Abstract
Tumor-associated macrophages (TAMs) are pivotal in the immunosuppressive tumor microenvironment of pancreatic ductal adenocarcinoma (PDAC). The efficacy of targeting the CSF-1/CSF-1R axis in PDAC remains uncertain. Using single-cell RNA sequencing on specimens from patients treated with Surufatinib plus chemotherapy, we identified a distinct subset of damage-associated macrophages (DAMs) characterized by high GPR34 expression. In Gpr34ΔLyz2 mouse models and in vitro co-cultures, GPR34+ macrophages responded to tissue damage by releasing lysophosphatidylserine (LysoPS), which enhanced CXCL16 secretion and efferocytosis. This efferocytosis promoted MHC-I degradation via the macrophage lysosomal pathway, leading to CD8+ T cell exhaustion. Combining a GPR34 antagonist with chemotherapy and surufatinib significantly enhanced anti-tumor responses in preclinical models. These findings identify GPR34 as a promising immune therapeutic target.
Similar content being viewed by others
Introduction
Pancreatic ductal adenocarcinoma (PDAC) stands as one of the most aggressive and lethal malignancies worldwide, presenting a formidable challenge in oncology due to its persistently poor prognosis.1 This dismal outcome is largely attributable to the complex and profoundly immunosuppressive tumor microenvironment (TME) that characterizes PDAC, a milieu that not only supports the initiation and progression of pancreatic cancer but also actively undermines therapeutic efforts.2,3 The immune landscape of PDAC is notoriously dense and inhibitory, populated by a diverse array of cellular components that collaborate to foster immune escape and confer resistance to treatment. This intricate ecosystem is filled with various inhibitory immune cells and components such as macrophages, T cell subsets, and cancer-associated fibroblasts (CAFs), all of which contribute to immune escape as well as chemotherapy resistance.4,5,6 Among these constituents, tumor-associated macrophages (TAMs) have emerged as particularly pivotal orchestrators of the immunosuppressive niche. TAMs play a central role in shaping the microenvironment by facilitating immune regulation, fostering tumor growth, and driving immune evasion as well as metastasis.7,8 Their recruitment, survival, and functional polarization are heavily influenced by specific signaling pathways, with the colony-stimulating factor-1 (CSF-1) and its receptor (CSF-1R) axis being a master regulator. The CSF-1/CSF-1R axis is a key regulator of macrophage recruitment and polarization.9 Activation of CSF-1R signaling cascades not only promotes the survival and proliferation of macrophages but also critically enhances the accumulation and functional skewing towards an M2-type, pro-tumorigenic phenotype within the PDAC immune microenvironment, thereby significantly amplifying the tumor’s immune suppressive effects and contributing to therapeutic failure.10,11 Therefore, given this central role in maintaining an immunosuppressive fortress, targeting the CSF-1R pathway has become one of the key research areas for developing novel therapeutic strategies aimed at reprogramming the TME and restoring anti-tumor immunity.
In this context, Surufatinib represents a promising multi-kinase inhibitor designed to simultaneously disrupt several pro-tumorigenic pathways. Surufatinib is a multi-target small molecule inhibitor that exerts anti-tumor effects by targeting vascular endothelial growth factor receptor (VEGFR), fibroblast growth factor receptor (FGFR), and CSF-1R, thereby modulating the TME.12 This triple-action mechanism allows it to attack the tumor on multiple fronts: inhibiting angiogenesis via VEGFR blockade, disrupting stromal signaling through FGFR inhibition, and directly targeting the immunosuppressive macrophage compartment via CSF-1R. The immune-regulatory properties of Surufatinib have been established in several preclinical and clinical studies, particularly highlighting its significant role in reshaping the function and composition of TAMs. By potently inhibiting CSF-1R signaling, Surufatinib effectively reduces the recruitment, survival, and accumulation of immunosuppressive M2-like macrophages, thereby potentially dismantling a key pillar of immune suppression and enhancing endogenous anti-tumor immunity. This macrophage-modulating effect is central to improving the overall immune landscape of tumors. In addition to its impact on macrophages, Surufatinib facilitates T cell infiltration into the tumor bed and concurrently inhibits tumor angiogenesis, a combination that can further enhance the immunogenicity of the tumor and create a more permissive environment for immune attack. These multifaceted actions provide a new and compelling therapeutic option for patients with solid tumors.13,14,15
Despite its demonstrated efficacy across multiple tumor types, including neuroendocrine tumors, the specific and detailed impact of Surufatinib on the uniquely hostile immune microenvironment of PDAC, particularly when used in combination with standard-of-care chemotherapy regimens, remains insufficiently characterized. Chemotherapy, such as gemcitabine-based protocols, itself can induce complex shifts in the TME,16,17 and how Surufatinib interacts with these changes is poorly understood. In particular, the nuanced influence of Surufatinib on distinct macrophage subsets, their polarization states, transcriptional programs, and its potential role in overcoming or modulating treatment resistance have yet to be fully elucidated. Given the critical, non-redundant role of macrophages in shaping and maintaining the immunosuppressive TME of PDAC, a deeper, high-resolution understanding of how Surufatinib modulates macrophage phenotypes and their multifaceted interactions with other immune and stromal cells within the TME is essential for refining and optimizing therapeutic strategies. Such knowledge could identify biomarkers of response and resistance, and guide rational combination therapies.
To address these gaps in knowledge, the primary objective of this study is to investigate the immunomodulatory effects of Surufatinib combined with chemotherapy on the immune microenvironment of PDAC and to explore the critical role of macrophages in this process. We employ a comprehensive, multi-faceted approach to dissect these complex interactions. Using single-cell RNA (scRNA) sequencing technology, we conducted a comprehensive analysis of immune cell subsets in surgical specimens from PDAC patients who received neoadjuvant or perioperative Surufatinib combined with chemotherapy, assessing the dynamic changes in macrophage subpopulations and their relationship with treatment response. This high-resolution analysis allows for an unbiased characterization of the immune compartment at a cellular level. Furthermore, we also explored the potential role of GPR34, a macrophage marker associated with immune suppressive functions, in mediating resistance and immune suppression, and validated the direct regulatory effects of Surufatinib on macrophage function through a series of complementary in vitro assays and in vivo mouse models. Ultimately, by integrating these findings, we aim to provide novel mechanistic insights into immune modulation strategies for pancreatic cancer and offer a robust theoretical basis for optimizing clinical treatment strategies to improve outcomes for patients with this devastating disease.
Results
Macrophage Reprogramming is Associated with Resistance to Surufatinib in Pancreatic Cancer
To evaluate the efficacy and safety of macrophage-targeted adjuvant therapy in PDAC, we conducted a phase II clinical trial (NCT05908747) in our center. Surufatinib, a multi-target tyrosine kinase inhibitor, is known to inhibit CSF-1R, FGFR1, and VEGFR. However, analysis of the expression of the above target transcripts in The Cancer Genome Atlas (TCGA) dataset revealed that among these targets, CSF1-CSF-1R is uniquely up-regulated in pancreatic tumor tissues, whereas FGFR1 and VEGFA are predominantly expressed in normal pancreatic tissues (Supplementary Fig. 1a-c). Further examination of publicly available scRNA sequencing datasets from the TISCH database confirmed that CSF1R is highly enriched in monocyte/macrophage populations across multiple PDAC datasets (Supplementary Fig. 1d). Given the marked overexpression of CSF-1R in pancreatic cancer patients, we propose that surufatinib may exert immunomodulatory effects by inhibiting the CSF-1R signaling pathway in macrophages.
To investigate the impact of CSF-1R inhibition on the tumor immune microenvironment and the mechanisms underlying resistance to Surufatinib, we performed scRNA sequencing on tumor samples from patients enrolled in the clinical trial. In addition, we performed validation in prospective and retrospective cohorts, mechanistic exploration in vivo and in vitro models, as well as translational studies in preclinical models (Fig. 1a). Patients were stratified into responder and non-responder groups based on radiologic criteria defined by RECIST v1.1: those achieving partial response (PR) or stable disease (SD) with tumor regression were classified as responders, while those with progressive disease (PD) or SD with tumor growth were considered non-responders (Fig. 1b). Following >4 weeks of treatment with Surufatinib in combination with gemcitabine and nab-paclitaxel, surgical resection was performed using a standardized operative protocol. Postoperative patients received standardized treatment procedures and long-term follow-up. To date, 33 patients have been enrolled, of whom 8 have PD, 11 have PR, and 14 have SD (Fig. 1c). Tumor tissues from 16 patients were subjected to scRNA sequencing. The resulting transcriptomes were analyzed for cell-type composition and transcriptional variability (Fig. 1d, Supplementary Fig. 1e).
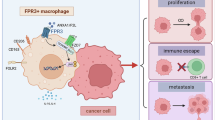
Single-cell RNA sequencing reveals the cellular composition of pancreatic cancer after chemotherapy combined with Surufatinib treatment. a Schematic diagram of the process of this study. First, we performed single-cell RNA (scRNA) sequencing of surgical specimens obtained from patients who were treated with gemcitabine, nab-paclitaxel, and surufatinib in the clinical trial. Patients were divided into responder group and non-responder group according to clinicopathological evaluation. Comparing the differences in macrophage subsets between the two groups, it was found that GPR34+ damage-associated macrophages (DAMs) were absolutely dominant in the treatment-resistant patients. Subsequently, GPR34+ DAMs function and underlying mechanisms were validated in retrospective and prospective pancreatic cancer patient cohorts and mouse models. Finally, targeting GPR34+ DAMs was demonstrated to enhance the efficacy of surufatinib in a preclinical model. Schematic diagram was created with BioRender.com. PDAC: pancreatic ductal adenocarcinoma, DAMs: damage associated macrophages, scRNA: single cell RNA. b Representative contrast-enhanced CT images of responder group (R) and non-responder group (NR) before and after treatment. Patients underwent contrast-enhanced CT and tumor assessment every 8 weeks, and two independent experienced radiologists determined the location and contour of the target lesion. The length of the tumor was measured and counted by pancreatic tumor specialists. The baseline level before treatment as well as the last evaluation before surgery were compared respectively. c Waterfall plot showing efficacy evaluation results from clinical trial patients, with upper dashed line indicating the cutoff for disease progression according to the RECIST (Response Evaluation Criteria in Solid Tumors) V1.1 standard, lower dashed line indicating the cutoff for PR. The final evaluation results are marked above and below each column of the waterfall plot. d UMAP plot displaying the cellular subtypes of all cells (n = 84380) in the surgical specimens after treatment. e Scale bar chart showing the cell type composition of all patient samples, and aggregated samples from non-responders and responders. f The box plot compared the proportion differences of each cell type between non-responders and responders. A beta regression analysis was used to examine the correlation between the proportions of various cell components and treatment efficacy. g UMAP plot showing macrophage subtypes clustering in all patient samples. mac macrophage, cl cluster. h Bar chart showing the composition of macrophage cluster in all patient samples and aggregated samples from non-responders and responders. i The box plot compared the proportion differences of each macrophage subcluster between non-responders and responders. A beta regression analysis was used to examine the correlation between the proportions of various subclusters and the treatment efficacy. j Heatmap displaying the expression of key functional genes across macrophage subclusters. The characteristic genes of each subpopulation were ranked by log2FC, and the top 50 genes with adjusted P-value < 0.05 for each subcluster were selected for visualization and gene set enrichment analysis (GSEA). The main enrichment pathways were selected for display. Each column represents a cell barcode, sorted by cell type, and each row represents a gene. k Bar plot showing the GSEA analysis results for the Mac_cl1 cluster. The analysis was based on the Hallmarks, GOBP and KEGG data sets, and all results were screened based on the scores of the data sets mentioned above. NS no significance, *P < 0.05
Dimensionality reduction and clustering revealed marked differences in immune and epithelial cell populations between responders and non-responders. Notably, the non-responder group exhibited higher proportions of macrophage (Fig. 1e, f). Given that Surufatinib has capacity of targeting macrophage CSF-1R, we further analyzed differences in macrophages between the efficacy groups. Macrophage clustering identified five macrophage subtypes (Fig. 1g, Supplementary Fig. 1f). Among them, the Mac_cl1 subset was significantly enriched in non-responders (Fig. 1h-i). Gene set enrichment analysis (GSEA) revealed that Mac_cl1 was highly enriched in lysosomes, damage response pathways, and apoptotic cell clearance pathways (Fig. 1j, k), suggesting a role in immunosuppressive remodeling. In addition, other macrophage clusters- including IL1β+ macrophages (Mac_cl2), immune-activated macrophages (Mac_cl3), protein kinase pathway-activated macrophages (Mac_cl4), and Ki67+ macrophages (Mac_cl5)- displayed transcriptional signatures associated with immune activation and cell cycle progression (Supplementary Fig. 1g–j, Table S1). For the significantly higher proportion of Mac_cl1 in the non-responder group and its overall infiltration proportion in macrophages (40-60%; Fig. 1i), we hypothesized that Mac_cl1 might be associated with resistance to surufatinib combined with chemotherapy.
Further analysis of T cell subsets revealed that non-responders had increased frequencies of naïve and dysfunctional cytotoxic T lymphocytes (CTLs), but reduced effector T cell populations (Supplementary Fig. 2a–d). While we did not annotate an exhausted T cell (Tex) cluster explicitly, CTL populations in non-responders expressed high levels of canonical exhaustion markers (e.g., NR4A1, TIGIT, BATF; Supplementary Fig. 2b, e–h), indicating a dysfunctional cytotoxic phenotype. Thus, T cells in non-responders exhibited higher exhaustion levels. In contrast, no significant changes were observed in the proportions of CAF subtypes-including myCAF, inflammatory CAF (iCAF), and antigen-presenting CAF (apCAF) -between responder groups and non-responder group (Supplementary Fig. 2i-j). Except for the higher proportion of apCAF in responder group, no significant differences were observed in the proportions of other subtypes between the two groups (Supplementary Fig. 2k-l). Collectively, these findings suggest that the Mac_cl1 macrophage population, enriched in lysosome- and efferocytosis-related pathways, is highly correlated with T-cell dysfunction and resistance to Surufatinib-based therapy in PDAC.
Chemotherapy-Induced Damage-Associated Macrophages (DAMs) Drive Immune Suppression and Therapeutic Resistance
scRNA sequencing analysis identified the Mac_cl1 population as a distinct subset of damage-associated macrophages (DAMs) enriched for transcriptional programs involved in tissue injury response and repair. This subset was characterized by robust complement activation, including elevated expression of C1q family genes (Fig. 1j–k), a hallmark of pro-resolving macrophage activity. Although C1q⁺ TAMs have been previously linked to wound healing and tumor-promoting inflammation, heir effector mechanisms in PDAC remain largely undefined.
To elucidate the functional landscape of Mac_cl1, we analyzed its expression of receptors involved in apoptotic cell clearance and damage recognition. Notably, Mac_cl1 exhibited upregulation of TREM2, CD163, ADORA3, SLAMF8, and CD180 and GPR34 (Supplementary Fig. 3a). Furthermore, Mac_cl1 primarily displayed an M2-like polarization pattern in classical macrophage functions (Supplementary Fig. 3b). Among these, GPR34, CD163, and TREM2 were markedly enriched in non-responder group (Fig. 2a). While TREM2 and CD163 were broadly expressed across multiple macrophage subsets, GPR34 expression was uniquely restricted to Mac_cl1, with minimal expression observed in other TAM populations or non-myeloid cell types (Fig. 2b, c, Supplementary Fig. 3a-b). In addition, Mac_cl1 is highly enriched in the classical M2 activation pathway (Supplementary Fig. 3c). Given that GPR34 is a known receptor for lysophosphatidylserine (LysoPS)-a phospholipid elevated in necrotic tissue and pancreatic tumor environments-we hypothesized that GPR34⁺ TAMs serve as sensors of tissue injury, integrating damage signals to promote immunosuppressive reprogramming and therapy resistance.
Cohort and in vitro studies reveal the role of GPR34 in macrophages. a Volcano plot showing differentially expressed genes between responder group and non-responder group from the macrophage subclusters of scRNA sequencing data. The horizontal dashed line represents the P-value cutoff (P < 10⁻⁵⁰), and the vertical dashed line represents the log2FC cutoff (-1 or 1). FC fold change, sig significance, R Responder, NR Non-responder. b, c UMAP plot showing the expression of GPR34 in all cells (b) and macrophages (c). Mac: Macrophage, cl cluster. d Representative mIF staining images (100x) of surgical specimens from responders and non-responders in the clinical trial (n = 26). Navy: GPR34, Magenta: CD68, Red: CD8, Green: Tim-3, Yellow: CK19, Blue: DAPI. White arrowheads: Tim-3+CD8+ exhausted T cells (Tex); White arrows: GPR34+CD68+ macrophages. White scale bar = 100 μm. e Bar plot comparing the proportion of CD68+ macrophages (top) and the proportion of GPR34+ cells among CD68+ macrophages (bottom) between responders and non-responders. Two-tailed unpaired t-test. Data are presented by mean ± SD. *P < 0.05. f Bar plot comparing the proportion of CD8+ T cells (top) and the proportion of Tim-3+ cells among CD8+ T cells (bottom) between responders and non-responders. Two-tailed unpaired t-test. Data are presented by mean ± SD. g Representative mIF staining images (200x) of surgical specimens from clinical trial patients (n = 26). Navy: GPR34, Magenta: CD68, Red: MPO, Green: CD3, Dark yellow: CD20, Yellow: CK19, Blue: DAPI. White scale bar = 50 μm. h Representative mIF staining images (200x) of clinical trial patient surgical specimens (n = 26). Navy: GPR34, Magenta: CD68, Red: α-SMA, Green: CD31, Dark yellow: CD117, Yellow: CD56, Blue: DAPI. White scale bar = 50 μm. i Bar plots showing the ratio of GPR34+ cells in different cell types (n = 26). One-way ANOVA with Dunnett’s test was used to compare other cell types versus macrophages. Data are presented by mean ± SD. j Bar plots showing the ratio of GPR34+ cells in different cell types (n = 26) examined by flow cytometry. One-way ANOVA with Dunnett’s test was used to compare other cell types versus macrophages. Data are presented by mean ± SD. k, l Kaplan-Meier curves for overall survival (OS) and Recurrence-free survival (RFS) in the prospective cohort (k) and retrospective cohort (l), comparing prognosis between groups with high versus low infiltration of GPR34+ macrophages. The cutoff for the proportion of GPR34+ cells among CD68+ cells were dichotomized using a 20%. Log-rank test was used for comparison. m Flow cytometry analysis of GPR34+ cells in tumor tissue versus adjacent non-tumorous tissue from prospective cohort patients (n = 42). Scatter plot shows the paired infiltration proportion of GPR34+ cells in tumor and normal tissue from the same patient. Two-tailed paired t-test was used. n Flow cytometry analysis of GPR34+ cells in tumor tissue from prospective cohort patients (n = 42). Bar plots show the infiltration proportions of CD45+ cells, CD8+ T cells, Tim-3+PD-1+ T cells, MRC1+ macrophages, and MHC-I+ macrophages in the low GPR34 ( ≤ 20%) versus high GPR34 ( > 20%) groups. Two-tailed unpaired t-test. Data are presented by mean ± SD. o BMDMs from C57BL/6 mice were cultured until day 5, stimulated with KPC cell TCM for 12 h, followed by treatment with Surufatinib (4 nM) or CSF-1R inhibitor (PLX3397, 20 nM) for 24 h, then analyzed by flow cytometry (n = 3). Bar plot compares the gMFI of GPR34 among groups. One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. NS no significance, * P < 0.05, **P < 0.01, *** P < 0.001
To validate the scRNA sequencing findings, we performed multiplex immunofluorescence (mIF) staining on the paraffin slice of surgical specimens from clinical trial patients. The results showed significantly higher infiltration of GPR34⁺CD68⁺ TAMs in non-responders compared to responders (Fig. 2d–f). In contrast, no significant difference was observed in TREM2⁺ macrophage infiltration between the two groups (Supplementary Fig. 3d–f), indicating that GPR34, rather than TREM2, plays a critical role in damage signal response within DAMs. Additionally, non-responders exhibited a significantly higher proportion of exhausted CD8⁺ T cells than responders (Fig. 2f). mIF analysis of patient samples confirmed that GPR34 was specifically highly expressed in macrophages, with significantly lower expression in other immune cells (including T cells, B cells, NK cells, and neutrophils) as well as in CAFs and vascular endothelial cells (Fig. 2g–i). Flow cytometric analysis of surgical specimens yielded consistent results (Supplementary Fig. 3g, Fig. 2j).
To further validate the presence and phenotype of GPR34+ TAMs in PDAC, we analyzed three independent external scRNA sequencing datasets (GSE155698, CRA001160, GSE205013). All datasets confirmed the existence of GPR34+ macrophage clusters (Supplementary Fig. 4a–c). These TAMs were implicated in lysosome and damage response signal (Supplementary Fig. 4d–f). Additionally, samples with a higher proportion of GPR34+ macrophages exhibited elevated frequencies of exhausted T cells (Tex) and naïve T cells (Supplementary Fig. 4g–m), consistent with our findings from the clinical trial cohort. These data suggest that GPR34+ TAMs are linked to tissue injury response, necrotic cell clearance, and immunosuppression.
Tissue microarrays (TMAs) analysis of a retrospective cohort (patients received upfront surgery, n = 109) and a prospective cohort (patients received neoadjuvant chemotherapy, n = 42) of PDAC patients in our center revealed that the proportion of GPR34+ macrophages and the proportion of macrophages were negatively correlated with CD8+ T cell infiltration, and positively associated with FOXP3+ regulatory T (Treg) cells infiltration, while the proportion of GPR34- macrophage have no significant correlation with CD8+ T cell and Treg. (Supplementary Fig. 5a–h). High infiltration of GPR34+ macrophages was associated with shorter recurrence-free survival (RFS) and overall survival (OS) (Fig. 2k, l). Multivariate Cox regression analysis indicated that high GPR34⁺ macrophage infiltration was an independent risk factor for both RFS and OS in the retrospectiveand perspective cohort (Supplementary Fig. 5i–l). Spatial distribution analysis indicated that GPR34+ TAMs were confined to tumor nests, with minimal infiltration into distal stromal areas, while CD8⁺ T cells were largely restricted to peripheral stromal regions (Supplementary Fig. 5m–o). Flow cytometry of freshly resected tumor samples from the prospective cohort confirmed GPR34⁺ TAMs were significantly more abundant in tumor tissues than in matched adjacent normal tissues (Fig. 2m, Supplementary Fig. 6a), and this was accompanied by increased CD8⁺ T cell exhaustion and M2-like TAMs (Fig. 2n, Supplementary Fig. 6b). Together, these results establish GPR34+ TAMs are highly associated with PDAC immunosuppression and poor prognosis.
To investigate the regulation of GPR34 expression, we stimulated bone marrow-derived macrophages (BMDMs) with tumor-conditioned media (TCM) and evaluated the effect of Surufatinib and a selective CSF-1R inhibitor. Neither compound alone induced GPR34, but both compounds, in combination with TCM significantly upregulated GPR34 expression in BMDMs (Fig. 2o, Supplementary Fig. 6c). Moreover, GPR34 expression was enhanced under M2-polarizing conditions (IL-4, IL-13) and in the presence of chronic inflammatory cytokines (IL-1β, IL-6) (Supplementary Fig. 6d). These findings suggest that CSF-1R blockade by Surufatinib potentiates GPR34 expression in macrophages within a tumor-associated cytokine milieu, and GPR34 had correlation with immune suppression and therapy resistance.
GPR34 promotes macrophage-mediated immunosuppression and efferocytosis
To elucidate the function of GPR34 in macrophages during pancreatic tumor progression, we generated a macrophage-specific conditioned knockout model (Gpr34ΔLyz2) by crossing Gpr34flox/flox mice with Lyz2-CreERT mice. Orthotopic pancreatic tumors were established using KPC-GFP cells, and tamoxifen was administered seven days prior to tumor implantation to induce. To simulate tissue damage signals, mice were treated with gemcitabine and nab-paclitaxel, and flow cytometry analysis of tumors was performed at the end of the experiment (Supplementary Fig. 7a, b, 8a). To extend these findings, we performed bone marrow transfer (BMT) model by transplanting GPR34−/− hematopoietic stem cells (HSCs) into KrasG12D/+; Trp53R176H/+; Pdx1-Cre (KPC) mice after bone marrow depletion, and mice were given chemotherapy after tumor formation (Supplementary Fig. 8b). Bioluminescence imaging revealed that tumor growth was comparable between Gpr34ΔLyz2 and Gpr34flox/flox mice. However, upon administration of chemotherapy, Gpr34ΔLyz2 mice exhibited significantly reduced tumor burden, suggesting that GPR34 promotes tumor tolerance in response to treatment-induced damage (Fig. 3a–c). Flow cytometric analysis confirmed GPR34 depletion in F4/80⁺ macrophages in Gpr34ΔLyz2 mice (Supplementary Fig. 8c). Immune profiling of tumors from Gpr34ΔLyz2 mice exhibited a marked reduction in immunosuppressive macrophages, accompanied by CTLs, and decrease in Tex (Fig. 3d). Further analysis revealed a shift toward an M1-polarized macrophage phenotype, reduced M2 macrophages, and expansion of intratumoral dendritic cells in Gpr34ΔLyz2 mice, consistent with enhanced antigen presentation (Supplementary Fig. 8d). In contrast, other immune cell populations— including myeloid-derived suppressor cells (MDSCs), Tregs, and monocytes—remained unchanged (Supplementary Fig. 8e). Peripheral blood mononuclear cells (PBMCs) showed no significant alterations in CD45+ cell subsets (Supplementary Fig. 8f).
Macrophage-specific GPR34 knockout improves chemotherapy outcomes in mice. a Orthotopic pancreatic injection of KPC-GFP-LUC cells into Gpr34flox/flox and Gpr34ΔLyz2 mice. After tumor formation, chemotherapy was administered to simulate tumor killing. Tumor bioluminescence intensity was dynamically monitored by bioluminescence imaging. Representative bioluminescence images show tumor growth in each group (n = 5). b Time-course curve of bioluminescence intensity in orthotopic tumor-bearing mice (n = 5). Two-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SEM. c Bar plot of tumor weight on day 21 post-implantation in orthotopic tumor-bearing mice (n = 5). One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. NS: no significance, *: P < 0.05. d Flow cytometry analysis of the proportions of CTL, Tex, M1 macrophages, and M2 macrophages in tumor tissues of orthotopic tumor-bearing mice (n = 5). Bar plots show the infiltration differences of immune cells among groups. One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. e KPC mice were irradiated with a total dose of 10 Gy, followed by tail vein injection of bone marrow cells from Gpr34+/+ or Gpr34−/− mice. After successful transplantation confirmed by flow cytometry. Tumor growth was monitored by B-ultrasound during chemotherapy simulating injury signals (n = 8–10). Representative B-ultrasound images compare tumor size among groups. Black dashed lines outline tumor boundaries. Tumor volume was calculated as 0.5 * long diameter * (short diameter) ². S: spleen; K: kidney. Black arrows point to the tumor. f Time-course plot of tumor volume monitored by B-ultrasound. Each solid line represents an individual mouse. Two-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. g Kaplan-Meier curve of KPC spontaneous tumor-bearing mice (n = 8–10). Log-rank test was used. h mIF staining of paraffin sections from orthotopic tumors in the bone marrow transplantation KPC mice (n = 6). Representative fluorescence images (top 100x, bottom 400x) show exhausted T cell infiltration among groups. Red: CD8α, Green: Tim-3, White: CK19, Blue: DAPI. White scale bar = 50 μm. i Bar plot showing the proportion of Tim-3+CD8+ T cells from mIF staining in KPC spontaneous tumor-bearing mice (n = 6). One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. j Fluorescence imaging of macrophages sorted by flow cytometry from tumors of KPC-GFP-LUC orthotopic injection mice model. Representative images show GFP phagocytosis by macrophages in each group (n = 6). Green: GFP, Blue: DAPI. White scale bar = 20 μm. k Bar plot showing the gMFI of GFP phagocytosis by flow-sorted macrophages from orthotopic tumors (n = 5). One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. l Bar plot of the gMFI of GFP in macrophages from KPC cell orthotopic tumor-bearing mice (n = 6). One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. m Bar plot showing the proportion of MerTK+ or AXL+ macrophage from flow cytometry analysis in KPC mice (n = 6). One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. n BMDMs from Gpr34+/+ and Gpr34−/− mice were cultured until day 5, cultured with TCM for 12 h, followed by co-culture with CD8+ T cells for 24 h. Flow cytometry detected the proportion of MerTK+ or AXL+ macrophage. Bar plots show differences between Gpr34+/+ and Gpr34−/− groups (n = 6). Two-tailed unpaired t-test was used. Data are presented by mean ± SD. NS no significance, *P < 0.05, **P < 0.01, ***P < 0.001
In BMT model, ultrasound imaging confirmed that GPR34 knockout significantly enhanced the chemotherapeutic efficacy against primary tumors (Fig. 3e, f) and prolonged mouse survival (Fig. 3g). Histological analysis revealed enhanced tumor cell apoptosis (TUNEL+) and reduced proliferation (Ki-67+) in Gpr34⁻/⁻ tumors compared to controls (Supplementary Fig. 8g, h). mIF stain further demonstrated a reduced proportion of Tim-3+ Tex in Gpr34−/− tumor microenvironment (Fig. 3h, i).
To assess the efferocytosis of GPR34⁺ macrophages, we performed flowcyte sorting on orthotopic tumors derived from KPC-GFP cells and conducted morphological analysis of sorted macrophages (CD45+CD11b+F4/80+). The results showed significantly impaired uptake of GFP⁺ apoptotic tumor cells by F4/80⁺ macrophages in Gpr34ΔLyz2 mice (Fig. 3j, k). Flow cytometry confirmed that the proportion of GFP⁺ macrophages was significantly lower in Gpr34ΔLyz2 mice compared to controls (Fig. 3l, Supplementary Fig. 8i). These findings were further validated by in vitro co-culture assays, in which BMDMs from Gpr34⁻/⁻ mice exhibited diminished phagocytosis of chemotherapy-induced apoptotic GFP⁺ tumor cells compared to Gpr34+/+ macrophages. Additionally, flow cytometry indicated that expression of efferocytosis receptors (MerTK, AXL) was significantly lower in macrophages from Gpr34ΔLyz2 mice (Fig. 3m, Supplementary Fig. 8j, k). Similar conclusions were drawn from a co-culture model of mouse BMDMs with GFP-labeled apoptotic tumor cells (Fig. 3n, Supplementary Fig. 8l–o).
Given the potential role of macrophage GPR34 in damage response, we compared Gpr34ΔLyz2 mice and Gpr34flox/flox mice in a cerulein-induced pancreatitis model. hematoxylin-eosin (HE) staining showed that macrophage-specific GPR34 knockout had no significant effect on Schmidt scores (Supplementary Fig. 9a, b). Moreover, Masson staining indicated that Gpr34 deletion did no significantly affect fibrosis in the inflammatory model (Supplementary Fig. 9c, d). Flow cytometric analysis of pancreatic tissues revealed no notable changes in macrophage or neutrophil infiltration upon Gpr34 knockout (Supplementary Fig. 9e–h). Additionally, no significant differences were observed in serum amylase or lipase levels (Supplementary Fig. 9i, j). Flow cytometric comparison of GPR34⁺ macrophage infiltration between inflammatory and pancreatic tumor models showed that the proportion of GPR34⁺ macrophages was significantly lower in the pancreatitis model than in the tumor model (Supplementary Fig. 9k, l). These results indicate that GPR34⁺ macrophages infiltrate predominantly in malignant pancreatic tissues, with minimal presence in pancreatitis. Therefore, macrophage-specific Gpr34 knockout does no significantly influence pancreatic inflammation levels.
Collectively, these results establish GPR34 as a critical mediator of macrophage-driven efferocytosis and immune suppression in pancreatic cancer. Loss of GPR34 reprograms the tumor microenvironment by attenuating immunosuppressive macrophage activity, promoting CD8⁺ T cell infiltration, and impairing apoptotic cell clearance. These data underscore the potential of GPR34 as a therapeutic target to remodel the myeloid compartment and enhance anti-tumor immunity in PDAC.
GPR34 Drives CD8⁺ T Cell Exhaustion via CXCL16 Secretion by Macrophages
To elucidate the underlying mechanisms by which GPR34⁺ TAMs mediate immunosuppression, we conducted depletion of CD8+ T cells and natural killer (NK) cells in Gpr34ΔLyz2 and Gpr34flox/flox mice (Supplementary Fig. 10a–c). Depletion of CD8+ T cell completely abrogated the chemo-sensitization effect observed in Gpr34ΔLyz2 mice (Fig. 4a–c), whereas NK cell depletion had no significant impact (Supplementary Fig. 10d–f), These results indicate that GPR34 primarily regulates tumor killing through CD8⁺ T cell-dependent mechanisms.
Validation of GPR34 function in macrophage and CD8+ T cell co-culture system. a Gpr34flox/flox and Gpr34ΔLyz2 mice were treated with anti-CD8α or IgG, followed by orthotopic pancreatic injection of KPC-LUC cells. After tumor formation, chemotherapy was administered to simulate an injury signal. Tumor bioluminescence was dynamically monitored. Representative bioluminescence images show tumor growth in different groups (n = 6). b Time-course curve of bioluminescence imaging for the KPC-LUC orthotopic model (n = 6). Two-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. NS no significance, ***P < 0.001. c Bar plot showing tumor weight on day 21 in the KPC-LUC orthotopic model (n = 6). One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. NS no significance. d, e BMDMs from Gpr34+/+ and Gpr34−/− mice were cultured until day 5, then co-cultured with TCM and KPC-GFP cells for 12 h. BMDMs were then isolated and co-cultured with CD8+ T cells for 24 hours. Flow cytometry analyzed the expression of functional molecules in BMDMs (d) and CD8+ T cells (e). Bar plots show levels in Gpr34+/+ vs Gpr34−/− groups (n = 3). Two-tailed unpaired Student’s t test was used. Data are presented by mean ± SD. f, g BMDMs from Gpr34+/+ and Gpr34−/− mice were cultured until day 5, pre-stimulated with SIINFEKL, then cultured with TCM for 12 h, followed by co-culture with CD8+ T cells from OT-1 mice for 24 h. Flow cytometry detected T cell-specific killing function (f) and BMDM antigen presentation function (g). Bar plots show differences between Gpr34+/+ and Gpr34−/− groups (n = 3). Two-tailed unpaired t-test was used. Data are presented by mean ± SD. h Violin-box plots of cytokine transcript expression in macrophage clusters from scRNA sequencing data. White dot and solid lines in boxes represent medians and quartiles. Two-tailed Wilcoxon test. i BMDMs from Gpr34+/+ and Gpr34−/− mice were cultured until day 5, then stimulated with TCM and chemotherapy-induced apoptotic KPC-GFP cells for 12 h. qPCR detected Cxcl16 transcript levels. Bar plot compares Cxcl16 transcripts between groups (n = 3). Two-tailed unpaired t-test was used. Data are presented by mean ± SD. j ELISA detection of cytokine secretion in supernatant from BMDMs stimulated with apoptotic KPC-GFP cells. Bar plot shows CXCL16 protein secretion levels from Gpr34+/+ and Gpr34−/− BMDMs (n = 10). Two-tailed unpaired t-test was used. Data are presented by mean ± SD. k, l BMDMs from C57BL/6 mice were cultured until day 5, transiently transfected with siRNA, then co-cultured with TCM, LysoPS and chemotherapy-induced apoptotic KPC-GFP cells for 12 h. BMDMs were then isolated and co-cultured with CD8+ T cells. Flow cytometry detected T cell exhaustion (k) and cytotoxicity levels (l) (n = 3). One-way ANOVA with Dunnett’s test compared siRNA groups versus control. Data are presented by mean ± SD. NS no significance, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001
To further dissect macrophage- T cell interactions, we employed a co-culture system comprising BMDMs and CD8⁺ T cells (Supplementary Fig. 10g). Flow cytometric analysis demonstrated that Gpr34−/− BMDM, stimulated with chemotherapy-conditioned tumor medium, exhibited reduced immunosuppressive activity diminished efferocytic capacity, and upregulated MHC-I expression (Fig. 4d, Supplementary Fig. 10h). These changes correlated with decreased Tex and enhanced CD8+ T cell cytotoxicity, and increased differentiation to effector T cells (Fig. 4e; Supplementary Fig. 10h). Notably, Gpr34−/− BMDMs potentiated the antigen-specific cytotoxic activity of OT-1 CD8+ T cells and reduced their exhaustion levels (Fig. 4f, g, Supplementary Fig. 10i, j). A T cell killing assay further confirmed that Gpr34⁻/⁻ BMDMs promoted specific killing of KPC-OVA cells by OT-1 CD8⁺ T cells (Supplementary Fig. 10k–l). Additionally, we evaluated the impact of macrophage Gpr34 knockout on non-specific antigen presentation and immune response in both a BMDM–CD8⁺ T cell co-culture system (Supplementary Fig. 11a–d) and the KPC–PBMC co-culture system (Supplementary Fig. 11e–h), and obtained consistent results. These data support the role of GPR34 in modulating macrophage-dependent antigen presentation and T cell dysfunction.
To identify the mediators of T cell exhaustion, we fractionated macrophage conditioned medium (MCM) using 3 kDa ultrafiltration and applied the resulting fractions to CD8⁺ T cells (Supplementary Fig. 12a). The small-molecule fractions failed to reproduce the exhaustion phenotype (Supplementary Fig. 12b, c), suggesting that macrophage-secreted proteins are the principal effectors. To validate these observations, we employed a patient-derived tumor organoid co-culture system with monocyte-derived macrophages (MDMs) and CD8+ T cells. Knockdown of GPR34 in MDMs significantly reduced CD8⁺ T cell exhaustion and enhanced cytotoxic function, confirming the clinical relevance of this pathway (Supplementary Fig. 12d–f). scRNA sequencing analysis of cell-cell communication revealed enriched interactions between GPR34+ TAMs and T cells via CXCL and CCL signaling pathways (Supplementary Fig. 13a). Among these, CXCL16 was the most significantly upregulated secreted protein transcript in GPR34⁺ macrophages (Fig. 4h; Supplementary Fig. 13b–d). Both transcriptional and secreted levels of CXCL16 and were significantly attenuated in Gpr34−/− BMDMs (Fig. 4i, j). Moreover, knockdown of Cxcl16 in BMDM rescued LysoPS (a GPR34 agonist)-induced CD8⁺ T cell exhaustion and restored cytotoxicity (Fig. 4k-l). Together, these findings demonstrate that GPR34 promotes CD8⁺ T cell exhaustion by inducing macrophage secretion of CXCL16, thereby reinforcing immunosuppression and fostering resistance to chemotherapy in the PDAC tumor microenvironment.
The LysoPS-GPR34 signaling axis regulates macrophage efferocytosis and cytokine secretion
As the sole known ligand for GPR34, LysoPS mediates classical GPCR signaling. Both LysoPS and its precursor PS are abundantly generated during tissue injury and apoptosis. To elucidate LysoPS-GPR34 signaling in macrophage efferocytosis, the co-cultured system of LysoPS-treated BMDMs with chemotherapy-induced apoptotic tumor cells was employed. LysoPS treatment markedly enhanced macrophage efferocytosis, an effect that was significantly attenuated in Gpr34−/− cells (Fig. 5a, Supplementary Fig. 14a). LysoPS activates the PI3K-AKT pathway (Fig. 5b, Supplementary Fig. 14b), consistent with established GPCR signaling mechanisms. LysoPS acts through GPR34 to promote macrophage immunosuppressive function and reduce MHC-I expression (Fig. 5c, Supplementary Fig. 14c). Importantly, PI3K/AKT inhibitor not only reduced macrophage efferocytosis (Fig. 5d) but also restored immune function suppressed by LysoPS (Fig. 5e, Supplementary Fig. 14d–f), demonstrating that the LysoPS-GPR34 axis mediates activation of immunosuppression and efferocytosis through PI3K-AKT signaling pathway.
LysoPS-GPR34 modulates macrophage efferocytosis and inflammatory cytokine secretion. a, b BMDMs from Gpr34+/+ and Gpr34−/− mice were cultured until day 5, co-cultured with TCM, LysoPS and chemotherapy-induced apoptotic KPC-GFP cells for 12 h. These were then co-cultured with CD8+ T cells for 24 hours. Flow cytometry detected GFP efferocytosis (a) and p-AKT levels (b) in BMDMs. Bar plots show differences among groups. One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. c BMDMs from Gpr34+/+ and Gpr34−/− mice were treated with LysoPS and co-cultured with CD8+ T cells. Flow cytometry detected MRC1, MHC-I on macrophages, and Tim-3, GZMB on CD8+ T cells. Bar plot shows gMFI differences among groups. One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. d, e BMDMs from C57BL/6 mice were treated with LysoPS and AKT inhibitor, then co-cultured with chemotherapy-induced apoptotic KPC-GFP cells for 12 h, followed by co-culture with CD8+ T cells for 24 h. Flow cytometry detected GFP uptake by BMDMs (d) and exhaustion/cytotoxicity levels of CD8+ T cells (e). One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. f UMAP plot showing relative expression of efferocytosis-related genes from scRNA sequencing analysis. g BMDMs from Gpr34+/+ and Gpr34−/− mice were cultured until day 5, co-cultured with TCM and either chemotherapy-induced apoptotic KPC-GFP cells or normal KPC-GFP cells for 12 h. After removing cells, RNA was extracted for qPCR. Bar plot compares transcript levels among groups. One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. h BMDMs from C57BL/6 mice were cultured until day 5, co-cultured with apoptotic KPC cells of negative control for 12 hours. RNA was extracted for qPCR. Bar plot compares transcript differences among groups. One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. i BMDMs from C57BL/6 mice were cultured until day 5, stimulated with TCM and LysoPS for 12 h. ELISA detected CXCL16 secretion in supernatant. Bar plot shows levels in LysoPS vs. control groups. Two-tailed unpaired Student’s t test was used. Data are presented by mean ± SD. NS no significance, *P < 0.05, **P < 0.01, ***P < 0.001
Analysis of scRNA sequencing data identified significant upregulation of efferocytosis-related genes in GPR34+ TAMs, including MERTK, AXL, TYRO3, and GAS6 (Fig. 5f). Gpr34 knocking out in macrophages rescued the upregulation of apoptotic cell recognition receptors. Given that both LysoPS and PS accumulate in apoptotic tissues, with PS being converted to LysoPS by phospholipase A2 in pancreatic tissue, we examined whether the LysoPS-GPR34 axis upregulated the efferocytosis-related molecules.18 In vitro experiments confirmed that LysoPS upregulated MERTK expression in an AKT-dependent manner (Fig. 5g, h). These results support a model wherein LysoPS-GPR34 signaling enhances macrophage efferocytosis and promotes clearance of apoptotic cells. Additionally, LysoPS induced expression of immunosuppressive cytokines CXCL16 in vivo (Fig. 5i). Collectively, our findings demonstrate that tissue injury-derived LysoPS activates macrophage GPR34 to upregulate MERTK expression via PI3K-AKT signaling, thereby enhancing PS-dependent efferocytosis, and directly promote secretion of CXCL16, which contribute to CD8+ T cell exhaustion.
Macrophage Efferocytosis Promotes MHC-I Degradation via the Lysosomal Pathway
Our findings have demonstrated that GPR34 expression in macrophages is associated with reduced MHC-I protein levels and impaired antigen presentation capacity (Fig. 4f, g). Initial experiments indicated that LysoPS treatment alone did not significantly alter MHC-I protein levels (Supplementary Fig. 15a). However, co-culture of macrophages with apoptotic tumor cells resulted in significant downregulation of MHC-I (Supplementary Fig. 15b), suggesting that MHC-I suppression is dependent on efferocytosis. This efferocytosis-mediated MHC-I reduction was reversed by MerTK inhibitor (Fig. 6a, b, Supplementary Fig. 15c). Notably, MerTK inhibitor enhanced macrophage antigen presentation, improved T cell cytotoxicity, and alleviated CD8+ T cell exhaustion (Fig. 6b, c; Supplementary Fig. 15d, e). Further analysis revealed that MerTK inhibitor reduced phagolysosome formation and restored MHC-I protein levels in macrophages (Fig. 6d–f, Supplementary Fig. 15f), indicating that efferocytosis promotes MHC-I degradation via lysosomal activation. scRNA sequencing analysis showed that GPR34+ TAMs exhibited increased expression of lysosomal enzymes and related genes (Figs. 1j, k, 6g). Genetic ablation of GPR34 impaired both lysosomal biogenesis and MHC-I degradation (Fig. 6h). Mechanistically, we identified upregulation of the lysosomal regulators TFEB and MITF in GPR34+ TAMs (Fig. 6i). Knockdown of either transcription factor prevented MHC-I degradation upon apoptotic cell uptake (Fig. 6j, Supplementary Fig. 15g) and enhanced CD8+ T cell cytotoxic function in co-culture assays (Supplementary Fig. 15h). Thus, the efferocytosis process promotes the clearance of apoptotic cells by upregulating the lysosomal pathway in macrophages. Furthermore, lysosomal inhibitors disrupted sustained macrophage efferocytosis and blocked MHC-I degradation via the lysosomal pathway (Fig. 6k–m; Supplementary Fig. 16a–d), thereby enhancing macrophage antigen presentation to CD8⁺ T cells (Fig. 6l–n, Supplementary Fig. 16e). Finally, using a BMDM-OT1 CD8⁺ T cell co-culture model, we demonstrated that LysoPS-GPR34 signaling promotes CD8⁺ T cell exhaustion by inducing CXCL16 secretion from macrophages, and facilitates MHC-I degradation via the lysosomal pathway through enhanced efferocytosis (Fig. 6o–q). Together, these data confirm that macrophage GPR34 drives immune exhaustion through dual mechanisms: secretion of the chronic inflammatory cytokine CXCL16 and impairment of antigen presentation.
Macrophage efferocytosis function influences antigen presentation ability through MHC-I. a BMDMs from C57BL/6 mice were cultured until day 5, co-cultured with TCM and chemotherapy-induced apoptotic KPC-GFP cells for 12 hours, then analyzed by flow cytometry for GFP uptake. Bar plot shows gMFI of GFP in BMDMs treated with MerTK inhibitor vs control (n = 3). One-way ANOVA with Dunnett’s test compared MerTKi groups to control. Data are presented by mean ± SD. b BMDMs from C57BL/6 mice were cultured until day 5, co-incubated with TCM and chemotherapy-induced apoptotic KPC-OVA-GFP cells for 12 hours, treated with MerTK inhibitor, then co-cultured with CD8+ T cells from OT1 mice for 24 hours. Flow cytometry detected MHC-I, SIINFEKL loading, CD80, CD86 on BMDMs. Bar plot shows differences between MerTK inhibitor and control groups (n = 3). One-way ANOVA with Dunnett’s test was used. Data are presented by mean ± SD. c Flow cytometry detection of Tetramer+, PD-1+, Tim-3+, and GZMB+ cells after co-culture of BMDMs with OT1 CD8+ T cells. Bar plot shows differences between MerTK inhibitor and control groups (n = 3). One-way ANOVA with Dunnett’s test was used. Data are presented by mean ± SD. d Apoptotic KPC cells induced by chemotherapy and labeled with Caspase3/7 green were co-cultured with BMDMs. Phagolysosome formation was detected using pHrodo red. Representative fluorescence microscopy images (1000x) show differences between MerTK inhibitor and control groups (n = 6). Green: Caspase3/7, Red: pHrodo, Blue: DAPI. White scale bar= 20 μm. e Bar plots show total pHrodo fluorescence intensity (left) and the number of Caspase3/7+ pHrodo+ vesicles per cell (right) in BMDMs after incubation with apoptotic cells (n = 6). One-way ANOVA with Dunnett’s test was used. Data are presented by mean ± SD. f Flow cytometry analysis of pHrodo gMFI in BMDMs after incubation with apoptotic cells. Bar plot shows pHrodo gMFI levels between MerTK inhibitor and control groups (n = 3). One-way ANOVA with Dunnett’s test was used. Data are presented by mean ± SD. g Violin-box plots of lysosome-associated gene transcript expression in macrophage subpopulations from scRNA sequencing data. Solid lines represent medians and quartiles. One-way ANOVA with Kruskal-Wallis H test compared groups (Mac_cl1 as reference). h, i BMDMs from Gpr34+/+ and Gpr34−/− mice were cultured until day 5, co-cultured with TCM and chemotherapy-induced apoptotic KPC cells for 12 hours. After removing apoptotic cells, RNA was extracted for qPCR. Bar plots show transcript differences between Gpr34+/+ and Gpr34−/− BMDMs (h) efferocytosis receptors, (i) lysosome-related/transcription factors, (n = 3). Two-tailed unpaired t-test was used. Data are presented by mean ± SD. j BMDMs from C57BL/6 mice were cultured until day 5, transiently transfected with siRNA, then co-incubated with TCM and chemotherapy-induced apoptotic KPC cells for 12 hours and analyzed by flow cytometry. Bar plot shows differences in MHC-I protein levels between knockdown and control groups (n = 3). One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. k BMDMs from C57BL/6 mice were cultured until day 5, co-incubated with TCM and chemotherapy-induced apoptotic KPC-OVA-GFP for 12 hours, treated with lysosomal inhibitor, then co-cultured with OT1 CD8+ T cells for 24 hours. Bar plot shows pHrodo gMFI in macrophages from flow cytometry, comparing lysosomal inhibitor group vs control. One-way ANOVA with Dunnett’s test was used. Data are presented by mean ± SD. l, m Flow cytometry detection of macrophage antigen presentation function (l) and CD8+ T cell specific killing capacity (m) in the BMDM-OT1 CD8+ T cell co-culture system. Bar plots show differences between lysosomal inhibitor and control groups. One-way ANOVA with Dunnett’s test was used. Data are presented by mean ± SD. n BMDMs from C57BL/6 mice were cultured until day 5, co-incubated with TCM, MerTK inhibitor/Lysosome inhibitor and chemotherapy-induced apoptotic KPC cells for 12 hours. Bar plot shows differences in MHC-I protein levels between different groups detected by flow cytometry (n = 3). One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. o–q BMDMs from Gpr34+/+ and Gpr34−/− mice were cultured until day 5, transiently transfected with Cxcl16 siRNA, co-incubated with TCM, LysoPS and chemotherapy-induced apoptotic KPC-OVA-GFP for 12 hours, treated with MerTK inhibitor, then co-cultured with OT1 CD8+ T cells for 24 hours. Flow cytometry detected CD8+ T cell specific killing function (o), cytotoxic function (p), and exhaustion levels (q). One-way ANOVA with Dunnett’s test was used. Data are presented by mean ± SD. NS no significance, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001
GPR34 Antagonist Enhances the Therapeutic Efficacy of Surufatinib and Chemotherapy
Given the critical role of LysoPS–GPR34 signaling in driving efferocytosis, immunosuppression, and resistance to surufatinib, we investigated the therapeutic potential of GPR34 antagonist in preclinical models. In orthotopic pancreatic tumor models, GPR34 antagonist markedly enhanced the antitumor efficacy of both standard chemotherapy and surufatinib (Fig. 7a–c). Notably, combination treatment with a GPR34 antagonist resulted in significant tumor growth suppression and prolonged survival in KPC mice (Fig. 7d–f). Additionally, TUNEL and Ki-67 staining results demonstrated that the GPR34 antagonist markedly promoted tumor apoptosis and inhibited proliferation (Supplementary Fig. 17a–c). Mechanistically, GPR34 antagonist inhibited efferocytosis and reprogrammed macrophages toward a more immunostimulatory phenotype, as evidenced by enhanced antigen presentation capacity (Supplementary Fig. 18a–d). In xenograft models, these macrophage alterations were accompanied by enhanced CD8⁺ T cell cytotoxic activity and decreased expression of exhaustion markers (Supplementary Fig. 18e–f). To further evaluate clinical relevance, we employed patient-derived organoid–PBMC co-culture systems. In this setting, GPR34 antagonist consistently improved the efficacy of both chemotherapy and surufatinib (Fig. 7g, h).
GPR34 Antagonist Sensitizes Chemotherapy and Surufatinib. a C57BL/6 mice underwent orthotopic pancreatic injection with KPC-LUC cells. After tumor formation, mice were treated with chemotherapy, Surufatinib, and GPR34 antagonist (GPR34a). Tumor dynamics were monitored by bioluminescence imaging. Representative bioluminescence images show dynamic changes in different treatment groups (n = 6). b Tumor growth curve from bioluminescence imaging of orthotopic tumor-bearing mice (n = 6). Two-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD at each time point. c Bar plot showing differences in orthotopic tumor weight after treatment (n = 6). One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. d KPC mice underwent B-ultrasound examination twice weekly during treatment. Representative B-ultrasound images show the last examination before experiment endpoint or death (n = 10). White dashed lines indicate tumor boundaries. S: spleen. K: kidney. e The fold change of the tumor volume at the last examination before death or euthanasia in the KPC spontaneous tumor-bearing model to the baseline level (n = 10). Two-way ANOVA with Kruskal-Wallis H test was used. Data are presented by mean ± SD. f Kaplan-Meier curves comparing survival of KPC mice receiving different treatments. The experimental endpoint was 102 days after treatment initiation. Log-rank test was used. g Organoids derived from patient tumor tissue were co-cultured with matched PBMCs and treated with different drugs. Apoptosis was labeled with caspase-3/7 probe and imaged (n = 6). Representative microscopy images show organoid morphology and apoptosis protein levels in different treatment groups. Green: caspase3/7. Black scale bar = 50 μm. h Bar plot showing the gMFI of caspase-3/7 in the organoid-PBMC co-culture model across treatment groups (n = 6). One-way ANOVA with post-hoc Tukey’s test was used. Data are presented by mean ± SD. NS no significance, *P < 0.05, **P < 0.01, ***P < 0.001
From a safety perspective, the combination of chemotherapy, Surufatinib, and the GPR34 antagonist was generally well-tolerated. H-E staining of major organs in tumor-bearing mice showed no significant pathological changes in the heart, lung, liver and kidney (Supplementary Fig. 19a). Furthermore, the combination of the GPR34 antagonist and Surufatinib did no significantly exacerbate chemotherapy-induced myelosuppression or hepatotoxicity (Supplementary Fig. 19b–e). Thus, the GPR34 antagonist, as an immune adjuvant, exhibited no obvious systemic side effects in mice.
Given that the LysoPS–GPR34 axis exerts immunosuppressive functions via CXCL16, we also evaluated the therapeutic potential of anti-CXCL16 as an immune target in preclinical models. However, anti-CXCL16 did not demonstrate therapeutic effects comparable to the GPR34 antagonist (Supplementary Fig. 20a–c). Moreover, anti-CXCL16 failed to significantly enhance T cell cytotoxicity (Supplementary Fig. 20d–g). Therefore, targeting CXCL16 did not markedly improve immune-mediated tumor killing in mice.
Collectively, these findings demonstrate that GPR34 antagonist can overcome immunosuppressive barriers and enhance therapeutic responses in pancreatic cancer. Targeting GPR34 thus represents a promising combinatorial strategy to augment the clinical effectiveness of surufatinib and chemotherapy.
Discussion
Targeting TAMs has emerged as a promising strategy in the immunotherapy of solid tumors. CSF-1R inhibitors modulate macrophage activity by suppressing TAM polarization and promoting dendritic cell differentiation.19,20, establishing CSF-1R as a compelling therapeutic target in oncology. CSF-1R inhibitors have demonstrated significant translational potential in glioblastoma.21,22,23,24 and have shown clinical benefit in colorectal cancer, lymphoma, and other malignancies.25,26,27 However, their efficacy in PDAC remains limited, and the mechanisms underlying this resistance are not fully understood. Recent studies have suggested that CSF-1R inhibition may upregulate PD-L1 expression on TAMs within the pancreatic cancer immune microenvironment, suggesting that CSF-1R blockade with PD-L1 in combination with immune checkpoint inhibitors may enhance antitumor immunity.28 Additionally, CSF-1R inhibitors have been reported to impair dendritic cell differentiation by inhibiting FLT3 signaling, which could contribute to their limited clinical efficacy. Our findings reveal that resistance to CSF-1R inhibition is associated with an increased proportion of GPR34+ macrophages, with CSF-1R blockade promoting GPR34 expression in the tumor microenvironment. Interestingly, prior research has suggested that GPR34 expression can be induced via alternative activation pathways such as m-CSF and IL-4 signaling.29,30 Our findings showed that resistance to CSF-1R inhibitor treatment is associated with an increased proportion of GPR34+ macrophages, and that CSF-1R inhibitors can promote the expression of GPR34 in the tumor microenvironment.
GPR34, a member of the G protein-coupled receptor family, is expressed across various immune cells population.29 Its ligand, LysoPS, is generated in large quantities during tissue injury,31 positioning GPR34 as a damage-associated molecular pattern receptor that responds to injury signals. Previous studies have shown that neutrophils in inflammatory bowel disease secrete LysoPS, which acts on GPR34 in type 3 innate lymphoid cells (ILC3s), promoting tissue repair via IL-22.32 In the context of cancer, LysoPS-GPR34 signaling has been implicated in suppressing ILC1 activation in colorectal cancer, thereby facilitating tumor progression.33
Our study identifies GPR34 as a predominant marker of TAMs in pancreatic cancer, with GPR34+ macrophages enriched in injury responses, efferocytosis, and T cell exhaustion. Transcriptomic analysis further revealed that GPR34+ macrophage subpopulations exhibit high expression of C1q (C1QA, C1QB, C1QC) and other complement-associated genes. While the precise function of C1q+ macrophages remains incompletely understood, prior studies suggest that C1q+ macrophages may arise from tissue-resident macrophages and are linked to T cell exhaustion and poor prognosis in multiple cancers.34,35,36 Notably, lipid metabolism and fatty acid uptake appear to be dominant pathways in C1q+ macrophages, facilitating the production of immunosuppressive cytokines that reinforce the tolerogenic tumor microenvironment.36 Similar to GPR34+ macrophages, C1q+ macrophages show a transcriptomic profile resembling microglia, further supporting the notion that this subpopulation belongs to tissue-resident macrophages.36,37,38 While complement activation represents a key effector mechanism in macrophage function, it is not the sole determinant of macrophage responses to tissue damage.34 While the concept of C1q+ macrophages has been widely accepted in research, our study primarily focuses on the molecules involved in macrophage injury responses and efferocytosis. Given the substantial release of LysoPS during tissue damage, and the presence of various lipases in pancreatic tissue that facilitate the breakdown of phosphatidylserine (PS) into LysoPS, we hypothesize that LysoPS acts on GPR34 in macrophages, activating downstream signaling to promote the expression of the PS receptor MERTK. This feedback loop enhances macrophage efferocytosis, facilitating the clearance of necrotic material and limiting inflammatory responses.
Furthermore, among the highly expressed genes in damage-associated macrophages, TREM2 has been extensively studied for its functional mechanisms. Previous research indicates that TREM2 activation generally exerts metabolic-regulating effects mainly promoting the lipid metabolism, resulting in the accumulation of lipids within the macrophages and the formation of foam-like macrophages.39,40,41 This mechanism can promote local fibrosis and the exhaustion of CD8+ T cells.42,43,44 Additionally, TREM2 can serve as a receptor for damage-associated molecules like phosphatidylserine, activating macrophage phagocytic function—a role similar to GPR34-mediated efferocytosis in our study.45 However, some studies have also demonstrated that TREM2 knockout promotes IL-1β secretion in macrophages, contributing to PDAC progression.46 Thus, the role of macrophage TREM2 in tumor immunity remains incompletely elucidated. We propose that TREM2, as a molecule regulating macrophage phagocytosis, operates within a complex immune signaling network. Early loss of pattern recognition receptors in macrophage activation may impair phagocytic and activation capacity, driving a pro-inflammatory state. In contrast, sustained efferocytosis in already activated macrophages may exhaust their phagocytic clearance and antigen presentation functions. Moreover, lysosomal digestion of phagocytosed material in the complex tumor microenvironment can lead to chronic cytokine production and immune exhaustion. This complexity underlies the diverse activation and polarization states of macrophages. Our data indicate that compared to TREM2, GPR34 is more highly expressed in mature and polarized macrophages and is strongly associated with surufatinib treatment. Therefore, targeting GPR34 may significantly mitigate the impact of damage-induced efferocytosis on macrophage function.
In contrast to GPR34⁺ DAMs, Mac_cl2 is characterized by NF-κB activation and secretion of inflammatory cytokines such as IL-1β. Previous studies have indicated that the impact of macrophage-derived inflammatory cytokines on the tumor immune microenvironment is dual-sided.47,48 On one hand, when macrophages mount an effective immune response, they can activate pathways like NF-κB to promote the secretion of chemokines such as CXCL9 and CXCL10, thereby enhancing CD8⁺ T cell activation.48 On the other hand, under conditions of immune tolerance, macrophages may produce chronic inflammatory factors like IL-1β, which can foster pancreatic cancer cell proliferation, metastasis, and immune suppression.48,49,50 Thus, the effects of inflammatory cytokine production by macrophages are highly complex and largely dependent on whether macrophages initiate an effective immune response. Furthermore, the types of inflammatory and chemotactic factors produced by macrophages are closely associated with upstream activating signals. For instance, TLR stimulation typically induces acute inflammatory cytokines such as IFN-γ,51 whereas CSF-1R activation by CSF-1 promotes a chronic inflammatory cytokine secretion profile, including IL-6.52 Our sequencing data revealed a significant reduction in the proportion of Mac_cl2 in non-responders (Fig. 1i). This phenomenon may be attributed to Surufatinib’s targeting of the CSF-1R pathway in macrophages, which potentially diminishes their differentiation toward an immunosuppressive phenotype. This would also explain the observed decrease in Mac_cl2 proportion among responders following treatment. Future studies should focus on elucidating the specific molecular mechanisms by which Surufatinib modulates macrophage polarization and cytokine secretion, which could clarify the role and regulatory pathways of macrophage-derived inflammatory factors in treatment response.
Efferocytosis, the process by which macrophages engulf and remove apoptotic cell debris, plays a crucial role in maintaining tissue homeostasis and limiting secondary inflammation.53 Impaired efferocytosis has been implicated in chronic inflammatory diseases.54 Emerging evidence suggests that efferocytosis also reprograms macrophage metabolism, as the degradation of apoptotic cells generates substantial metabolic byproducts—including amino acids, lipids, and nucleotides—that require efficient processing. For instance, in atherosclerosis, efferocytosis drives a metabolic shift toward cholesterol catabolism in foam cells.55,56 Our study demonstrated that GPR34 promotes immune suppression by downregulating MHC-I expression in macrophages, a process mediated through efferocytosis. Although the lipid metabolism pathway is significantly upregulated in GPR34+ macrophages, there is no direct evidence suggesting that metabolic alterations influence macrophage antigen presentation. Instead, recent studies have suggested that the functional state of macrophages following efferocytosis is influenced by the type of cells they ingest, and that the ingestion of apoptotic tumor cells promotes macrophage polarization toward an M2-like phenotype.57 Consistently, our in vivo data show that macrophage MHC-I downregulation correlates significantly with the uptake of fluorescently labeled apoptotic tumor cells.
Prior studies have shown that blocking TAM efferocytosis in a gallbladder cancer animal model enhance MHC-II expression on TAMs,58 while inhibition of macrophage efferocytosis may activate the STING pathway, increasing tumor immunogenicity and boosting antitumor immunity.59 Additionally, macrophage efferocytosis is known to promote lysosomal degradation of necrotic fragments via pathways, sustaining the efferocytosis process.60 Several studies have shown that activating macrophage efferocytosis can alter macrophage function and promote immune suppression through metabolic reprogramming and polarization pathways.61,62,63 Our findings align with this model, suggesting that tissue damage activates macrophage efferocytosis through the GPR34-MERTK pathway. To process engulfed cell debris, macrophages activate lysosomal gene expression, leading to MHC-I degradation via the lysosomal pathway. These findings are consistent with previous studies. Undoubtedly, GPR34 plays a central role as an immune-modulatory molecule in macrophages, sensing injury signals, activating efferocytosis, and regulating CD8+ T cell responses. Targeting GPR34 has also shown promising results in preclinical models. Therefore, GPR34 may serve as a critical target for future immunotherapy in PDAC.
CXCL16 is a chemokine secreted by macrophages, dendritic cells, and non-immune cells such as epithelial cells and fibroblasts.64,65 It acts on CD8⁺ T cells and NK cells, exerting typical chemotactic effects.66,67 Furthermore, CXCL16 can promote the exhaustion of CD8⁺ T cells and enhance the proliferation and invasion of tumor cells.66,68,69 Consequently, CXCL16 exhibits a dual role in immune regulation, and its function may vary depending on its cellular origin.64 We propose that this complexity might explain why anti-CXCL16 treatment did not demonstrate immune-activating effects in our mouse models. Our data suggest that, compared to the CXCL16 pathway, GPR34-mediated downregulation of MHC-I protein levels—leading to reduced CD8⁺ T cell activation and ultimately immune tolerance—represents a more fundamental mechanism underlying CD8⁺ T cell dysfunction. Our findings further validate that the loss of immunogenicity is a more critical determinant of immune response efficacy than immunoreactivity.
In summary, transcriptomic analysis of surgical specimens from patients treated with Surufatinib identifies GPR34+ macrophages as a distinct immunosuppressive subset associated with treatment resistance. Mechanistically, macrophage GPR34 responds to LysoPS, a damage-associated molecule, in tumor tissues. Through the PI3K-AKT pathway, macrophages secrete inflammatory cytokines such as CXCL16. LysoPS-GPR34 signaling further activates efferocytosis via the AKT-MERTK pathway. After efferocytosis activation, macrophages upregulate lysosomal genes, and MHC-I is degraded via the lysosomal pathway, impairing macrophage antigen presentation to T cells. Targeting GPR34 significantly enhances the antitumor response and improves the efficacy of Surufatinib in pancreatic cancer.
Methods
Clinical trial design overview and sample collection
The clinical trial design follows the standardized protocol outlined in the study “Efficacy and Safety of Surufatinib Combined with Gemcitabine and Albumin-bound Paclitaxel in the Peri-operative Treatment of Pancreatic Cancer” (NCT05908747, https://clinicaltrials.gov/study/NCT05908747).
Sample Collection and Processing: Peripheral blood samples (5–10 mL) were collected using EDTA anticoagulant tubes and centrifuged within 2 h (1,500 × g, 10 min, 4 °C) to separate plasma and peripheral blood mononuclear cells (PBMCs), which were then stored at −80 °C and in liquid nitrogen, respectively. Tumor tissue samples were obtained through surgical biopsy. Freshly resected tumor tissue was digested into a single-cell suspension. Flow cytometry was employed to analyze immune cell composition using the following antibody panels: Myeloid panel: CD45, CD11b, F4/80, CD68, GPR34, MHC-I, MHC-II, CD80, CD86, MRC1 (CD206), MerTK, AXL. Lymphocyte panel: CD45, CD3ε, CD4, CD8α, CD19, NK1.1, Tim-3, PD-1, LAG-3, CTLA-4, GZMB, Ki-67, FOXP3. The gating strategy for identifying cell populations is detailed in Supplementary Fig. 7. Data were acquired on a BD LSRFortessa and analyzed with FlowJo v.10.
scRNA sequencing of patient surgical samples
Fresh tumor samples (~50-100 mg) were dissected into <1 mm³ pieces and incubated with 0.125 mg/mL collagenase IV(Sigma), 0.125 mg/mL Dispase II (Sigma) and Soybean Trypsin Inhibitor (MCE) at 37 °C for 2 h for tissue digestion. Subsequently, cells were filtered through a 70 μm mesh filter and collected. To remove dead cells, SYTOX Blue (Thermo Fisher, Cat.No.S7020) staining was applied to ensure only viable cells were used in subsequent analyses. Single-cell libraries were constructed using the 10x Genomics Chromium Single Cell 3’ GEM, Library & Gel Bead Kit v3.1 (10x Genomics, PN-1000121) targeting 10,000 cells per sample. Sequencing was performed on an Illumina NovaSeq 6000 platform to a median depth of 50,000 reads per cell. Raw sequencing data were initially processed with CellRanger v3.0.2 (10x Genomics) and aligned to the human genome (GRCh38). Downstream analysis was performed using the Seurat package (v5.0.1) in R. for normalization, dimensionality reduction, and clustering. Genes detected in fewer than 20 cells were excluded, and mitochondrial and ribosomal genes were removed. Cells of low quality, defined as having fewer than 200 or more than 6000 genes, or more than 10% mitochondrial gene content, were also filtered out.
scRNA sequencing data underwent quality control (retaining cells with mitochondrial gene percentage < 20% and detected gene counts > 200 and < 5,000), followed by selection of the top 2,000 most highly variable genes for data normalization. Principal component analysis (PCA) was performed using the highly variable gene expression matrix, with the top 30 principal components used for subsequent nonlinear dimensionality reduction and graph-based clustering. Cell clustering was based on shared nearest neighbor graphs (SNN, k. param = 20) and modularity optimization. After batch effect correction, to ensure data quality, the “decontX” function from the celda (v1.26.0) package was used to remove potential contamination from unwanted cell populations. Additionally, to eliminate the influence of homologous doublets, DoubletFinder (v2.0.3) was employed to detect and remove any identified doublets. After testing resolution parameters (0.2–1.2), a resolution of 1 was selected to balance biological rationality and computational efficiency. Cluster results were validated through UMAP visualization, showing good separation between clusters. Differential expression analysis was performed using the Wilcoxon rank-sum test with thresholds of log2FC > 0.5 and adjusted P-value < 0.01. Dimensionality reduction based on the Harmony results was followed by cell clustering using the Seurat functions FindNeighbors and FindClusters. Following clustering, cell type annotation was performed by cross-referencing cluster-specific differentially expressed genes (Wilcoxon rank-sum test, log₂FC > 0.5, adjusted p-value < 0.01) with canonical marker genes: KRT8, KRT18, KRT19 (epithelial cells); CD3D, CD3E, (T cells); CD79A, MS4A1 (B cells); JCHAIN, CD79A (plasma cell); CD14, HLA-DRA, APOE (monocytes/macrophages); FCGR3B (neutrophils); DCN, LUM (fibroblast); CDH5, VWF, PLAVP (endothelial cells); RGS5, PDGFRB (pericyte); CPA3, TPSAB1 (mast cell). A dot plot containing the expression of marker genes for all clusters is provided in Supplementary Fig. 1e. Functional characteristics of the cell types were further validated by their gene expression profiles. To resolve functional heterogeneity within key immune compartments, we performed secondary subclustering and analysis on extracted T cells and macrophages. For each resulting subpopulation, functional annotation was further informed by Gene Ontology (GO) Biological Process and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of their respective marker genes, which corroborated and refined the identity assigned by canonical markers. Characteristic genes for each cell subtype are listed in Table S1.
Gene Set Enrichment Analysis (GSEA) was conducted using the clusterProfiler (v4.10.0) package. Differentially expressed genes were extracted from the differential expression analysis, ranked by log2 fold change, and enriched against Hallmarks, KEGG, and GO gene sets. Enrichment results were visualized using enrichplot, with bar plots showing significantly enriched gene sets and their associated biological functions.
Data were first reduced in dimensionality and clustered based on Seurat clustering results. Pseudotime analysis was then performed using reduce Dimension function (with the DDRTree method) of Monocle 3 (v 0.2.3) followed by orderCells to arrange cells along a differentiation trajectory. The pseudotime analysis results were visualized using UMAP, which displays the position and trajectory of cells during differentiation. The dynamic tracking of cells through different stages of differentiation was used to analyze their differentiation trends.
Intercellular communication analysis was performed using the CellChat (v1.6.1) package. The computeCommunProb function was utilized to calculate the communication probabilities between cell populations, and the resulting signaling pathways were visualized using the netVisual_aggregate function. Based on cell type annotations, further analysis was conducted to explore the interactions and communication networks between different macrophage subpopulations and T cell subpopulations.
Animal Models
Macrophage Gpr34 conditional knockout mouse model (Gpr34ΔLyz2) was constructed by Gpr34flox/flox (Shanghai Model Organisms Center, Inc. Cat.No.NM-CKO-233912) and Lyz2-CreERT mice (B6.129P2-Lyz2tm1(cre)Ifo/J, Jackson Laboratories) were generated by crossing. Cre recombinase is specifically expressed in Lyz2-expressing monocyte-macrophages and induces Gpr34 gene knockout. Three weeks after modeling, tamoxifen (Sigma-Aldrich, Cat. No. T5648) was given to activate Cre recombinase and induce Cre gene deletion. Tamoxifen was dissolved in corn oil (Solarbio, Cat.No.C7030) at a concentration of 20 mg/mL and administered by intraperitoneal injection in a volume of 100 μL/20 g mouse body weight daily for 5 days to ensure effective induction of Cre recombinase activity in macrophages. After the model was established, efficient deletion (>90% reduction in GPR34 protein) in F4/80⁺ tumor-associated macrophages was confirmed by flow cytometry 10 days after the last tamoxifen injection (Supplementary Fig. 8c). Flow cytometry analysis showed that Gpr34 expression was significantly down-regulated in macrophages, confirming the successful construction of Gpr34 CKO model.
Gpr34 global knockout (Gpr34−/−) mice were generated by crossing Gpr34flox/flox mice with Dppa3-Cre mice (C57BL/6Smoc-Dppa3em1(IRES-Cre) Smoc, Shanghai Model Organisms Center, Inc. Cat. No. NM-KI-00040). The Dppa3-Cre transgene activates Cre recombinase in fertilized eggs, enabling global deletion of the Gpr34 gene. Gpr34 gene knockout was detected by next-generation sequencing, and changes in immune cell population were detected by flow cytometry, which confirmed the loss of Gpr34 in immune cells.
KPC spontaneous model: C57BL/6Smoc-Trp53em4(R172H) Krasem4(LSL-G12D) Tg (Pdx1-Cre)Smoc (KrasG12D/+; Trp53R172H/+; Pdx1-Cre, Shanghai Model Organisms Center, Inc. Cat.No.NM-KI-210096) spontaneously developed into pancreatic cancer in a natural background. In these mice, transformation of pancreatic tissue occurs naturally and tumors develop without external intervention. Tumor growth and volume changes were monitored by routine imaging using high-frequency ultrasound (Vevo 2100, VisualSonics). Tumor size and growth trends in the pancreatic region were assessed by ultrasound at early stages of tumor formation. Tumor size was monitored every 7 days to evaluate its growth rate and morphological characteristics. Ultrasound images were processed using VevoLAB software, and tumor volume was calculated using the formula V = 0.5 * (length) * (width2).
OT-1 Mice: C57BL/6-Tg (TcraTcrb)1100Mjb/J mice (Jackson Laboratories, Stock No: 003831) were used as a source of OVA257-264 (SIINFEKL)-specific CD8⁺ T cells.
Mouse CD8+ T cells/ NK cells were depleted using anti-mouse anti-CD8α (Bioxcell, Cat.No.BE0061)/anti-mouse NK1.1 antibody (Bioxcell, Cat.No.BE0036). First, mice were given an intraperitoneal injection of 200 μg. Depletion effects were assessed by looking at surface markers associated with CD8+ T cells /NK cells (Supplementary Fig. 10b, c). After injection, blood and spleen cells were analyzed by flow cytometry to ensure complete depletion of CD8+ T/NK cells. Subsequently, 200 μg/dose was maintained every 5 days. Treatment usually begins one week after intratumoral injection in mice, or when the tumor in KPC mice reaches 150 mm3. The dosage is as follows: Gemcitabine (Selleck, CAT.NO. S4419) 25 mg/kg, twice a week; Nab-paclitaxel (Selleck, E1068, CAT.NO. S4419),10 mg/kg, i.p., once a week; Surufatinib: 50 mg/kg, once every other day, i.p., for a total of 3 times. Anti-CXCL16 (InVivoMAb anti-mouse CXCL16 BioXcell, CAT.NO.BE0450) 5 mg/kg, i.p., 3 times a week. GPR34 antagonist (Compound D2, MCE, CAT. NO. HY-138501), 25 mg/kg, i.p., daily. The treatment cycle for the in-situ injection tumor model is 3 weeks, and the treatment duration for KPC mice is 8 weeks.
Establishment of the Mouse Model of Pancreatitis. Age-matched (8–10 weeks) Gpr34flox/flox or Gpr34ΔLyz2mice pre-treated with tamoxifen as described, received 12 hourly i.p. injections of cerulein (50 µg/kg, Selleck, S9690). Mice were euthanized 48 hours after the first injection for tissue collection and analysis.
Cell line and macrophages isolation
Generation of the KPC Cell Line. KPC mice (KrasG12D/+; Trp53R176H/+; Pdx1-Cre) aged 8–10 weeks were used for tumor isolation. When tumor volume reached approximately 200–300 mm³, tumors were excised under sterile conditions using aseptic techniques. Excised tumors were transferred to sterile glass culture flasks and finely minced into 1–2 mm³ tissue fragments. These fragments were then enzymatically digested in a solution containing collagenase IV (2 mg/mL, Sigma, Cat.No.9001-12-1), soybean trypsin inhibitor (MCE, Cat. No. HY-126388), and DNase I (50 µg/mL, SparkJade, Cat.No.AC1711) at 37 °C for 30 min. The cell suspension was filtered, washed, and plated in DMEM (Gibco, C11995500BT) with 10% FBS (Gibco) and 1% Penicillin/Streptomycin (Solarbio, P7630). After five passages to enrich for epithelial cells, adherent cells were collected.
The established cell lines (designated KPC-0117, used throughout this study) were validated by: a) Morphology: Spindle-shaped epithelial morphology under phase-contrast microscopy. b) Immunophenotype: >95% positive for the epithelial marker CK19 (Cytofix/Cytoperm kit, BD, 554714) by flow cytometry. c) Genotype: Sanger sequencing confirmed the presence of the KrasG12D and Trp53R172H mutations. d) Pathogen Testing: Confirmed negative for mycoplasma (MycoAlert PLUS, Lonza, LT07-710).
For imaging, cells were transduced with lentivirus carrying GFP or firefly luciferase (Luc). For antigen presentation assays, cells were transduced with lentivirus encoding ovalbumin (OVA). Stable pools were selected with puromycin (2 µg/mL).
Isolation and Differentiation of Bone Marrow-Derived Macrophages (BMDMs). BMDMs were isolated from the femur and tibia of 6-week-old C57BL/6, Gpr34−/−, and OT-1 mice following euthanasia. The metaphysis of the bones was removed under sterile conditions, and the bone marrow was flushed out using a 1 mL sterile syringe containing PBS with 2% FBS. The collected bone marrow cells were subjected to erythrocyte lysis and then resuspended in DMEM supplemented with 10% FBS and 1% penicillin/streptomycin. Cells were seeded into 10 cm² culture dishes and differentiated into macrophages by adding 20 ng/mL monocyte colony-stimulating factor (M-CSF, Proteintech, Cat.No.003831). After five days of culture, non-adherent cells were removed, and the adherent BMDMs were maintained in culture. Prior to co-culture experiments, BMDMs were stimulated with tumor-conditioned medium (TCM) derived from KPC cells for 24 h to mimic the tumor microenvironment and activate macrophages.
Preparation of Tumor-Conditioned Medium (TCM). KPC cells were grown to 70-80% confluence in T75 flasks, washed with PBS, and then cultured in serum-free DMEM for 48 h. The supernatant was collected, centrifuged at 2000 × g for 10 min, filtered through a 0.22 µm filter, and stored at −80 °C. This TCM was used at a 1:1 dilution with fresh complete medium for macrophage stimulation.
Human Cell Lines and Monocyte-Derived Macrophages (hMDMs). Human PDAC cell lines (Panc-1, BxPC-3, AsPC-1) were obtained from the Chinese Academy of Sciences Cell Bank and cultured per supplier instructions. hMDM were isolated from PBMCs obtained from healthy donors. CD14⁺ monocytes were purified using CD14 MicroBeads (Miltenyi Biotec, Cat.No.130-050-201) according to the manufacturer’s protocol. Briefly, PBMCs were incubated with CD14 antibody-conjugated magnetic beads for 20 min at 4 °C with gentle mixing. CD14⁺ cells were then separated from unlabeled cells using magnetic separation (BioLegend, Cat.No.480173). Following isolation, CD14⁺ cells were washed twice with PBS to remove unbound magnetic beads and contaminating cells. The purity of CD14⁺ monocytes was assessed by flow cytometry. Purified CD14⁺ cells were then cultured in RPMI-1640 medium (Gibco, Cat.No. C11875500BT) supplemented with human M-CSF (20 ng/mL, Proteintech, Cat.No. 300-25) and 10% FBS. The culture medium was replaced every two days, and after five days of differentiation, macrophages were used for downstream experiments.
Orthotopic Tumor Model Construction and In vivo Imaging
Pancreatic cancer cells (KPC cells) from KPC mice were transduced with lentivirus carrying luciferase (Luc) plasmids. These stably transduced KPC-Luc cells were then orthotopically injected into the pancreas of recipient mice (Gpr34ΔLyz2 and C57BL/6 strains) to establish an in situ tumor model. In the anesthetized mice, a longitudinal incision was made in the left upper abdominal area to expose the spleen and exteriorize the pancreas. KPC-Luc cells were injected into the pancreas parenchyma (40 μL, 1 × 10⁵ cells). After the surgery, the mice were monitored for 7–10 days, and when the tumors bioluminescence reached approximately 105–106, imaging was performed to monitor tumor growth and metastasis.
Monitoring tumor growth and metastasis: The IVIS Spectrum in vivo imaging system (PerkinElmer) was used for bioluminescence imaging to assess tumor size and location. Mice injected with KPC-Luc cells were given intraperitoneal injections of D-luciferin (150 mg/kg body weight) as a substrate. After 10 min, images were captured using the IVIS Spectrum system. Tumor growth dynamics were quantified by measuring the bioluminescence intensity within the tumor region. During the experiment, tumor growth and the overall health status of the mice were monitored and recorded.
Total body irradiation and bone marrow reconstitution experiments
Total body irradiation was used to ablate blood cells such as hematopoietic stem cells (HSCs) from the bone marrow of mice. The radiation dose was 10 Gy was used to ensure effective destruction of the hematopoietic system. Following irradiation, mice were maintained under standard conditions for 24 hours to allow for the full effect of irradiation on bone marrow depletion. Bone marrow was collected as described previously. The harvested cells were treated with red blood cell lysis buffer (Servicebio, Cat. No. R1010) to remove red blood cells. The cells were then resuspended in DMEM medium and cultured at 37 °C in the presence of 5% CO₂. After bone marrow clearance, Total bone marrow cells were prepared for transplantation. Total bone marrow cells derived from the donor mice were transplanted into recipient mice via tail vein injection. Each recipient mouse received approximately 5 × 106 total bone marrow cells which were resuspended in 200 µL PBS and injected slowly through the tail vein. After injection, mice were placed under standard rearing conditions for recovery. Bone marrow reconstitution was assessed approximately 2 weeks after transplantation. Staining was performed with an anti-mouse CD45 antibody (BD Biosciences, Cat.No.566439) and flow cytometry were used to detect the proportion of CD45+ cells and their immunophenotype in peripheral blood to confirm whether the donor hematopoietic cells were successfully reconstructed. Approximately 2 weeks after transplantation, bone marrow reconstitution was assessed. The peripheral blood was analyzed using flow cytometry to detect the proportion of CD45+ cells and their immune phenotype, confirming whether there was successful reconstitution of hematopoietic cells from the donor.
Tissue Dissociation and Single Cell Suspension Preparation Tissue
Dissociation: Tumors were excised and placed in medium containing PBS and 1% penicillin/streptomycin. The tissue was minced and passed through a 70 µm filter to remove debris to form a single cell suspension. The tissue suspension was then incubated in a digestive enzyme solution containing 0.1% soy trypsin inhibitor and 0.125 mg/mL collagenase IV for 30 min at 37 °C to further digest the tissue and obtain single-cell populations. The digestion was terminated by adding RPMI-1640 medium, and the undigested tissue blocks were removed by centrifugation to obtain the final single-cell suspension.
Preparation of single cell suspensions: Cell suspensions were filtered and centrifuged and washed with PBS. Cell concentrations were determined using a hemocytometer and adjusted as needed to the desired cell density. The cells were resuspended in serum-free RPMI-1640 medium.
Flow Cytometry Analysis
The single cell suspension was resuspended in MACS buffer (1% BSA, 2 mM EDTA, PBS). If it is for detecting the secreted proteins of T cells from PBMCs (such as IFN-γ), the medium should also contain phorbol myristate acetate (PMA, 20 ng/mL; Selleck, Cat. No. S7791) / Ionomycin (1 μg/mL; Selleck, Cat.No.S7074) and Golgi blocker Brefeldin A (50 ng/mL; Selleck, Cat.No.S7046) and incubated for 4 hours at 37 °C with 5% CO₂. To block Fc receptor binding, cells were incubated with anti-mouse CD16/CD32 antibody (BD Biosciences, Cat.No.553141) for 10 minutes on ice. Then, according to experimental requirements, cells were then stained with antibodies against specific surface markers (see in Table S2), followed by two washes to remove unbound antibodies. For analysis of secreted factors and transcription factors, we used a Cytoperm Fixation/Permeabilization Kit (BD Biosciences, Cat.No.554714), and cells were permeabilized and stained according to the manufacturer’s instructions. Subsequently, according to experimental requirements, cells were stained with antibodies against specific transcription factors or intracellular factors (see in Table S2). Finally, the samples were fixed with 1% paraformaldehyde (PFA, Servicebio, Cat. No. G1101). Flow cytometry data were acquired using BD LSRFortessa (BD Biosciences) equipment, and data were processed using FACSDiva software (BD Biosciences). FlowJo v.10.8.1 software was used for data analysis and cell populations were classified according to the fluorescence intensity of specific laser channels. For flow cytometry analysis, appropriate gating strategies were used to exclude debris and doublets. In the analysis of fibroblasts, we excluded EPCAM+, CD31+, and CD45+ cell subsets, gated the Lineage-negative (Lin-) cell population, and used PDPN for positive gating to identify fibroblasts. Myeloid cells were identified by gating on CD45+CD11b+ cells, and lymphocytes were identified by gating on CD45+CD3ε+CD8α+ cells. In specific analyses, we further quantified the immunophenotype, activation status, and cytokine secretion levels of the cell populations by analyzing the corresponding markers.
Macrophage Conditioned Medium (MCM) Preparation
Macrophage-conditioned medium (MCM) was prepared by culturing macrophages under the desired experimental conditions. After the cells were cultured, the medium was replaced, and the macrophages were incubated for an additional 48 hours. The collected medium was filtered through a 0.45 µm filter to remove cell debris, resulting in the macrophage-conditioned medium (MCM). This MCM was used for subsequent co-culture experiments or downstream analyses. To separate MCM from small molecular materials, the supernatant was added to the upper chamber of a 3 kDa molecular weight cutoff protein concentrator (MilliporeSigma, Cat.No.ACK5003PA). The concentrator was placed at 4 °C and centrifuged at 5,000 × g for 1 hour. After centrifugation, approximately 80–90% of the solution was transferred from the upper chamber to the lower chamber. Larger proteins (>3 kDa) were retained in the upper chamber, while small molecules (<3 kDa) were collected in the lower chamber. The concentrated MCM ( > 3 kDa) from the upper chamber was diluted to its original volume with fresh serum-free RPMI 1640 medium, while 10% FBS was added to the MCM ( < 3 kDa) collected in the lower chamber. Both fractions were then used for subsequent experiments as required.
Co-culture of BMDMs and T Cells
BMDMs were pre-stimulated with TCM (50% v/v) for 24 hours. For antigen-specific assays, BMDMs were pulsed with 1 µg/mL SIINFEKL peptide (MCE, HY-P1489) for 2 hours. CD8⁺ T cells were isolated from spleens of OT-1 or C57BL/6 mice using the human CD8+ T Cell Isolation Kit (Negative Isolation) (Miltenyi, Cat. No. 130-104-075). BMDMs and T cells were co-cultured at a 1:5 ratio in RPMI-1640 with 10% FBS. For polyclonal T cell activation, plates were pre-coated with Dynabeads™ mouse T activator CD3/CD28 for T cell activation/expansion kit (Thermo Fisher Scientific, Cat. No. 11452D) according to the instructions provided by the manufacturer. After 48–72 h, cells were harvested for flow cytometric analysis of activation/exhaustion markers and cytokines.
For BMDM and OT-1 CD8+ T cell co-culture, BMDMs from OT-1 mice (8 weeks) were isolated and cultured according to standard methods, and the stimulation reached No. At 5 days, BMDMs were stimulated with OVA peptide (257-264) (SIINFEKL, 1 μg/mL, MCE, HY-P1489) added to TCM for 24 h to induce antigen presentation. After stimulation, medium was replaced. BMDMs were co-cultured with OT-1 mouse-derived CD8+ T cells at a ratio of 1:5 (BMDM:CD8+ T cells) for 24 h. After co-culture, cells were harvested and subjected to surface staining. First, CD8+ T cells were labeled with anti-CD8α antibodies. Then, Tetramer staining was performed using PE-H-2Kb/OVA (SIINFEKL) MHC Tetramer (Creative Biolabs) following the manufacturer’s protocol. For BMDM, Anti-Mouse H-2Kb/SIINFEKL antibody (BD Pharmingen™, 569791) and other indicator antibodies were added. Staining reactions were performed for 30 min on ice. After staining, cells were washed and resuspended in flow cytometry buffer and tested by flow cytometry.
Organoid Construction and In Vitro PBMC Co-culture
Human PDAC tissues were acquired from patients undergoing surgery for pancreatic cancer who consented to donate their samples for research. Briefly, fresh PDAC tissues were collected during surgery operation and immediately washed three times with PBS to remove blood clots, dead tissues and other connective tissues. PDAC tissues were minced and digested with a mix of 5 mL enzymes buffer containing 1 mg/mL collagenase (Sigma-Aldrich, Cat.No.C2799), 2.5 U/mL hyaluronidase (Sigma-Aldrich, Cat.No.H3506) and 0.1 mg/mL DNase (Sigma-Aldrich, Cat. No. DN25) in 37 °C water bath for 15 min. The mixture was then filtered through a 40 μm strainer to obtain single cell suspension. The single cells were embedded in Growth Factor reduced (GFR) Matrigel (Corning, Cat.No.356231) and cultured in complete medium containing Advanced DMEM/F-12 (Gibco, Cat.No.12634-010), 10 mM HEPES pH=7.2–7.5 (Solarbio, Cat.No.H1080), 1 × GlutaMAX Supplement (Invitrogen, Cat.No.35050061), 100 µg/mL Primocin (Invivogen, Cat.No. ant-pm-2), 10 mM Nicotinamide (Sigma-Aldrich, Cat.No.N0636), 1 × Wnt3a-Conditioned Medium, 1 × R-spondin1-Conditioned Medium, 100 ng/mL mNoggin (R&D, Cat.No. 6997-NG-025), 1 × B27 Supplement (Thermo Fisher Scientific, Cat.No.17504044), 100 ng/mL hFGF-10 (Peprotech, Cat.No.100-26), 1.25 mM N-acetylcysteine (Sigma-Aldrich, Cat.No.A9165), 1 µM Prostaglandin E2 (R&D, Cat.No.2296/10), 50 ng/mL hEGF (Peprotech, Cat.No.AF-100-15), 10 µM Rho Kinase inhibitor Y27632 (Sigma-Aldrich), 10 nM hGastrin Ⅰ (Sigma-Aldrich, Cat.No.G9145) and 500 nM A 83-01(R&D, Cat.No.2939-10). PDAC organoids (3 × 103 cells/well) were seeded in 96-well plates. The medium was changed approximately every 3 days and organoids were passaged approximately every 7 days according to their growth conditions.
PBMCs were isolated from the peripheral blood of healthy donor. PBMCs were co-cultured with organoids at a ratio of 5:1 (PBMCs: organoid). CD8+ T cells were enriched from PBMCs using magnetic bead-based isolation to ensure sufficient immune cells in the co-culture system. To activate T cells, Dynabeads Human T-Activator (Gibco, Cat.No.11163) was added to the medium. During co-culture, IL-2 (20 IU/mL, PeproTech, Cat.No.200-02) was added as a stimulant, along with drugs and other reagents. After 36–48 hours of co-culture, organoid morphology, proliferation, and immune responses of the cells were evaluated by immunohistochemistry and flow cytometry. Apoptosis levels were assessed using the caspase 3/7 fluorescence probe (Beyotime, Cat.No.C1073S), and images were captured using both optical and fluorescence microscopy for statistical analysis.
Multi-Immunofluorescence (mIF) Staining of Paraffin and Frozen Sections
Formalin-fixed, paraffin-embedded (FFPE) tissue sections (4 µm) were deparaffinized, rehydrated, and subjected to antigen retrieval in EDTA buffer (pH 9.0) using a pressure cooker. Autofluorescence was quenched with TrueVIEW Autofluorescence Quenching Kit (Vector Labs, SP-8400). Sections were blocked and incubated with primary antibodies overnight at 4 °C (antibody details in Supplementary Table S2). Opal™ 7-Color Manual IHC Kit (Akoya Biosciences, NEL811001KT) was used for tyramide signal amplification with fluorophores Opal 520, 570, 620, 690, and 780. Nuclei were counterstained with DAPI. Slides were scanned using a Vectra Polaris imaging system (Akoya Biosciences) at 20x magnification. Image analysis and cell segmentation were performed using inForm® software (Akoya Biosciences) and QuPath.
Enzyme-Linked Immunosorbent Assay (ELISA)
CXCL16 levels in cell culture supernatants were measured using the Mouse CXCL16 DuoSet ELISA (Biotechwell, Cat. No. EM30142) according to the manufacturer’s instructions. Absorbance was read at 450 nm with correction at 570 nm on a BioTek Synergy MX plate reader.
pHrodo Red Dye and GFP Uptake Assay for Macrophage Phagocytosis Activity
Prior to the experiment, the pHrodo Red AM Ester (Thermo Fisher Scientific, Cat. No. P36600) was equilibrated to room temperature. The working staining solution was prepared as follows: 10 μL of pHrodo Red AM Ester is added to 100 μL of PowerLoad concentrate (provided with the kit), mixed thoroughly, and then diluted with 10 mL of PBS. On day 5 of culture, BMDMs from different mouse strains were treated with TCM and co-cultured with chemotherapy-induced apoptotic tumor cells for 12 h. After removal of apoptotic cells and culture medium, the cells were washed once with PBS. The pHrodo Red AM Ester staining solution is then added, and the cells are incubated at 37 °C with 5% CO₂ for 30 minutes. After incubation, cells are washed with PBS to remove any unbound dye. Stained cells are observed for phagocytosis activity using an integrated fluorescence microscopy imaging system (Ex/Em: 560/585 nm), or the fluorescence signal is quantitatively measured by flow cytometry.
BMDMs were co-cultured with apoptotic KPC-GFP cells (effector: target = 1:5) for 4 hours. After extensive washing to remove non-ingested cells, macrophages were analyzed by flow cytometry for GFP signal and by immunofluorescence microscopy to visualize internalized GFP⁺ material.
Immunofluorescence
Cell coverslips were first immersed in 75% ethanol and washed with PBS, then placed into a 24-well plate, with approximately 2 × 10⁴ cells seeded per well. Cells were cultured for 24 hours. The following day, cells were washed with PBS 5–10 minutes × 3, fixed with 4% paraformaldehyde for 10 minutes, and washed again with PBS 5–10 min × 3. If permeabilization is required, cells were treated with 0.1% Triton X-100 (Solarbio, Cat. No. T8200) for 10 minutes, followed by PBS washing 5–10 min × 3. Then, cells were blocked with 3% BSA for 1 hour at room temperature and washed once with PBS for 10 minutes. Primary antibodies were incubated overnight at 4 °C, protected from light, at a dilution of 1:200 in a humidified chamber. On the third day, cells were brought to room temperature for 1 hour, washed with PBS 5–10 minutes × 3, and then incubated with secondary antibody (1:400, 30 minutes, Thermo fisher scientific, A-11008/ A-11001/ A-21070) protected from light. Cells are washed with PBS 10 minutes × 3. Finally, 20 μL of mounting medium containing DAPI was added to each coverslip. The slides were allowed to dry in the dark before being stored at 4 °C. Images were captured using a fluorescence microscope.
Small interfering RNA (siRNA) transient transfection
After hMDM or BMDM was stimulated with h-mcsf or m-CSF (20 ng/mL) until day 5, siRNA transient transfection was performed. First, siRNA (Shanghai Sangon Inc, see in Table S3) was mixed with Advanced DNA RNA Transfection Reagent (zeta-life, Cat.No.AD600150) in a 1:1 ratio and incubated for 20 minutes at room temperature. The mixture was then added to the macrophage culture medium at a 1:200 ratio (transfection reagent to medium). Transfection was terminated by changing the medium 24 hours after transfection. The transfected cells were used for subsequent experiments.
Cellular RNA extraction and Quantitative real-time PCR (qRT-PCR)
Total RNA was extracted from cultured cells using TRIzol reagent (Epizyme, Cat. No. YY101) according to the manufacturer’s instructions. Briefly, cells were collected from culture dishes and gently washed with PBS to remove residual medium. The cells were then lysed in TRIzol reagent to ensure complete disruption and homogenization, with RNA stabilizers included in the lysate to prevent degradation. Chloroform was added to the lysate, followed by thorough mixing and centrifugation at 12,000 × g for 15 minutes at 4 °C to achieve phase separation. The aqueous phase containing RNA was carefully transferred to a fresh RNase-free tube, and an equal volume of isopropanol was added to facilitate RNA precipitation. After incubation at 4 °C for 10 minutes, the RNA was pelleted by centrifugation at 12,000 × g for 10 minutes at 4 °C. The resulting pellet was washed with 75% ethanol, centrifuged, and air-dried before being dissolved in RNase-free water. The concentration and purity of the extracted RNA were assessed using a spectrophotometer.
For cDNA synthesis, a StarScript III All-in-one RT Mix with gDNA Remover (GenStar, Cat.No.A240) was used according to the manufacturer’s protocol. A defined amount of total RNA was reverse-transcribed into cDNA, and inactivation was performed at the end of the reaction. qRT-PCR was performed using a one-step qRT-PCR detection kit (Genstar, Cat. No. A301-10) in a reaction system containing the cDNA template, SYBR Green fluorescent dye, specific primers (see in Supplementary table 2), and qPCR Master Mix. The PCR program consisted of an initial denaturation at 95 °C for 5 minutes, followed by 40 cycles of denaturation at 95 °C for 15 s, annealing at 60 °C for 30 s, and extension at 72 °C for 30 s. The relative expression levels of target genes were analyzed using the ΔΔCt method, with β-actin serving as an internal reference.
Statistical Analysis
Data from at least three independent samples or biological replicates are presented as mean ± standard deviation. Two-tailed Student’s t test was used to compare the data between the two groups. One-way analysis of variance (ANOVA) followed by post-hoc Tukey’s or Dunnett’s tests, or the Kruskal-Wallis H test was used to test P-values for data comparison between more than two groups. For Kaplan-Meier survival analysis, P-values were calculated using the Log-rank (Mantel-Cox) test and multivariate COX regression was used to analyze the influence of various clinical factors to patients’ survival. Use the β test (β regression) to analyze the correlation between the cell proportions in scRNA sequencing data and the clinical treatment effect. Sequencing data were compared relative gene expression using a two-sided Wilcoxon test. *P < 0.05, **P < 0.01, ***P < 0.001.
Data availability
All data are accessible in NODE (https://www.biosino.org/node) with the accession number OEP00006844 or through the URL: https://www.biosino.org/node/project/detail/OEP00006844. External data were obtained from GEO database and GSA database, including GSE155698, GSE205013 and CRA001160.
References
Siegel, R. L., Kratzer, T. B., Giaquinto, A. N., Sung, H. & Jemal, A. Cancer statistics, 2025. CA-Cancer J. Clin. 75, 10–45 (2025).
Polak, R., Zhang, E. T. & Kuo, C. J. Cancer organoids 2.0: modelling the complexity of the tumour immune microenvironment. Nat. Rev. Cancer 24, 523–539 (2024).
Yasuda, T. & Wang, Y. A. Gastric cancer immunosuppressive microenvironment heterogeneity: implications for therapy development. Trends Cancer 10, 627–642 (2024).
Falcomatà, C. et al. Context-Specific Determinants of the Immunosuppressive Tumor Microenvironment in Pancreatic Cancer. Cancer Discov. 13, 278–297 (2023).
Mao, X. et al. Crosstalk between cancer-associated fibroblasts and immune cells in the tumor microenvironment: new findings and future perspectives. Mol. Cancer 20, 131 (2021).
Rebelo, R., Xavier, C. P. R., Giovannetti, E. & Vasconcelos, M. H. Fibroblasts in pancreatic cancer: molecular and clinical perspectives. Trends Mol. Med 29, 439–453 (2023).
Mantovani, A. et al. Macrophage diversity in cancer dissemination and metastasis. Cell. Mol. Immunol. 21, 1201–1214 (2024).
Vázquez-Bellón, N., Martínez-Bosch, N., García de Frutos, P. & Navarro, P. Hallmarks of pancreatic cancer: spotlight on TAM receptors. EBioMedicine 107, 105278 (2024).
Wang, Q., Wang, J., Xu, K. & Luo, Z. Targeting the CSF1/CSF1R signaling pathway: an innovative strategy for ultrasound combined with macrophage exhaustion in pancreatic cancer therapy. Front. Immunol. 15, 1481247 (2024).
Ho, W. J. et al. Macrophage-Targeting by CSF1/1R Blockade in Pancreatic Cancers. Cancer Res 81, 6071–6073 (2021).
Cannarile, M. A. et al. Colony-stimulating factor 1 receptor (CSF1R) inhibitors in cancer therapy. J. Immunother. Cancer 5, 53 (2017).
Syed, Y. Y. Surufatinib: First Approval. Drugs 81, 727–732 (2021).
Ali, M. A. et al. Efficacy and toxicity of surufatinib in neuroendocrine tumors: A systematic review and meta-analysis. J. Neuroendocrinol. 34, e13149 (2022).
Xu, J. et al. Current treatments and future potential of surufatinib in neuroendocrine tumors (NETs). Ther. Adv. Med. Oncol. 13, 17588359211042689 (2021).
Xu, J. et al. Surufatinib in advanced extrapancreatic neuroendocrine tumours (SANET-ep): a randomised, double-blind, placebo-controlled, phase 3 study. Lancet Oncol. 21, 1500–1512 (2020).
Harris, M. A. et al. Towards targeting the breast cancer immune microenvironment. Nat. Rev. Cancer 24, 554–577 (2024).
Natu, J. & Nagaraju, G. P. Gemcitabine effects on tumor microenvironment of pancreatic ductal adenocarcinoma: Special focus on resistance mechanisms and metronomic therapies. Cancer Lett. 573, 216382 (2023).
Zhao Zhao, Y., Hasse, S. & Bourgoin, S. G. Phosphatidylserine-specific phospholipase A1: A friend or the devil in disguise. Prog. Lipid Res. 83, 101112 (2021).
Holmgaard, R. B. et al. Targeting myeloid-derived suppressor cells with colony stimulating factor-1 receptor blockade can reverse immune resistance to immunotherapy in indoleamine 2,3-dioxygenase-expressing tumors. EBioMedicine 6, 50–58 (2016).
Achkova, D. et al. Role of the colony-stimulating factor (CSF)/CSF-1 receptor axis in cancer. Biochem Soc. Trans. 44, 333–341 (2016).
Butowski, N. et al. Orally administered colony stimulating factor 1 receptor inhibitor PLX3397 in recurrent glioblastoma: an Ivy Foundation Early Phase Clinical Trials Consortium phase II study. Neuro Oncol. 18, 557–564 (2016).
Fermi, V. et al. Effective Reprogramming of Patient-Derived M2-Polarized Glioblastoma-Associated Microglia/Macrophages by Treatment with GW2580. Clin. Cancer Res 29, 4685–4697 (2023).
Pyonteck, S. M. et al. CSF-1R inhibition alters macrophage polarization and blocks glioma progression. Nat. Med. 19, 1264–1272 (2013).
Quail, D. F. et al. The tumor microenvironment underlies acquired resistance to CSF-1R inhibition in gliomas. Science 352, aad3018 (2016).
Fujiwara, T. et al. CSF1/CSF1R Signaling Inhibitor Pexidartinib (PLX3397) Reprograms Tumor-Associated Macrophages and Stimulates T-cell Infiltration in the Sarcoma Microenvironment. Mol. Cancer Ther. 20, 1388–1399 (2021).
Lv, Q. et al. CSF1R inhibition reprograms tumor-associated macrophages to potentiate anti-PD-1 therapy efficacy against colorectal cancer. Pharm. Res 202, 107126 (2024).
Yin, M. et al. Potent BRD4 inhibitor suppresses cancer cell-macrophage interaction. Nat. Commun. 11, 1833 (2020).
Zhu, Y. et al. CSF1/CSF1R blockade reprograms tumor-infiltrating macrophages and improves response to T-cell checkpoint immunotherapy in pancreatic cancer models. Cancer Res 74, 5057–5069 (2024).
Schöneberg, T., Meister, J., Knierim, A. B. & Schulz, A. The G protein-coupled receptor GPR34 - The past 20 years of a grownup. Pharmacol. Ther. 189, 71–88 (2018).
Heinrich, F. et al. Morphologic, phenotypic, and transcriptomic characterization of classically and alternatively activated canine blood-derived macrophages in vitro. PLoS One 12, e0183572 (2017).
Izume, T. et al. Structural basis for lysophosphatidylserine recognition by GPR34. Nat. Commun. 15, 902 (2024).
Wang, X. et al. GPR34-mediated sensing of lysophosphatidylserine released by apoptotic neutrophils activates type 3 innate lymphoid cells to mediate tissue repair. Immunity 54, 1123–1136 (2021).
Yan, J. et al. GPR34 is a metabolic immune checkpoint for ILC1-mediated antitumor immunity. Nat. Immunol. 25, 2057–2067 (2024).
Bohlson, S. S. et al. Complement, c1q, and c1q-related molecules regulate macrophage polarization. Front. Immunol. 5, 402 (2014).
Revel, M., Sautès-Fridman, C., Fridman, W. H. & Roumenina, L. T. C1q+ macrophages: passengers or drivers of cancer progression. Trends Cancer 8, 517–526 (2022).
Zhang, S. et al. C1q(+) tumor-associated macrophages contribute to immunosuppression through fatty acid metabolic reprogramming in malignant pleural effusion. J. Immunother. Cancer 11, e007441 (2023).
Dong, L. et al. The loss of RNA N(6)-adenosine methyltransferase Mettl14 in tumor-associated macrophages promotes CD8(+) T cell dysfunction and tumor growth. Cancer Cell 39, 945–957 (2021).
Pulanco, M. C. et al. Complement Protein C1q Enhances Macrophage Foam Cell Survival and Efferocytosis. J. Immunol. 198, 472–480 (2017).
Jaitin, D. A. et al. Lipid-Associated Macrophages Control Metabolic Homeostasis in a Trem2-Dependent Manner. Cell 178, 686–698 (2019).
Patterson, M. T. et al. Trem2 promotes foamy macrophage lipid uptake and survival in atherosclerosis. Nat. Cardiovasc. Res. 2, 1015–1031 (2023).
Yu, W. et al. Myeloid Trem2 ameliorates the progression of metabolic dysfunction-associated steatotic liver disease by regulating macrophage pyroptosis and inflammation resolution. Metabolism 155, 155911 (2024).
Tan, J. et al. TREM2(+) macrophages suppress CD8(+) T-cell infiltration after transarterial chemoembolisation in hepatocellular carcinoma. J. Hepatol. 79, 126–140 (2023).
Zhong, J. et al. Distinct roles of TREM2 in central nervous system cancers and peripheral cancers. Cancer Cell 42, 968–984 (2024).
Cui, H. et al. TREM2 promotes lung fibrosis via controlling alveolar macrophage survival and pro-fibrotic activity. Nat. Commun., 16, 1761 (2025).
Wang, X. et al. Prolonged hypernutrition impairs TREM2-dependent efferocytosis to license chronic liver inflammation and NASH development. Immunity 56, 58–77 (2023).
Yang, D. et al. TREM2 Depletion in Pancreatic Cancer Elicits Pathogenic Inflammation and Accelerates Tumor Progression via Enriching IL-1β(+) Macrophages. Gastroenterology 168, 1153–1169 (2025).
Liu, C. et al. SPP1+ macrophages promote head and neck squamous cell carcinoma progression by secreting TNF-α and IL-1β. J. Exp. Clin. Cancer Res 43, 332 (2024).
Liu, X. et al. Context-dependent activation of STING-interferon signaling by CD11b agonists enhances anti-tumor immunity. Cancer Cell 41, 1073–1090 (2023).
Caronni, N. et al. IL-1β(+) macrophages fuel pathogenic inflammation in pancreatic cancer. Nature 623, 415–422 (2023).
Sattiraju, A. et al. Hypoxic niches attract and sequester tumor-associated macrophages and cytotoxic T cells and reprogram them for immunosuppression. Immunity 56, 1825–1843 (2023).
Sacristan Santos V. et al. Reprogramming tumor-associated macrophages using STING or TLR agonists: a promising strategy to enhance immunotherapy in hormone-dependent cancers. J. Immunother. Cancer 13, e010950 (2025).
LI, Y. et al. Nicotinamide N -methyltransferase promotes M2 macrophage polarization by IL6 and MDSC conversion by GM-CSF in gallbladder carcinoma. Hepatology 78, 1352–1367 (2023).
Doran, A. C. et al. Efferocytosis in health and disease. Nat. Rev. Immunol. 20, 254–267 (2020).
Mehrotra, P. & Ravichandran, K. S. Drugging the efferocytosis process: concepts and opportunities. Nat. Rev. Drug Discov. 21, 601–620 (2022).
Cui, D. et al. Pivotal advance: macrophages become resistant to cholesterol-induced death after phagocytosis of apoptotic cells. J. Leukoc. Biol. 82, 1040–1050 (2007).
Moore, K. J. & Tabas, I. Macrophages in the pathogenesis of atherosclerosis. Cell 145, 341–355 (2011).
Raymond, M. H. et al. Live cell tracking of macrophage efferocytosis during Drosophila embryo development in vivo. Science 375, 1182–1187 (2022).
Cheng, M. et al. CD276-dependent efferocytosis by tumor-associated macrophages promotes immune evasion in bladder cancer. Nat. Commun. 15, 2818 (2024).
Zhou, Y. et al. Blockade of the Phagocytic Receptor MerTK on Tumor-Associated Macrophages Enhances P2X7R-Dependent STING Activation by Tumor-Derived cGAMP. Immunity 52, 357–373 (2020).
Tufan, T. et al. Rapid unleashing of macrophage efferocytic capacity via transcriptional pause release. Nature 628, 408–415 (2024).
Hong, S. M. et al. NAMPT-Driven M2 Polarization of Tumor-Associated Macrophages Leads to an Immunosuppressive Microenvironment in Colorectal Cancer. Adv. Sci. (Weinh.) 11, e2303177 (2024).
Hou, Z. et al. Loss of Annexin A1 in macrophages restrains efferocytosis and remodels immune microenvironment in pancreatic cancer by activating the cGAS/STING pathway. J. Immunother. Cancer 12, e009318 (2024).
Wei, Y. T. et al. Thymosin α-1 Reverses M2 Polarization of Tumor-Associated Macrophages during Efferocytosis. Cancer Res. 82, 1991–2002 (2022).
Argyris, D. G. et al. An emerging paradigm of CXCL16 involvement in cancer progression. Cytokine Growth Factor Rev. 84, 87–100 (2025).
Liu, Y. et al. High metastatic tumor-derived CXCL16 mediates liver colonization metastasis by inducing Kupffer cell polarization via the PI3K/AKT/FOXO3a pathway. Neoplasia 65, 101174 (2025).
Srivastava, N. et al. CXCL16-dependent scavenging of oxidized lipids by islet macrophages promotes differentiation of pathogenic CD8(+) T cells in diabetic autoimmunity. Immunity 57, 1629–1647 (2024).
Korbecki, J. et al. The Role of CXCL16 in the Pathogenesis of Cancer and Other Diseases. Int. J. Mol. Sci. 22, 3490 (2021).
Ge, X. et al. Metabolic reprogramming through histone lactylation in microglia and macrophages recruits CD8(+) T lymphocytes and aggravates spinal cord injury. Neuron 113, 2280–2296 (2025).
Jiang, W. et al. CD8 T cells induce the peritubular capillary rarefaction during AKI to CKD transition. Int. J. Biol. Sci. 20, 2980–2993 (2024).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2021YFA1201100), the National Natural Science Foundation of China (grants 82430057, 82030092, 82473338), Tianjin Natural Science Foundation (24JCYBJC00580). Tianjin Key Medical Discipline (Specialty) Construction Project (TJYXZDXK-009A). Collaborative Principal Investigator Program inBasic Sciences, Clinical Tianjin Medical and University Cancer Institute & Hospital (20250105).
Author information
Authors and Affiliations
Contributions
Conception and design: Xiaofan Guo, Jihui Hao, Song Gao, Xuan Zhou and Jun Yu. Development of methodology: Xiaofan Guo, Yuxiao Liu and Song Gao Investigation: Xiaofan Guo, Yuxiao Liu, Xiaopeng An, Tianchen Li, Yuning Song, Peijun Xu, Jing Huang, Yiping Zou, Bohang Xu, Yongjie Xie, Zekun Li. Data curation: Xiaofan Guo, Yuxiao Liu and Song Gao. Manuscript writing, review, and revision: Xiaofan Guo, Yuxiao Liu, Song Gao and Jihui Hao. Funding acquisition: Jihui Hao. Study supervision: Jihui Hao. All authors have read and approved the review article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All human sample manipulation involved in this study was in accordance with the Declaration of Helsinki and related ethical norms, and was approved by the Ethics Committee of Tianjin Medical University Cancer Hospital (approval number: EK20240448). All participants signed informed consent forms. All animal procedures involved in this study followed the international guidelines for the care and use of laboratory animals and were approved by the Laboratory Animal Ethics Committee of Tianjin Medical University Cancer Institute and Hospital (approval number: 2021045).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guo, X., Liu, Y., Li, T. et al. Targeting GPR34 in damage-associated macrophages enhances anti-tumor immunity and the efficacy of Surufatinib in pancreatic cancer. Sig Transduct Target Ther 11, 156 (2026). https://doi.org/10.1038/s41392-026-02641-4
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41392-026-02641-4