Abstract
Circulating levels of uric acid are influenced by a complex mix of intrinsic and environmental factors, including genetics, diet, and drugs. We analyzed levels of uric acid in a recent phase 3 clinical trial of patients with bipolar mania treated with 24 mg/day of the antipsychotic iloperidone or placebo. Initial results revealed that iloperidone treatment was associated with increases in uric acid from baseline (LS mean change (SE) of 27.2 (-4.93) μmol/L, compared with a change of 0.1 (-4.77) μmol/L for placebo group (LS mean difference (95% CI) = 27.1 (14.94, 39.20), p = <0.0001). Similar results were further observed in a previous phase 3 study of iloperidone treatment of schizophrenia. Pharmacogenetic analysis examining the urate transporter SLC2A9 revealed that iloperidone associated increases were linked to a genetic variant (rs7442295), correlating with both urate levels at baseline and in interaction with iloperidone vs placebo, and a pronounced increase of 35.9 μmol/L (0.67 mg/dL) was seen in iloperidone-treated patients homozygous for the for the rs7442295 (G) allele at the SLC2A9 gene, compared to a decrease of -16.5 (0.31 μmol/L in the corresponding GG placebo group (LS mean difference (95% CI) = 40.79 (14.61, 66.96, p = 0.0024). Further investigation suggested potentially clinically relevant sex differences associated with this variant. Specifically, male GG genotype patients exhibiting more frequent shifts from above the upper limit of normal for iloperidone-treated patients in comparison to female, AG/AA, and placebo groups. Overall, the mechanism of this iloperidone-induced increase in serum urate levels is likely due to decrease in clearance of urate through interaction with the SLC2A9 urate transporter protein. These results may hold clinical significance for patients treated with iloperidone.
Similar content being viewed by others
Introduction
Uric acid is the end product of purine metabolism in humans and primates, who lack the enzyme uricase to further oxidize the heterocyclic compound. Under normal conditions the urate anion is generated after breakdown of excess purine nucleosides in the liver and the majority of this moiety is then excreted through the kidneys via urine. Circulating levels of uric acid the blood (i.e., blood uric acid or serum urate) correlate with disease states including high blood pressure, metabolic syndrome, diabetes, gout, and cardiovascular disease [1]. Accordingly, serum uric acid testing is routinely conducted in clinical settings where the practice facilitates diagnosis of hyperuricemia, renal or kidney impairment, and rare inherited conditions [2].
The concentration of serum urate in an individual is influenced by multiple overlapping factors, including diet, medications, and genetic polymorphisms. For example, mutations identified in renal solute transporters have been identified as a major contributor to an individual’s baseline serum urate levels due to loss or gain of function in uric acid reabsorption [3, 4], whereas anti-tubercular drugs such as pyrazinamide increase reabsorption while decreasing secretion of uric acid, which can cause hyperuricemia and gout [5].
Several pharmacological interventions have been linked to clinically relevant changes in serum urate, for instance, use of low-dose acetylsalicylic acid (aspirin) on two consecutive days is associated with an increased risk of recurrent gout attacks [6]. In another example, diuretics, particularly of the thiazide class, are associated with an increase in serum urate levels, despite being indicated for treatment of hypertension [6]. These drug interactions represent direct mechanisms through which transport of uric acid can be impacted by drug interactions with various transporters and have led to inclusions of warnings and recommendations in physician prescribing guidelines regarding those drug’s risks for hyperuricemia and gout during treatment.
One critical player mediating circulating levels of uric acid is the integral membrane protein GLUT9, a high-capacity hexose-urate transporter encoded by the gene SLC2A9. GLUT9 is primarily expressed in the basolateral membrane kidneys, where it mediates reabsorption of uric acid from the proximal tubule into the bloodstream [7]. Genome-wide association studies previously identified associations between SLC2A9 and serum urate concentrations [8], including loss of function variants in SLC2A9 associated with elevated levels of serum urate [9]. Access to commercial genetic testing has consistently expanded, facilitating diagnosis of renal pathology in clinical settings [10], including for SLC2A9 variants associated with hyperuricemia [11]. Accordingly, understanding iatrogenic hyperuricemia in the context of such polymorphisms in these transporters can enable health care providers to tailor treatment to individual patients and improve outcomes through precision medicine.
Iloperidone is an antipsychotic medication approved by the FDA for the treatment of schizophrenia in 2009. Previous reports have documented increases in serum urate levels following treatment with drugs of the same pharmacological class, including olanzapine [12,13,14] and risperidone [15]. Here, we report analyses of blood uric acid concentrations obtained as part of routine clinical laboratory evaluations in two large, blinded, phase 3, randomized clinical trials of iloperidone treatment in two psychiatric patient populations (schizophrenia and bipolar mania, refer to Cutler et. al. 2008 [16] and Torres et. al. 2024 [17]). Iloperidone was associated with increases in serum urate compared to placebo or active comparator treatments in both studies. This observation led to investigation of genetic associations for this phenomenon and revealed iloperidone associated increases in serum urate are greatest in patients who were homozygous (GG) for the rs7442295 (G) allele at the SLC2A9 gene. In particular, iloperidone treated male patients with the GG genotype were observed to have serum urate concentrations above the upper limit of normal at study endpoint more frequently compared placebo group. Ultimately, these results support sex and rs7442295 genotype can be used define subgroups of patients with differential propensities for iatrogenic elevations in serum urate following iloperidone treatment, which may be clinically relevant for individuals with hyperuricemia.
Results
Iloperidone treatment was associated with serum urate increases in two placebo controlled phase 3 clinical studies
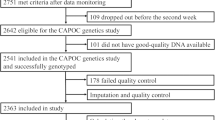
Clinical laboratory evaluations are routinely conducted during clinical trials and are relevant in populations of patients with neuropsychiatric disorders who are at increased risk of disorder associated comorbidities (e.g., obesity, diabetes, and metabolic syndrome) [18]. To evaluate the efficacy of the antipsychotic iloperidone, we conducted two separate, four-week, double-blinded, randomized, placebo controlled studies: one trial in acutely ill patients with schizophrenia [16] (hereafter, Study 3101) and, more recently, one trial in acutely ill patients with bipolar mania [17] (hereafter, Study 3201) (Fig. 1A). In Study 3101, eligible patients diagnosed with schizophrenia (n = 593) were randomized per protocol to iloperidone 24 mg/d given as 12 mg twice daily (n = 295), ziprasidone 160 mg/d given as 80 mg twice daily as an active control (n = 149), or placebo (n = 149). Overall, 193 (65%) iloperidone patients, 98 (66%) ziprasidone patients, and 90 (60%) placebo patients completed the study. Separately, in Study 3201, 414 participants diagnosed with bipolar I disorder were randomized and administered at least 1 dose of study medication (iloperidone 24 mg/d given as 12 mg twice a day, n = 206; placebo, n = 208). Overall, 139 (67.1%) iloperidone patients and 153 (72.9%) placebo patients completed the study. For demographic information of patients analyzed in each study, refer to Supplementary Table 1.
a Schematic diagram depicting the 4-week, randomized, blinded, clinical trial design/schema generalizable to both studies. b Mean change from baseline for serum urate for observed cases in Study 3201 and Study 3101. Error bars represent standard error of the mean (SEM). c Least squares (LS) mean difference for change in serum urate and p-values for ANCOVA between iloperidone and placebo. Refer to Tables 1 and 2 for a tabular presentation of numeric results of LS mean P-values, and 95% CIs.
Both trials had similar fixed designs: patients were titrated over approximately 1 week to a dose of 12 mg twice daily (24 mg/day) of iloperidone or placebo and treated at this dose for 3 consecutive weeks (Fig. 1A). Iloperidone showed a statistically significant benefit compared to placebo in each study’s respective primary endpoints, which measured reduction in symptoms from baseline associated schizophrenia (PANSS) or bipolar mania (YMRS). For the complete disposition, methods, and associated topline efficacy and safety results for each study, refer to Cutler et. al. 2008 [16] and Torres et. al. 2024 [17], respectively).
Analysis of clinical laboratory results for serum chemistry revealed iloperidone treated patients in both studies had statistically significant increases in serum urate from baseline to endpoint (day 28) as compared to placebo (and also for active control ziprasidone for Study 3101). (Fig. 1B and C, and Tables 1 and 2). Refer to the methods section of this work for relevant details on serum laboratory chemistry.
In Study 3201 iloperidone patients had 4-fold greater increases in serum urate from baseline to endpoint compared to placebo groups (33.4 μmol/L change for iloperidone and 4.2 μmol/L in placebo groups, respectively) (Table 1). This finding was replicated in Study 3101, with a roughly 4-fold greater increase in blood levels of uric acid for iloperidone compared to placebo treatment groups, and more than twice the change from baseline observed for iloperidone versus ziprasidone treated individuals (28.0 μmol/L, 4.2 μmol/L, and 13.1 μmol/L for iloperidone, placebo, and ziprasidone groups, respectively) (Table 2).
In both studies, significant differences in uric acid for iloperidone vs placebo groups were detectable as early as 14 days (Tables 1 and 2). At endpoint (day 28) in Study 3201, blood uric acid LS mean change was 27.2 μmol/L for iloperidone groups and 0.1 μmol/L for placebo groups (ANCOVA, p < 0.0001, adjusting for covariates) (Fig. 1C, Table 1). Similarly, LS mean change in Study 3101 was 28.0 μmol/L for iloperidone treatment group compared to 4.2 μmol/L in placebo group (p = 0.0014, Fig. 1C, Table 2), providing further confirmatory evidence that acute iloperidone treatment is associated with increases in circulating levels of uric acid.
SLC2A9 variant rs7442295 was associated with different baseline levels of serum urate in two separate psychiatric patient populations
We routinely collect genetic information during our clinical trials for exploratory pharmacogenetic (PG) analysis, and patients in both studies provided blood samples for DNA extraction and genetic analysis. In Study 3201, whole blood PG samples were collected with written consent, and whole genome sequencing was conducted for 367/414 (88.6%) of patients with bipolar I disorder in the safety population.
Genetic variants in renal transport proteins can affect rates of uric acid reabsorption and secretion, impacting levels of serum urate in various genetic subgroups in the general population [8]. We hypothesized that genes encoding transporter proteins may be associated with observed baseline levels of serum urate in our patients as well.
To characterize the relationship between patient genetic compositions and serum urate levels prior to treatment, we analyzed the results from Study 3201 using linear regression model for serum urate, adjusting for age, sex, and principal components. One of the most significant variants identified was rs7442295 (p-value 10-5 (beta = -23.12) (Fig. 2A). A similar relationship was generally observed for association of this allele with baseline uric acid levels in Study 3101 (Supplementary Figure 1), in which we acquired PG blood samples with written consent and determined genotypes for this variant via 500 K SNP Gene Chip Human Mapping array.
Genotype status of urate transporter SLC2A9 variant RS7442295 (a) Baseline uric acid by RS7442295 genotype in Study 3102. b Baseline uric acid by RS7442295 sex and genotype in Study 3102. Scatter plot dots show baseline uric acid concentrations of individuals, and median is depicted by red line. Results were generally similar in Study 3101 (refer to Supplementary Figure 1). c Major allele frequencies for SLC2A9 variant rs7442295 according to genetically determined racial groups. Calculated from data retrieved from GnomeAD.
Overall, our results are consistent with prior reports associating variant rs7442295 and baseline serum urate levels, including the observation of an intermediary phenotype in heterozygous individuals [9].
Interestingly, male carriers in Study 3201 were broadly observed to have a higher baseline level of uric acid compared to females (beta = 24.08 per G allele, p = 0.04) (Fig. 2B), consistent with prior work supporting a sex-specific effect for this variant [19].
Additionally, we performed a full GWAS on these samples, and SLC2A9 variants were amongst the strongest loci detected.
The rs7442295 variant has a global MAF (GnomAD) of 0.24 with the highest MAF reported in patients of African/African American ancestry (MAF = 0.40), which was nearly twice the MAF reported for non-Finnish Europeans of (MAF = 0.20) (Fig. 2C). Variant rs7442295 is in a strong LD with a coding nonsynonymous variant rs16890979 (Val253Ile), shown in which was previously associated with serum uric acid levels where the minor allele, rs16890979 T, was associated with a decrease of 0.47 mg/dl in the uric acid level (CI: 0.31-0.63, p = 1.43 x 10e-11) [20]. Refer to Supplementary Figure 2 for a PCA plot depicting ancestry of studied cases as shown on the first 2 PC components.
Variant rs7442295 was associated with changes in serum urate levels in iloperidone treated individuals
Applying pharmacogenetic analysis to predict the clinical effects of drugs in different patient subgroups remains a critical goal in pharmaceutical research and development. Accordingly, based on the strong associations detected for SLC2A9 variants at baseline, we hypothesized that there may be a relationship between carrier status of variant rs7442295 and serum urate changes observed during iloperidone treatment. We analyzed clinical laboratory results according to genotype status in Study 3201 for variant rs7442295. Remarkably, for iloperidone-treated patients homozygous for the rs7442295 (G) allele in Study 3201, we observed a pronounced increase of 40.9 μmol/L (0.674 mg/dL) compared to a decrease of -16.86 μmol/L in the corresponding GG placebo group (Fig. 3A Fig. 3B and Table 3).
A Violin plot showing individual and mean (SEM) change from baseline according to carrier status for variant rs7442295 and treatment group in Study 3201. B Mean (SEM) serum urate levels at baseline and EOS (day 28) according to carrier status for variant rs7442295 and treatment group in Study 3201. C Scatter plot showing individual and median serum urate levels at baseline (black symbols) and EOS (day 28) (white symbols) for patients with GG genotype for variant rs7442295, according to sex, and treatment group in Study 3201. Median depicted by red lines.
Given we observed a sex effect for genotype and baseline serum urate levels, we further analyzed the effect of this variant on serum urate change from baseline in Study 3201, using linear regression models stratified by sex and treatment. Changes from baseline in serum urate in individuals with the GG genotype were particularly pronounced in the male subgroup (Fig. 3C and Table 4), with a 65.61 μmol/L increase in serum urate in iloperidone treated GG males compared to a -37.35 μmol/L decrease in the corresponding placebo group, further supporting a sex effect for males with the GG genotype.
To address the extent to which iloperidone treatment interacts with these different subgroups, we analyzed the effect of this variant on serum urate change from baseline using linear regression models stratified by sex and treatment. Among male placebo group, each additional G allele increased the serum urate change by 1.82 μmol/L. This effect was dramatically larger in the iloperidone treated subgroup, with an increase of 27.64 μmol/L (p-value = 1.54e-3).
Since sex was not balanced in this study, we conducted sample matching to fit a linear regression model in iloperidone group with a gene-by-sex interaction term. In iloperidone group males, each additional rs7442295 G allele increased the urate change by 29.13 (p-value = 5.83e-3).
Clinically relevant serum urate levels following iloperidone treatment, according to genotype, drug, and sex
Females have lower circulating levels of uric acid compared to males; a phenomenon hypothesized to be driven by differences in the sex hormone estradiol [21], and sex differences in serum urate are reflected in laboratory ranges used to evaluate clinical laboratory results. The lab specified upper limits of normal (ULN) were >453 μmol/L (7.62 mg/dL) for males and >394 μmol/L (6.46 mg/dL) for female patients in Study 3201, in alignment with clinical guidelines for interpreting uric acid blood tests for hyperuricemia defined as approximately 6 mg/dL for women and 7 mg/dL for men [22]. A key goal of clinical research is to characterize the safety profile of medications studied, including which patient subgroups have differential risk for adverse events. Accordingly, to further characterize the clinical relevance of serum urate changes following iloperidone treatment, we evaluated the proportions of patients who had concentrations of serum urate exceeding ULN at baseline compared to endpoint in Study 3201 according to treatment, sex, and genotype (Table 5). Subgroups of iloperidone treated patients had concentrations of serum urate above the upper level of normal range more frequently than placebo.
Consistent with the results for a strong interaction with G-allele and male subgroups, iloperidone-associated increase in serum urate above the upper limit normal (ULN) was largest in males with homozygous (GG) genotype for rs7442295 (Table 5). Treatment with iloperidone resulted in an approximately 4-fold increase in the proportions of GG males with lab results above the ULN for serum urate (baseline, 8.3%; EOS, 33.3%, respectively). Conversely, the percentage of placebo treated GG males above the ULN decreased (baseline, 14.3%; EOS, 0.0%, respectively). Consistent with the compounding effect for G allele for the variant observed in previous sections, proportions of patients with laboratory values above the ULN increased for iloperidone treated with AG genotype (baseline, 2.4% and EOS ~ 13.5%), while the proportion of patients in the corresponding placebo group remained essentially the same (baseline, 14.9%; EOS, 15.5%). Analysis using Fisher’s exact test combining GG + AG genotypes vs baseline was significant for iloperidone versus placebo groups, (p-value = 0.03, OR = 5.73), supporting that GG patients are more likely to be above the ULN following four weeks of treatment.
Discussion
We report here that iloperidone treatment was associated with statistically significant increases in uric acid compared to placebo groups in two independent clinical trials, conducted in two different psychiatric patient populations.
In both studies, participants received 24 mg/day (12 mg b.i.d.) iloperidone, the highest therapeutic dosage currently approved for acute and maintenance treatment of schizophrenia [23]. Increases in serum urate levels were nearly twice as large in iloperidone-treated patients homozygous for the common variant rs7442295 (G) in the SLC2A9 transporter gene. The GG genotype is carried by 7.6% of the general population, where it is most common in African/African Americans (16.5%) and in Latino/Admixed Americans (14.2%), while being less common non-Finnish Europeans (3.9%), and nearly absent in South and East Asians (0%).
Based on the existing study data this observed effect of increased serum urate in the GG genotype population in association with iloperidone is particularly pronounced in males. Male patients are observed to exceed the upper limit normal after iloperidone treatment more frequently for individuals that are homozygous (GG) for the rs7442295 (G) allele at the SLC2A9 gene and had 4-fold higher increases in serum urate compared to placebo treated groups.
We observe a baseline effect consistent with the effect previously reported in literature by Wallace et al. where in 1900 hypertensive Caucasian Europeans, the more common allele rs7442295(A) allele was associated with higher serum urate and hyperuricemia with a reported odds ratio of 1.89 (CI: 1.36-2.61, p = 5 ×10-5) [24]. Overall, 79% of Europeans carry one or two alleles leading to higher serum urate levels. Each copy leads to an average increase of 20 μmol/L, and an approximate doubling of risk for hyperuricemia [8].
There are many factors which influence reabsorption and excretion of uric acid in the blood, including the GLUT9 protein. Wu et al., 2024 reported reduced absorption of uric acid in kidney organoids with rs16890979 mutants; conversely, uric acid absorption was consistently increased in organoids with SLC2A9 overexpression and decreased in those with SLC2A9 knockdown [20]. These results support that GLUT9 is responsible for uric acid absorption into kidney cells and highlights rs16890979 (and rs7442295 in strong LD) as notable variants that may compromise mechanisms of uric acid transport in the renal system. Based on our findings, iloperidone-induced increase in serum urate levels is likely due to a decrease in clearance of urate through interaction with the SLC2A9 gene encoding for this urate transporter protein.
A notable limitation of this work is incomplete elucidation of a molecular mechanism through which iloperidone impacts these targets (including following exposure beyond the acute 4-week treatment evaluated in these studies). Additionally, we note results for Study 3101 are in part limited by the unavailability of whole genome sequencing at the time of the study conduct and an imbalance in the ratio of male/female patients enrolled. Future work should explore potential drug-protein interactions between iloperidone and GLUT9 isoforms based on this variant, as well as investigation of the long term impacts of iloperidone exposure on serum urate levels in patients. In particular, in-vitro evaluation of iloperidone treatments on iPSC based model systems and spheroid organoids may provide further insights into the molecular underpinnings of the drug’s role in this system.
Commercial tests are available to determine patient genotypes for this variant and could support physician decisions in individuals with preexisting hyperuricemia. Ultimately, our results hold clinical relevance for patients receiving iloperidone treatment – persons with a history of gout and carriers of the variants of interest can be monitored for serum urate levels, as these patients may need to increase the doses of their urate lowering treatments.
Materials and methods
Study VP-VYV-683-3201 (bipolar mania)
ClinicalTrials.gov identifier: NCT04819776; EudraCT: 2020–000405-83
Study 3201 was a phase 3, randomized, double-blind, placebo-controlled study conducted at 27 US and international sites between April 2021 and September 2022. The study’s objective was to evaluate the efficacy and safety of a fixed dose of iloperidone in patients with bipolar I disorder experiencing symptoms of mania. The study enrolled acutely ill male and female patients between 18–65 years of age who had a diagnosis of bipolar I disorder, with or without mixed features, in accordance with DSM-5 criteria, as confirmed by the Structured Clinical Interview for Diagnostic and Statistical Manual of Mental Disorders, Fifth Edition. Participants were randomized 1:1 to iloperidone (24 mg/d given twice daily) or placebo for 4 weeks. In accordance with US Prescribing Information for FDA the approved medication Fanapt (iloperidone) oral tablets, participants who were poor metabolizers of CYP2D6 received iloperidone 12 mg/d given twice daily.
The study protocol and all amendments were reviewed by the Independent Ethics Committee or Institutional Review Board for each center. The study was conducted according to the ethical principles of the Declaration of Helsinki. Informed consent was obtained from each patient in writing before any study-specific procedures were performed.
For the complete methods, safety, and efficacy results, refer to:
Torres, R., Czeisler, E. L., Chadwick, S. R., Stahl, S. M., Smieszek, S. P., Xiao, C., … & Polymeropoulos, M. H. (2024). Efficacy and Safety of Iloperidone in Bipolar Mania: A Double-Blind, Placebo-Controlled Study. The Journal of Clinical Psychiatry, 85(1), 51087.
Serum chemistry sample collection (Bipolar Mania Study 3201)
Serum urate was measured as part of routine safety testing by a central laboratory. Using kits provided by a central laboratory, blood samples for serum chemistry analyses were collected on day 0 (baseline), days 7, 14, 21, and on day 28 (endpoint) of the study. Serum samples for analytical chemistry was obtained via venipuncture and collection of whole blood into 7.5 ml serum separation tubes (SSTs). Samples were inverted 8 times and allowed to clot for 30 to 60 min, after which samples were immediately centrifuged for 10 min at 1300 g or until a gel barrier had formed. Serum samples were shipped to the central laboratory at ambient temperature on the same day where feasible, otherwise samples were stored refrigerated until shipping at ambient temperature on the following business day.
Pharmacogenetic sample collection and whole genome sequencing (Bipolar Mania Study 3201)
Samples for pharmacogenetic analysis were obtained with written consent during screening visits using kits provided by a central laboratory. Whole blood samples were obtained via venipuncture and collection of whole blood into a 10 ml K2 EDTA tube, inverted 8 times, and immediately frozen and stored at -70 deg C until shipping on dry ice to the central laboratory. Following shipping to the central pharmacogenetics laboratory, genotype status was obtained through whole genome sequencing, using NovaSeq X performed by New York Genome Center (NYGC) in accordance with Good Clinical Laboratory Practices (GCLP) standards. Following isolation of DNA/RNA for downstream sequencing applications, nucleic acid samples were quantified using fluorescent-based assays (PicoGreen) to accurately determine whether sufficient material was available for library preparation and sequencing. DNA sample size distributions were profiled by a Fragment Analyzer (Advanced Analytics) or BioAnalyzer (Agilent Technologies), to assess sample quality and integrity.
Whole genome sequencing (WGS) libraries were prepared using the Truseq DNA PCR-free Library Preparation Kit. Paired-end 150 bp reads were aligned to the GRCh37 human reference (BWA-MEM v0.7.8) and processed with GATK best-practices workflow (GATK v3.4.0). The mean coverage was 35.8, it reflects the samples average.
A genotype missing rate threshold of 0.1 was applied to filter out low-quality samples, but no individuals were removed. Variants were filtered based on a MAF threshold of 0.01 and a Hardy-Weinberg exact test p-value cutoff of 1e-7. Between-sample relatedness was assessed using KING software, and individuals with a relatedness degree of second-degree or closer were excluded from the analysis. 9642385 variants and 446 samples remain after QC. PCA were then performed and a plot of PC1 and PC2 can be found in the Supplementary Figures. All the QC and PCA were done using PLINK2.
All high quality variants obtained from GATK were annotated for functional effects (intronic, intergenic, splicing, nonsynonymous, stopgain and frameshifts) based on RefSeq transcripts using Annovar 31. Additionally, Annovar was used to match general population frequencies from public databases (Exac, gnomAD, ESP6500, 1000 g).
Linear models adjusted for 3 PCs, age and sex were conducted in PLINK. Linear regression was used for all genetic association analysis, with adjustment of participants’ age, sex, and genetically derived principal components (PCs), wherever they are available. Genotypes were additively coded as the count of the alternative allele to represent a dosage effect. Gene-by-sex and gene-by-treatment analyses were conducted by adding a genotype-sex or genotype-treatment 2-way interaction term into the regression model. All regression analyses were performed using R statistical language (version 4.3.2).
Since sex ratio was not well balanced in this study, a sample matching by sex was performed prior to the analysis stratified by treatment. The sample matching step was done by using R package Matchit with nearest neighbor matching method and a ratio of 1 to ensure a balanced sex ratio.
Study VP-VYV-683-3101 (Schizophrenia)
ClinicalTrials.gov identifier: NCT00254202
Study 3101 was a phase 3, randomized, double-blind, placebo-controlled study conducted at 35 centers in the United States and 9 in India between November 2005 and May 2007. The study’s objective was to evaluate the efficacy and safety of a fixed dose of iloperidone in patients with acute exacerbations of schizophrenia. The study enrolled acutely ill male and female patients between 18-65 years of age who had a diagnosis of schizophrenia according to Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition (DSM-IV). Participants were randomized to receive iloperidone 24 mg/d given twice daily, ziprasidone 160 mg/d given twice daily as an active control, or placebo in a 2:1:1 ratio.
Study protocol was approved by appropriate institutional review boards or independent ethics committees. This study was conducted in accordance with the Declaration of Helsinki, the US Code of Federal Regulations governing the protection of human subjects and obligations of clinical investigators, and Good Clinical Practice and International Conference on Harmonisation (ICH) guidelines. Voluntary written informed consent was obtained from each patient before the performance of any study procedures.
For the complete methods, safety, and efficacy results, refer to:
Cutler, A. J., Kalali, A. H., Weiden, P. J., Hamilton, J., & Wolfgang, C. D. (2008). Four-week, double-blind, placebo-and ziprasidone-controlled trial of iloperidone in patients with acute exacerbations of schizophrenia. Journal of clinical psychopharmacology, 28(2), S20-S28.
Serum chemistry sample collection (Schizophrenia Study 3101)
Serum urate was measured as part of routine safety testing by using kits provided by a central laboratory. Blood samples for serum chemistry analyses were collected on day 0 (baseline), day 14, and on day 28 (endpoint) of the study. Serum samples for analytical chemistry was obtained via venipuncture and collection of whole blood into 6 or 10 ml serum separation tubes (SSTs) and allowed to clot for 30 to 60 min, after which samples were immediately centrifuged for 10 to 15 min at approximately 1000 g. Serum samples were aliquoted into 2 ml tubes and shipped to the central laboratory at ambient temperature on the same day where feasible, otherwise samples were stored refrigerated until shipping at ambient temperature on the following business day.
Pharmacogenetic sample collection and genotyping (Schizophrenia Study 3101)
Samples for pharmacogenetic analysis were collected on an optional basis and separate written informed consent was obtained from participants prior to collection of PG samples. PG samples were typically obtained during baseline visits but could be collected on a subsequent visit if PG written consent was obtained. Using kits provided by a central laboratory, whole blood samples were obtained via venipuncture and collection of whole blood into a 3 ml K2 EDTA tube, inverted 10 times, and immediately frozen and stored at -80 deg C (or -20 deg C only if -80 deg C was unavailable) until shipping on dry ice to the central laboratory. Patient were genotyped using a Affymetrix 500k SNP Gene Chip Human Mapping Array.
Statistical Analysis
Analysis of covariance (ANCOVA) was conducted using SAS. LS means, CIs, and p-values were based on ANCOVA mixed effects model with main effects of treatment group and pooled site, and baseline as a covariate. All p values represent the difference between iloperidone and placebo.
Structural 3D Representation of GLUT9 Protein
Structural representation of the impact of rs16890979 mutation on the GLUT9 protein was created using 3DVizSNP [25] and iCn3D [26] to predict and render images in (Supplementary Figure 1).
Data availability
Vanda is committed to sharing with qualified external researchers access to patient- level data and supporting clinical documents from eligible studies. All data provided are anonymized to respect the privacy of patients who have participated in the trial in line with applicable laws and regulations. Contact the corresponding author regarding data requests.
References
Richette P, Bardin T. Gout. Lancet. 2010;375:318–28. https://doi.org/10.1016/S0140-6736(09)60883-7
Mandell BF. Clinical manifestations of hyperuricemia and gout. Cleve Clin J Med. 2008;75:S5–8. https://doi.org/10.3949/ccjm.75.suppl_5.s5
Dalbeth N, Stamp LK, Merriman TR. The genetics of gout: Towards personalised medicine? BMC Med. 2017;15:108 https://doi.org/10.1186/s12916-017-0878-5
Sun HL, Wu YW, Bian HG, Yang H, Wang H, Meng XM, et al. Function of Uric Acid Transporters and Their Inhibitors in Hyperuricaemia. Front Pharmacol. 2021;12:667753 https://doi.org/10.3389/fphar.2021.667753
Ben Salem C, Slim R, Fathallah N, Hmouda H. Drug-induced hyperuricaemia and gout. Rheumatology (Oxford). 2017;56:679–88. https://doi.org/10.1093/rheumatology/kew293
Raja R, Kavita F, Amreek F, Shah A, Sayeed KA, Sehar A. Hyperuricemia Associated with Thiazide Diuretics in Hypertensive Adults. Cureus. 2019;11:e5457 https://doi.org/10.7759/cureus.5457
Chung S, Kim GH. Urate Transporters in the Kidney: What Clinicians Need to Know. Electrolyte Blood Press. 2021;19:1–9. https://doi.org/10.5049/EBP.2021.19.1.1
Köttgen A, Albrecht E, Teumer A, Vitart V, Krumsiek J, Hundertmark C, et al. Genome-wide association analyses identify 18 new loci associated with serum urate concentrations. Nat Genet. 2013;45:145–54. https://doi.org/10.1038/ng.2500
Tin A, Schlosser P, Matias-Garcia PR, Thio CHL, Joehanes R, Liu H, et al. Epigenome-wide association study of serum urate reveals insights into urate co-regulation and the SLC2A9 locus. Nat Commun. 2021;12:7173 https://doi.org/10.1038/s41467-021-27198-4
Aron AW, Dahl NK, Besse W. A Practical Guide to Genetic Testing for Kidney Disorders of Unknown Etiology. Kidney360. 2022;3:1640–3. https://doi.org/10.34067/KID.0007552021
Li L, Zhang Y, Zeng C. Update on the epidemiology, genetics, and therapeutic options of hyperuricemia. Am J Transl Res. 2020;12:3167–81. PMID: 32774692
Tohen M, Kryzhanovskaya L, Carlson G, DelBello M, Wozniak J, Kowatch R, et al. Olanzapine versus placebo in the treatment of adolescents with bipolar mania. Am J Psychiatry. 2007;164:1547–56. https://doi.org/10.1176/appi.ajp.2007.06111932
Tohen M, Vieta E, Goodwin GM, Sun B, Amsterdam JD, Banov M, et al. Olanzapine versus divalproex versus placebo in the treatment of mild to moderate mania: a randomized, 12-week, double-blind study. J Clin Psychiatry. 2008;69:1776–89. https://doi.org/10.4088/JCP.v69n1113
Kryzhanovskaya L, Schulz SC, McDougle C, Frazier J, Dittmann R, Robertson-Plouch C, et al. Olanzapine versus placebo in adolescents with schizophrenia: a 6-week, randomized, double-blind, placebo-controlled trial. J Am Acad Child Adolesc Psychiatry. 2009;48:60–70. https://doi.org/10.1097/CHI.0b013e3181900404
Vanwong N, Srisawasdi P, Ngamsamut N, Nuntamool N, Puangpetch A, Chamkrachangpada B, et al. Hyperuricemia in Children and Adolescents with Autism Spectrum Disorder Treated with Risperidone: The Risk Factors for Metabolic Adverse Effects. Front Pharmacol. 2016;7:527 https://doi.org/10.3389/fphar.2016.00527
Cutler AJ, Kalali AH, Weiden PJ, Hamilton J, Wolfgang CD. Four-week, double-blind, placebo- and ziprasidone-controlled trial of iloperidone in patients with acute exacerbations of schizophrenia. J Clin Psychopharmacol. 2008;28:S20–S28. https://doi.org/10.1097/JCP.0b013e318169d4ce
Torres R, Czeisler EL, Chadwick SR, Stahl SM, Smieszek SP, Xiao C, et al. Efficacy and Safety of Iloperidone in Bipolar Mania: A Double-Blind, Placebo-Controlled Study. J Clin Psychiatry. 2024;85:23m14966 https://doi.org/10.4088/JCP.23m14966
Forty L, Ulanova A, Jones L, Jones I, Gordon-Smith K, Fraser C, et al. Comorbid medical illness in bipolar disorder. Br J Psychiatry. 2014;205:465–72. https://doi.org/10.1192/bjp.bp.114.152249
Döring A, Gieger C, Mehta D, Gohlke H, Prokisch H, Coassin S, et al. SLC2A9 influences uric acid concentrations with pronounced sex-specific effects. Nat Genet. 2008;40:430–6. https://doi.org/10.1038/ng.107
Wu S, Li C, Li Y, Liu J, Rong C, Pei H, et al. SLC2A9 rs16890979 reduces uric acid absorption by kidney organoids. Front Cell Dev Biol. 2023;11:1268226 https://doi.org/10.3389/fcell.2023.1268226
Wang Y, Charchar FJ. Establishment of sex difference in circulating uric acid is associated with higher testosterone and lower sex hormone-binding globulin in adolescent boys. Sci Rep. 2021;11:16983 https://doi.org/10.1038/s41598-021-96959-4
George C, Leslie SW, Minter DA. Hyperuricemia. StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2023. Available from: https://www.ncbi.nlm.nih.gov/books/NBK459218/
Vanda Pharmaceuticals Inc. Fanapt (iloperidone) tablets [package insert]. 2024. https://www.accessdata.fda.gov/drugsatfda_docs/label/2024/022192s023lbl.pdf
Wallace C, Newhouse SJ, Braund P, Zhang F, Tobin M, Falchi M, et al. Genome-wide association study identifies genes for biomarkers of cardiovascular disease: serum urate and dyslipidemia. The American Journal of Human Genetics. 2008;82:139–49. https://doi.org/10.1016/j.ajhg.2007.11.001
McLaren W, Gil L, Hunt SE, Riat HS, Ritchie GRS, Thormann A, et al. The Ensembl Variant Effect Predictor. Genome Biol. 2016;17:122 https://doi.org/10.1186/s13059-016-0974-4
Wang J, Youkharibache P, Zhang D, Lanczycki CJ, Geer RC, Madej T, et al. iCn3D, a web-based 3D viewer for sharing 1D/2D/3D representations of biomolecular structures. Bioinformatics. 2020;36:131–5. https://doi.org/10.1093/bioinformatics/btz502
Acknowledgements
Vanda Pharmaceuticals and the authors thank the patients, study sites, and investigators who participated in these clinical trials. Poster presented at the ASHG meeting, Washington, DC, September 2023 and ASCPP Meeting, Miami, May 2024
Funding
This study was funded by Vanda Pharmaceuticals, Inc. (Washington, DC).
Author information
Authors and Affiliations
Contributions
S.S made the initial discovery. S.S., S.C., E.C., H.B., and C.X. analyzed the data. S.S. and S.C. created the figures and wrote the manuscript with input from the other authors. R.T. and E.C. conceived and conducted Study 3201 with input from C.P., G.B., and M.P., who oversaw the research and study conduct. All authors were involved in writing and editing the manuscript. S.S. and S.C. contributed equally to the manuscript.
Corresponding author
Ethics declarations
Competing interests
Authors are employees of Vanda Pharmaceuticals Inc.
Ethics
The study protocols and all amendments were reviewed by the Independent Ethics Committee or Institutional Review Board for each center in each study. The studies from which samples were obtained was conducted according to the ethical principles of the Declaration of Helsinki and all methods were carried out in accordance with relevant guidelines and regulations. Informed consent was obtained from all participants and/or their legal guardian in writing before any study-specific procedures were performed.
Role of the sponsor
The sponsor was responsible for the design, analysis, interpretation, and publication of the clinical trials. Final approval for the decisions to submit the manuscript was the sole decision of the authors.
Relevant financial relationships
Drs Smieszek, Chadwick, Bai, Torres, Xiao, C. Polymeropoulos, M. Polymeropoulos, Mr Birznieks and Ms Czeisler are full time employees of Vanda Pharmaceuticals Inc. and may hold company stock and/or stock options.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Smieszek, S.P., Chadwick, S.R., Czeisler, E.L. et al. A potential association of SLC2A9 variant rs7442295 with uric acid at baseline and in interaction with iloperidone. Pharmacogenomics J 26, 10 (2026). https://doi.org/10.1038/s41397-026-00402-8
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41397-026-00402-8