Abstract
Hypoxia drives diabetic kidney disease (DKD) progression through Hypoxia Inducible Factor (HIF) signaling. The kidney’s cellular heterogeneity and complex architecture pose challenges for directly assessing the pharmacologic effects on kidney oxygenation and hypoxia-responsive pathways in vivo, such as treatment with SGLT2 inhibitors (SGLT2i), presumed to impact kidney oxygenation. Using single-cell transcriptional profiling of kidney tissue from youth with type 2 diabetes (T2D) who showed minimal clinical evidence of DKD, we identified cell type enrichment of HIF-regulated genes, findings that replicated in people with later-stage DKD in the Kidney Precision Medicine Project (KPMP). Using conserved transcription factor (TF) binding motifs, higher-order promoter regulatory structures identified potential cooperating TFs that explained the cell type enrichment pattern. From these promoter elements, 7 interconnected regulatory pathways were identified, comprising a network of 237 genes. Analysis of multiome data from reference tissue in KPMP demonstrated that 80% of the network genes resided in accessible chromatin. Expression of network genes increased significantly in the late compared to the early stage DKD and was validated in a hypoxic human organoid model system. Kidney tissue from individuals with T2D treated with SGLT2i demonstrated reversal of the accumulated changes in the HIF network compared to those not treated with SGLT2i. Most high-confidence genes showed concordant differential expression in spatial transcriptomics from individuals with T2D. Hypoxic kidney organoids treated with SGLT2i confirmed these protective effects. Our promoter-anchored HIF regulatory network provides a multi-component read-out that captures disease progression and quantifies therapeutic response to SGLT2i.
Similar content being viewed by others
Introduction
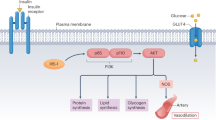
Diabetic kidney disease (DKD) is a leading cause of kidney failure globally and is associated with high morbidity and mortality.1 Hypoxic stress contributes to the progressive damage seen in the tubular compartments of the kidneys that is characteristic of DKD in type 2 diabetes (T2D).2 The response to hypoxia is transcriptionally regulated by the transcription factor (TF) hypoxia inducible factor-1A (HIF1A).3 In normoxia, HIF1A protein is targeted for degradation by cytosolic prolyl hydroxylases, while in hypoxia, these prolyl hydroxylases are inhibited, allowing HIF1A to translocate to the nucleus, dimerize with HIF1β protein, and the dimer then binds to hypoxia response elements to modulate transcription of target genes, prompting changes in metabolism and angiogenesis to promote survival.3 In murine models, alterations in the HIF-hypoxia response axis contribute to the pathogenesis of progressive DKD.4,5,6,7,8,9 However, confirming the relevance of this mechanism to human kidney disease remains challenging.10,11 This is due to the complex transcriptional cascade of HIF-driven events superimposed on the intricate anatomy and inherent hypoxia-related physiology of the kidney with highly variable oxygen demand and supply along the nephron.12 Moreover, much of the prior knowledge regarding HIF signaling has been generated from malignant tissue.
One long-standing hypothesis, the “chronic hypoxia” hypothesis, posits that damage in the glomerular capillary bed or peri-tubular capillaries forces the tubulointerstitial compartment of the kidney to experience chronic, ongoing hypoxia, and the response to these insults is at least partially regulated by HIF activity.6,7,13 Non-invasive measurements of oxygen tension of the kidney, such as BOLD-MRI, have demonstrated that kidney hypoxia is associated with kidney disease progression.11 In the kidney, HIF activity is increased by multiple stimuli, including hypoxia, inflammatory cytokines and angiotensin-II.14,15 Regardless of its initial stimulus, animal models show that HIF has competing effects in kidney tissue and vary by cell type, injury type and stage.4,16,17,18,19,20,21,22 Increased HIF activity in tubular epithelial cells prior to the onset of a toxic or ischemic renal insult appears to diminish the severity of tubular injury, which should be a beneficial effect, but increased expression of HIF in tubular epithelial cells has also been shown to trigger interstitial fibrosis, which is linked to loss of kidney function.17,18,19,20,23 Data from cell cultures suggest that HIF1A helps mediate the fibrotic response of tubular epithelial cells to albumin.24 However, there are limited human data available to assess the impact of HIF pathway activity on patient outcomes.10,23,25 One cross-sectional study involving a cohort of patients with diverse causes of CKD showed that increased HIF pathway activity in the glomerular compartment was more common in patients with nephrosclerosis and was associated with decreased estimated glomerular filtration rate.10 Moreover, since HIF1A is mostly translationally regulated, it has been difficult to quantify its overall downstream impact.
Therapeutic agents that potentially influence either oxygen tensions in the kidney or HIF activity are now available for patients with DKD. For example, SGLT2 inhibitors (SGLT2i), which slow progression of CKD in T2D,26,27,28 have been hypothesized to attenuate hypoxia within the kidney cortex partially by decreasing energy demand in the proximal tubule (PT) and potentially increasing energy demand in the medulla.2,29 Conversely, prolyl hydroxylase inhibitors directly increase HIF protein levels in the kidney, which can increase erythropoietin expression and treat anemia of CKD.30,31 Their uptake into clinical practice has been limited partly due to safety concerns, although some mechanistic studies suggest that prolyl hydroxylase inhibitors may be protective in mouse models of DKD.32,33 Thus, due to the complexity of the HIF signaling cascade and intricacies of nephron anatomy, substantial uncertainty remains regarding the impact of available treatments on HIF signaling and whether such alterations are beneficial to patients with DKD.
Identification of a hypoxia-associated regulatory network in the kidney could act as a target platform to evaluate the potential effects of therapeutic agents on this key pathophysiological pathway, resulting in progressive DKD. The advent of single-cell resolution transcriptomic technologies offers an opportunity to gain unparalleled insight into how this complex regulatory network and various pharmacological therapies affect the individual cell types of the nephron. This study aims to identify a HIF regulatory network that is reflective of nephron anatomy and physiology and serves as a useful tool to interrogate kidney disease progression and response to therapies. Accordingly, we leveraged single-cell transcriptional profiling of kidney biopsies from multiple human cohorts to construct a HIF regulatory network using comparative promoter analysis. Next, we validated the network with bulk transcriptional data, single-cell transcriptional data and epigenetic data using kidney tissue from independent human cohorts. We first examined whether the expression of network genes was perturbed in altered cell states and with the progression of DKD in humans. We then demonstrated concordant regulation of the network using a model system of hypoxia in kidney organoids. Finally, the activity of the network in relation to SGLT2i was tested in people with DKD treated with SGLT2i and similarly treated hypoxic organoids (Fig. 1).
Study Overview. In Phase 1, the HIF regulatory network was developed using comparative promoter analysis and then validated using snATACseq data, RNA sequencing data and a model system of hypoxic organoids. In Phase 2, the HIF regulatory network was used to test the impact of SGLT2i treatment using human kidney biopsy data and a hypoxic organoid model system. Created partly in BioRender. Subramanian, L. (2026) https://BioRender.com/vgyyur8
Results
HIF-responsive gene expression shows cell type enrichment in kidney tissue
To assess the transcriptional response associated with the change in oxygen availability, we created a curated list of HIF response genes (HRG). A comprehensive literature review identified 1,666 unique transcripts associated with an upregulated transcriptional response to hypoxia or experimental alterations in HIF activity.10,34,35,36,37,38,39,40 Of these 1,666 genes, 272 were replicated in more than one data source and were used for further study (Supplementary Table 1). This gene set was further filtered to 181 genes, rejecting those expressed in fewer than 10% of cells of any kidney cell type in single cell RNA sequencing (scRNAseq) data from kidney tissue of youth with T2D and minimal evidence of DKD (Supplementary Fig. 1). Functional enrichment analysis of the 181 genes appropriately represented the established biologic pathways that are regulated by HIF, including glycolysis, angiogenesis and apoptosis (Supplementary Table 2). Notably, EPO, the gene that encodes erythropoietin, which is a well-characterized target gene of HIFs, was not included in our analysis as expression was not detected in the single-cell transcriptional profiles.
In youth with T2D not treated with SGLT2i (T2Di(-)) and the Kidney Precision Medicine Project (KPMP) participants with adjudicated DKD (KPMP-DKD), a few nephron segments and cell types differed in their patterns of HRG expression. However, the overall expression patterns were similar between T2Di(-) and KPMP-DKD, since the cell type with maximal expression of a gene was the same for 85% of the genes between the two cohorts (Supplementary Table 1).
HIF cooperates with distinct TF families in the kidney within defined hierarchical promoter structures
As a combination of multiple interacting TFs is needed for the expression of a specific gene transcript, we hypothesized that there were TF families that cooperated with HIF to dictate the synchronized expression of genes linked to hypoxic stress. By examining the promoter regions of HRG, we identified potential HIF-associated higher-order promoter regulatory structures, known as Multiple Organized Regulatory Elements (MORE cassettes). MORE cassettes reflect specific patterns of TF binding motifs based on their conserved distance, orientation and spacing (Fig. 2a). As this approach identifies binding sites for a TF family, our approach could not distinguish between the binding of HIF1A, HIF2A or HIF3A. Two TF families were overrepresented in the HRG-associated MORE cassettes, identifying them as potential HIF cooperating TF families: Kruppel-like Factor (KLF) and E26 Transformation Specific (ETS) factor (Fig. 2b). The presence of these MORE cassettes was enriched by 2 to 30-fold in promoter regions of HIF regulated transcripts relative to all promoters across the human genome (Fig. 2c). One specific MORE cassette identified from the descending thin limb was overrepresented 30-fold, while the greatest number of MORE cassettes were identified from macrophages and principal cells (PC) (Fig. 2c).
Identification of MORE cassettes. a Schematic of how MORE cassettes are identified by comparing promoter sets for enriched syntactic patterns of TF binding sites. b KLF and ETS TF families were the most frequently identified TF families cooperating with HIF. c MORE cassettes identified from comparative promoter analysis were over-represented compared to the approximately 140,000 promoters in the human genome. MORE cassettes are shown in the nephron segment where they were over-represented
Development of the HIF regulatory network
The genome-wide MORE cassette search was a discovery approach of observed genomic features and identified additional potential hypoxia-associated genes outside the original list, resulting in a raw and highly redundant list of several thousand genes due to the presence of multiple different MORE cassettes in matching gene promoters. Therefore, we employed enrichment analysis of this gene list for signaling pathways, which automatically purged redundant genes. We focused on top-scoring pathways and scrutinized these pathways further by checking their association with HIF factors, kidney diseases and/or hypoxia. Seven pathways (Fig. 3a, b, unique color for each pathway, Supplementary Fig. 2 and Supplementary Table 3) fulfilled all these criteria and included 1) HIF1A transcription factor network (pink), 2) hypoxic and oxygen hemostasis regulation of HIF1A (black), 3) Vascular Endothelial Growth Factor (VEGF) hypoxia and angiogenesis (blue), 4) HIF2A transcription factor network (green), 5) signaling by Transforming Growth Factor-β (TGFB) receptor complex (brown), 6) signaling by Platelet Derived Growth Factor (PDGF, yellow), and 7) hypoxia response by HIF activation pathways (orange). These pathway networks are regulated by HIF1A as the central pathway (Fig. 3c and Supplementary Fig. 3). The total nonredundant gene number of all 7 pathways was 237, several of which were present in more than one pathway. This pathway gene list formed the basis of the hypoxia- associated network constructed based on published pathway graphs (Supplementary Table 3 and Supplementary Figs. 4–8). Due to the inclusion of all genes from the 7 pathways, where not all genes contained a HIF-MORE cassette (108 of the 237 genes contained at least 1 HIF-MORE cassette), the network was thus expanded beyond the results of promoter analysis to a functional pathway network. Specifically, 206 of our network genes were not in the initial literature-derived list (Supplementary Table 3).
Creation of the HIF regulatory network. a Schematic of regulatory network with genes belonging to different pathways in different colors, transcription factors highlighted with open black circles around red dots. Gene names can be viewed in Supplementary Fig. 2. b Schematic of regulatory network with all 7 pathways represented in different colors (pink = HIF1A transcription factor network, orange = hypoxia response via HIF activation, black = hypoxic and oxygen homeostasis regulation of HIF1A, blue = Vascular Endothelial Growth Factor (VEGF) hypoxia and angiogenesis, green = HIF2A transcription factor network, yellow = signaling by PDGF, brown=signaling by TGF-beta.) c Schematic of regulatory network showing that the HIF1A pathway is a central portion of the network (pink lines). d 80% of the network promoters were present in open chromatin (represented by light blue circles)
Essentially, by using information from regulatory elements in promoter regions, we were able to find a network of transcriptionally linked genes that both reflects the known biology of HIF and expands upon the literature-derived lists. While the MORE cassettes were identified based on genes enriched in specific kidney cell types, the underlying promoter structure is present in the genome of every cell. Nevertheless, the MORE cassettes were identified in promoters of genes encoding transcripts that were expressed in kidney tissue; therefore, we would expect that genes in the reconstructed network reside in open chromatin regions in kidney tissue.
Using available multiome data from reference kidney tissue from KPMP, we demonstrated that 80% (189/237) of the network genes have at least one promoter in open chromatin (Fig. 3d). Taken together, our bioinformatic approach identified a network of genes involved in HIF biology that are accessible for transcriptional expression in the human kidney, supporting an environment inherently responsive to hypoxic conditions.
Gene expression in the HIF regulatory network is perturbed in adaptive PT cells and progressive DKD
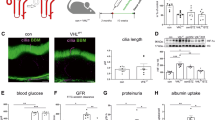
As we hypothesized that disease progression is associated with altered expression of genes in the HIF regulatory network, we first sought to examine perturbations of the expression of network genes at the earliest signs of disease. We focused on adaptive proximal tubular (aPT) cells since increased numbers of these cells were previously linked with poor long-term outcomes.41 We compared the differences in the HIF regulatory network between healthy proximal tubular cells (PT, n = 13,818) and aPT cells (n = 2157) in the KPMP reference tissue multiome data. Only a small proportion of open chromatin regions (peaks) were differentially accessible across PT and aPTs (n = 164/4310; 3.8%). However, those chromatin regions with altered accessibility were frequently associated with genes within the regulatory network (n = 97/237); thus, 41.0% of network genes were annotated with at least one differentially accessible region (Supplementary Table 4). Furthermore, most of the differentially accessible regions were near promoters (n = 101/164 peaks; n = 83/97 genes), so the altered accessibility may perturb gene expression. In fact, nearly half of the HIF regulatory network genes (n = 104/237; 43.9%) had different expression levels between aPT and PT (Supplementary Table 5). Of those transcripts, 49 (47%) coincided with altered accessibility at the promoter, creating a high-confidence set of HIF regulatory network transcripts that are altered in a cell type that has been previously associated with poor long-term outcomes (Fig. 4a). An example of altered chromatin accessibility and gene expression is shown for TGFBR2 in Fig. 4b.
HIF regulatory network is relevant in early and late DKD. a Orange rings represent network genes with 5% of expression for both snATAC and snRNASeq (n = 44 genes) and differential chromatin accessibility and gene expression when comparing aPT cells to PT cells (unadjusted p-value < .05) using publicly available data from KPMP. b Aggregated read pileup tracks across all individuals within the KPMP multiome data showing higher expression (top) and more accessibility (middle) for TGFBR2 in adaptive (blue) compared to healthy (red) PT cells. Annotation tracks of all nearby genes with RNA expression (bottom) highlight promoter regions for each transcript. Transcription start sites (TSS) are also indicated with a vertical red line in the middle accessibility plot. CPM = counts per million. SPMR = signal per million reads. c In bulk RNAseq data, early DKD was associated with upregulation (green) and downregulation (blue) of transcripts from network genes compared to living donor controls (unadjusted p-value < 0.05). d In bulk RNAseq data, late DKD was associated with a larger number of upregulation (green) and downregulation (blue) of transcripts of network genes compared to living donor controls (p-value < 0.05). e Healthy (red) and injured (blue) PT spots near an immune infiltration (top) with log-normalized TGFBR2 expression on the same tissue region (bottom; darker shades of orange indicate more expression)
We then examined the network genes in diseased kidney tissue by interrogating RNA sequencing data from cortical kidney tissue from two cohorts with early and advanced DKD. Since the kidney cortex is about 65% PT cells, this would largely reflect changes in PT cells. We found that 49 genes were differentially expressed when comparing early DKD to reference (Fig. 4c; Supplementary Table 6) while 163 genes were differentially expressed when comparing late DKD to reference (Fig. 4d; Supplementary Table 6) (p-value < 0.05 for both early DKD to reference and late DKD to reference). Similar results were obtained when comparing tissue transcriptomic data for people with DKD from the European Renal cDNA Bank (ERCB) cohort42,43 to living transplant kidney donors (Supplementary Table 7).
Expression of high-confidence genes is dysregulated in injured PT cells by spatial transcriptomics
In VisiumHD spatial transcriptomics (ST) data from four individuals with T2D, we identified PT spots (n = 58,349) and spots sharing features of adaptive and degenerative PT (aPT/dPT; n = 6552). Both groups of spots expressed more CUBN, a PT marker gene, than spots not labeled as PT or aPT/dPT, and the aPT/dPT spots had higher signals of cellular injury (HAVCR1, PROM1, VIM, VCAM1) than spots annotated as healthy proximal tubules (unadjusted i-values < 0.05; Supplementary Figs. 9 and 10). As further validation, we selected five genes from the HIF-network with concordant altered expression between healthy and disease states within multiome measurements and both bulk DKD datasets, to explore differential expression in situ within VisiumHD PT spot categories. Of the selected genes, three (AKT3, CREB1, and TGFBR2) were upregulated, and two (VEGFA and HNF4A) lost expression within diseased or damaged tissue. Within the ST data, four of the five high confidence genes (AKT3, CREB1, TGFBR2, and VEGFA) matched the pattern of expression between healthy and damaged PT spots observed in the other transcriptomic data (unadjusted p’s < 0.05; Supplementary Figs. 9 and 10, Supplementary Table 8). On the tissue, spots labeled as aPT/dPT co-localized near immune infiltrations and, as a highlighted example, showed higher expression of TGFBR2 (Fig. 4e).
Expression of genes in the HIF regulatory network is altered in hypoxic organoids and functionally dependent on HIF1A and HIF2A in cell culture
After demonstrating that expression of genes of the HIF regulatory network is enhanced in altered cell states, including aPT and progressive DKD, we determined the effect of hypoxia in an experimental platform. Since promoter regulatory elements are not always conserved across animal species,44 we utilized single-cell transcriptional profiling data from kidney organoids generated from human pluripotent stem cells and exposed to 1% O2 for 24 h.45 Interrogation of transcriptomes of organoid PT cells (Fig. 5a) revealed that 60 genes were differentially expressed under hypoxic conditions out of the 159 genes that were expressed in this cell type (Fig. 5b and Supplementary Fig. 11). Direction of differential expression was about equally split between increased and decreased, experimentally confirming that alterations in expression of the HIF-network genes are associated with decreased oxygen availability (Supplementary Fig. 11). There were no significant differences in the HIF network between Hypoxia vehicle control (DMSO) versus Hypoxia (no DMSO)(Supplementary Fig. 11).
HIF regulatory network validated in a kidney organoid model, which reflects SGLT2i treatment effects. a UMAP of hypoxic, normoxic and SGLT2i-treated organoids. b Log-fold change of 6 exemplar genes out of 60 genes significantly regulated in hypoxic compared to normoxic organoids that are reversed by dapagliflozin (FDR < 0.05). c Schematic representation of transcriptional changes in HIF regulatory network genes in the PT cells of youth with T2D treated with SGLT2i compared to healthy controls (HC), p-value < 0.05. Of the genes differentially expressed between both comparisons (T2Di(-) vs. HC and T2Di(+) vs. T2Di(-)), about half of the genes reverted closer to the HC state (n = 35; green), with 28 of the genes moving farther from the HC state (blue)
We additionally validated that expression of genes in the HIF network is functionally dependent on HIF1A and HIF2A using previously published cell culture data from hypoxic HK2 cells in which there was knockdown of HIF1A and HIF2A.10 After adjustment for multiple testing, 106 of 233 genes expressed in HK2 cells were significantly (FDR < 5%) differentially expressed with hypoxia (59 up- and 47 down-regulated). As expected, HIF1A and HIF2 (EPAS1) were both sharply down-regulated under their respective knockdowns. Of the 18 genes with greater than 0.4 log2 fold-change under hypoxia, all were significant at FDR < 1% and generally showed attenuation of the hypoxia response under HIF knockdown (Supplementary Fig. 12).
Expression of genes in the HIF regulatory network is altered with SGLT2i treatment
SGLT2i has been proposed to mitigate hypoxic stress in the PT epithelial compartment in T2D.2,8,29,46 Accordingly, we hypothesized that treatment with SGLT2i would attenuate gene expression changes in the HIF regulatory network that are associated with progressive T2D. Comparison of gene expression in PTs from T2D +/- SGLT2i treatment and healthy controls identified 83 genes from the network with differential expression in T2Di(-) compared to healthy controls. Of those, 35 reverted closer to the baseline state with SGLT2i and another 28 moved farther from HC expression levels following treatment (Fig. 5c, p-value < 0.05; Supplementary Table 9). In PTs of hypoxic organoids, SGLT2 inhibition reversed the expression of key nodes in the network, including HIF1A, prolyl hydroxylases (EGLN1 and EGLN3), SMAD7, which regulates TGF-β, and RORA (Fig. 5b). Taken together, these data show that SGLT2i modulates the expression of HIF1A, HIF-network and hypoxia-related genes in PT cells. For a visual summary of all the data mapped onto the network, please see Supplementary Movie 1.
Discussion
Chronic hypoxia and a perturbed hypoxia-HIF axis with associated downstream changes have long been hypothesized to contribute to the pathogenesis of DKD.6,7 Here we have shown that HIF-regulated transcripts exhibit cell type enrichment within the kidney. Our in-silico analysis predicted that KLF and ETS family were the most likely HIF-cooperating TFs within a defined framework, which is a potential cause for the observed gene coregulation. The conserved promoter regulatory structures were sufficient to find and expand an HIF regulatory network, and promoters of this network were predominantly located in open chromatin in reference samples. Engagement of this HIF regulatory network was detectable in individuals with T2D without overt clinical evidence of DKD, while expression of additional key nodes of the regulatory network was further perturbed in later stages of DKD, suggesting that regulation of the network transcripts is a component of both disease initiation and progression. These findings were further supported by spatial transcriptomic data from an independent cohort of individuals. Furthermore, we experimentally confirmed that the network is active in hypoxic conditions by demonstrating hypoxia-induced network gene expression changes in PT cells of kidney organoids. The network is functionally dependent on HIF1A and HIF2A, as demonstrated by HIF1A/HIF2A knockdown in HK2 cells.10 Moreover, in an independent experiment that identified HIF1A peaks using Chip-Seq in promoters of hypoxic HK2 cells,47 there was extensive overlap between the portions of the network that were functionally dependent on HIF1A or HIF2A and the occurrence of a Chip-Seq peak in the promoter region, which supports that the HIF TF family is binding to the relevant portions of the network. Portions of the hypoxia-induced expression network changes were rescued by treatment with SGLT2i in kidney tissue and further validated in the hypoxic organoid model, supporting previously published hypotheses.
Hyperfiltration in the earliest stages of DKD is thought to increase sodium reclamation in the PT and subsequently increase oxygen utilization, while damage to the glomeruli diminishes blood supply to the peritubular capillaries and lowers oxygen supply, resulting in mismatched oxygen supply and demand.2 Though physiology suggests that there may be chronic hypoxia in DKD, model systems demonstrate that the HIF response is impaired in diabetes. For example, reactive oxygen species in DKD impair the binding of p300, an essential cofactor for HIF1A, resulting in impaired HIF1A transcriptional response.48
Prior investigations in this area were hindered by the complex anatomy and physiology of the human kidney.5,16 Additionally, transcriptional analysis of human kidney tissue was hampered by bulk tissue processing, which limits the ability to examine individual cell types.10 The cell type enrichment pattern, which we replicated in two independent cohorts, would have been impossible to appreciate without single-cell resolution transcriptomic data. Previous studies using RNA sequenced microdissected tissue were able to provide insight into compartment-level regulatory aspects of HIF regulation in the kidney, but were unable to capture patterns specific to nephron segments.5,10,16
While HIF has been previously shown to cooperate with multiple other TF families,49 our results revealed that HIF likely cooperates with KLF and ETS TF families at promoter sites, providing a foundation for a kidney-specific enrichment pattern. As the kidney medulla operates at a lower oxygen tension than the kidney cortex,50 the enrichment pattern may serve a more specific biologic function, which may be revealed by a more extensive evaluation of cell types that cross the corticomedullary junction, such as PT cells and thick ascending limb cells.
Our studies indicate that the gene expression pattern of this network is perturbed in aPT cells, which is an intriguing finding since published findings demonstrated that aPT cells are associated with poor long-term outcomes.41 Indeed, perturbations in the network gene expression were already detectable in seemingly healthy reference tissue without structural changes, indicating that these changes preceded histologic damage. The network of genes was also altered in acutely hypoxic organoids, experimentally validating HIF involvement. Further perturbation of the network was appreciated in two independent cohorts of early, but morphologically evident DKD and further validated in spatial transcriptomic data. These results indicate that alterations in the HIF-regulated transcripts precede structural damage in kidney tissue. More extensive gene perturbations were apparent in later-stage DKD, which likely reflects more profound structural changes, including interstitial fibrosis and tubular atrophy.
However, these results do not conclusively prove that hypoxia is the inciting HIF-related factor driving DKD progression. There are several alternative possibilities for alterations in HIF activity in DKD apart from hypoxia. HIF activity is regulated post-translationally by tri-carboxylate acid (TCA) cycle metabolites and mTORC activity.51,52,53 Transcripts of the TCA cycle are increased in T2D compared to controls and suppressed with SGLT2i,54 and TCA cycle flux is increased in the kidneys of db/db mouse models of diabetes.55 Thus, it is plausible that the HIF regulatory network is deranged not primarily due to hypoxia but rather is a secondary consequence of metabolic changes in diabetes and DKD.
Importantly, while expression of network genes was perturbed in DKD, SGLT2i treatment was associated with diminished expression perturbations in this network, suggesting that components of this network are potentially amenable to targeted pharmacologic therapy. Of note, individuals treated with SGLT2i had minimal structural changes in their kidney tissue apart from a mild increase in glomerular volume.54 Despite minimal structural changes, the scRNAseq data identified that 36% of the network genes had altered expression in the diabetic state. This is a higher number of altered transcripts than were identified in the bulk tissue, which may be related to differences in scRNAseq versus bulk RNAseq.
Our results suggest that this network is likely more suitable to monitor the impact of changes in oxygen supply or therapy that modulates the hypoxia-HIF axis than a single read-out of HIF activity. Accordingly, we propose that our HIF regulatory network provides a robust multi-component read-out reflective of disease stage and suitable for investigating and monitoring treatment effects in human kidney tissue related to SGLT2i or prolyl hydroxylase inhibitors. Our results also provide critical information to guide future research related to HIF activity in the kidney. We identified derangements in HIF activity occurring before structural damage and persisting throughout DKD disease progression, demonstrating the necessity of examining tissues across the spectrum of disease. Additionally, our identification of cell-type enrichment regarding HIF response in kidney tissue demonstrates the necessity to leverage single-cell resolution molecular technologies.
While our study has several strengths, including that the network was validated in multiple human cohorts using RNA and epigenetic sequencing technologies and experimental validation in organoids, there are important limitations. While validation in the organoid system demonstrates that changes in oxygenation are sufficient to alter expression in the network, our analysis did not definitively identify the mechanism that is responsible for altering the expression of the HIF regulatory network in human DKD. Moreover, while the organoid model system was suitable to evaluate the impact of SGLT2i on PT cells, our prior work suggests that SGLT2i impact the remainder of the nephron and is partially contingent on nephron anatomy.54 Thus, the organoid model system is not ideally suited to examine SGLT2i on other nephron cell types. Additionally, the multiome data were derived from reference samples and not from individuals with diabetes, although we would anticipate that diseased tissue would experience greater hypoxia and thus have even greater chromatin accessibility of the HIF regulatory network. We chose to use a model system of hypoxic human organoids rather than a murine model because promoter regulatory elements are not always conserved across species.44 However, hypoxic organoids do also have limitations; the kidney cell types were immature relative to adult human kidney tissue, and the extent of hypoxia in the organoid model was acute and profound, which provides a limited view of the complex physiology of chronic hypoxia seen in DKD.
In this study, we present an extensively validated HIF regulatory network that is present in open chromatin, relevant in early and late stage DKD, where expression of network genes is altered in hypoxic organoids. While these data show that expression of the network genes is altered with an increase in HIF activity that can be reverted closer to a healthy state with SGLT2i treatment, further work is needed to determine the inciting causes of changes in HIF network activity in DKD.
Materials and methods
Cell type enrichment
To define genes that are regulated by HIF, we identified differentially expressed genes in the setting of hypoxia or experimental alterations in HIF activity in non-malignant tissue and replicated in more than one published study. Since most of the literature reported HIF response genes (HRG) were not derived from kidney tissue, we evaluated the HRG in kidney single-cell RNA sequencing (scRNAseq) data from youth with T2D without SGLT2i treatment (T2Di(-)) and kidney samples with an adjudicated diagnosis of DKD from the Kidney Precision Medicine Project (KPMP).41,54 We defined a gene as sufficiently expressed in kidney tissue if more than 10% of one cell type expressed the transcript of interest. Each transcript was assigned to a cell type annotation based on the cell type in which it exhibited the highest average maximum expression in the pseudobulk scRNAseq transcriptional data. This produced a list of transcripts that are cell-type enriched and their associated cell type (Supplementary Table 1 and Supplementary Fig. 1).
Identification of higher-order gene promoter structures linked to HIF
Interacting TFs within gene promoters are necessary for the expression of a gene. Within promoters, TF binding sites conserved in spacing and orientation help coordinate the co-expression of genes whose products functionally interact within a biological context44,56, such as the coordinated response to hypoxic stress. We identified candidate potential cooperating TFs within higher-order promoter structures using a well-established in-silico promoter analysis using the Genomatix software installed on a local server.57,58,59,60,61 The premise is that HIF acts within a framework of other TFs simultaneously binding to the promoters of target genes. Changing the cofactors very often modifies the expression pattern of the corresponding genes.44,56 Provided that there is a sub-group of genes responding to the same TF combinations, candidates for potential co-factors within promoter structures can be deduced through comparative promoter analysis of this subgroup of target genes via the identification of common groups of the respective TF-binding sites (TFBS), termed Multiple Organized Regulatory Elements (MORE) cassettes (Fig. 2a).
The approach works as follows: after identification of cell-type-enriched HRG, the kidney-enriched promoters associated with each HRG are extracted (with at least one HIF TFBS as a mandatory element). These promoter sets were analyzed for enriched syntactic patterns of TFBSs (order, orientation, distances, etc., similar to sentences in a language analysis). We then examined if the resultant candidate promoter models were also present in the promoters of genes which are enriched in kidney cells to identify an expanded predicted gene promoter network. Next, the selected TFBS patterns (MORE cassettes) were subjected to an enrichment analysis in HRG promoters versus all promoters in the human genome ( > 140,000) to evaluate if they are selective for HRG. The overrepresentation of the MORE cassettes in promoters of HRG (499 promoters) was calculated by comparing to the total matches in the HRG promoters with the expected value based on the random distribution of all gene promoter matches in the human genome. The list of all co-factors present in the final selection of overrepresented MORE cassettes was then verified for expression of the respective binding TFs as deduced from the scRNAseq data for each cell line. All MORE-cassettes containing TFBSs for co-factors without expressed TF were subsequently purged.
Derivation of a HIF regulatory network
To find a network based on the MORE cassettes and expand upon the list of literature-derived genes. Using a signaling pathway enrichment analysis on the resulting gene list we determined which pathways were significantly associated with genes containing at least one hypoxia associated HIF- MORE cassette (Supplementary Fig. 1). Top pathway hits were then further scrutinized for association with HIF factors, kidney diseases and/or hypoxia resulting in a list of seven most relevant pathways comprising a total of 237 genes all together (Supplementary Figs. 4–8). These genes form a densely connected network with 40 genes appearing in multiple pathways (some in up to 5). As a result, simply overlaying the pathway graphs is not optimal. Therefore, the network topology was manually arranged to allow for a two-dimensional representation with no genes obscuring other genes.
Multiome analysis
Previously published multiome data were evaluated to see if the genes in the HIF regulatory network were overrepresented in open, accessible chromatin.62 Peaks from the original manuscript (number of peaks = 336,500) were annotated to gene bodies and nearby (−100 bp to +1000 bp) promoters (GENCODE hg38 version 104). We then filtered down to peaks within healthy PT cells as defined by KPMP and adaptive proximal tubular cells (aPT), which is an altered cell state in the kidney. To inspect these cell types, we aggregated counts within healthy PT clusters (n = 13,818 cells) and aPT cells (n = 2157). Those cells comprised more than a third of the total cells from the original publication (n = 15,975/47,217 cells; 33.8%) with non-zero reads in almost every open chromatin region (n = 336,488/336,500). Peaks with fragments in at least 5% of PT and aPT cells that intersected HIF regulatory network promoter regions were taken as evidence of open, accessible chromatin. Next, we evaluated whether the peaks and genes associated with the HIF regulatory network were differentially accessible and expressed between PT and aPT cells. Genes and open chromatin regions with reads in at least 5% of both cell types (n = 4310 peaks; 237 genes) were tested for differential regulation using the Wilcoxon rank test within scanpy (v1.10.1). Genes with an unadjusted p-value < 0.05 were considered differentially regulated. Differentially expressed genes with concordant differentially accessible open chromatin at the promoter created a high-confidence set of genes with altered regulation in early disease states.
Analysis of progressive DKD
Previously published kidney transcriptomic data from two cohorts with progressive DKD were analyzed to see if/how the network is altered in progressive DKD.63,64 Of the network components that were both differentially accessible and expressed between healthy PT and aPT, we examined if these genes were differentially expressed in the two cohorts: 1) between DKD and living donors from the European Renal cDNA Biobank (ERCB) cohort (GSE104954)65,66 and 2) between early versus late DKD from previously published data (GSE142025).64,67 In the ERCB cohort, kidney biopsy tissue procurement and gene expression profiling were performed on Affymetrix GeneChip Array Human Genome series U133A and Plus 2.0 (Affymetrix, Inc., Santa Clara, CA). Affymetrix image files were obtained, processed, normalized, and batch corrected as previously described.63 A differential analysis using the LIMMA method was applied on the tubular transcriptome profiles from DKD (n = 17) and living donor (LD, n = 30).
For early and late DKD, kidney cortex transcriptomic data were downloaded from Gene Expression Omnibus (GEO).63,64 Gene-level expression quantification was performed using HTSeq, and the resultant count data were normalized in voom. PCA and hierarchical clustering were used to identify and remove samples with abnormal expression profiles due to technical issues, and the mapping statistics were obtained from STAR. Twenty-seven samples passed the quality control criteria and were divided into two groups, early (n = 6) and advanced DKD (n = 21). Early DKD was defined as a urine albumin/creatinine (uACR) between 30 and 300 mg/g, estimated glomerular filtration rate (eGFR) > 90 ml/min/1.73 m2, whereas advanced DKD was defined as uACR>300 mg/g, eGFR < 90 ml/min/1.73 m2. Control samples (n = 9) were obtained from the unaffected portion of tumor nephrectomies.64 A limma analysis was performed to identify differentially regulated HRG genes between the early and late DKD compared to control samples. In both analyses, genes were considered differentially expressed if p < 0.05.
Validation of highly concordant genes in spatial transcriptomics data
The study, “Molecular Mechanisms of SGLT2 Inhibition in Diabetic Kidney Disease,” was approved by the Colorado Multiple Institutional Review Board and conducted in accordance with ethical standards. Written informed consent was obtained from all participants prior to any study visits or procedures, including kidney biopsies. Formalin fixed paraffin embedded (FFPE) biopsy cores from four individuals with T2D not treated with SGLT2i were sectioned, placed on a slide, and run through the VisiumHD platform (v1; 10x). Each slide is comprised of a grid of 8 μm by 8 μm squares, and each square or spot is covered in a lawn of one unique barcode identifier that anneals to any RNA within that grid space. Probes covering nearly the entire transcriptome (Visium Human Transcriptome Probe Set v2.0) are then applied to the tissue to pair transcript expression with the spatial barcode identifier for each molecule. Thus, each measured transcript can be mapped to specific tissue coordinates, allowing for an exploration of RNA expression in native tissue context. After sequencing of the transcripts and accompanying barcodes, transcript-based reads were aligned against the h38 genome (GRCh38-2020-A) within spaceranger (v3.1.1) to build count matrices of gene expression for each 8 μm × 8 μm spot. Spots with low counts ( < 20) were removed using Seurat v5. Then the filtered matrices were normalized, scaled, reduced to low-dimensional embeddings, and subsequently integrated using the Harmony algorithm to create a unified representation of the data. The integrated dataset was then clustered, and highly upregulated genes in each cluster (identified by the Wilcoxon rank sum test in Seurat), along with the spatial context of the cluster’s location in situ, were used to annotate the predominant cell type in each spot. Spots within PT labeled clusters had high expression of CUBN compared to other clusters (unadjusted p < 0.05), and injured PT were identified by upregulated HAVCR1, VIM, VCAM1, and PROM1 expression (unadjusted p < 0.05) as well as physical proximity to immune infiltrations. Validation of four genes showing consistent altered expression (log fold change in the same direction and unadjusted p’s <0.05) across other data types was also assessed with the Wilcox sum rank test (unadjusted p < 0.05).
Kidney organoid single-cell transcriptional data
Single-cell transcriptional data from hypoxic kidney organoids generated from UM77-2 human embryonic stem cells were treated with dapagliflozin as described45 and detailed in the supplementary methods. Briefly, kidney organoids were generated from UM77-2 human embryonic stem cells (NIHhESC-14-0278) and incubated with 1% hypoxia or normoxia for 24 h +/− dapagliflozin 10 mM or DMSO as vehicle control (VC, <0.1%). Whole well samples were collected on ice, dissociated into single cells and submitted for scRNAseq. scRNAseq transcriptome data from the 4 samples were then processed, aligned, clustered and annotated manually. For this study, the proximal tubular cell cluster was identified (based on enhanced expression of SLC3A2, EPCAM, CUBN and LRP2). Differential expression was carried out on proximal tubular cells using the Seurat FindMarkers function, comparing multiple treatment groups. Only genes detected to be expressed in at least ten percent of the cells in either group were analyzed.
HIF1A and HIF2A knockdown in hypoxic HK2 cells
In a previous study,10 immortalized human proximal tubular epithelial (HK2) cells with and without stable HIF1A and/or HIF2A (EPAS1) suppression were subjected to normoxic (20% O2) or hypoxic (1% O2) conditions for 24 h. Gene expression was subsequently measured by microarray (data are available in GEO under accession GSE99324). To evaluate differential expression (DE) attributable to hypoxia, HIF1 and HIF2 (EPAS1) knockdown, a trivariate model was applied using limma.
Expression of network genes is altered with SGLT2i therapy
Using previously published samples,54 we compared expression of the network genes between individuals with T2D treated with SGLT2i (T2Di(+)) and those without SGLT2i (T2Di(-)). To identify therapy alterations, we completed a reversal analysis as was done previously.54 Specifically, we compared gene expression from the T2Di(-) group to a group of healthy controls (HC) to identify disease-related alterations. Subsequently, we compared gene expression between T2Di(+) to T2Di(-) to identify SGLT2i-related effects. Gene expression with a log fold change (log2FC) in opposite directions between the two groups and an unadjusted p-value < 0.05 in both comparisons were considered significantly reversed with SGLT2i treatment (closer to healthy control (HC)) Similarly, genes with an altered log2FC in the same direction between groups and unadjusted p-values <0.05 in both comparisons were categorized as farther from HC.
Study approvals
Transcriptomic data used in these analyses were obtained from previously published sources.41,54 The Molecular Mechanisms of SGLT2 Inhibition in Diabetic Kidney Disease study (COMIRB 22-0668), the Renal-HEIR study (COMIRB 16-1752) and the IMPROVE-T2D (COMIRB18-0704) were reviewed and approved by the Colorado Multiple Institutional Review Board. The KPMP study is approved by the University of Washington Institutional Review Board (IRB 20190213).
Data availability
Transcriptomic data used in this study are available from GSE104954, GSE142025, GSE220939, GSE99340 and GSE183279. The expression matrices for the multiome are available at https://doi.org/10.48698/HHE6-YV15. The Seurat object is available in Zenodo at https://doi.org/10.5281/zenodo.8029990. The organoid data are available from GSE306041. The VisiumHD data can be accessed from GSE317226.
References
Tuttle, K. R. et al. Moving from evidence to implementation of breakthrough therapies for diabetic kidney disease. Clin. J. Am. Soc. Nephrol. 17, 1092–1103 (2022).
Hesp, A. C. et al. The role of renal hypoxia in the pathogenesis of diabetic kidney disease: a promising target for newer renoprotective agents including SGLT2 inhibitors? Kidney Int. 98, 579–589 (2020).
Schödel, J. & Ratcliffe, P. J. Mechanisms of hypoxia signalling: new implications for nephrology. Nat. Rev. Nephrol. 15, 641–659 (2019).
Haase, V. H. Pathophysiological consequences of HIF activation: HIF as a modulator of fibrosis. Ann. N. Y Acad. Sci. 1177, 57–65 (2009).
Rosenberger, C. et al. Adaptation to hypoxia in the diabetic rat kidney. Kidney Int. 73, 34–42 (2008).
Fine, L. G., Orphanides, C. & Norman, J. T. Progressive renal disease: the chronic hypoxia hypothesis. Kidney Int. Suppl. 65, S74–S78 (1998).
Fine, L. G. & Norman, J. T. Chronic hypoxia as a mechanism of progression of chronic kidney diseases: from hypothesis to novel therapeutics. Kidney Int. 74, 867–872 (2008).
Layton, A. T., Vallon, V. & Edwards, A. Predicted consequences of diabetes and SGLT inhibition on transport and oxygen consumption along a rat nephron. Am. J. Physiol. Ren. Physiol. 310, F1269–F1283 (2016).
Franzén, S., Pihl, L., Khan, N., Gustafsson, H. & Palm, F. Pronounced kidney hypoxia precedes albuminuria in type 1 diabetic mice. Am. J. Physiol. Ren. Physiol. 310, F807–F809 (2016).
Shved, N. et al. Transcriptome-based network analysis reveals renal cell type-specific dysregulation of hypoxia-associated transcripts. Sci. Rep. 7, 8576 (2017).
Pruijm, M. et al. Reduced cortical oxygenation predicts a progressive decline of renal function in patients with chronic kidney disease. Kidney Int. 93, 932–940 (2018).
Hansen, J. et al. A reference tissue atlas for the human kidney. Sci. Adv. 8, eabn4965 (2022).
Nangaku, M. Chronic hypoxia and tubulointerstitial injury: a final common pathway to end-stage renal failure. J. Am. Soc. Nephrol. 17, 17–25 (2006).
Page, E. L., Robitaille, G. A., Pouyssegur, J. & Richard, D. E. Induction of hypoxia-inducible factor-1alpha by transcriptional and translational mechanisms. J. Biol. Chem. 277, 48403–48409 (2002).
Zhou, J., Fandrey, J., Schumann, J., Tiegs, G. & Brune, B. NO and TNF-alpha released from activated macrophages stabilize HIF-1alpha in resting tubular LLC-PK1 cells. Am. J. Physiol. Cell Physiol. 284, C439–C446 (2003).
Rosenberger, C. et al. Expression of hypoxia-inducible factor-1alpha and -2alpha in hypoxic and ischemic rat kidneys. J. Am. Soc. Nephrol. 13, 1721–1732 (2002).
Hill, P. et al. Inhibition of hypoxia inducible factor hydroxylases protects against renal ischemia-reperfusion injury. J. Am. Soc. Nephrol. 19, 39–46 (2008).
Weidemann, A. et al. HIF activation protects from acute kidney injury. J. Am. Soc. Nephrol. 19, 486–494 (2008).
Bernhardt, W. M. et al. Preconditional activation of hypoxia-inducible factors ameliorates ischemic acute renal failure. J. Am. Soc. Nephrol. 17, 1970–1978 (2006).
Kimura, K. et al. Stable expression of HIF-1alpha in tubular epithelial cells promotes interstitial fibrosis. Am. J. Physiol. Ren. Physiol. 295, F1023–F1029 (2008).
Tanaka, T., Miyata, T., Inagi, R., Fujita, T. & Nangaku, M. Hypoxia in renal disease with proteinuria and/or glomerular hypertension. Am. J. Pathol. 165, 1979–1992 (2004).
Tanaka, T. & Nangaku, M. The role of hypoxia, increased oxygen consumption, and hypoxia-inducible factor-1 alpha in progression of chronic kidney disease. Curr. Opin. Nephrol. Hypertens. 19, 43–50 (2010).
Higgins, D. F. et al. Hypoxia promotes fibrogenesis in vivo via HIF-1 stimulation of epithelial-to-mesenchymal transition. J. Clin. Invest. 117, 3810–3820 (2007).
Hu, J. et al. Hypoxia inducible factor-1alpha mediates the profibrotic effect of albumin in renal tubular cells. Sci. Rep. 7, 15878 (2017).
Neusser, M. A. et al. Human nephrosclerosis triggers a hypoxia-related glomerulopathy. Am. J. Pathol. 176, 594–607 (2010).
Heerspink, H. J. L. et al. Dapagliflozin in patients with chronic kidney disease. N. Engl. J. Med. 383, 1436–1446 (2020).
Perkovic, V. et al. Canagliflozin and renal outcomes in type 2 diabetes and nephropathy. N. Engl. J. Med. 380, 2295–2306 (2019).
Wanner, C. et al. Empagliflozin and progression of kidney disease in type 2 diabetes. N. Engl. J. Med. 375, 323–334 (2016).
Layton, A. T., Vallon, V. & Edwards, A. Modeling oxygen consumption in the proximal tubule: effects of NHE and SGLT2 inhibition. Am. J. Physiol. Ren. Physiol. 308, F1343–F1357 (2015).
Chen, N. et al. Roxadustat treatment for anemia in patients undergoing long-term dialysis. N. Engl. J. Med. 381, 1011–1022 (2019).
Chen, N. et al. Roxadustat for anemia in patients with kidney disease not receiving dialysis. N. Engl. J. Med. 381, 1001–1010 (2019).
Ku, E. et al. Novel anemia therapies in chronic kidney disease: conclusions from a Kidney Disease: Improving Global Outcomes (KDIGO) Controversies Conference. Kidney Int. 104, 655–680 (2023).
Hasegawa, S. et al. The oral hypoxia-inducible factor prolyl hydroxylase inhibitor enarodustat counteracts alterations in renal energy metabolism in the early stages of diabetic kidney disease. Kidney Int. 97, 934–950 (2020).
Schödel, J. et al. High-resolution genome-wide mapping of HIF-binding sites by ChIP-seq. Blood 117, e207–e217 (2011).
Leonard, M. O., Cottell, D. C., Godson, C., Brady, H. R. & Taylor, C. T. The role of HIF-1 alpha in transcriptional regulation of the proximal tubular epithelial cell response to hypoxia. J. Biol. Chem. 278, 40296–40304 (2003).
Elvidge, G. P. et al. Concordant regulation of gene expression by hypoxia and 2-oxoglutarate-dependent dioxygenase inhibition: the role of HIF-1alpha, HIF-2alpha, and other pathways. J. Biol. Chem. 281, 15215–15226 (2006).
Xin, J. et al. Chromatin accessibility landscape and regulatory network of high-altitude hypoxia adaptation. Nat. Commun. 11, 4928 (2020).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) Hallmark Gene Set Collection. Cell Syst. 1, 417–425 (2015).
Kim, H. et al. Analysis of the effect of aging on the response to hypoxia by cDNA microarray. Mech. Ageing Dev. 124, 941–949 (2003).
Manalo, D. J. et al. Transcriptional regulation of vascular endothelial cell responses to hypoxia by HIF-1. Blood 105, 659–669 (2005).
Lake, B. B. et al. An atlas of healthy and injured cell states and niches in the human kidney. Nature 619, 585–594 (2023).
Schmid, H. et al. Modular activation of nuclear factor-kappaB transcriptional programs in human diabetic nephropathy. Diabetes 55, 2993–3003 (2006).
Yasuda, Y., Cohen, C. D., Henger, A. & Kretzler, M. Gene expression profiling analysis in nephrology: towards molecular definition of renal disease. Clin. Exp. Nephrol. 10, 91–98 (2006).
Fessele, S., Maier, H., Zischek, C., Nelson, P. J. & Werner, T. Regulatory context is a crucial part of gene function. Trends Genet. 18, 60–63 (2002).
Minakawa, A. et al. Dapagliflozin mitigates hypoxia-induced metabolic stress, kidney tubular cell death and fibrosis. bioRxiv https://doi.org/10.64898/2025.12.13.693777 (2025)
Layton, A. T., Laghmani, K., Vallon, V. & Edwards, A. Solute transport and oxygen consumption along the nephrons: effects of Na+ transport inhibitors. Am. J. Physiol. Ren. Physiol. 311, F1217–f1229 (2016).
Kushida, N. et al. Hypoxia-Inducible Factor-1α Activates the Transforming Growth Factor-β/SMAD3 Pathway in Kidney Tubular Epithelial Cells. Am. J. Nephrol. 44, 276–285 (2016).
Thangarajah, H. et al. The molecular basis for impaired hypoxia-induced VEGF expression in diabetic tissues. Proc. Natl. Acad. Sci. USA 106, 13505–13510 (2009).
Zhang, Y., Wang, S., Hu, H. & Li, X. A systematic study of HIF1A cofactors in hypoxic cancer cells. Sci. Rep. 12, 18962 (2022).
Aukland, K. & Krog, J. Renal oxygen tension. Nature 188, 671 (1960).
Hudson, C. C. et al. Regulation of hypoxia-inducible factor 1alpha expression and function by the mammalian target of rapamycin. Mol. Cell Biol. 22, 7004–7014 (2002).
You, Y. H., Quach, T., Saito, R., Pham, J. & Sharma, K. Metabolomics reveals a key role for fumarate in mediating the effects of NADPH oxidase 4 in diabetic kidney disease. J. Am. Soc. Nephrol. 27, 466–481 (2016).
Selak, M. A. et al. Succinate links TCA cycle dysfunction to oncogenesis by inhibiting HIF-alpha prolyl hydroxylase. Cancer Cell 7, 77–85 (2005).
Schaub, J. A. et al. SGLT2 inhibitors mitigate kidney tubular metabolic and mTORC1 perturbations in youth-onset type 2 diabetes. J. Clin. Invest. 133, e164486 (2023).
Sas, K. M. et al. Tissue-specific metabolic reprogramming drives nutrient flux in diabetic complications. JCI Insight 1, e86976 (2016).
Gerstein, M. B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature 489, 91–100 (2012).
Martini, S. et al. From single nucleotide polymorphism to transcriptional mechanism: a model for FRMD3 in diabetic nephropathy. Diabetes 62, 2605–2612 (2013).
Rozman, J. et al. Identification of genetic elements in metabolism by high-throughput mouse phenotyping. Nat. Commun. 9, 288 (2018).
Cohen, C. D. et al. Comparative promoter analysis allows de novo identification of specialized cell junction-associated proteins. Proc. Natl. Acad. Sci. USA 103, 5682–5687 (2006).
Stein, M. F. et al. Multiple interferon regulatory factor and NF-κB sites cooperate in mediating cell-type- and maturation-specific activation of the human CD83 promoter in dendritic cells. Mol. Cell Biol. 33, 1331–1344 (2013).
Werner, T., Fessele, S., Maier, H. & Nelson, P. J. Computer modeling of promoter organization as a tool to study transcriptional coregulation. Faseb J. 17, 1228–1237 (2003).
Gisch, D. L. et al. The chromatin landscape of healthy and injured cell types in the human kidney. Nat. Commun. 15, 433 (2024).
Cohen, C. D., Frach, K., Schlöndorff, D. & Kretzler, M. Quantitative gene expression analysis in renal biopsies: a novel protocol for a high-throughput multicenter application. Kidney Int. 61, 133–140 (2002).
Fan, Y. et al. Comparison of kidney transcriptomic profiles of early and advanced diabetic nephropathy reveals potential new mechanisms for disease progression. Diabetes 68, 2301–2314 (2019).
Alakwaa, F. et al. Leveraging complementary multi-omics data integration methods for mechanistic insights in kidney diseases. JCI Insight 10, e186070 (2025).
Grayson, P. C. et al. Metabolic pathways and immunometabolism in rare kidney diseases. Ann. Rheum. Dis. 77, 1226–1233 (2018).
Fan, Y. et al. Erratum. Comparison of kidney transcriptomic profiles of early and advanced diabetic nephropathy reveals potential new mechanisms for disease progression. Diabetes 2019;68:2301-2314. Diabetes 69, 797 (2020).
Acknowledgements
We would like to thank Drs. Robert Nelson and Helen Looker for their assistance with this project. This work was supported by the Michigan George M. O’Brien Kidney National Resource Center (U54DK137314). We also thank the University of Michigan Medical School Central Biorepository (RRID:SCR_026845) for providing biospecimen storage, management, and distribution services in support of the research reported in this study. We also thank the Renal Precompetitive Consortium (RPC2) collaboration, which is jointly funded by the participating members, for the kidney samples used for spatial transcriptomics. This study was supported, in part, by the George M. O’Brien Michigan Kidney Translational Resource Center (MKTC), funded by NIH/NIDDK grant U54DK137314, Michigan Diabetes Research Center, and the Michigan Diabetes Translational Research Center. JAS was funded for this work through grants from the NIH/NIDDK K08DK124449, R03DK138962. JLH was funded by NIH UH3 TR003288 and Breakthrough T1D Center of Excellence (5-COE-2019-861-S-B). PB was funded by NIDDK (R01 DK129211, R01 DK132399, R01 DK137844, R01 DK138915, U01 DK142249, U01 DK144965), NHLBI (R01 HL165433) and Breakthrough T1D (3-SRA-2025-1786-M-B). The Kidney Precision Medicine Project (KPMP) is supported by the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) through the following grants: U01DK133081, U01DK133091, U01DK133092, U01DK133093, U01DK133095, U01DK133097, U01DK114866, U01DK114908, U01DK133090, U01DK133113, U01DK133766, U01DK133768, U01DK114907, U01DK114920, U01DK114923, U01DK114933, U24DK114886, UH3DK114926, UH3DK114861, UH3DK114915, and UH3DK114937. MTE, DLG and SJ were supported by U54DK134301. JAB was funded by NIH K08 DK125776. We gratefully acknowledge the essential contributions of our patient participants and the support of the American public through their tax dollars. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
V.N., C.C.B., A.M., C.S., J.A.S., M.K., and T.W. conceived and designed this study. P.B., M.K., J.L.H., J.A.S., E.A.O., R.M., D.F., Y.J.C., L.P., F.E., T.W., A.M., J.L.H. acquired the data. A.M., C.S., V.N., C.C.B., J.H., R.M., Y.J.C., L.P., D.F., F.E., J.L.H., F.A., J.E.S., and T.W. analyzed the data. V.N., D.L.G., M.T.E., S.J., M.P.R., L.S., J.E.S., J.B.H., F.C.B., J.A.B., P.B., P.J.N., J.L.H., T.W., M.K., and J.A.S. played an important role in presenting and interpreting the results. J.A.S., V.N., C.S., L.S., F.C.B., and M.K. drafted and revised the manuscript. All authors have read and approved the article, and the corresponding authors agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Corresponding authors
Ethics declarations
Competing interests
P.B. reports serving or having served as a consultant for AstraZeneca, Bayer, Bristol-Myers Squibb, Boehringer Ingelheim, Eli-Lilly, LG Chemistry, Sanofi, Novo Nordisk, and Horizon Pharma. P.B. also serves or has served on the advisory boards and/or steering committees of AstraZeneca, Bayer, Boehringer Ingelheim, Novo Nordisk, Lilly, and XORTX. P.B. serves as DMC Chair for Bayer and a DMC member for Cytel. P.B. reports grant funding from AstraZeneca, Novo Nordisk, Eli Lilly, Boehringer-Ingelheim, Merck, and Amgen/Horizon Pharma. M.K. reports grants and contracts through the University of Michigan outside of this work from Chan Zuckerberg Initiative, AstraZeneca, NovoNordisk, Eli Lilly, Boehringer-Ingelheim, European Union Innovative Medicine Initiative, Certa Therapeutics, RenalytixAI, Regeneron, Novo Nordisk, Sanofi, Dimerix, Travere and Vera Therapeutics. M.K. has received consulting fees through the University of Michigan from NovoNordisk, Alexion, Novartis, Roche Diagnostics and Vera Therapeutics. In addition, M.K. and V.N. have a patent, PCT/EP2014/073413 “Biomarkers and methods for progression prediction for chronic kidney disease” licensed. All other authors do not have conflicts to report.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nair, V., Minakawa, A., Smith, C. et al. Hypoxia inducible factor network reflects kidney disease progression in diabetes and sodium-glucose co-transporters inhibition. Sig Transduct Target Ther 11, 144 (2026). https://doi.org/10.1038/s41392-026-02653-0
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41392-026-02653-0