.e1 (2008).
Volpe, J. J. Neurology of the Newborn. 5th edn, (Saunders, 2008)
Dyet, L. E. et al. ‘Natural history of brain lesions in extremely preterm infants studied with serial magnetic resonance imaging from birth and neurodevelopmental assessment’. Pediatrics 118, 536–548 (2006).
Jeon, T. Y. et al. ‘Neurodevelopmental outcomes in preterm infants: comparison of infants with and without diffuse excessive high signal intensity on MR images at near-term-equivalent age’. Radiology 263, 518–526 (2012).
Lawrie, C. H. et al. ‘MicroRNA expression distinguishes between germinal center B cell-like and activated B cell-like subtypes of diffuse large B cell lymphoma’. Int. J. Cancer. 121, 1156–1161 (2007).
Abdelrahman, A. H. et al. ‘Evaluation of circulating miRNAs and mRNAs expression patterns in autism spectrum disorder’. Egypt. J. Med. Hum. Genet. 22, 1–10 (2021).
Kuwabara, Y. et al. ‘Increased microRNA-1 and microRNA-133a levels in serum of patients with cardiovascular disease indicate myocardial damage’. Circ. Cardiovasc. Genet. 4, 446–454 (2011).
Cortez, M. A. et al. ‘MicroRNAs in body fluids-the mix of hormones and biomarkers’. Nat. Rev. Clin. Oncol. 8, 467–477 (2011).
Cuellar, T. L. et al. ‘Dicer loss in striatal neurons produces behavioral and neuroanatomical phenotypes in the absence of neurodegeneration’. Proc. Natl. Acad. Sci. USA 105, 5614–5619 (2008).
Dugas, J. C. & Notterpek, L. ‘MicroRNAs in oligodendrocyte and Schwann cell differentiation’. Dev. Neurosci. 33, 14–20 (2011).
Doculara, L. et al. ‘Circulating tumor DNA in pediatric cancer’. Front. Mol. Biosci. 9, 885597 (2022).
Janssen, F. W. et al. A comprehensive overview of liquid biopsy applications in pediatric solid tumors’. npj Precis. Oncol. 8, 172 (2024).p.
Markou, A. et al. ‘Prognostic value of mature microRNA-21 and microRNA-205 overexpression in non-small cell lung cancer by quantitative real-time RT-PCR’. Clin. Chem. 54, 1696–1704 (2008).
Cissell, K. A. & Deo, S. K. ‘Trends in microRNA detection’. Anal. Bioanal. Chem. 394, 1109–1116 (2009).
Xing, C., Di, X., Qi, Z. & Zhu-Hong, Y. MicroRNAs and complex diseases: from experimental results to computational models. Briefings in Bioinformatics 20, 515–539 (2019).
Ye, J., Xu, M., Tian, X., Cai, S. & Zeng, S. ‘Research advances in the detection of miRNA’. J. Pharm. Biomed. Anal. 9, 217–226 (2019).
Iroanya, O. O. et al. ‘Stability of selected microRNAs in human blood, semen and saliva samples exposed to different environmental conditions’. Forensic Sci. Int. 336, 111338 (2022).
Mariana Lagos-Quintana et al. Identification of novel genes coding for small expressed RNAs. Science 294, 853–858 (2001).
Schmittgen, D. T., Jiang, J., Liu, Q. & Yang, L. A high-throughput method to monitor the expression of microRNA precursors. Nucleic Acids Res. 32, e43 (2004).
Saez-Rodriguez, J. et al. A logical model provides insights into T cell receptor signaling’. PLoS Comput. Biol. 3, e163 (2007).
Balaskas, N. et al. ‘Gene regulatory logic for reading the Sonic Hedgehog signaling gradient in the vertebrate neural tube’. Cell. 148, 273–284 (2012).
Steinway, S. N. et al. ‘Combinatorial interventions inhibit TGFβ-driven epithelial-to-mesenchymal transition and support hybrid cellular phenotypes’. npj Syst. Biol. Appl. 1, 15014 (2015).
miRNet 2.0 (no date). Available at: https://www.mirnet.ca/miRNet/home.xhtml (accessed: 1 2025).
Chang, L., & Xia, J. MicroRNA regulatory network analysis using miRNet 2.0. in Transcription Factor Regulatory Networks. (eds Song, Q., Tao, Z.) 2594 (Humana, 2023).
miRPathDB (no date). Available at: https://mpd.bioinf.uni-sb.de/overview.html (accessed: 1 2025).
Kehl, T. et al. ‘miRPathDB 2.0:a novel release of the miRNA pathway dictionary database’. Nucleic Acids Res. 48, D142–D147 (2020).
Back, S. A. ‘Perinatal white matter injury:the changing spectrum of pathology and emerging insights into pathogenetic mechanisms’. Ment. Retard. Dev. Disabil. Res. Rev. 12, 129–140 (2006).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677 (2025).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Milacic, M. et al. The reactome pathway knowledgebase 2024. Nucleic Acids Res. 52, D672–D678 (2024).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
The Gene Ontology Consortium. The Gene Ontology knowledgebase in 2023. Genetics 224, iyad031 (2023).
Huang, D., Sherman, B. & Lempicki, R. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50, W216–W221 (2022).
Altounian, M., Bellon, A. & Mann, F. ‘Neuronal miR-17-5p contributes to interhemispheric cortical connectivity defects induced by prenatal alcohol exposure’. Cell Rep. 42, 113020 (2023).
Qiu, S. et al. ‘The therapeutic potential of microRNAs to ameliorate spinal cord injury by regulating oligodendrocyte progenitor cells and remyelination’. Front. Cell. Neurosci. 18, 1404463 (2024).
Baumann, N. & Pham-Dinh, D. ‘Biology of oligodendrocyte and myelin in the mammalian central nervous system’. Physiol. Rev. 81, 871–927 (2001).pp.
Leong, S. Y. et al. ‘Heterogeneity of oligodendrocyte progenitor cells in adult human brain’. Ann. Clin. Transl. Neurol. 1, 272–283 (2014).
Galloway, D. A. & Moore, C. S. ‘MiRNAs as emerging regulators of oligodendrocyte development and differentiation’. Front. Cell Dev. Biol. 4, 59 (2016).
Zhao, X. et al. ‘MicroRNA-mediated control of oligodendrocyte differentiation’. Neuron 65, 612–626 (2010).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Xie, Z. et al. Gene set knowledge discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Sherman, S. M. ‘The function of metabotropic glutamate receptors in thalamus and cortex’. Neuroscientist 20, 136–149 (2014).
Amer, A. A. A. et al. ‘Relationship between white matter integrity and plasma Leptin levels in drug-naïve and medicated patients with major depressive disorder’. Front. Neurosci. 13, 707 (2019).
Lee, W. onS. uk, Won-Ha Lee, Y. ongC. hulB. ae & Suk, K. youngho “Axon guidance molecules guiding neuroinflammation.”. Exp. Neurobiol. 28, 311–319 (2019).
Chédotal, A. lain “Roles of axon guidance molecules in neuronal wiring in the developing spinal cord”. Nat. Rev. Neurosci. 20, 380–396 (2019).
Harrington, E. P. et al. ‘Oligodendrocyte PTEN is required for myelin and axonal integrity, not remyelination’. Ann. Neurol. 68, 703–716 (2010).
Budde, H. et al. ‘Control of oligodendroglial cell number by the miR-17-92 cluster’. Development 137, 2127–2132 (2010).
Fancy, S. P. J. et al. ‘Overcoming remyelination failure in multiple sclerosis and other myelin disorders’. Exp. Neurol. 225, 18–23 (2010).
Saez-Rodriguez, J. et al. ‘Discrete logic modelling as a means to link protein signalling networks with functional analysis of mammalian signal transduction’. Mol. Syst. Biol. 5, 331 (2009).
Li, Y. et al. ‘M2 microglia-derived extracellular vesicles promote white matter repair and functional recovery via miR-23a-5p after cerebral ischemia in mice’. Theranostics 12, 3553–3573 (2022).
Nissimov S., Sibrecht G., Weerasekara I., Bartocci M., Bruschettini M. Minimizing blood sampling in preterm infants. Cochrane Database of Systematic Reviews 11, CD016077 (2024).
Mak, I. W., Evaniew, N. & Ghert, M. ‘Lost in translation: animal models and clinical trials in cancer treatment’. Am. J. Transl. Res. 6, 114–118 (2014).
Hu, Z. et al. miRNA-17 promotes nasopharyngeal carcinoma radioresistance by targeting PTEN/AKT. Int. J. Clin. Exp. Pathol. 12, 229–240 (2019).
Rao, E. et al. The miRNA-17∼92 cluster mediates chemoresistance and enhances tumor growth in mantle cell lymphoma via PI3K/AKT pathway activation. Leukemia 26, 1064–1072 (2012).
Zhiheng, D. et al. lncRNA GAS5 restrains CCl4-induced hepatic fibrosis by targeting miR-23a through the PTEN/PI3K/Akt signaling pathway. Am. J. Physiol.-Gastrointest. Liver Physiol. 316, G539–G550 (2019).
Stoll, G. et al. ‘MaBoSS 2.0: an environment for stochastic Boolean modeling’. Bioinformatics 33, 2226–2228 (2017).
Roche Sequencing and Life Science. 2022. Top tips for RNA-sequencing that involves degraded inputs. News-Medical, viewed 03 2025, https://www.news-medical.net/whitepaper/20220802/Top-tips-for-RNA-sequencing-that-involves-degraded-inputs.aspx.
Barberán-Soler, S. et al. ‘Decreasing miRNA sequencing bias using a single adapter and circularization approach’. Genome Biol. 19, 105 (2018).
Kok, M. G. M. et al. ‘Small sample sizes in high-throughput miRNA screens: A common pitfall for the identification of miRNA biomarkers’. Biomol. Detect. Quantif. 15, 1–5 (2018).
Acknowledgements
We thank the families who participated in this research and acknowledge Genecore, Germany, and Dr. Vladimir’s lab for their generosity in offering their NGS sequencing services of our samples.
Funding
Lolia Ala Ibanibo, an early stage researcher of the PARENT Project, received funding from the European Union’s Horizon 2020 research and innovation programme under the Maria Sklodowska-Curie − Innovative Training Network 2020, Grant Agreement N° 956394. Raúl Montañez Martínez and Simón Lubián López received funding from Andalusian Ministry of Health and Families, Spain-FEDER funds (PIGE-0026-2020).
Author information
Authors and Affiliations
Contributions
L.A.I., R.M.M., and S.L.L designed and conceptualized the study. I.B.F. and S.L.L. supported participant recruitment. M.L.G. and I.B.F performed clinical data annotation, including Kidokoro scoring. L.A.I. conducted the experimentation and investigation. A.O.L. analyzed the NGS data. L.A.I. completed the primary data analysis and figure creation. L.A.I. drafted the manuscript with all other authors providing substantive feedback and final approval. R.M.M., I.B.F., and S.L.L. acquired funding. R.M.M. and S.L.L. supervised the project. I.B.F. and S.L.L. administered the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
ETHICAL CONSIDERATIONS
Informed consent was obtained from the parents or legal guardians of all participants included in the study. The study was conducted in accordance with the Declaration of Helsinki and was approved by the Research and Ethics Committee (Protocol number: PIGE-0026-2020; code: 117.21).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ala Ibanibo, L., Montañez-Martínez, R., Ortega Leon, A. et al. Circulating microRNAs in preterm white matter injury: a systems biology/qPCR-based pilot study. Pediatr Res (2026). https://doi.org/10.1038/s41390-026-04854-3
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41390-026-04854-3