Abstract
Dendritic cells (DCs) regulate both innate and adaptive immunity during sepsis. Prostaglandins (PGs), small lipid molecules derived from arachidonic acid via COX enzymes, are crucial regulators of immune homeostasis and inflammation. However, their role in sepsis pathogenesis remains poorly defined. In this study, we identified a significant negative correlation between DC depletion and disease severity in patients with sepsis. Thromboxane (TX) A2 receptor (TP) expression was markedly reduced in the blood DCs of patients with sepsis. Patients with low DC-TP expression presented increased blood neutrophil counts and worsened disease severity. In murine models of sepsis induced by cecal ligation and puncture and lipopolysaccharide challenge, DC-specific TP deficiency exacerbated sepsis by promoting S100a8/a9-mediated neutrophil recruitment and, subsequently, neutrophil extracellular trap (NET) formation and lung injury. Genetic and pharmacological inhibition of the S100a8/a9-TLR4 axis protected TP-deficient mice from fatal sepsis. Mechanistically, TP signaling suppressed S100a8/a9 expression in DCs via PKCδ-Stat1 signaling, thereby restricting neutrophil infiltration and NET formation. Finally, the targeted activation of TP in DCs via the nanodrug DCpep-U-46619 effectively alleviated sepsis-induced lung injury in mice. These findings establish TP as a critical immunoregulatory receptor in DCs, highlighting its potential as a therapeutic target for sepsis.
Similar content being viewed by others
Introduction
Sepsis, a life-threatening condition triggered by pathogenic infection, primarily affects the respiratory or gastrointestinal system and leads to multiorgan dysfunction.1 With approximately 50 million cases reported annually, sepsis remains a major global health burden and one of the leading causes of mortality in intensive care units (ICUs).2 Current clinical management relies largely on supportive measures, including early diagnosis and fluid resuscitation, the rapid delivery of effective antibiotics, and lung protective ventilation. However, therapeutic strategies aimed at broadly normalizing physiological parameters—such as gas exchange, glucose control, or oxygen delivery—have proven largely ineffective or, in some cases, detrimental.3 Consequently, there remains a critical lack of targeted therapies, particularly small-molecule drugs, highlighting an urgent unmet need for precision interventions to improve outcomes in sepsis.
Despite advances in critical care and immunology, the pathogenesis of sepsis remains incompletely understood, especially the complex regulatory networks governing immune cell function. A central pathological feature of sepsis is an excessive or dysregulated immune response to infection, ultimately leading to multiple organ dysfunction.4 However, anti-TNF therapy increases mortality in septic shock patients5 and the benefits of corticosteroids remain controversial.6 These outcomes highlight the need for more precise immunomodulatory strategies that restore immune balance rather than indiscriminately suppress inflammation. Dendritic cells (DCs), as antigen-presenting cells bridging innate and adaptive immunity, play important and context-dependent roles in sepsis. On one hand, DCs initiate immune responses through recognition of pathogen- and damage-associated molecular patterns; on the other, their dysfunction or exhaustion contributes to immunosuppression and disease progression.7 Clinical and experimental evidence demonstrates profound DC impairment during sepsis. Patients with sepsis exhibit increased apoptosis of splenic DCs, reduced circulating DC numbers, and diminished antigen-presenting capacity, collectively compromised antimicrobial immunity.8 In murine models, functionally paralyzed DCs emerge in the lungs following viral or bacterial pneumonia, producing excessive TGF-β that promotes regulatory T cell accumulation and establishes a highly immunosuppressive microenvironment.9 Similarly, DCs in the spleen and lymph nodes during sepsis fail to fully mature—characterized by reduced expression of CD80, CD86, and CD40—and secrete more IL-10, thereby impairing antigen presentation and T-cell priming.7 Newly generated DCs after sepsis also exhibit a persistently hypoimmunogenic phenotype, with reduced capacity for antigen uptake, processing, and T-cell activation.10 Importantly, DC dysfunction in sepsis extends beyond adaptive immune suppression and may also impair innate immune activation, including natural killer (NK) cell responses to bacterial infection.7 Despite the central role of DCs in sepsis-associated immune dysregulation, the mechanisms underlying DC functional impairment in sepsis remain poorly understood. In particular, the molecular pathways that drive DC paralysis—and whether this state is reversible—have yet to be defined.
Prostanoids are bioactive lipid mediators derived from arachidonic acid through the sequential actions of cyclooxygenases (COXs) and specific prostaglandin (PG) synthases. Five major prostanoids—PGE₂, PGI₂, PGD₂, PGF₂α, and thromboxane A₂ (TXA₂)—exert distinct biological effects through binding to a diverse family of G protein–coupled prostanoid receptors. Among these, TXA2 is best known for its role in platelet activation, vasoconstriction, and angiogenesis via signaling through thethromboxane-prostanoid (TP) receptor.11,12 The clinical efficacy of low-dose aspirin, which irreversibly inhibits platelet COX-1–derived TXA₂, underscores the importance of this pathway in cardiovascular disease.13 However, antiplatelet therapies have failed to significantly improve disease severity or outcomes in sepsis.14,15,16 Emerging evidence suggests that the TXA₂/TP axis also exerts immunomodulatory functions: TP agonist enhances chemokinesis of naive but not memory T cells, impairs DC-T cell adhesion, and inhibits DC-dependent proliferation of T cells. Consistently, immune responses to foreign antigens are enhanced in TP-deficient mice17; Mechanistically, TXA₂ selectively disrupts low-avidity—but not high-avidity—interactions between DCs and CD4+ T cells, and this tonic suppression of weak CD4+ T cell-DC interactions by TXA2-TP signaling has been proposed to improve the overall quality of adaptive immune responses18; In addition, TXA2 can act directly on T cells to trigger an immunosuppressive pathway dependent on the guanine exchange factor ARHGEF1, thereby suppressing T cell receptor-driven kinase signaling, proliferation and effector functions.19 Nevertheless, the role of TXA₂-TP signaling in regulating immune responses during sepsis remains largely unexplored.
Here, we revealed a significant reduction in TP expression in the DCs of patients with sepsis and CLP-challenged mice, which was negatively correlated with disease severity. Mice lacking TP in their DCs had worse sepsis outcomes, including higher mortality rates, more severe inflammation, increased neutrophil infiltration, neutrophil extracellular trap (NET) formation, and lung injury. TP signaling in DCs suppressed the expression of S100a8/a9 via the protein kinase Cδ (PKCδ)-signal transducer and activator of transcription (Stat)-1 pathway, reducing neutrophil recruitment. Blocking S100a8/a9 or its receptor, toll-like receptor-4 (TLR4), protects TP-deficient mice from severe sepsis by reducing inflammation and NET formation. Finally, the activation of TP in DCs via a DC-targeting nanodrug (DCpep-U-46619) reduced neutrophil recruitment and improved survival in sepsis. These findings suggest that TP in DCs plays a role in regulating the immune balance in sepsis and may be a potential therapeutic target.
Results
TP receptor expression in DCs is negatively correlated with sepsis severity
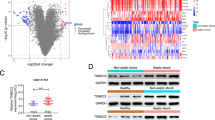
To comprehensively analyze immune alterations during sepsis, we analyzed a publicly available single-cell RNA sequencing dataset of peripheral blood mononuclear cells (PBMCs) from 19 healthy controls and 29 patients with sepsis.20 After removing doublets and low-quality cells, a total of 92,761 cells with 22,858 detected genes were analyzed. Unsupervised downscaling and T-distributed stochastic neighbor embedding (t-SNE) clustering revealed 22 cell clusters that were visualized separately in patients with sepsis and healthy controls (Fig. 1a). On the basis of the expression of specific markers, cells are classified into five major subtypes: monocytes (LYZ and FCN1), DCs (JCHAIN, LILRA4, CLEC9A and CD1C), natural killer (NK) cells (KLRD1 and NKG7), T cells (CD2 and CD3D), and B cells (MS4A1 and CD79A).20,21 Notably, we observed a substantial reduction in the proportions of DCs, NK cells, and T-cell subsets in patients with sepsis, while the monocyte count was significantly increased (Fig. 1b). Furthermore, correlation analysis revealed a significant negative association between the proportion of DCs and disease severity in patients with sepsis, suggesting a decrease in DCs with increasing disease severity (Fig. 1c and Supplementary Fig. 1). These findings highlight the profound effects of sepsis on peripheral immune cell populations, particularly DCs. PGs are critical mediators of immune responses and inflammation. According to the Cell Gene database, PG receptors are widely expressed in normal human peripheral blood DCs (Fig. 1d). To investigate the potential involvement of PG signaling in DC dysfunction during sepsis, we analyzed the expression of various PG receptors in DCs in the blood of patients with sepsis. Among these genes, TP expression was significantly negatively correlated with disease severity (Fig. 1e and Supplementary Fig. 2). In addition, TP expression in DCs was markedly lower in patients with sepsis than in healthy controls (Fig. 1f). Consistent with the human data, DCs isolated from the spleens of lipopolysaccharide (LPS)- and cecal ligation and puncture (CLP)-treated mice (Fig. 1g) exhibited a significant decrease in TP expression in both sepsis models (Fig. 1h, i). To further explore the functional role of TP in DCs during sepsis, we performed adoptive transfer experiments in which DCs from TP-deficient (TP−/−) mice were transferred into wild-type (WT) recipients. Notably, the transfer of TP−/− DCs exacerbated the severity of sepsis in CLP-treated WT mice, as evidenced by decreased survival rates (Fig. 1j), increased serum inflammatory cytokine levels (Fig. 1k, l), lung permeability (Fig. 1m), lung edema (Fig. 1n, o), neutrophil infiltration (Fig. 1p), and lung injury scores (Fig. 1q, r) in these mice. Collectively, these findings suggest that TP downregulation in DCs may impair their function and contribute to the exacerbation of sepsis severity.
TP receptor expression in DCs is negatively correlated with sepsis severity. a scRNA-seq analysis (SCP548) of immune cells in PBMCs from healthy controls (n = 19) and patients with sepsis (n = 29). b Percentages of immune cell populations in each cluster for both groups. c Correlation analysis between the percentage of DCs and the SOFA score in patients with sepsis (n = 19 [control] and n = 29 [sepsis]). d scRNA-seq analysis of PG receptor expression in DCs from the PBMCs of healthy individuals. e Correlation analysis between TP receptor expression in DCs and SOFA score in sepsis patients (n = 10 [control] and n = 24 [sepsis]). f Expression of TP in DCs from the PBMCs of controls and patients with sepsis (n = 13 [control] and n = 24 [sepsis]). g Schematic representation of the sepsis model induced via lipopolysaccharide (LPS) and cecal ligation and puncture (CLP). h, i TP expression in primary DCs isolated from the spleens of LPS-treated and CLP-challenged mice (n = 7–8 mice per group). j Survival curves of CLP-challenged mice after adoptive transfer of WT DCs or TP−/− DCs (n = 28 mice per group). k, l Plasma concentrations of proinflammatory (IL-1β) and anti-inflammatory (IL-10) cytokines in CLP-challenged mice after adoptive transfer of WT DCs or TP−/− DCs (n = 8 mice per group). m Lung vascular permeability detected by EBD leakage in CLP-challenged mice after adoptive transfer of WT DCs or TP−/− DCs (n = 6 mice per group). Lung wet/dry weight ratio (n) (n = 8 mice per group) and BALF protein levels (o) (n = 8 mice per group) in CLP-challenged mice after adoptive transfer of WT DCs or TP−/− DCs. p Quantitative analysis of neutrophil infiltration via measurement of MPO activity in lung tissue from CLP-challenged mice after adoptive transfer of WT DCs or TP−/− DCs (n = 6 mice per group). q Representative histological images of lungs from CLP-challenged mice after adoptive transfer of WT DCs or TP−/− DCs. Scale bars, 50 μm. r Quantification of the histology scores for lung injury in (q) (n = 9 mice per group). Statistical significance was evaluated via Spearman’s correlation coefficient (c, e), Mann–Whitney U tests (f, h, i, k–p, r), or the log-rank test (j)
TP ablation in DCs exacerbates CLP- and LPS-induced sepsis in mice
To investigate the role of TP in the contribution of DCs to sepsis pathogenesis, we performed CLP in DC-conditional TP knockout mice (TPflox/floxCD11cCre) and their littermate controls (Supplementary Fig. 3). Notably, TP deficiency in DCs resulted in significantly higher mortality rates in mice with sepsis than in control mice (Fig. 2a). This increased susceptibility was associated with elevated levels of interleukin (IL)-1β (Fig. 2b) and IL-10 in the blood (Fig. 2c) and increased lung permeability (Fig. 2d), edema (Fig. 2e, f) and neutrophil infiltration (Fig. 2g). Moreover, histological analysis revealed that, compared with control mice, TPflox/floxCD11cCre mice presented more severe lung injury, as evidenced by worse lung injury scores and increased perivascular neutrophil enrichment (Fig. 2h, i and Supplementary Fig. 4). Similar findings were observed in an LPS-induced sepsis model, where TP deficiency in DCs exacerbated disease severity, increased inflammatory cytokine levels, and caused increased neutrophil infiltration and severe lung damage (Fig. 2j–r). Thus, TP deletion in DCs aggravated lung injury in CLP- and LPS-challenged mice.
TP ablation in DCs exacerbates CLP- and LPS-induced sepsis in mice. a Survival analysis of CLP-challenged TPflox/flox (n = 20 mice) and TPflox/floxCD11cCre (n = 27 mice) mice. b, c Plasma concentrations of proinflammatory (IL-1β) and anti-inflammatory (IL-10) cytokines in CLP-challenged TPflox/flox and TPflox/floxCD11cCre mice (n = 8 mice per group). d Lung vascular permeability detected by EBD leakage in TPflox/flox and TPflox/floxCD11cCre mice at 0, 1, and 3 days post-CLP (n = 6 mice per group). Lung wet/dry weight ratio (e) (n = 6–8 mice per group) and BALF protein levels (f) (n = 6–8 mice per group) in lungs from TPflox/flox and TPflox/floxCD11cCre mice at 0, 1, and 3 days post-CLP. g Quantitative analysis of neutrophil infiltration via measurement of MPO activity in lung tissue from TPflox/flox and TPflox/floxCD11cCre mice at 0, 1, and 3 days post-CLP (n = 6 mice per group). h Representative histological images of lungs from CLP-challenged (0, 1, and 3 days) TPflox/flox and TPflox/floxCD11cCre mice. Scale bars, 50 μm. i Quantification of histology scores for lung injury in (h) (n = 6–9 mice per group). j Survival analysis of LPS-treated TPflox/flox (n = 15 mice) and TPflox/floxCD11cCre (n = 15 mice) mice. k, l Plasma cytokine levels of proinflammatory (IL-1β) and anti-inflammatory (IL-10) markers in LPS-treated TPflox/flox and TPflox/floxCD11cCre mice (n = 8 mice per group). m Lung vascular permeability detected by EBD leakage in LPS-treated (0, 8, and 24 h) TPflox/flox and TPflox/floxCD11cCre mice (n = 6 mice per group). Lung wet/dry weight ratio (n) (n = 6 mice per group) and BALF protein levels (o) (n = 5–6 mice per group) in LPS-treated (0, 8, and 24 h) TPflox/flox and TPflox/floxCD11cCre mice. p Quantitative analysis of neutrophil infiltration via measurement of MPO activity in lung tissue from LPS-treated (0, 8, and 24 h) TPflox/flox and TPflox/floxCD11cCre mice (n = 6 mice per group). q Histological analysis of lung sections from LPS-treated (0, 8, and 24 hours) TPflox/flox and TPflox/floxCD11cCre mice. Scale bars, 50 μm. r Lung histological scores corresponding to (q) (n = 4 mice at 0 h, 6–8 mice at 8 and 24 h). Statistical significance was evaluated via two-way ANOVA followed by Sidak’s test for multiple comparisons (b‒g, i, k‒p, r) or the log-rank test (a and j)
TP deficiency in DCs aggravates CLP-induced sepsis in mice through the recruitment of hyperactivated neutrophils
To elucidate the molecular mechanisms by which TP in DCs influences sepsis progression, we performed single-cell RNA sequencing (scRNA-seq) of total immune cells (CD45+) isolated from the spleens of mice subjected to CLP. Following stringent quality control, 54,774 cells were analyzed, resulting in the identification of 27,236 genes. The cells were analyzed and visualized via t-SNE to assess qualitative changes in the immune cell composition within the spleen (Fig. 3a). Clustering analysis identified 10 distinct immune cell types on the basis of the expression of canonical marker genes: B cells (Ms4a1, Cd79a, and Cd19), CD4+ T cells (Cd2, Cd3d, and Cd4), CD8+ T cells (Cd2, Cd3d, and Cd8a), NK cells (Klrd1, Nkg7, Klrb1c, and Ccl5), eosinophils (Prss34), basophils (Prg2), DCs (Fscn1, Apol7c, Cd74, Naaa, and Siglech), macrophages (C1qa and C1qb), monocytes (Csf1r and Fn1) and neutrophils (Lcn2, Ltf, and Camp) (Fig. 3b). Interestingly, the loss of TP in DCs did not significantly affect the total percentage or number of DCs in the spleen during sepsis (Supplementary Fig. 5a, b). As DCs are highly efficient APCs involved in T-cell proliferation and activation, we further analyzed adaptive immune responses and detected no significant changes in T-cell subsets or B cells in TPflox/floxCD11cCre mice compared with those in the controls (Supplementary Fig. 5c–p). However, TP deficiency resulted in a significant increase in the proportion of neutrophils in mice after CLP (Fig. 3c–e). To investigate whether DCs from CLP mice directly influence neutrophil recruitment, we conducted a coculture assay to assess DC-induced neutrophil chemotaxis (Fig. 3f). Our results demonstrated that DCs from mice with sepsis presented increased neutrophil recruitment, which was further exacerbated by TP deficiency (Fig. 3g). Notably, a significant negative correlation was observed between TP expression in DCs and the neutrophil ratio in patients with sepsis (Fig. 3h), suggesting that the loss of TP in DCs may exacerbate sepsis by promoting neutrophil infiltration.
TP deficiency enhances neutrophil recruitment by increasing S100a8/a9 expression. a scRNA-seq analysis of CD45+ immune cells isolated from the spleens of TPflox/flox and TPflox/floxCD11cCre (n = 2 mice per group) mice at 3 days post-CLP. b Dot plots showing the expression level of the canonical annotation marker used for cluster annotation. c Proportions of immune cell clusters in the spleens of TPflox/flox and TPflox/floxCD11cCre mice. Percentages (d) and numbers (e) of neutrophils (CD45+CD3−Ly6G+) in the spleens of TPflox/flox and TPflox/floxCD11cCre mice at 0, 1, and 3 days post-CLP. (n = 4–6 mice per group). f Schematic of the neutrophil transwell migration assay. CFSE-labeled neutrophils from bone marrow were placed in the upper chamber, whereas DCs from TPflox/flox or TPflox/floxCD11cCre mice were seeded in the lower chamber. Neutrophil migration was assessed after 2 h (Created with BioRender.com). g Quantification of neutrophil migration toward DCs from TPflox/flox and TPflox/floxCD11cCre mice at 0 and 3 days post-CLP. (n = 6 mice per group). h The correlation between TP receptor expression in blood DCs and neutrophil percentages in patients with sepsis (n = 12 [control] and n = 23 [sepsis]) was analyzed. i Volcano plot of differentially expressed genes in DCs between TPflox/flox and TPflox/floxCD11cCre mice (p.adjust < 0.05, Log2FC ≥ 0.5). j Top 10 enriched GO pathways associated with DEGs in TPflox/floxCD11cCre vs TPflox/flox DCs. The mRNA expression of S100a8 (k) and S100a9 (l) in primary DCs from the spleens of CLP-challenged mice (n = 4–6 mice per group). Plasma concentrations of S100a8 (m) and S100a9 (n) in CLP-treated (0 and 3 days) TPflox/flox and TPflox/floxCD11cCre mice (n = 8 mice per group). o, p Correlation analysis between TP and S100A8/A9 expression in blood DCs from patients with sepsis (n = 12 [control] and n = 17 [sepsis]). q Neutrophil migration assay using DCs from TPflox/flox and TPflox/floxCD11cCre mice post-CLP (3 days), with or without paquinimod (n = 6–9 mice per group). r Schematic illustration showing that the deletion of TP promotes DC-mediated neutrophil recruitment through increased S100a8/a9 expression (Created with BioRender.com). Statistical significance was evaluated via Spearman’s correlation coefficient (h, o, p) and two-way ANOVA, followed by Sidak’s test (d, e) or Tukey’s test (g, k–n, q) for multiple comparisons
To further investigate the mechanisms underlying increased neutrophil recruitment in TP-deficient mice, we compared the transcriptional profiles of DCs from control (TPflox/flox) and TP-deficient (TPflox/floxCD11cCre) mice following CLP. Differential gene expression analysis revealed 85 genes whose expression was significantly altered in TP-deficient DCs (Fig. 3i). Gene ontology enrichment analysis revealed that many of these genes, including S100a8 and S100a9, which were significantly upregulated in TP-deficient DCs, were associated with leukocyte migration (Fig. 3j). The increased expression of S100a8 and S100a9 was further validated at the transcriptional level via reverse transcription‒polymerase chain reaction (RT‒PCR) (Fig. 3k, l). Moreover, plasma analysis revealed significantly elevated levels of S100a8 and S100a9 in TP-deficient mice during sepsis (Fig. 3m, n). In patients with sepsis, TP expression was significantly negatively correlated with S100A8/A9 expression in blood DCs (Fig. 3o, p). To assess the functional significance of S100a8/a9 upregulation, we treated DCs with paquinimod, a potent S100a8/a9 antagonist. Paquinimod treatment significantly attenuated the increase in neutrophil recruitment by TP-deficient DCs (Fig. 3q), further supporting the essential role of S100a8/a9 in mediating neutrophil recruitment by TP-deficient DCs (Fig. 3r). In addition, neutrophil subpopulation analysis revealed that TP deletion significantly increased the percentage of the hyperactivated subpopulation (N2) during sepsis, which was consistent with previous findings (Supplementary Fig. 6a, b).22 This subpopulation was enriched in gene ontology terms associated with neutrophil activation and killing of cells of another organism (Supplementary Fig. 6c), as indicated by the upregulation of genes involved in neutrophil activation, degranulation, and NETosis (Supplementary Fig. 6d), and N2 was a more mature or terminal cell state derived from N1 (Supplementary Fig. 6e). Moreover, assessment of neutrophil deposition and neutrophil extracellular trap (NET) formation revealed a marked increase in NET-related protein expression in the lungs of TPflox/floxCD11cCre mice following CLP challenge compared with that in the lungs of littermate controls (Supplementary Fig. 6f–i). Collectively, these results suggest that TP activation in DCs mitigates the severity of sepsis by downregulating S100a8/a9 expression, thereby reducing neutrophil recruitment and tissue damage.
Blockade of the S100a8/a9-TLR4 axis abrogates neutrophil recruitment and alleviates sepsis in TP-deficient mice
To explore the role of TP-deficient DC-derived S100a8/a9 in neutrophil recruitment in sepsis, we generated DC-specific S100a9-deficient (S100a9flox/floxCD11cCre) and TP and S100a9 double-knockout (TPflox/floxS100a9flox/floxCD11cCre) mice. Remarkably, the deletion of S100a9 in DCs completely slowed the increased mortality rate (Fig. 4a) and cytokine production (Fig. 4b, c) observed in TP-deficient CLP mice, indicating that DCs are a key source of S100a8/a9 in sepsis. Importantly, S100a9 deficiency did not affect the DC count (Fig. 4d–f) but significantly reduced neutrophil infiltration in both S100a9flox/floxCD11cCre and TPflox/floxS100a9flox/floxCD11cCre mice (Fig. 4g–i). Moreover, NET formation (Fig. 4j–n) and lung permeability, edema, and injury (Fig. 4o–s), which were exacerbated by TP deficiency, were significantly alleviated by the deletion of S100a9.
Deletion of S100a9 in DCs alleviates CLP-induced sepsis in mice. a Survival analysis of CLP-challenged CD11cCre, TPflox/floxCD11cCre, S100a9flox/floxCD11cCre, and TPflox/floxS100a9flox/floxCD11cCre mice (n = 20–26 mice per group). b, c Plasma cytokine levels of proinflammatory (IL-1β) and anti-inflammatory (IL-10) cytokines in CLP-challenged CD11cCre, TPflox/floxCD11cCre, S100a9flox/floxCD11cCre, and TPflox/floxS100a9flox/floxCD11cCre mice (n = 8 mice per group). d Representative flow plots showing the percentages of DCs in the spleens of CLP-challenged CD11cCre, TPflox/floxCD11cCre, S100a9flox/floxCD11cCre, and TPflox/floxS100a9flox/floxCD11cCre mice. Quantification of DC percentages (e) and absolute numbers (f) in the spleens of CLP-challenged mice from (d) (n = 6–8 mice). g Representative flow plots showing the percentages of neutrophils in the spleens of CLP-challenged CD11cCre, TPflox/floxCD11cCre, S100a9flox/floxCD11cCre, and TPflox/floxS100a9flox/floxCD11cCre mice. Quantification of neutrophil percentages (h) and absolute numbers (i) in the spleens of CLP-challenged mice from (g) (n = 6–8 mice). j Representative immunofluorescence images showing the expression of neutrophil elastase (NE) and Ly6G in lung sections from CLP-challenged CD11cCre, TPflox/floxCD11cCre, S100a9flox/floxCD11cCre, and TPflox/floxS100a9flox/floxCD11cCre mice. Scale bars, 50 and 25 μm. k Quantification of NE-positive neutrophils in the lungs of (j) (n = 7–8 mice per group). l Western blot analysis of citH3 (NET marker) in the lungs of CD11cCre, TPflox/floxCD11cCre, S100a9flox/floxCD11cCre, and TPflox/floxS100a9flox/floxCD11cCre mice after CLP challenge. m Quantification of protein levels in (l) (n = 7 per group). n Quantitative analysis of neutrophil infiltration via measurement of MPO activity in lung tissue from CLP-challenged CD11cCre, TPflox/floxCD11cCre, S100a9flox/floxCD11cCre, and TPflox/floxS100a9flox/floxCD11cCre mice (n = 6 mice per group). o Lung vascular permeability detected by EBD leakage in CLP-challenged CD11cCre, TPflox/floxCD11cCre, S100a9flox/floxCD11cCre, and TPflox/floxS100a9flox/floxCD11cCre mice (n = 6 mice per group). Lung wet/dry weight ratio (p) (n = 8 mice per group) and BALF protein levels (q) (n = 8 mice per group) in CLP-challenged CD11cCre, TPflox/floxCD11cCre, S100a9flox/floxCD11cCre, and TPflox/floxS100a9flox/floxCD11cCre mice. r Representative histology images of lung tissue from CLP-challenged CD11cCre, TPflox/floxCD11cCre, S100a9flox/floxCD11cCre, and TPflox/floxS100a9flox/floxCD11cCre mice. Scale bars, 50 μm. s Lung histology scores from (r) (n = 7–9 mice per group). t Schematic illustration showing that ablation of S100a9 abolishes the enhanced neutrophil recruitment driven by TP-deficient DCs (Created with BioRender.com). Statistical significance was evaluated by one-way ANOVA followed by Tukey’s test for multiple comparisons (b, c, e, f, h, i, k, m–q, s) or the log-rank test (a)
To further investigate whether pharmacological blockade of S100a8/a9 could alleviate sepsis in mice with TP-deficient DCs, we administered paquinimod (10 mg/kg/day) via oral gavage for 3 days to mice subjected to CLP (Supplementary Fig. 7a). Treatment with paquinimod effectively mitigated the increased mortality rate observed in both WT and TP-deficient mice (Supplementary Fig. 7b) and reduced circulating cytokine levels (Supplementary Fig. 7c, d). Notably, no significant differences in the number of DCs were detected between the TPflox/floxCD11cCre mice and their controls (Supplementary Fig. 7e–g). However, paquinimod treatment significantly decreased neutrophil abundance in the spleen (Supplementary Fig. 7h–j) and reduced the formation of NETs (Supplementary Fig. 7k–n) in the lungs of TPflox/floxCD11cCre mice. Histological analysis of lung injury indices confirmed that paquinimod effectively alleviated lung damage in both TPflox/floxCD11cCre and control mice (Supplementary Fig. 7o–r). These results provide compelling evidence that TP signaling in DCs inhibits neutrophil recruitment by suppressing S100a8/a9 expression during sepsis (Fig. 4t).
To evaluate the potential therapeutic targeting of the S100a8/a9 pathway, we examined whether blocking its receptor could mitigate the severity of sepsis. Given that S100a8/a9 activates both the toll-like receptor 4 (TLR4) pathway and the receptor for advanced glycation end products (RAGE) pathway,23 CLP model mice were treated with the TLR4 inhibitor Resatorvid or the RAGE inhibitor FPS-ZM1 (Supplementary Fig. 8a). Interestingly, Resatorvid, but not FPS-ZM1, significantly improved survival (Supplementary Fig. 8b) and decreased neutrophil counts in both the spleen and lungs (Supplementary Fig. 8c, d), indicating the crucial role of TLR4 signaling in sepsis. To further confirm the involvement of TLR4 in mediating S100a8/a9-related signaling downstream of the TP receptor, we treated WT neutrophils with Resatorvid and performed a coculture experiment with DCs. Treatment with Resatorvid significantly attenuated neutrophil recruitment induced by both control and TP-deficient DCs (Supplementary Fig. 8e). Additionally, the administration of Resatorvid to CLP-induced TPflox/floxCD11cCre and control mice significantly reduced the mortality rate (Fig. 5a) and cytokine levels (Fig. 5b, c) without affecting the DC count (Fig. 5d–f). However, neutrophil infiltration in the spleen (Fig. 5g–i) and NET formation in the lungs (Fig. 5j–m) were markedly reduced in both groups following Resatorvid treatment, and lung injury was significantly ameliorated (Fig. 5n–q). Collectively, these findings suggest that blockade of the S100a8/a9 receptor TLR4 effectively counteracts TP-deficient DC-induced neutrophil recruitment and alleviates sepsis (Fig. 5r).
Blocking the S100a8/a9 receptor (TLR4) alleviated sepsis. a Survival analysis of CLP-challenged TPflox/flox and TPflox/floxCD11cCre mice with or without Resatorvid treatment (n = 21–26 mice per group). b, c Plasma cytokine levels of proinflammatory (IL-1β) and anti-inflammatory (IL-10) cytokines in CLP-challenged TPflox/flox and TPflox/floxCD11cCre mice with or without Resatorvid treatment (n = 8 mice per group). d Representative flow plots showing the percentages of DCs in the spleens of TPflox/flox and TPflox/floxCD11cCre mice with or without Resatorvid treatment following CLP challenge. Quantification of the percentages (e) and absolute numbers (f) of splenic DCs in (d) (n = 7–8 mice per group). g Representative flow plots showing the percentages of neutrophils in the spleens of TPflox/flox and TPflox/floxCD11cCre mice with or without Resatorvid treatment following CLP challenge. Quantification of splenic neutrophil percentages (h) and numbers (i) in (g) (n = 6–8 mice per group). j Representative immunofluorescence images showing the expression of NE and Ly6G in lung sections from CLP-challenged TPflox/flox and TPflox/floxCD11cCre mice with or without Resatorvid treatment. Scale bars, 50 and 25 μm. k Quantification of NE-positive neutrophils in the lungs of (j) (n = 6–8 mice per group). l Western blot analysis of citH3 in the lungs of TPflox/flox and TPflox/floxCD11cCre mice with or without Resatorvid treatment after CLP challenge. m Quantification of protein levels in (l) (n = 7 mice per group). n Representative histology images of lung tissues from CLP-challenged TPflox/flox and TPflox/floxCD11cCre mice with or without Resatorvid treatment. Scale bars, 50 μm. o Lung injury score quantification based on histological evaluation of the data in (n) (n = 4–6 mice per group). Lung wet/dry weight ratio (p) and BALF protein levels (q) in CLP-challenged TPflox/flox and TPflox/floxCD11cCre mice with or without Resatorvid treatment (n = 8 mice per group). r Schematic illustration showing that blockade of the S100a8/a9 receptor TLR4 negates the increased neutrophil recruitment induced by TP-deficient DCs (Created with BioRender.com). Statistical significance was evaluated via two-way ANOVA followed by Tukey’s test for multiple comparisons (b, c, e, f, h, i, k, m, o–q) or the log-rank test (a)
TP regulates Stat1-mediated S100a8/a9 expression in DCs via the PKCδ signaling pathway
To further elucidate the molecular mechanism by which TP regulates S100a8/a9 expression, we performed SCENIC analysis to identify key transcriptional regulators involved in the TP/S100a8/a9 signaling pathway in DCs. TP deficiency in DCs significantly altered the activity of 166 transcription factors. Analysis via the PROMO database predicted 73 transcription factors that could bind to the S100a8/a9 promoter region. By integrating the results from both the SCENIC analysis and the PROMO database, we identified 21 transcription factors that may regulate S100a8/a9 expression in DCs (Fig. 6a). Notably, the activity of Stat1 was markedly reduced in TP-deficient mice (Fig. 6b), whereas its abundance remained unchanged (Fig. 6c, d). Knockdown of Stat1 via siRNA (Fig. 6e) abolished the TP agonist-induced suppression of S100a8/a9 expression (Fig. 6f, g), suggesting that TP may exert its regulatory effects on S100a8/a9 expression in DCs through Stat1.
TP activation inhibits S100a8/a9 transcription via PKCδ/Stat1 signaling. a Venn diagram illustrating the transcription factors (TFs) predicted to regulate S100a8/a9 expression identified through SCENIC analysis and PROMO database screening. b Heatmap displaying the activity levels of 21 transcription factors in control and TP-deficient DCs. c Violin plots showing the mRNA expression levels of four TFs (Rela, Stat1, Stat5a, and Atf1) in control and TP-deficient DCs from scRNA-seq. d Quantification of Stat1 expression in DCs from TPflox/flox and TPflox/floxCD11cCre mice (n = 5 mice per group). e Efficiency of siRNA-mediated Stat1 knockdown in control and U-46619-treated DC2.4 cells (n = 6 wells per group). Relative mRNA levels of S100a8 (f) and S100a9 (g) in control and U-46619-treated DC2.4 cells after Stat1 knockdown (n = 6–8 wells per group). h Western blot analysis of Stat1 and phosphor-Stat1 in spleen DCs from TPflox/flox and TPflox/floxCD11cCre mice after CLP challenge. Effects of SQ29548 (i) and U-46619 (j) on S100a8 and S100a9 expression in DCs. k Effects of fludarabine on S100a8 and S100a9 expression in DCs with or without U-46619 treatment. l Western blot analysis of PKCβ, phosphor-PKCβ, PKCδ, and phosphor-PKCδ in spleen DCs from TPflox/flox and TPflox/floxCD11cCre mice after CLP challenge. m Effect of rottlerin on Stat1 phosphorylation in DCs with or without U-46619 treatment. n, o Coimmunoprecipitation analysis of the interaction of PKCδ with Stat1 in DCs. p Effects of U-73122 on PKCδ and Stat1 phosphorylation in DCs treated with or without U-46619. q Effects of YM-254890 on phosphorylated PKCδ and phosphorylated Stat1 levels in DCs treated with or without U-46619. r Schematic illustration summarizing the TP-mediated inhibition of S100a8/a9 transcription via the Gαq/PKCδ/Stat1 signaling pathway (Created with BioRender.com). Statistical significance was evaluated by the Mann–Whitney U test (d) and two-way ANOVA followed by Tukey’s test for multiple comparisons (e–g)
Stat1 typically requires phosphorylation to regulate the transcription of its target genes. Western blot analysis revealed a significant reduction in Stat1 phosphorylation in TP-deficient DCs following CLP (Fig. 6h). Treatment with a TP antagonist significantly inhibited Stat1 phosphorylation (Fig. 6i), whereas stimulation with a TP agonist increased Stat1 phosphorylation (Fig. 6j). Furthermore, fludarabine, a potent Stat1 antagonist, blocked TP agonist-induced Stat1 phosphorylation and subsequent downregulation of S100a8/a9 (Fig. 6k). Stat1 is regulated by multiple signaling pathways, including Janus kinase (JAK), mitogen-activated protein kinase, phosphoinositide-3-kinase-Akt, and PKC. Single-cell database analysis revealed that kinase genes, such as Jak1, Jak2, Akt1, PKCβ, PKCδ, Erk, and p38, were expressed in DCs (Supplementary Fig. 9a). Western blot analysis further confirmed that TP deficiency resulted in a significant decrease in PKCδ activity (Fig. 6l) but not in Jak, p38, Erk, or Akt activity (Supplementary Fig. 9b, c). Additionally, pretreatment with the PKCδ inhibitor rottlerin abolished TP agonist-induced Stat1 phosphorylation (Fig. 6m). Coimmunoprecipitation assays confirmed a direct interaction between PKCδ and Stat1 (Fig. 6n, o). Finally, we investigated the involvement of the Gαq/phospholipase C (PLC)β/diacylglycerol (DAG) signaling pathway, which is a classical upstream signaling pathway of PKCδ.24 Inhibition of PLCβ or Gαq abolished the TP agonist-induced phosphorylation of both PKCδ and Stat1 (Fig. 6p, q). Thus, TP activation suppresses S100a8/a9 transcription via the Gαq/PKCδ/Stat1 signaling pathway (Fig. 6r).
Specific activation of TP on DCs alleviates CLP-induced sepsis in mice
To explore whether the targeted activation of TP could alleviate sepsis after CLP induction, we aimed to develop a more specific approach to avoid the adverse effects commonly associated with TP agonists. For this purpose, we synthesized DCpep, a dodecapeptide (FYPSYHSTPQRP) conjugated to the NBD that specifically targets DCs.25 The self-assembly process of DCpep-U-46619, a complex containing a TP agonist (Fig. 7a), was characterized via transmission electron microscopy (TEM). In the presence of monomers, no well-defined structures were observed, indicating a random coil conformation. However, after undergoing the heating and cooling assembly process, the monomers formed needle-like nanofibers. Upon loading with a TP agonist (U-46619), morphology further transformed into a denser network of entangled, highly ordered nanofibers (Supplementary Fig. 10a). This self-assembly process was further validated via fluorescence spectroscopy (Supplementary Fig. 10b). Circular dichroism (CD) and Fourier transform infrared (FTIR) spectroscopy confirmed the formation of a parallel β-sheet secondary structure in DCpep-U-46619 (Supplementary Fig. 10c, d). Notably, zeta potential analysis revealed no significant changes in surface charge upon self-assembly (Supplementary Fig. 10e), suggesting the stability of the assembled complex. The biocompatibility of DCpep-U-46619 was assessed via a 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2H-tetrazolium bromide assay on DC2.4 cells, which revealed no significant cytotoxicity at concentrations of approximately 400 μM (Supplementary Fig. 10f). Additionally, comprehensive in vivo safety evaluations confirmed the biocompatibility of the construct without inducing systemic toxicity (Supplementary Fig. 10g–q).
Specific activation of TP in DCs attenuates CLP-induced sepsis in mice. a Schematic illustration of the synthesis of DCpep-U-46619 (Created with BioRender.com). b Schematic illustration of the experimental design of DCpep-U-46619 treatment in CLP-induced sepsis. c Representative flow plots of spleen DCs and non-DC cells from mice treated with or without DCpep-U-46619. d Quantification of the percentages of DCpep-labeled DCs and non-DC cells from mice treated with or without DCpep-U-46619 (c) (n = 3–5 mice per group). e Survival analysis of CLP-challenged wild-type (WT) mice following treatment with DCpep-U-46619 (n = 20–24 mice per group). f, g Plasma cytokine concentrations of proinflammatory (IL-1β) and anti-inflammatory (IL-10) cytokines in CLP-challenged WT mice receiving PBS, DCpep, U-46619, or DCpep-U-46619 treatment (n = 8 mice per group). h Representative flow plots showing the percentages of DCs in the spleens of CLP-challenged WT mice subjected to different treatments. Quantification of the percentages (i) and absolute numbers (j) of spleen DCs in the mice from (h) (n = 5–7 mice per group). k Representative flow plots showing the percentages of neutrophils in the spleens of CLP-challenged WT mice subjected to different treatments. Quantification of the percentages (l) and absolute numbers (m) of splenic neutrophils in the mice in (k) (n = 4–6 mice per group). n Representative immunofluorescence images showing the expression of NE and Ly6G in lung sections from CLP-challenged WT mice subjected to different treatments. Scale bars, 50 and 25 μm. o Quantification of NE-positive neutrophils in the lungs of (n) (n = 6–7 mice per group). p Western blot analysis of citH3 in the lungs of CLP-challenged WT mice subjected to different treatments. q Quantification of protein levels in (p) (n = 6–7 per group). r Quantitative analysis of neutrophil infiltration via measurement of MPO activity in lung tissue from CLP-challenged WT mice subjected to different treatments (n = 6 mice per group). s Lung vascular permeability detected by EBD leakage in CLP-challenged WT mice subjected to different treatments (n = 6 mice per group). Lung wet/dry weight ratio (t) (n = 6 mice per group) and BALF protein levels (u) (n = 6 mice per group) in CLP-challenged WT mice subjected to different treatments. v Representative histological images of lungs from CLP-challenged WT mice subjected to different treatments. Scale bars, 50 μm. w Quantification of lung injury scores on the basis of histological images from (v) (n = 4–6 mice per group). Statistical significance was evaluated via one-way ANOVA followed by Tukey’s test for multiple comparisons (f, g, i, j, l, m, o, q–u, w) or two-way ANOVA followed by Tukey’s test for multiple comparisons (d) and the log-rank test (e)
To evaluate the effect of DCpep-U-46619 on sepsis, the mice were administered DCpep-U-46619 (i.p., 5 μg/kg/day) beginning at 2 h after CLP induction, with continuous treatment for 3 days (Fig. 7b). Flow cytometry demonstrated that DCpep-U-46619 labeled with carboxytetramethylrhodamine (TAMRA) selectively targeted DCs but not non-DCs in vivo (Fig. 7c, d). Compared with no treatment, treatment with DCpep-U-46619 significantly improved mouse survival outcomes (Fig. 7e) and reduced the levels of inflammatory cytokines, including IL-1β and IL-10 (Fig. 7f, g). However, these beneficial effects were not observed in TP-deficient mice (Supplementary Fig. 11a–c). Interestingly, DCpep-U-46619 treatment did not alter the number of DCs in the spleen (Fig. 7h–j and supplementary Fig. 11d–f), indicating that the observed effects were likely due to functional modulation rather than increased recruitment of DCs. Importantly, the administration of DCpep-U-46619 significantly reduced neutrophil infiltration in the spleen (Fig. 7k–m) and NET formation in the lungs of WT mice (Fig. 7n–r), whereas no such effect was observed in TP-deficient mice (Supplementary Fig. 11g–m). Furthermore, DCpep-U-46619 treatment effectively alleviated lung injury in WT mice, as evidenced by improvements in lung permeability, wet‒dry weight ratios, BALF protein levels, and injury scores (Fig. 7s–w and supplementary Fig. 11n–q). For the post sepsis therapeutic regimen, the mice were administered DCpep-U-46619 (i.p., 5 μg/kg/day) starting 24 h after CLP induction, with continuous treatment for 3 days. Consistently, DCpep-U-46619 effectively alleviated sepsis severity in the mice (Supplementary Fig. 12a–j). Thus, the specific targeting of TP receptors on DCs via DCpep-U-46619 offers a promising therapeutic strategy to mitigate the severity of sepsis by modulating neutrophil recruitment and systemic inflammation.
Discussion
DCs are the central regulators of the immune response; however, their functional impairment in sepsis remains poorly understood. In this study, we identified a significant reduction in the DC population in patients with sepsis, which was correlated with disease severity. Notably, TP was markedly downregulated in the DCs of patients with sepsis and CLP-challenged mice. TP deficiency in DCs exacerbates sepsis by promoting S100a8/a9-mediated neutrophil recruitment and, subsequently, neutrophil extracellular trap-mediated lung injury. TP signaling limits neutrophil recruitment by suppressing S100a8/a9 expression via the PKCδ-Stat1 pathway. Blockade of S100a8/a9 or its receptor, TLR4, mitigated sepsis progression in mice lacking TP in DCs, whereas targeted TP activation in DCs via DCpep-U-46619 improved the survival of septic mice. These findings reveal a previously unrecognized protective role of TP in DCs, highlighting its potential as a therapeutic target for modulating immune dysregulation in sepsis.
One of our major findings is that dysregulated DC signaling promotes S100A8/A9-dependent neutrophil recruitment, resulting in excessive neutrophil accumulation and activation that amplifies tissue inflammation and aggravates sepsis. During sepsis, excessive neutrophil activation and NET release amplify endothelial inflammation, driving cytokine storms and severe tissue damage.26 Neutrophil mobilization from the bone marrow is largely mediated by chemokine–receptor pairs such as CXCL1–CXCR2.27 However, in lethally irradiated WT mice reconstituted with Cxcr2⁻/⁻ bone marrow, neutrophil recruitment to the lungs was reduced by only ~50% after aerosolized bacterial LPS inhalation,28 suggesting additional chemotactic pathways. Recent studies have identified S100a8/a9 as a DAMP that promotes sepsis severity by recruiting and activating neutrophils.29 However, the cellular source of S100a8/a9 remains unclear. In this study, we identify dendritic cells as a major source of S100a8/a9 during sepsis and demonstrate that DC-derived S100a8/a9 drives excessive neutrophil accumulation and activation. Despite being highly expressed in neutrophils and monocytes, DCs exhibit markedly increased baseline and inducible S100a8/a9 expression.30,31 We show that DC-derived S100a8/a9 substantially contributes to serum S100a8/a9 and that the inhibition of S100a8/a9 or its receptor TLR4 significantly reduces neutrophil infiltration and ameliorates sepsis severity. In addition to their antigen-presenting functions, DCs are potent cytokine producers that orchestrate immune activation.32 For example, DCs regulate neutrophil recruitment to inflamed skin by secreting vascular endothelial growth factor-α during cutaneous bacterial infection.32 Our findings establish the DC-derived S100a8/a9-TLR4 axis as a key pathway for neutrophil recruitment in sepsis. Given that neutrophil-derived S100a8/a9 also exacerbates tissue injury,33,34 a vicious feedforward loop of DC-derived S100a8/a9-neutrophils-S100a8/a9 may be formed during sepsis.
A particularly intriguing finding in our study was that the loss of the TXA2 receptor TP markedly increased S100a8/a9 expression in DCs, leading to severe neutrophil infiltration and NET formation during sepsis. Under steady-state conditions, DCs constitutively produce substantial amounts of TXA₂,17 enabling TXA₂ to regulate DC function in an autocrine manner. Notably, previous studies, including ours, have shown that TP signaling does not affect the antigen-presenting capacity of DCs toward T cells.17 Sepsis, however, is characterized by elevated plasma TXA₂ levels and an increased risk of thrombosis.35 TXA₂ has been shown to induce endothelial intracellular adhesion molecule-1 expression, facilitating neutrophil recruitment.36 Beyond endothelial activation, we and others have demonstrated that TXA2 directly suppresses DC-derived S100a8/a9 or counteracts the IL-10-induced upregulation of S100a8/a9 in DCs,37 thereby restraining excessive neutrophil recruitment and limiting NET-driven tissue injury. When TP expression is downregulated in DCs, this balance is lost, leading to uncontrolled neutrophil infiltration and exacerbated tissue injury. In parallel, endothelial cell death and macrophage-mediated inflammation also contribute to sepsis-related pathology.38,39,40 Elevated S100a8/a9 leads to mitochondrial complex I deficiency and mtDNA release, subsequently activating ZBP1-mediated PANoptosis in endothelial cells and aggravating vascular injury during sepsis.34 High S100a8/a9 level also drives monocytes towards M2-like macrophage differentiation,41 and an imbalance between M1-like and M2-like macrophages can precipitate the onset and exacerbate the progression of sepsis.42 Collectively, these findings suggest that TP−/− DC-derived S100a8/a9 may amplify sepsis-related tissue injury through coordinated crosstalk among DCs, neutrophils, endothelial cells, and macrophages. Consequently, targeting the TP agonist U-46619 specifically to DCs via a DC-binding peptide (DCpep-U-46619) markedly reduced neutrophil infiltration and NET formation, thereby effectively alleviating lung injury in septic mice. Recent advances have demonstrated that targeting DCs via specific surface receptors holds considerable promise in contexts such as cancer and infectious diseases.43,44 However, the translation of DC-targeted strategies into widespread clinical application remains under active development.
During DC maturation, signaling shifts from constitutive STAT6 activation to predominant STAT1 activity,45 which suppresses the expression of S100a8/a9.46 Although JAK1 is the primary upstream regulator of STAT1 in DCs,47 we found that JAK1 activity was nearly undetectable in DCs under septic conditions, likely reflecting the markedly reduced plasma interferon (IFN)-γ levels observed in patients with sepsis and mice challenged with CLP.48,49 In this context, alternative kinases such as PKCδ may compensate for the ability of JAK1 to phosphorylate and activate Stat1.50 Given that TP primarily couples with Gq proteins, which activate PLC to hydrolyze phosphatidylinositol 4,5-bisphosphate into DAG, leading to PKCδ activation,24 TP signaling can suppress S100a8/a9 expression via PKCδ–Stat1. Accordingly, the loss of TP signaling may relieve this suppression, promoting S100a8/a9 expression. In contrast, the PGE2‒EP2‒EP4 axis mediates immunosuppression by inducing the expression of chemokines such as Ccl22 and Ccl17 in DCs.51 Together, these findings highlight that distinct PG pathways exert divergent and nonredundant effects on host immune responses during infection. This finely tuned PG signaling network underscores the importance of selectively targeting specific prostanoid receptors, rather than broadly suppressing inflammatory pathways, as a strategy for precision immunomodulation in sepsis.
In summary, we identified a previously unrecognized role of TP in regulating DC-mediated neutrophil recruitment and tissue damage during sepsis. TP signaling constrained DC-driven neutrophil infiltration through the PKCδ/Stat1/S100a8/a9 axis, thereby conferring protection against sepsis (Supplementary Fig. 13). These findings suggest that the targeted activation of TP in DCs may serve as a potential therapeutic strategy for sepsis intervention.
Methods
Human participants
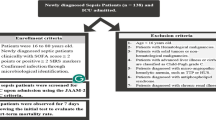
Sepsis patients and controls were recruited at Tianjin Medical University General Hospital. Sepsis was diagnosed according to the 2016 Sepsis-3 criteria, defined as suspected or confirmed infection (evidenced by clinical signs of infection or receipt of antibiotics for > 48 h) with a sequential organ failure assessment (SOFA) score ≥ 2. All participants were recruited from the intensive care unit (ICU). The inclusion criteria for sepsis patients were (1) confirmed diagnosis of sepsis, (2) first ICU admission, and (3) aged > 18 years. The exclusion criteria were as follows: (1) received immunosuppressive therapies; (2) were pregnant or lactating; (3) had a history of malignant tumors or hematological disorders; (4) had psychiatric conditions; and (5) had incomplete clinical documentation. The inclusion criteria for the control group were as follows: (1) normal serum levels of C-reactive protein (CRP) and procalcitonin (PCT) with no fever within the previous week, and (2) aged ≥ 18 years. The exclusion criteria for controls were as follows: (1) immunosuppressive therapy; (2) body temperature > 37.3 °C at recruitment; (3) positive blood culture; and (4) abdominal surgery within the past 48 hours. Overall, 24 patients with sepsis and 13 healthy controls were enrolled in this study (Supplementary Table 1 and Table 2). This study was approved by the Institutional Ethics Committee of Tianjin Medical University General Hospital (IRB2021-YX-258-01), and written informed consent was obtained from all participants.
Cells
DC2.4 cells (Cellverse Co., Cat# iCell-m016) were maintained in our laboratory. DC2.4 cells were cultured in RPMI-1640 medium (Gibco, Cat# 11875093) supplemented with 10% fetal bovine serum (FBS) (Gibco, Cat# 10099141 C), 10 ng/mL murine granulocyte macrophage-colony stimulating factor (GM-CSF) (PeproTech, Cat# 315-03-20UG), and 100 U/mL penicillin‒streptomycin (Gibco, Cat #15140122). All the cells were incubated at 37 °C in 5% CO2.
Mice
C57BL/6 J mice (8–10 weeks) were purchased from Beijing Vital River Laboratory Animal Technology Co., Ltd. (Beijing, China). The TP−/−, TPflox/flox, and CD11cCre mice were bred in the laboratory. S100a9flox/flox mice were obtained from Cyagen Bioscience, Inc. (Jiangsu, China). CD11cCre mice were crossed with TPflox/flox or S100a9flox/flox mice to generate TPflox/floxCD11ccre or S100a9flox/floxCD11cCre mice. TPflox/floxS100a9flox/floxCD11cCre mice were generated by crossing TPflox/floxCD11cCre and S100a9flox/flox mice. The genetic background of all the mice was C57BL/6J. Male mice were used to avoid hormonal effects.52 The mice were maintained in an environment with a temperature of 22 °C ± 1 °C and a humidity of 50 ± 5% on a 12:12 h light/dark cycle, with free access to sterile food and water. All animal experiments were performed with the approval of the Laboratory Animal Management and Use Committee of Tianjin Medical University.
Induction of murine sepsis
The mice were assigned a numerical code and randomly assigned to either the experimental treatment group or the control group via a random table. Sepsis was induced in mice via the CLP model, which was performed as previously described.53 Briefly, the mice were anesthetized with isoflurane, the abdomen was disinfected, and a midline incision was made to expose the cecum. The cecum was ligated below the ileocecal valve and punctured with a 16-gauge needle, and a small amount of fecal content was extruded by gentle compression. The incision was closed, and the animals received 1 mL of 0.9% saline subcutaneously for fluid resuscitation. Postoperative analgesia was provided by buprenorphine (0.05 mg/kg, s.c.). Sham-operated mice underwent cecal exposure without ligation or puncture. For the alternative endotoxemia model, the mice received an intravenous injection of LPS (10 mg/kg, Sigma‒Aldrich, Cat# L2880). The animals were euthanized with CO₂ at the indicated time points for subsequent assays. To evaluate the therapeutic effect of S100a8/a9 inhibition, the mice were orally administered paquinimod (10 mg/kg/day, MCE, Cat# HY-100442) for 3 consecutive days following CLP. To assess the impact of S100a8/a9 receptor inhibition, the mice were intraperitoneally injected with either Resatorvid (3 mg/kg/day, MCE, Cat# HY-11109) or FPS-ZM1 (10 mg/kg/day, MCE, Cat# HY-19370) for 3 days post-CLP. Additionally, to investigate the therapeutic potential of TP receptor activation, the mice were intraperitoneally injected with DCpep-U-46619 (5 μg/kg/day) beginning at 2 h or 24 h after CLP induction, with continuous treatment for 3 days.
Cell isolation, culture, and transfection
PBMCs were isolated from human venous blood samples via Ficoll‒Hypaque density gradient centrifugation (Cytiva, Cat# 17144002). A single-cell suspension from a mouse spleen was obtained via mechanical dissociation through a 100 μm sieve. The DCs were purified via CD11c UltraPure microbeads (Miltenyi, Cat# 130-125-835), achieving a purity of > 85%, as determined via flow cytometry. The DCs were cultured in RPMI-1640 medium supplemented with 1% penicillin‒streptomycin and 10% FBS. Bone marrow single-cell suspensions were prepared by flushing the bone marrow through a 100 μm sieve. Bone marrow-derived neutrophils were isolated via Percoll gradient centrifugation (Solarbio, Cat# P8370) following a previously described protocol.54
DC2.4 cells were transfected with siRNA via TransIntroTM EL transfection reagent (TransGen Biotech, Cat# FT201-01). Briefly, approximately 2 × 104 DCs were seeded into 24-well plates. In a separate microfuge tube, 50 µL of Opti-MEM was mixed with 2 µL of EL transfection reagent and 50 pmol of siRNA, followed by incubation at room temperature for 20 min. The Opti-MEM-EL-siRNA mixture was then added to the cells, and the 24-well plate was gently shaken. Twenty-four hours post-transfection, the cells were harvested for mRNA expression analysis.
Adoptive transfer of DCs
To evaluate the role of TP in DCs during sepsis, WT/TP−/− DCs were isolated from mouse spleens via CD11c UltraPure microbeads. Approximately 1 × 106 DCs were injected intravenously via the caudal vein into WT recipient mice 1 h before CLP challenge.
Flow cytometry
PBMCs and single-cell suspensions from the spleen or lung were incubated at 4 °C for 30–40 min in phosphate-buffered saline (PBS) containing 1% FBS and fluorochrome-conjugated antibodies. Cell identification by flow cytometry: Human DCs: CD3⁻CD11c⁺HLA-DR⁺. Human neutrophils: CD3⁻CD11b⁺CD15⁺. Mouse DCs: CD3⁻CD11c⁺HLA-DR⁺. Mouse neutrophils: CD45⁺CD3⁻CD11b⁺Ly6G⁺. Mouse B cells: CD45⁺CD3⁻B220⁺. Mouse CD8+ T cells: CD45⁺CD3⁺CD8a⁺. Mouse T helper subsets: TH1: CD45⁺CD3⁺CD4⁺IFN-γ⁺; TH2: CD45⁺CD3⁺CD4⁺IL-4⁺; TH17: CD45⁺CD3⁺CD4⁺IL-17A⁺. Mouse regulatory T cells (Tregs): CD45⁺CD3⁺CD4⁺Foxp3⁺. Flow cytometry was performed using a BD LSRFortessa Fortessa Cell Analyzer (BD Biosciences). Data acquisition and analysis were conducted via FlowJo 8.3.3 software (TreeStar). The cells were sorted via a BD FACSAria Fusion flow cytometer (BD Biosciences).
Enzyme-linked immunosorbent assay (ELISA)
To assess cytokine levels in septic mice and cell culture supernatants, an ELISA was performed according to the manufacturer’s instructions. The cytokine kits and their detection range were as follows: IL-1β (31.3–2000 pg/mL) (DAKEWE, Cat# 1210122), IL-10 (62.5–4000 pg/mL) (DAKEWE, Cat# 1211002), S100a8 (2.7–2000 ng/mL) (CUSABIO, Cat# CSB-EL020641MO), and S100a9 (0.45–30 ng/mL) (CUSABIO, Cat# CSB-EL020642MO). The cytokine concentration in the serum was detected without dilution. Cytokine levels in the serum were expressed as absolute concentrations (pg/mL or ng/mL).
Histological analysis
The mice were euthanized, and the lungs, hearts, spleens, livers, and kidneys were harvested and fixed in 4% paraformaldehyde for 48 h. The tissues were then paraffin-embedded, sectioned at a thickness of 8 μm, and stained with hematoxylin and eosin for histological examination. Lung injury severity was quantitatively assessed via a lung injury scoring system as previously described.55
Lung vascular permeability measurements
Pulmonary vascular permeability was assessed via Evans blue–albumin (EBA) extravasation as previously described.39,56 Briefly, anesthetized mice were injected intravenously with 150 μL of EBA solution (1% Evans blue dye and 4% albumin in PBS) and allowed to circulate for 30 min. Mice were then euthanized, and the lungs were perfused with 20 mL of PBS. Lung tissues were excised, weighed, homogenized in 0.5 mL of PBS, and incubated in 1 mL of formamide at 60 °C for 24 h. The Evans blue concentration in the lung homogenate supernatants was quantified spectrophotometrically at 620 nm. The extravasated dye content was expressed as μg of Evans blue dye per g of fresh lung tissue, which was calculated against a standard curve.
Myeloperoxidase (MPO) assay
MPO activity was measured via a colorimetric assay kit (Solarbio, #BC5715) according to the manufacturer’s instructions. Briefly, lung tissues were harvested, homogenized in cold PBS, and centrifuged at 12,000 × g for 10 min at 4 °C to remove insoluble material. The supernatants were collected and assayed directly. MPO activity was quantified spectrophotometrically at 460 nm and expressed as units per gram of tissue (U/g).
Chemotaxis assay
Neutrophils were labeled with carboxyfluorescein succinimidyl ester (CFSE) (BioLegend, Cat# 423801) and resuspended at a concentration of 1 × 105 cells/100 μL in RPMI-1640 medium containing 1% FBS. CFSE-labeled neutrophils were added to the upper chamber of a 24-well transwell insert (Corning, Cat# 3421). The DCs (5 × 103 cells in 200 μL) were placed in the lower chamber. The Transwell plate was incubated at 37 °C with 5% CO2 for 2 h. After incubation, neutrophil migration was assessed by imaging via a Zeiss LSM900 confocal microscope.
Immunofluorescence
After CLP induction, the lungs were excised and embedded in optimal cutting temperature (OCT) compound (SAKURA, Cat# 4583) before being quickly frozen in liquid nitrogen. The OCT-grade tissue was sectioned at a thickness of 8 μm. The sections were fixed in cold acetone and washed with PBS. After incubation with 5% bovine saline albumin in PBS for 30 min, the samples were incubated with primary antibodies against Ly6G/Ly6C (Invitrogen, Cat# 14-5931-82) and neutrophil elastase (ABclonal, Cat# A13015) for 12 h at 4 °C, followed by washing with PBS and incubation with Alexa Fluor 594- (Invitrogen, Cat# A-21207) or Alexa Fluor 488- (Invitrogen, Cat# A11008) for 2 h in the dark at room temperature. The nuclei were labeled with 4′,6-diamidino-2-phenylindole (DAPI; Solarbio, Cat# C0060) and examined via a Zeiss LSM900 confocal microscope (Carl Zeiss).
RNA extraction and real-time polymerase chain reaction (PCR)
RNA was isolated from cells or tissues via TRIzol reagent (Invitrogen, Cat# 15596018CN) according to the manufacturer’s instructions. Purity and concentration were measured via a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific). The RNA was reverse transcribed to cDNA via Hifair III 1st Strand cDNA Synthesis SuperMix for qPCR (Yeason, Cat# 11141ES10). The cDNA was amplified for 40 cycles by using Hieff UNICON® Universal Blue qPCR SYBR Green Master Mix (Yeason, Cat# 11184ES03) on a Roche LightCycler 480 Instrument II (Roche). The primers used are listed in Supplementary Table 3.
Western blotting
Cellular proteins were extracted via lysis buffer containing protease inhibitors. The protein concentration was determined via a BCA protein assay kit (Thermo Fisher Scientific, Cat# 23225). An equal amount of total protein (5 μg) was loaded into each lane to ensure consistency. Lysates containing equal amounts of protein were separated via 15% sodium dodecyl sulfate‒polyacrylamide gel electrophoresis and transferred onto polyvinylidene difluoride membranes (Millipore, Cat# IPVH00010). The membrane was incubated with 5% skim milk (Solarbio, Cat# D8340) for 2 h and then incubated with primary antibodies at 4 °C overnight. The following primary antibodies were used: rabbit polyclonal anti-S100a8 (Proteintech, Cat# 15792-1-AP), rabbit polyclonal anti-S100a9 (Proteintech, Cat# 26992-1-AP), rabbit monoclonal anti-Stat1 (Cell Signaling Technology, Cat# 9172), rabbit polyclonal anti-pStat1 (Cell Signaling Technology, Cat# 9167), rabbit polyclonal anti-PKCβ (ABclonal, Cat# A21241), rabbit polyclonal anti-pPKCβ (ABclonal, Cat# AP1375), rabbit polyclonal anti-PKCδ (ABclonal, Cat# A7778), rabbit polyclonal anti-pPKCδ (ABclonal, Cat# AP0776), rabbit polyclonal anti-P38 (Cell Signaling Technology, Cat# 9212S), rabbit polyclonal anti-pP38 (Cell Signaling Technology, Cat# 9216S), rabbit polyclonal anti-erk (Cell Signaling Technology, Cat# 9102S), rabbit polyclonal anti-perk (Cell Signaling Technology, Cat# 9101S), rabbit polyclonal anti-Jak1 (Cell Signaling Technology, Cat# 3332S), rabbit polyclonal anti-pJak1 (Cell Signaling Technology, Cat# 3331S), rabbit polyclonal anti-Jak2 (Cell Signaling Technology, Cat# 3230S), rabbit polyclonal anti-pJak2 (Cell Signaling Technology, Cat# 3776S), rabbit polyclonal anti-Akt (Cell Signaling Technology, Cat# 4691S), rabbit polyclonal anti-pAkt (Cell Signaling Technology, Cat# 4060S), rabbit monoclonal anti-Citrullinated Histone H3 (Cell Signaling Technology, Cat# 97272), and rabbit polyclonal anti-β-actin (Abclonal, Cat# AC026). The membranes were then incubated with a horseradish peroxidase-labeled secondary antibody (Cell Signaling Technology) in blocking buffer at room temperature for 2 h. The blots were developed via an enhanced chemiluminescence reagent (Thermo Fisher Scientific). All commercial antibodies have been validated by manufacturers or in published studies. β-actin was used as the loading control in all the experiments. The blots were scanned via a Tanon imaging system (Tanon-5200Multi). Band intensities were quantified with ImageJ software. The grayscale intensity of each target protein was normalized to that of β-actin, and normalized values for the treatment groups were expressed as fold changes relative to the mean of the control group (set to 1.0).
Preparation of the DCpep peptide derivative
The DCpep peptide derivative was synthesized via Fmoc-based solid-phase peptide synthesis with 2-chlorotriphenylmethyl chloride resin. The synthesis process involved the sequential addition of N-terminal Fmoc-protected amino acids corresponding to the target sequence. Benzotriazole-N,N,N′,N′-tetramethylurea hexafluorophosphate was used as a coupling agent to facilitate peptide bond formation with the next free amino group. After the synthesis was completed, the peptide was cleaved from the resin and deprotected with trifluoroacetic acid. To enable fluorescence tracking of the nanoagent, 4-chloro-7-nitro-2,1,3-benzoxadiazole was conjugated at the N-terminus of the peptide. The crude product obtained from the cleavage reaction was further purified via high-performance liquid chromatography to obtain the final pure DCpep peptide derivative. To assess the biological safety of DCpep-U-46619, mice received a single dose (5 μg/kg/day) and were examined on days 1 and 14 postadministration. Blood samples and major organs were collected for analysis. Hematological and serum biochemical tests were conducted to evaluate safety, and tissue sections were fixed, paraffin embedded, and subjected to H&E staining for histopathological assessment.
Transmission electron microscopy (TEM)
For TEM imaging, 10 µL of a solution containing DCpep assemblies or coassemblies of DCpep with the TP agonist U-46619 were carefully deposited onto a carbon-coated copper mesh grid. Excess liquid was removed via filter paper, and the samples were subjected to negative staining with uranyl acetate to enhance contrast and improve the visualization of the nanostructures. The prepared grids were subsequently imaged at 200 kV via a Tecnai G2 F20 transmission electron microscope, which allowed high-resolution observation of the self-assembled nanostructures.
Fluorescence spectrum analysis
To assess the fluorescence properties of DCpep-U-46619, 100 µL of DCpep-U-46619 solution was added to a 96-well plate both before and after the heating/cooling self-assembly process. The fluorescence spectra were recorded using a BioTek SynergyTM 4 Hybrid Microplate Reader at an excitation wavelength of 470 nm. This analysis was performed to monitor the structural and conformational changes in the DCpep-U-46619 assemblies.
Circular dichroism spectrum
To analyze the secondary structural composition of DCpep and its coassembled complex, DCpep-U-46619, three sample solutions were prepared: a stable DCpep solution, a self-assembled DCpep solution after the heating‒cooling process, and a coassembled DCpep-U-46619 solution. For the measurements, a special light-transmitting stone clip was used for circular dichroism chromatography. A 200 µL aliquot of the prepared solution was carefully placed onto the clip, ensuring that no bubbles or impurities were introduced. A second clip was gently attached to the edge to secure the sample. CD spectra were recorded via a MOS-450 (Biologic) spectrometer at 180–250 nm. The raw data were smoothed and processed via the DICHROWEB online analysis tool (http://dichroweb.cryst.bbk.ac.uk/html/sendform.shtml) to obtain the detailed secondary structure composition. The structural elements—including α-helices, β-sheets, β-turns, and random coils—were quantitatively determined via the standard CONTIN algorithm.
Single-cell RNA sequencing and analysis
Publicly available human scRNA-seq datasets were downloaded from the single-cell portal (SCP548). Low-quality cells were removed on the basis of the following criteria: cells expressing between 200 and 2500 genes were retained, whereas cells with > 10% mitochondrial gene expression were excluded. The data were normalized via the LogNormalize function in the Seurat (v4.4.0) package. Principal component analysis (PCA) was conducted via the RunPCA function, and batch effects were corrected via the Harmony (v1.2.0) package. The cells were clustered on the basis of the top 30 principal components (PCs) and visualized via t-SNE via the RunTSNE, FindNeighbors, and FindClusters functions in Seurat, with a resolution parameter of 0.8 applied to optimize cluster identification. Clusters were annotated by matching them with known cell-type marker genes and classifying them into five major immune cell populations.
Sorted splenic DCs from two mice per group (TPflox/flox and TPflox/floxCD11cCre) were subjected to droplet-based single-cell RNA-seq via the 10X Genomics platform. Single-cell RNA libraries were generated via a single-cell 3 v2 kit (10X Genomics) following the manufacturer’s protocol. Libraries were sequenced on an Illumina NovaSeq 6000 instrument. The raw sequencing data were processed via the Cell Ranger pipeline (v7.2.0, 10X Genomics) for alignment, barcode assignment, and UMI quantification. Low-quality cells were removed on the basis of the following criteria: retained cells expressing between 200 and 7,500 genes; cells with > 20% mitochondrial gene expression; and doublets identified and filtered via the DoubletFinder package. The data were normalized via the LogNormalize function in the Seurat (v4.4.0) package. PCA was conducted via the RunPCA function, and batch effects were corrected via the Harmony (v1.2.0) package. The cells were clustered on the basis of the top 20 PCs and visualized via t-SNE via the RunTSNE, FindNeighbors, and FindClusters functions in Seurat. A resolution parameter of 0.3 was applied to optimize distinct cluster identification. Clusters were annotated by matching them with known cell-type marker genes and classifying them into ten immune cell populations.
Differentially expressed genes (DEGs) were identified via FindMarker in Seurat software. The selection criteria for significant DEGs were as follows: adjusted p value < 0.05 and absolute log2-fold change (|log2FC|) ≥ 0.5. For functional enrichment analysis, gene ontology analysis was conducted via the enrichGO function from the clusterProfiler package (v4.6.2), with a focus on biological processes enriched in significantly upregulated genes.
RNA velocity analysis
Spliced and unspliced transcript counts for each gene in every cell were quantified via Velocyto v0.17.17 in Python. RNA velocity analysis was then performed with scVelo v0.3.2. Genes expressed in at least 20 cells were retained, and the top 2000 highly variable genes were selected via scv.pp.filter_and_normalize. Duplicate cells were removed with scv.pp.remove_duplicate_cells. The neighborhood graph was computed on the basis of 30 principal components and 30 nearest neighbors via scv.pp.moments. Cellular transcriptional dynamics were inferred with the dynamical model, and RNA velocities were estimated and projected onto the tSNE embedding. The resulting velocity vectors were used to construct the velocity graph with scv.tl.velocity_graph and to visualize transcriptional trajectories as continuous streamlines via scv.pl.velocity_embedding_stream.
Single-cell regulatory network inference and clustering (SCENIC) analysis
To predict single-cell gene regulatory networks, we performed SCENIC analysis via the SCENIC (v1.3.1) package. The raw count expression matrix was processed via GENIE3 (v1.20.0) to infer the initial coexpression networks between transcription factors (TFs) and their potential target genes. Transcription factor motifs were identified via RcisTarget (v1.18.2), which incorporates regulatory motif databases for mice (mm9-tss-centered-10 kb and mm9-500 bp-upstream) to refine the TF-target interactions. The final regulon activity scores were computed for each cell via AUCell (v1.20.2), allowing the identification of key transcription factors that drive cellular states in sepsis.
Statistical analysis
Sample size determination and power analysis were performed via G*Power software (version 3.1.9.6). All the data are expressed as the means ± standard errors of the means. The data were analyzed via Prism 8 (GraphPad Prism Software). The Mann–Whitney U test was used to compare two independent groups. Two-way analysis of variance followed by Tukey’s and Kruskal–Wallis tests and Dunn’s post hoc test were used for multiple comparisons. Survival curves were used for survival analysis. Statistical significance was set at P < 0.05. Data collection and analysis were performed blindly, without disclosing the group name to the investigator.
Data availability
No code was generated for this study. All data necessary to understand and evaluate the conclusions of this study are provided in the article and Supplementary Information. Sequencing data are deposited under the NCBI GEO accession number, GSE315040.
References
Singer, M. et al. The third international consensus definitions for sepsis and septic shock (sepsis-3). JAMA 315, 801–810 (2016).
Rudd, K. E. et al. Global, regional, and national sepsis incidence and mortality, 1990-2017: analysis for the Global Burden of Disease Study. Lancet 395, 200–211 (2020).
Gotts, J. E. et al. Sepsis: pathophysiology and clinical management. BMJ 353, i1585 (2016).
Giamarellos-Bourboulis, E. J. et al. Precision immunotherapy to improve sepsis outcomes: the immunoSep randomized clinicaltrial. JAMA 8, e2524175 (2025).
Fisher, C. J. Jr et al. Treatment of septic shock with the tumor necrosis factor receptor: Fc fusion protein. The Soluble TNF Receptor Sepsis Study Group. N. Engl. J. Med 334, 1697–1702 (1996).
Patel, G. P. et al. Systemic steroids in severe sepsis and septic shock. Am. J. Respir. Crit. Care Med. 185, 133–139 (2012).
Kumar, V. Dendritic cells in sepsis: potential immunoregulatory cells with therapeutic potential. Mol. Immunol. 101, 615–626 (2018).
Poehlmann, H. et al. Phenotype changes and impaired function of dendritic cell subsets in patients with sepsis: a prospective observational analysis. Crit. Care 13, R119 (2009).
Roquilly, A. et al. Local modulation of antigen-presenting cell development after resolution of pneumonia induces long-term susceptibility to secondary infections. Immunity 47, 135–147 (2017).
Roquilly, A. et al. The role of dendritic cell alterations in susceptibility to hospital-acquired infections during critical-illness related immunosuppression. Mol. Immunol. 68, 120–123 (2015).
Nakahata, N. Thromboxane A2: physiology/pathophysiology, cellular signal transduction and pharmacology. Pharm. Ther. 118, 18–35 (2008).
Harizi, H. et al. Arachidonic-acid-derived eicosanoids: roles in biology and immunopathology. Trends Mol. Med. 14, 461–469 (2008).
Otto, G. P. et al. Effects of low-dose acetylsalicylic acid and atherosclerotic vascular diseases on the outcome in patients with severe sepsis or septic shock. Platelets 24, 480–485 (2013).
Al Harbi, S. A. et al. Association between aspirin therapy and the outcome in critically ill patients: a nested cohort study. Bmc Pharm. Toxico 17, 5 (2016).
Valerio-Rojas, J. C. et al. Outcomes of severe sepsis and septic shock patients on chronic antiplatelet treatment: a historical cohort study. Crit. Care Res. Pract. 2013, 782573 (2013).
Wiewel, M. A. et al. Chronic antiplatelet therapy is not associated with alterations in the presentation, outcome, or host response biomarkers during sepsis: a propensity-matched analysis. Intens. Care Med. 42, 352–360 (2016).
Kabashima, K. et al. Thromboxane A2 modulates interaction of dendritic cells and T cells and regulates acquired immunity. Nat. Immunol. 4, 694–701 (2003).
Moalli, F. et al. Thromboxane A2 acts as tonic immunoregulator by preferential disruption of low-avidity CD4+ T-cell-dendritic cell interactions. J. Exp. Med. 211, 2507–2517 (2014).
Yang, J. et al. Aspirin prevents metastasis by limiting platelet TXA2 suppression of T-cell immunity. Nature 640, 1052–1061 (2025).
Reyes, M. et al. An immune-cell signature of bacterial sepsis. Nat. Med. 26, 333–340 (2020).
Yao, R. Q. et al. Single-cell transcriptome profiling of sepsis identifies HLA-DRlowS100Ahigh monocytes with immunosuppressive function. Mil. Med. Res. 10, 27 (2023).
Xiao, K. et al. A pan-immune panorama of bacterial pneumonia revealed by a large-scale single-cell transcriptome atlas. Signal. Transduct. Target Ther. 10, 5 (2025).
Guo, Q. et al. Induction of alarmin S100A8/A9 mediates activation of aberrant neutrophils in the pathogenesis of COVID-19. Cell Host Microbe 29, 222–235 (2021).
Black, J. D. et al. PKCα and PKCδ: friends and rivals. J. Biol. Chem. 298, 102194 (2022).
Xiao, Y. et al. Low cost delivery of proteins bioencapsulated in plant cells to human non-immune or immune modulatory cells. Biomaterials 80, 68–79 (2016).
Zhang, H. et al. Neutrophil, neutrophil extracellular traps and endothelial cell dysfunction in sepsis. Clin. Transl. Med. 13, e1170 (2023).
Paudel, S. et al. CXCL1 regulates neutrophil homeostasis in pneumonia-derived sepsis caused by Streptococcus pneumoniae serotype 3. Blood 133, 1335–1345 (2019).
Reutershan, J. et al. Critical role of endothelial CXCR2 in LPS-induced neutrophil migration into the lung. J. Clin. Investig. 116, 695–702 (2006).
Wang, Q. et al. S100A8/A9: an emerging player in sepsis and sepsis-induced organ injury. Biomed. Pharmacother. 168, 115674 (2023).
Averill, M. M. et al. S100A9 Differentially modifies phenotypic states of neutrophils, macrophages, and dendritic cells implications for atherosclerosis and adipose tissue inflammation. Circulation 123, 1216–1226 (2011).
Austyn, J. M. Dendritic cells in the immune system-history, lineages, tissues, tolerance, and immunity. Microbiol Spectr. 4, 6 (2016).
Janela, B. et al. A subset of type I conventional Dendritic cells controls cutaneous cacterial infections through VEGFα-mediated recruitment of neutrophils. Immunity 50, 1069–1083 (2019).
Li, X. X. et al. Neutrophil-derived exosomal S100A8 aggravates lung injury in sepsis by inducing pyroptosis. Mol. Immunol. 181, 29–39 (2025).
Wang, Y. H. Z. et al. S100A8/A9hi neutrophils induce mitochondrial dysfunction and PANoptosis in endothelial cells via mitochondrial complex I deficiency during sepsis. Cell Death Dis. 15, 462 (2024).
Carmona, R. H. et al. The role of prostacyclin and thromboxane in sepsis and septic shock. Arch. Surg. 119, 189–192 (1984).
Rossaint, J. et al. Directed transport of neutrophil-derived extracellular vesicles enables platelet-mediated innate immune response. Nat. Commun. 7, 13464 (2016).
Kumar, A. et al. Interleukin-10 influences the expression of MRPS and MRP14 on human dendritic cells. Int. Arch. Allergy Imm 132, 40–47 (2003).
Cheng, K. T. et al. Caspase-11-mediated endothelial pyroptosis underlies endotoxemia-induced lung injury. J. Clin. Investig. 127, 4124–4135 (2017).
Xiong, S. Q. et al. IL-1β suppression of transcription underlies sepsis-induced inflammatory lung injury. J. Clin. Investig. 133, e169500 (2023).
Di, A. K. et al. The TWIK2 potassium efflux channel in macrophages mediates NLRP3 inflammasome-induced inflammation. Immunity 49, 56–65 (2018).
Van Kooten, N. J. T. et al. S100A8/A9 drives monocytes toward M2-like macrophage differentiation and associates with M2-like macrophages in osteoarthritic synovium. Rheumatology 64, 332–343 (2024).
Wang, Z. Y. et al. The role of macrophages polarization in sepsis-induced acute lung injury. Front. Immunol. 14, 1209438 (2023).
Mamuti, M. et al. A polyvalent peptide CD40 nanoagonist for targeted modulation of Dendritic cells and amplified cancer immunotherapy. Adv. Mater. 34, 2109432 (2022).
Yin, D. et al. Dendritic-cell-targeting virus-like particles as potent mRNA vaccine carriers. Nat. Biomed. Eng. 9, 185–200 (2025).
Jackson, S. H. et al. Dendritic cell maturation requires STAT1 and is under feedback regulation by suppressors of cytokine signaling. J. Immunol. 172, 2307–2315 (2004).
Capitini, C. M. et al. Absence of STAT1 in donor-derived plasmacytoid dendritic cells results in increased STAT3 and attenuates murine GVHD. Blood 124, 1976–1986 (2014).
Vogel, A. et al. JAK1 signaling in dendritic cells promotes peripheral tolerance in autoimmunity through PD-L1-mediated regulatory T-cell induction. Cell Rep. 38, 110420 (2022).
Gatica-Andrades, M. et al. WNT ligands contribute to the immune response during septic shock and amplify endotoxemia-driven inflammation in mice. Blood Adv. 1, 1274–1286 (2017).
Romero, C. R. et al. The role of interferon-γ in the pathogenesis of acute intra-abdominal sepsis. J. Leukoc. Biol. 88, 725–735 (2010).
DeVries, T. A. et al. Protein kinase Cδ regulates apoptosis via activation of STAT1. J. Biol. Chem. 279, 45603–45612 (2004).
Thumkeo, D. et al. PGE2-EP2/EP4 signaling elicits immunosuppression by driving the mregDC-Treg axis in inflammatory tumor microenvironment. Cell Rep. 39, 110914 (2022).
Lakbar, I. et al. Interactions between gender and sepsis-implications for the future. Microorganisms 11, 746 (2023).
Rittirsch, D. et al. Immunodesign of experimental sepsis by cecal ligation and puncture. Nat. Protoc. 4, 31–36 (2009).
Oh, H. et al. Neutrophil isolation protocol. J. Vis. Exp. 23, 745 (2008).
Matute-Bello, G. et al. An official American Thoracic Society workshop report: features and measurements of experimental acute lung injury in animals. Am. J. Respir. Cell Mol. Biol. 44, 725–738 (2011).
Radu, M. et al. An in vivo assay to test blood vessel permeability. J. Vis. Exp. 16, e50062 (2013).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2025YFE0126700), the National Natural Science Foundation of China (82321001, 82530018, 82225002, and 82430003), and the Natural Science Foundation of Tianjin, China (24ZXZSSS00430). Y. Yu is a fellow of the Jiangsu Collaborative Innovation Center for Cardiovascular Disease Translational Medicine, China.
Author information
Authors and Affiliations
Contributions
Y. Yu and Y. Shen designed the study; R. Du, Q. Liu, X. Tao, S. Guo and D. Tian performed the experiments and provided experimental assistance; R. Du, Y. Fan and K. Xie collected clinical patient samples; R. Du and T. Pan assisted in the bioinformatics analysis; R. Du and Y. Wang performed the material experiments; R. Du and T. Pan analyzed the data; R. Du wrote the paper. Y. Yu, Y. Shen, J. Zhou, and Roger S.-Y. Foo critically revised the manuscript for key intellectual content. All the authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Du, R., Pan, T., Wang, Y. et al. Thromboxane receptor activation in dendritic cells mitigates sepsis by suppressing S100a8/a9-mediated neutrophil recruitment. Sig Transduct Target Ther 11, 75 (2026). https://doi.org/10.1038/s41392-026-02592-w
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41392-026-02592-w