Abstract
Transcriptomic studies of post-mortem brain samples in schizophrenia (SCZ) and bipolar disorder (BD) have primarily focused on messenger RNAs (mRNAs) but have given limited attention to small non-coding RNAs (sncRNAs). In this study, we present our analyses of sncRNA profiles from the prefrontal cortex of SCZ and BD cases and controls (53 SCZ cases, 40 BD cases, 77 controls), which we sourced from the Icahn School of Medicine at Mount Sinai and the NIMH Human Brain Collection Core brain banks. Corresponding mRNA-seq data were obtained from the CommonMind Consortium. Using a state-of-the-art pipeline, we mapped reads and determined differentially abundant and co-expressed sncRNAs and mRNAs, adjusting for known and hidden confounders. Across samples, 98% of all sncRNAs comprised miRNA isoforms (60.6%), tRNA-derived fragments (17.8%), rRNA-derived fragments (11.4%), and Y RNA-derived fragments (8.3%). In SCZ, 15% of the identified sncRNAs exhibited significant fold changes (FCs), with many also altered in BD, albeit to a lesser extent. For miRNAs, the FCs correlated strongly with the presence of non-templated nucleotides to their 3′-ends, independently of miRNA identity or locus of origin. Disease- and age-associated sncRNAs and mRNAs revealed accelerated aging in both SCZ and BD. Co-expression analyses also revealed, for the first time, disease-independent associations of many isomiRs, tRFs, rRFs, and yRFs with critical brain processes. These findings suggest complex and previously uncharacterized roles for novel classes of regulatory sncRNAs in synaptic signaling, neurogenesis, memory, behavior, and cognition.
Similar content being viewed by others
Introduction
Schizophrenia (SCZ) and bipolar disorder (BD) are complex neuropsychiatric disorders that significantly impact cognitive and emotional functioning. Transcriptomic studies of post-mortem brain tissues have been instrumental in highlighting alterations in gene expression in affected brains, particularly in pathways associated with synaptic signaling, neurogenesis, and neurotransmitter systems [1, 2]. Additionally, genome-wide association studies (GWAS) identified numerous risk loci associated with these disorders, underscoring the polygenic nature of their etiology [3]. Despite these advances, the heritability explained by GWAS is limited.
Small non-coding RNAs (sncRNAs) have regulatory roles and are a sizable component of an organism’s transcriptome [4]. Even though the universe of known sncRNAs outnumbers that of protein-coding genes by two orders of magnitude, sncRNAs remain largely uncharacterized outside the cancer context [5]. MicroRNAs (miRNAs) are the best-studied sncRNA class. A mere 18–22 nucleotides (nts) in length, miRNAs bind their mRNA or long non-coding RNA targets in a sequence-dependent manner [4, 6].
Several previous SCZ studies used microarrays or PCR array analyses to identify differentially abundant (DA) miRNAs between post-mortem brain samples of cases and controls [7,8,9,10,11,12,13] and the reader is referred to recent excellent reviews on this topic [14, 15]. Those studies showed limited concordance, presumably due to factors that can affect miRNA expression analysis, including differences in profiling platforms [16], batch effects [17], differing clinical and demographic variables (e.g., ethnicity, sex, age) [18, 19], and differences in the analyzed brain regions [20]. RNA-seq has superseded these earlier methods, offering improved sensitivity, specificity, data-driven normalization, and an unbiased characterization of a sample’s transcriptome [21,22,23].
An early success of RNA-seq was the discovery of miRNA isoforms (isomiRs) [24], which are co-expressed mature miRNAs arising from the same miRNA arm and having distinct sequences [4]. IsomiRs typically differ from one another, and from the miRbase [25] “reference” sequence, by a few nts at either their 5´-end or 3´-end. IsomiRs that match the source genome exactly are called “templated” and produced by a systematic and regulated cleavage of miRNA precursor hairpins by DROSHA and DICER [26]. “Non-templated” isomiRs are a rich emerging category produced through post-transcriptional nucleotide additions to their 3´-ends [26, 27]. IsomiRs from the same miRNA locus, whether templated or non-templated, can target different genes [26, 28, 29] and exhibit distinct subcellular localizations [30].
The universe of regulatory sncRNAs expanded in recent years with the inclusion of the tRNA-derived fragments (tRFs) [31], the rRNA-derived fragments (rRFs) [32], and the Y RNA-derived fragments (yRFs) [4, 33], all of which have yet to be characterized systematically. We previously showed that the abundance of these classes depends on “context” (e.g., tissue type, disease) [33,34,35] and “personal attributes” (e.g., ancestry and sex) [18, 36, 37], making these sncRNAs ideal candidates for exploration of their contributions to health disparities.
In this study, we comprehensively characterized sncRNAs in post-mortem brain samples of SCZ cases, BD cases, and controls. We ensured robustness by including 170 brain samples from two independent brain banks. We also identified which sncRNAs are differentially abundant (DA) between cases and controls. Finally, through co-expression analysis that followed our regressing out the disease dimension, we identified specific classes of sncRNAs that form co-expression modules with protein-coding genes from critical brain processes.
Materials and methods
We provide a summary here. For more details, please refer to the Supplement.
Brain samples
We analyzed 53 SCZ, 40 BD, and 77 control subjects who met the relevant DSM-IV diagnostic criteria. Tissue donors were sourced from the Icahn School of Medicine at Mount Sinai (MSSM) and the NIMH Human Brain Collection Core (HBCC). See Supplementary Table S1 for sample-level information.
RNA isolation, library preparation, sequencing, and mapping
We isolated RNA using miRNeasy kits (Qiagen) and assessed it using a 2200 TapeStation (Agilent Technologies). All samples had RIN ≥ 6 (Supplementary Table S1). We prepared sequencing libraries and sequenced all samples on an Illumina NextSeq 500, as previously described [30]. We used isoMiRmap [38] and MINTmap [39] to profile isomiRs and tRFs, a brute-force search [33] to profile rRFs, yRFs, and other repetitive fragments (rpFs), and bowtie v1.3.1 [40] to place any remaining unmapped reads to the reference GRCh38.p14 genome.
mRNA-seq data
We obtained the corresponding transcript-level read count data from the CommonMind Consortium [2], discarding samples with RIN values less than 6, which left 161 samples for mRNA analysis.
Differential abundance, enrichment, and co-expression analysis
We used DESeq2 v1.44 to identify sncRNAs and mRNAs that were DA between cases and controls [41]. We normalized the counts from the HBCC and MSSM samples separately using DESeq2’s algorithm, and winsorized them to remove the impact of outliers. Because we found significant correlations primarily associated with disease status, age at death, PMI, and RIN (Supplementary Fig. S1), we adjusted for confounders before the DA analysis – see Supplement. We controlled for potential hidden confounding factors using the SVA v3.52 package [42]. We controlled for false discovery using the Benjamini-Hochberg procedure. We generated gene set enrichments with GSEA [43]. For the co-expression analysis, we removed known and hidden confounders with CorrAdjust [44], which outperforms previous state-of-the-art tools. Supplementary Fig. S2 shows the example of two genes, KDM5D and RPS4Y1, whose correlation changes dramatically when adjusted for sex, which acts as a confounder variable – see Supplement for more information.
Results
IsomiRs, tRFs, rRFs, and yRFs abound in the brains of cases and controls
We generated sncRNA sequencing data from post-mortem brain samples of 53 SCZ and 40 BD cases, and 77 controls. We mapped and comprehensively annotated all sequenced reads using a state-of-the-art analytical pipeline (see Methods). First, we profiled isomiRs, tRFs, rRFs, yRFs, and repetitive fragments (rpFs), enforcing exact matching. Next, we identified those remaining reads whose Levenshtein distance (LD) from at least one annotated sncRNA was ≤ 2. Finally, we mapped all remaining reads, allowing at most one replacement but no insertions or deletions.
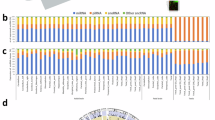
Our analysis only considered the 7,473 mapped sncRNAs whose abundance is ≥ 10 RPM in ≥ 25% of the samples. Figure 1A shows the distribution of the identified sncRNAs in each category. An additional entry comprises unmapped molecules. Individual molecules within each sncRNA type are sorted by mean abundance. The X-axis captures the molecules’ relative ranking within their own group, whereas the Y-axis shows their cumulative abundance (mean ± SD). We analyzed all the samples together since the total abundance of each sncRNA class showed no statistically significant differences between cases and controls.
A Within each molecule type, sncRNAs are ranked by mean RPM across 170 samples (rank is captured by increasing X values), whereas the cumulative mean (solid line) ± standard deviation (dashed line) are shown on the Y-axis. “Other” molecules include rpFs (see Methods). B Histograms of sncRNA lengths for each molecule type (% values sum up to 100% within each molecule type). IsomiRs, tRFs, rRFs, and yRFs include wild-type and LD ≤ 2 sncRNAs.
By profiling isomiRs, tRFs, rRFs, yRFs, and rpFs, we can annotate 99.9% of all sncRNAs whose abundance is ≥ 10 RPM. IsomiRs are most prevalent (60.6%), followed by tRFs (17.8%), rRFs (11.4%), yRFs (8.3%), and other sncRNAs (1.7%). Figure 1B shows the length distributions for the sncRNAs in each class. Most isomiRs are 22 nts long. Most tRFs are 30–36 nts long, suggesting they are primarily tRNA halves [45]. yRFs have a length distribution similar to tRFs. rRFs fall into two groups, 16–25 and 43–45 nts. The other sncRNAs have a “flatter” distribution.
Non-canonical and non-templated isomiRs abound
Consistent with our previous reports [33, 38, 46], most isomiRs have canonical 5´-ends matching the 5´ ends of the miRBase reference sequences. Supplementary Fig. S3 shows detailed statistics on isomiR types found in the brain samples. Several highly abundant isomiRs have shifted “seed” regions, meaning they target different genes than the reference isomiR, as we showed previously [28]. Also mirroring our previous findings, most non-canonical 5´-end isomiRs originate from the 3p arms of the miRNA precursors (Supplementary Fig. S3B) [33, 46].
On the other hand, more than half of all isomiRs have non-canonical 3´-ends, with ≥ 20% having non-templated nucleotide additions (Supplementary Fig. S3C-D). The identities of the non-templated 3´-end nucleotides differ between the 5p and 3p miRNA arms: adenylation is most frequent among isomiRs from the 5p arm (Supplementary Fig. S3E) and uridylation among isomiRs from the 3p arm (Supplementary Fig. S3F). Notably, most isomiRs with a guanylated 3´-end have higher abundance in SCZ (multivariable regression p-value < 3.3e-05, Supplementary Fig. S3E-F). See also below.
Many tRFs with sequence mismatches abound
Figure 1A shows an unexpectedly high abundance of molecules with LD ≤ 2 from known tRFs (7.3% of total sncRNA-ome), almost as many as the annotated tRFs (10.5%). For simplicity, we analyzed molecules with LD = 1, which comprise 84% of the LD ≤ 2 category total abundance (Supplementary Table S2), and identified two groups. The first (48% of all LD = 1 reads) comprises tRFs, primarily 5´-tRNA halves and 5´-tRFs, with a non-templated 3´-end nucleotide addition. They include tRFs from tRNAGlyCCC/GlyGCC, tRNAGluCTC, tRNALysCTT, and tRNAHisGTG. The distribution of the non-templated nucleotides (Supplementary Fig. S4A) closely resembles that for 3p miRNA arms (Supplementary Fig. S3F), including the significant (p-value = 1.9e-07) elevation of guanylated tRFs in SCZ. The second group comprises extremely abundant ( > 1,000 RPM) 5´-tRNA halves and 5´-tRFs from tRNAGluCTC and tRNAGlyCCC/GlyGCC with deletions or substitutions at position 6 of the parental tRNA (Supplementary Fig. S4B).
The sncRNA-omes of cases differ significantly from those of controls in SCZ and BD
We first present our analyses of cases and controls from the HBCC brain bank, because it contains both SCZ and BD cases that span a wide age range. Using a multivariable DESeq2 model adjusted for known and hidden confounding variables (see Methods), we found 1,127 sncRNAs that differ significantly (FDR < 0.05) in abundance between SCZ cases and controls (Fig. 2A-E, Supplementary Table S3A). Most sncRNAs have modest FC values: median |log2 FC | = 0.58 across 1,127 DA sncRNAs, consistent with the polygenic inheritance of SCZ [47]. At a |log2 FC | = 0.4 threshold (filled markers in Fig. 2A-E), the number of DA sncRNAs is 761.
(A-E) Volcano plots for different sncRNA types. Significantly differentially abundant sncRNAs (FDR < 0.05) are highlighted with color and are filled if |log2 FC | ≥ 0.4. F Mutual distribution of FCs from SCZ vs. control and BD vs. control comparisons. The diagonal dotted line corresponds to equal FCs (Y = X). Only molecules associated with SCZ or BD (likelihood-ratio test FDR < 0.05) are shown. Markers are colored and/or filled if the same FDR and FC thresholding criteria are met in either of the two comparisons. IsomiRs, tRFs, rRFs, and yRFs include wild-type and LD ≤ 2 molecules. “Other” molecules include rpFs and additional genomic mappings (see Methods).
The effect in BD is weaker, with only 104 DA sncRNAs between cases and controls, and FDR values in [0.01, 0.05] (Supplementary Table S3B). We also ran DESeq2 with the likelihood-ratio test to identify sncRNAs that are significantly associated with either SCZ or BD (Supplementary Table S3C). For these sncRNAs, the FCs between SCZ and controls are highly correlated with those from the BD vs. control comparison (Pearson’s r = 0.94, p-value < 1e-100, Fig. 2F). The absolute values of the FCs in BD are systematically lower than SCZ (Fig. 2F, Supplementary Fig. S5).
Highly abundant miRNAs increase further in abundance in SCZ
The top part of Fig. 2A shows several isomiRs with a median abundance in SCZ or control samples ≥ 1,000 RPM – all are upregulated in SCZ. Supplementary Fig. S6A shows boxplots for these isomiRs in SCZ cases and controls. Only two isomiRs have log2 FC above 0.4: miR-126-3p | 0 | 0 and miR-451a | 0 | −1. The previously-mentioned non-canonical miR-126-3p | +1 | 0 is also upregulated but has much lower abundance (Fig. 2A). The remaining isomiRs include miR-99a-5p, miR-125b-5p, miR-29a-3p, and let-7 family members (let-7b-5p, let-7c-5p, let-7i-5p). The loci producing miR-99a-5p, let-7c-5p, and miR-125b-5p are clustered on chromosome 21 and are co-transcribed [48]. These isomiRs are very abundant; thus, despite their modest FCs, a small change in their abundance can have a considerable impact.
Differentially abundant tRFs, rRFs, and yRFs exhibit biases in the sign of their change
The tRFs, rRFs, and yRFs we identified exhibit strong DA signatures and a clear bias for decreased abundance in the SCZ samples (Fig. 2B-E). A subset of highly abundant DA tRFs ( ≥ 1,000 RPM) is shown in Supplementary Fig. S6B. Nonetheless, two groups of tRFs are markedly upregulated in SCZ. The first group consists of highly abundant 5´-tRNA halves (three exceed 5,000 RPM in SCZ), including the above-mentioned tRNAGluCTC 5´-tRNA halves with a deletion at position 6. The second group includes tRFs with non-templated 3´ guanylation that resembles the isomiRs’ pattern. In complete analogy with isomiRs, several sibling tRFs produced from the same parental tRNA have opposite FC signs. For example, a 34-nt 5´-tRNA half derived from tRNAGluCTC increases in abundance in SCZ (log2 FC = 0.72, FDR = 0.01), whereas a 28-nt 5´-tRF from the same tRNA is less abundant in SCZ compared to control samples (log2 FC = −1.09, FDR = 0.0014). DA rRFs are typically less abundant in SCZ (Fig. 2C), with several clear exceptions: a group of i-rRFs that are produced from different regions of 28S rRNA are more abundant in SCZ samples, whereas other 28S i-rRFs are less abundant in SCZ. The most abundant yRFs originate from two parental Y RNA molecules, RNY1 and RNY4, with virtually all DA yRFs being depleted in SCZ (Fig. 2D). The FCs and abundances of other types of sncRNAs are shown in Fig. 2E. Of note, a 26-nt fragment of the 7SL signal recognition particle RNA (srpRNA) has the highest absolute FC across all sncRNAs (log2 FC = 2.23, FDR = 2e-05), but a low average abundance (median across SCZ samples = 10.2 RPM).
GWAS-identified miR-137-3p and miR-2682-5p are upregulated in SCZ but have low abundance
Several GWAS SCZ studies found a significant group of SNPs adjacent to miRNAs miR-137-3p and miR-2682-5p [3, 49]. Two isomiRs of miR-137-3p, miR-137-3p | 0 | −1 and miR-137-3p | 0 | 0, increase in SCZ (Fig. 2A) but the effect size is limited: log2 FC = 0.29 with FDR = 0.041, and log2 FC = 0.27 with FDR = 0.082, respectively. The less studied miR-2682-5p also increases: log2 FC = 0.45 with FDR = 0.0061. However, even when combining all DA isomiRs from these three loci, the resulting abundance is < 50 RPM, suggesting that their impact on their targets may not be consequential.
Non-templated isomiRs differ between SCZ cases and controls
Figure 3A shows the results of a gene set enrichment analysis (GSEA) analysis that assesses the enrichment of different non-templated nucleotides among DA sncRNAs. The X-axis shows the ranked list of all sncRNAs with abundance ≥ 10 RPM. The top left panel shows running enrichment scores corresponding to different non-templated isomiRs. The middle-left panel shows the type of the non-templated isomiR using tick marks at the corresponding positions of the ranked list. Most-upregulated sncRNAs in SCZ are on the left side and most-downregulated on the right (bottom left panel). IsomiRs with non-templated 3´-end runs of cytosine (“WT + C” isomiRs, p-value = 8e-14) and uridine (“WT + T” isomiRs, p-value = 2e-03), respectively, are prevalent among the isomiRs whose abundance decreases in SCZ. Indeed, these two groups account for 80% of the downregulated isomiRs with log2 FC ≤ −0.4. IsomiRs with non-templated 3´-end runs of guanine (“WT + G” isomiRs) are prevalent among the isomiRs whose abundance increases in SCZ (p-value = 9e-27): the WT + G group accounts for 54% of all upregulated isomiRs with log2 FC ≥ 0.4.
A GSEA plot for SCZ vs. control comparison in the HBCC brain bank. All abundant sncRNAs are ranked according to the log10 p-value multiplied by the fold change sign. Vertical bars in the middle panel indicate the presence of an isomiR with non-templated nucleotide addition. B Comparison of nuclear and cytoplasmic abundances of non-templated isomiRs in rat primary cortical neurons [50]. Non-templated isomiRs of 35 selected miRNA arms are shown (see also text). The X and Y axes show the ratio of reads mapped to a particular non-templated isomiR and the whole miRNA arm. The dotted diagonal line corresponds to equal FCs (Y = X). Only miRNAs whose sequences are identical in human and rat were considered.
We also found sibling isomiRs whose FC increases or decreases depending on the identity of their non-templated nucleotide addition. For example, miR-181a-5p | 0 | −1( + 1 G) increases in SCZ compared to controls (FDR = 0.0095, log2 FC = 0.46), whereas the 1-nt shorter miR-181a-5p | 0 | −2( + 1 C) decreases (FDR = 0.036, log2 FC = −0.42).
WT + G isomiRs were originally reported in a previous study of nuclear and cytoplasmic fractions of rat cortical neurons. It was shown that they are enriched in the nuclear fractions [50]. Given the findings of Fig. 3A, we re-analyzed the data of that earlier study, focusing on WT + G isomiRs with an abundance ≥ 10 RPM from the 35 miRNA arms whose sequences are the same in humans and rats and expressed in rat neurons (Supplementary Table S4). Figure 3B shows the abundances of the non-templated rat isomiRs originating from these 35 miRNA arms in the nuclear and cytoplasmic fractions. The WT + G rat isomiRs are highly enriched in the nuclear fraction (p-value = 2e-07), whereas the WT + C (p-value = 2e-06) and WT + A (p-value = 2e-05) isomiRs are enriched in the cytoplasmic fraction. In addition to WT + G isomiRs, WT + G tRFs are also enriched in the nuclear fraction (data not shown).
The changes of mRNAs in SCZ and BD mirror the changes of sncRNAs
We also analyzed the previously reported mRNA sequencing data [2] from the same samples. We considered only the 15,414 mRNA genes whose abundance is ≥ 1 transcript-per-million (TPM) in at least 25% of all samples (cases and controls). Of the 1,442 DA genes between SCZ and control, only 155 have |log2 FC | ≥ 0.4 (Supplementary Fig. S7A, Supplementary Table S5A). As with sncRNAs, the FCs in SCZ and BD are highly correlated (Pearson’s r = 0.9, p-value < 1e-100), with weaker effects in BD (Supplementary Fig. S7B, Supplementary Table S5B-C). We applied GSEA with the reference list of DA genes generated by the PsychENCODE Consortium [1] to compare the changes with previously reported mRNA-level data. The analysis shows clear, highly significant (p-value < 1e-200) enrichment with matching FC signs between our SCZ analysis and reference PsychENCODE SCZ data (Supplementary Fig. S7C). A similar, highly-significant concordance (p-value < 1e-60) exists between our BD FCs and PsychENCODE BD data. Gene Ontology (GO) term enrichment analysis with GSEA shows enrichment of pathways related to synaptic signaling, neuron and axon development among downregulated genes in SCZ, and translation-related pathways among upregulated genes (Supplementary Fig. S7D-E, Supplementary Table S6).
The basic trends are replicated in the MSSM brain bank with weaker effect sizes
We independently ran DA tests on sncRNAs and mRNAs of the MSSM brain bank. We found only five sncRNAs (Supplementary Table S3D) and one mRNA (Supplementary Table S5D) with an FDR ≤ 0.05. To compare the sign of molecule changes between the two brain banks, we used GSEA on the unthresholded MSSM-ranked sncRNAs list using statistically significant HBCC changes as a reference list (Supplementary Fig. S8A). Both brain banks show significant concordance (p-value = 2e-11), and an agreement in their FCs signs (Supplementary Fig. S8A). All the trends with non-templated isomiRs also replicate at the level of GSEA, with WT + G isomiRs showing significant abundance increases in SCZ and WT + A/WT + C/WT + T isomiRs showing significant abundance decreases (Supplementary Fig. S8B). GSEA also shows a substantial agreement of the DA mRNA-seq results from MSSM and HBCC (Supplementary Fig. S8C), and between MSSM and PsychENCODE (Supplementary Fig. S8D). The enrichments for the PsychENCODE reference list are lower in the MSSM samples (Supplementary Fig. S8D) compared to the HBCC ones (Supplementary Fig. S7C).
The sncRNA and mRNA expression profiles independently suggest accelerated aging in SCZ and BD
The differences in DA signal strength between the HBCC and MSSM brain banks may be due to differences in the age distribution (MSSM individuals are significantly older) or systematic batch effects (e.g., differences in sample collection and processing). We tested the interaction between disease status and age through separate DA analyses in two groups of samples from the HBCC brain bank: those aged < 45 years old and those aged ≥ 45 years old. We chose the cutoff point to balance the number of cases and controls in the age groups. We also used randomized down-sampling to ensure that sample size does not affect the analysis. The younger group (age < 45) is responsible for the SCZ DA mRNA signal, in concordance with the previous studies. In contrast, the older group shows a total absence of DA mRNAs (Fig. 4A, “horizontal” comparisons). Hundreds of mRNAs are DA between younger and older controls, but there are almost no differences between younger and older SCZ cases (Fig. 4A, “vertical” comparisons). The same holds for the BD cases (Supplementary Fig. S9A).
(A, D) The number of significantly differentially abundant mRNAs/sncRNAs (FDR < 0.05) between age-restricted SCZ vs. control comparisons (horizontal dimension) and condition-restricted age ≥ 45 vs. age < 45 comparisons (vertical dimension). To remove the effect of sample size on the reported numbers, we show median numbers of differentially abundant molecules derived from 100 random down-samplings (see Supplement). (B, E-I) Mutual distribution of FCs from SCZ vs. control and age ≥ 45 vs. age < 45 comparisons. The diagonal dotted line corresponds to equal FCs (Y = X). Only molecules associated with disease status or age group (likelihood-ratio test FDR < 0.05) are shown. Significantly differentially abundant mRNAs/sncRNAs are highlighted with color if FDR < 0.05 in either of the two comparisons. Highlighted markers are filled if |log2 FC | ≥ 0.4 in either of the two comparisons. IsomiRs, tRFs, rRFs, and yRFs include wild-type and LD ≤ 2 molecules. “Other” molecules include rpFs. C GSEA of differentially abundant mRNAs in SCZ in the HBCC brain bank with the GTEx aging signature as a reference gene set.
Figure 4B shows the joint distribution of FCs from two contrasts of the same multivariable models: SCZ cases vs. controls on the X-axis and age ≥ 45 vs. age < 45 individuals on the Y-axis. Only mRNAs significantly associated with SCZ or age (likelihood-ratio test) are included (Supplementary Table S5E-F). We see a striking positive correlation of 0.78 (p-value < 1e-100), suggesting changes in protein-coding genes common to SCZ and natural aging. We observed the same correlation when conducting GSEA on the HBCC SCZ changes with a brain aging gene signature derived by another group from the independent GTEx project [51] (p-value < 1e-16, Fig. 4C). These observations explain the effects from Fig. 4A and the weak effect sizes in the MSSM samples. Specifically, the impact of SCZ on the brain samples resembles the effect of natural aging, which minimizes the differences between SCZ and controls in older individuals. The high correlation between aging and disease FCs is also present in BD (Supplementary Fig. S9B-C).
A more complex picture exists at the sncRNA level. Figure 4D shows how many DA sncRNAs correspond to the interaction between SCZ status and age and leads to the same conclusions as the mRNAs (Fig. 4A). However, the correlations between SCZ and age group FCs are weaker and follow complex, molecule-type-specific patterns (Fig. 4E-I, Supplementary Table S3E-F). For example, there is a modest, significant, positive FC-FC correlation of isomiR (Fig. 4E) and yRF abundance (Fig. 4H), whereas the FC-FC correlation for rRFs is negative (Fig. 4G). Supplementary Fig. S9D-I, the BD counterpart of Fig. 4D-I, supports similar conclusions. The only exception is Supplementary Fig. S9D, which shows almost no DA sncRNAs between BD and controls in the age < 45 group.
Differentially abundant sncRNAs and mRNAs form distinct co-expression modules
To gain insight into the interrelations of DA molecules, we first regressed out known factors such as disease status, sex, age, and RIN, then computed all-vs-all Pearson correlations between sncRNAs and mRNAs using CorrAdjust [44]. Doing so ensures that any observed correlations are not driven by disease or other major hidden confounders captured by a subset of principal components from the expression data.
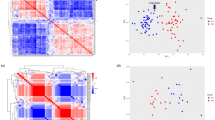
Figure 5 shows correlation heatmaps with (unsupervised) hierarchical clustering of sncRNAs (Fig. 5A) and mRNAs (Fig. 5B). This clustering remains highly similar when using only the control samples (cophenetic correlation = 0.64 for sncRNAs and 0.6 for mRNAs; 100,000 permutations p-value < 1e-5 in both cases). Since we regressed out the disease status, these linkages reflect putative modules of orchestrated expression involving sncRNAs and critical processes in the normal brain. Even though the clusters were computed after the disease status was regressed out, they agree exceptionally well with the SCZ FC signs, suggesting that the implicated sncRNAs and mRNAs (Supplementary Table S7) are also relevant for studying this disease.
Hierarchical clustering (using Ward's linkage) of sncRNAs (A) and mRNAs (B) using the samples from the HBCC brain bank. Correlations were computed after regressing out known and hidden confounders from the abundance levels (see Methods).. Only sncRNAs/mRNAs that exhibited differential abundance (FDR < 0.05) between SCZ cases and controls (HBCC brain bank) were considered for the clustering. Annotations of mRNA clusters were based on Gene Ontology enrichment analysis. Arrows between sncRNA and mRNA clusters show correlations between eigen-molecules below −0.5 or above 0.5 (see text). IsomiRs, tRFs, rRFs, and yRFs include wild-type and LD ≤ 2 molecules. “Other” molecules include rpFs.
The first sncRNA cluster includes sncRNAs whose abundance decreases in SCZ, i.e., virtually all yRFs, multiple tRFs, and 5S-rRNA-derived rRFs. The second cluster is enriched in non-templated WT + G isomiRs from multiple miRNA loci that cluster separately from their templated siblings. The third cluster comprises 5´-tRNA halves, and the fourth comprises various isomiRs upregulated in SCZ. The fifth cluster comprises i-rRFs derived from 28S rRNA. The remaining two clusters comprise isomiRs with increased and decreased abundance in SCZ.
Five of the six mRNA clusters show clear and specific Gene Ontology (GO) pathway enrichments (Supplementary Table S8 and Fig. 5B). The most intriguing findings correspond to gene groups downregulated in SCZ, including synaptic signaling, memory, behavior, cognition, neurogenesis, and ion transport. The other two clusters comprise genes upregulated in SCZ and include translation, stress response, response to Zn and Cu ions, response to cytokines, and regulation of cell death.
SncRNA clusters are wired with specific mRNA clusters
For each cluster in Fig. 5, we computed an “eigen-molecule” by computing the first principal component based on the expression of the cluster’s genes. Then, we calculated correlations between all possible eigen-molecules to link sncRNA clusters with mRNA clusters.
Four sncRNA-mRNA cluster pairs have absolute correlation values exceeding 0.5 and are depicted by arrows connecting the cluster pair’s members. The first sncRNA cluster (yRFs, tRFs, and rRFs downregulated in SCZ) is positively correlated with the cluster of mRNAs downregulated in SCZ (synaptic signaling, memory, behavior, cognition). It is also negatively correlated with the cluster of mRNAs upregulated in SCZ (response to ions/cytokines). The sncRNA cluster of 5´-tRNA halves whose abundance increases in SCZ is negatively correlated with the cluster of mRNAs that are downregulated in SCZ (synaptic signaling, neurogenesis). Finally, the sncRNA cluster of 28S i-rRFs upregulated in SCZ is positively correlated with the cluster of mRNAs upregulated in SCZ (response to ions/cytokines).
Specific isomiRs potentially cause dysregulation of specific mRNAs in SCZ
Without additional information, it is unclear whether the relationships of Fig. 5 between isomiRs and their potential mRNA targets are causal. To answer this question, we examined computationally-predicted and experimentally-validated isomiR targets to determine whether they are enriched among those mRNAs that are anticorrelated with the isomiRs. We used a) RNA22 [52] to generate sequence-based predictions, and b) TarBase [53] to identify previously-reported experimentally-validated targets from various tissues and cell types. For parity, we predicted targets with RNA22 only for the miRBase canonical reference miRNAs since these are the only ones included in TarBase. Of the analyzed 1,501 isomiRs, RNA22 predictions for 480 (32%) are significantly enriched among the 5% of mRNAs with the most negative correlations (FDR ≤ 0.05). Also, 790 (53%) have significantly enriched TarBase entries among the mRNAs in the same set (Supplementary Table S9). For simplicity, the rest of the analysis was based on the TarBase-derived findings.
We analyzed whether TarBase-identified targets that are anticorrelated with DA isomiRs tend to change in the opposite direction in SCZ. Because we regressed out the disease status from the expression values (see above), such a trend is not necessarily guaranteed. For 41 upregulated and four downregulated isomiRs in SCZ, the number of DA anti-correlated target genes that changed in the opposite direction is higher than is expected by chance (FDR < 0.05). See Supplementary Table S10A-B. The strongest enrichment signal pertains to 13 isomiRs from the let-7 family, including the canonical reference miRNAs for let-7b-5p, let-7c-5p, and let-7i-5p, whose abundance is ≥ 1,000 RPM (Fig. 2A). The highly abundant miR-29a-3p | 0 | −1 and other isomiRs from the same family also show statistically significant enrichment.
The gene targets of these isomiRs are spread across mRNA clusters 2, 3, and 4, which contain genes with decreased expression in SCZ (Fig. 5B, Supplementary Table S10C). In concordance with the mRNA enrichments of Fig. 5B, several critical pathways are highly enriched in the downregulated target gene sets, including synaptic signaling, neurogenesis, behavior, memory, cognition, and others (Supplementary Table S10D).
Discussion
We generated and analyzed the largest collection, to date, of small RNA-seq datasets from post-mortem brain samples of SCZ and BD cases and controls. We annotated virtually all sncRNAs with RPM ≥ 10, of which most (98%) consist of isomiRs (60.6%), tRFs (17.8%), rRFs (11.4%), or yRFs (8.3%). We also combined our findings with mRNA-level abundances from the CommonMind Consortium [2] for the same samples. Lastly, we identified disease-independent co-expression networks that implicate isomiRs, tRFs, rRFs, and yRFs for the first time in the regulatory control of mRNAs belonging to critical brain processes.
Our work goes beyond the three previous studies of sncRNAs in post-mortem brain samples of SCZ cases. One analyzed miRNA profiles in 34 SCZ cases and 102 controls [54], and found only two with elevated expression in SCZ, miR-936 and miR-3162. However, the study neither reported baseline abundances for these miRNAs nor explicitly stated abundance cutoffs. Another study compared sncRNAs from 22 SCZ cases and 22 matched controls, and found no DA molecules [55]. A third study compared 13 cases and 14 controls and found 17 DA miRNAs, with only miR-451a among our DA miRNAs [56]. None of the three studies examined isomiRs, or any other sncRNA types.
Most of the isomiRs we identified are 5´ or 3´ non-canonical (Supplementary Fig. S3A-D) and mirror our previous observations from other cell types and tissues [18, 33, 34, 36, 46]. Notably, miR-126-3p has two sets of abundant isomiRs: those with a canonical 5´-end and non-canonical 3´-ends that increase the number of gene targets [28, 57]; and those whose 5´-end lacks the first nucleotide, mirroring findings in megakaryocytes and platelets [33].
Four very abundant ( ≥ 1,000 RPM) isomiRs have the strongest links to gene targets, and their abundance increases further in SCZ: let-7b-5p, let-7c-5p, let-7i-5p, and miR-29a-3p. The let-7 and miR-29 families comprise essential regulators in the nervous system [58,59,60,61,62], and were previously discussed in the context of SCZ [8]. Regarding miR-137-3p and miR-2682-5p [3, 49], which are proximal to one of the strongest SCZ-associated genomic loci, their isomiRs increase in abundance in SCZ, but have a low combined abundance ( < 50 RPM), and weak effect sizes (FCs ≤ 0.45). While this suggests little to no impact on downstream mRNA targets, the loci might still have essential roles in SCZ in other brain regions or cell types.
Numerous isomiRs are non-templated (Supplementary Fig. S3) and enriched among sncRNAs whose abundance increases in SCZ (Fig. 3A). These isomiRs cluster separately (Fig. 5A) from templated and non-canonical isomiRs from the same loci (Supplementary Fig. S10). Non-templated, guanylated isomiRs were previously reported in the nuclei of rat cortical neurons [50]. In those datasets, we sought SCZ miRNAs whose sequences are identical in human and rat and found them to be ~10x enriched in the nuclear fraction, suggesting they are nuclearly localized in SCZ.
Other sncRNA types, especially tRFs and yRFs (Figs. 2B and D), are also DA, with many having higher abundances and stronger FCs than isomiRs. Co-expression analysis showed strong associations between clusters of tRFs/yRFs and clusters of genes associated with critical nervous system pathways (Fig. 5). This represents a significant advance because it directly implicates tRFs and yRFs, whose mode of regulatory action is largely unknown, in specific brain processes. Previous studies did not report links of tRFs to processes, only associations between altered abundance and neurodegenerative disorders or ischemic stroke response [63]. However, the tRFs’ and yRFs’ exact mechanism of action in this context remains uncharacterized.
The discovered tRFs included many unusual ones that have not been previously reported [31]. Nearly half of them, primarily 5´-tRNA halves and 5´-tRFs, had a non-templated addition to their 3´-ends (Supplementary Fig. S4A) whose distribution mirrors that at the 3p arm of isomiRs (Supplementary Fig. S3F). Another unusual tRF type comprises a deletion at position 6 of tRNAGluCTC, or a deletion/substitution at the same position of tRNAGlyCCC/GlyGCC (Supplementary Fig. S4B). While these changes may reflect sequencing errors of decorated tRNA bases, we are unaware of any modifications at this position. This second group of tRFs is DA in syncytiotrophoblast extracellular vesicles in preeclampsia [64].
Several non-templated sncRNAs exhibited an intriguing behavior in disease. For example, the guanylated miR-181a-5p | 0 | −1( + 1 G) significantly increased in SCZ, whereas the cytosylated miR-181a-5p | 0 | −2( + 1 C), decreased. As another example, the 34-nt 5´-tRNA half from tRNAGluCTC (cluster 3) is upregulated in SCZ, whereas the 28-nt 5´-tRF (cluster 4) from the same tRNA is downregulated, a finding we recently reported in other settings (colon and breast cancer specimens, and multiple cancer cell lines) [45].
When comparing the HBCC SCZ cases with controls, we found 1,127 sncRNAs (Figs. 2A-E) and 1,442 mRNAs (Supplementary Fig. S7A) with significant DA (FDR ≤ 0.05). Their FCs are modest and agree with previous reports [1, 47]. While we found the same DA sncRNAs and mRNAs in BD, again in concordance with earlier reports [1, 65], they had lower FCs and only a few satisfied the significance threshold (Fig. 2F, Supplementary Fig. S7B).
Dichotomizing the HBCC samples by age using 45 years as a threshold showed that most of the differential signal originates from the younger group (Figs. 4A, D). This agrees with the existing model of accelerated aging induced by SCZ and BD [66,67,68,69,70] transcriptomes. Moreover, mRNA changes in disease are highly correlated with changes due to aging (Fig. 4B). We also observed a high correlation with an independent aging signature derived from GTEx (Fig. 4C), which confirms the robustness of the finding. The sncRNA correlations were lower and depended on sncRNA type (Fig. 4E-I). The MSSM SCZ cases and controls showed a significant correlation of FCs for sncRNAs (Supplementary Fig. S8A) and mRNAs (Supplementary Fig. S8C) with the HBCC samples. However, only a handful of molecules were significant and had limited FCs.
Co-expression analyses between sncRNAs and mRNAs in the HBCC samples showed sncRNAs and mRNAs clustering into distinct modules, mainly containing molecules with the same SCZ FC sign (Fig. 5). This is another notable finding because it emerged after our explicit residualization of disease and other confounders. It implicates isomiRs, tRFs, rRFs, and yRFs in fundamental, disease-independent brain processes, including synaptic signaling, neurogenesis, behavior, memory, and cognition (Supplementary Table S10C-D). For isomiRs whose mode of regulation is well understood, we generated additional corroborating evidence: experimentally validated miRNA targets were enriched among the mRNAs that were significantly anti-correlated with isomiRs (Fig. 5, Supplementary Tables S9 and S10A). This is a non-trivial result because we residualized the disease contribution from the sncRNAs’ and mRNAs’ abundance before computing the correlations.
Our study has several limitations. First, while we accounted for known covariates in the co-expression analyses, factors may still remain that could be influencing the findings (e.g., antipsychotic treatment or cell heterogeneity). Second, it is likely that some of the identified linkages between sncRNAs and mRNAs may not be causal. Third, replication of the study on an independent cohort and targeted experiments are in order. However, the latter escapes the scope of this study because such experiments would need to overcome considerable limitations of the available technology:
-
Standard qRT-PCR cannot profile sncRNAs at a single-nucleotide resolution [71], requiring custom, time-consuming approaches [72] or expensive small RNA-seq.
-
Because the biogenesis and function of tRFs, rRFs, and yRFs are largely unknown, experiments overexpressing individual sncRNAs from these classes would not be a prudent choice.
-
We are not aware of any methods that can silence a single sncRNA but no other co-expressed sncRNAs from the same locus.
-
SncRNAs from the same locus generally have different subcellular localizations [30].
We view the results we reported as forming a hypothesis-generating framework that can facilitate future targeted experiments to unravel aspects of gene expression regulation in SCZ and BD.
Data availability
Raw data (FASTQ files) and processed data (read count matrices, metadata) are available in Synapse under synID syn63862703.
References
Gandal MJ, Zhang P, Hadjimichael E, Walker RL, Chen C, Liu S, et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science. 2018;362:eaat8127.
Hoffman GE, Bendl J, Voloudakis G, Montgomery KS, Sloofman L, Wang YC, et al. CommonMind Consortium provides transcriptomic and epigenomic data for Schizophrenia and Bipolar Disorder. Sci Data. 2019;6:180.
Trubetskoy V, Pardinas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604:502–8.
Rigoutsos I, Londin E, Kirino Y. Short RNA regulators: the past, the present, the future, and implications for precision medicine and health disparities. Curr Opin Biotechnol. 2019;58:202–10.
Nemeth K, Bayraktar R, Ferracin M, Calin GA. Non-coding RNAs in disease: from mechanisms to therapeutics. Nat Rev Genet. 2024;25:211–32.
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136:215–33.
Perkins DO, Jeffries CD, Jarskog LF, Thomson JM, Woods K, Newman MA, et al. microRNA expression in the prefrontal cortex of individuals with schizophrenia and schizoaffective disorder. Genome Biol. 2007;8:R27.
Beveridge NJ, Gardiner E, Carroll AP, Tooney PA, Cairns MJ. Schizophrenia is associated with an increase in cortical microRNA biogenesis. Mol Psychiatry. 2010;15:1176–89.
Kim AH, Reimers M, Maher B, Williamson V, McMichael O, McClay JL, et al. MicroRNA expression profiling in the prefrontal cortex of individuals affected with schizophrenia and bipolar disorders. Schizophr Res. 2010;124:183–91.
Moreau MP, Bruse SE, David-Rus R, Buyske S, Brzustowicz LM. Altered microRNA expression profiles in postmortem brain samples from individuals with schizophrenia and bipolar disorder. Biol Psychiatry. 2011;69:188–93.
Pietersen CY, Mauney SA, Kim SS, Lim MP, Rooney RJ, Goldstein JM, et al. Molecular profiles of pyramidal neurons in the superior temporal cortex in schizophrenia. J Neurogenet. 2014;28:53–69.
Santarelli DM, Beveridge NJ, Tooney PA, Cairns MJ. Upregulation of dicer and microRNA expression in the dorsolateral prefrontal cortex Brodmann area 46 in schizophrenia. Biol Psychiatry. 2011;69:180–7.
Smalheiser NR, Lugli G, Zhang H, Rizavi H, Cook EH, Dwivedi Y. Expression of microRNAs and other small RNAs in prefrontal cortex in schizophrenia, bipolar disorder and depressed subjects. PLoS One. 2014;9:e86469.
Martinez B, Peplow PV. MicroRNAs as potential biomarkers for diagnosis of schizophrenia and influence of antipsychotic treatment. Neural Regen Res. 2024;19:1523–31.
Smigielski L, Jagannath V, Rossler W, Walitza S, Grunblatt E. Epigenetic mechanisms in schizophrenia and other psychotic disorders: a systematic review of empirical human findings. Mol Psychiatry. 2020;25:1718–48.
Mestdagh P, Hartmann N, Baeriswyl L, Andreasen D, Bernard N, Chen C, et al. Evaluation of quantitative miRNA expression platforms in the microRNA quality control (miRQC) study. Nat Methods. 2014;11:809–15.
Ibing S, Michels BE, Mosdzien M, Meyer HR, Feuerbach L, Korner C. On the impact of batch effect correction in TCGA isomiR expression data. NAR Cancer. 2021;3:zcab007.
Loher P, Londin ER, Rigoutsos I. IsomiR expression profiles in human lymphoblastoid cell lines exhibit population and gender dependencies. Oncotarget. 2014;5:8790–802.
Fehlmann T, Lehallier B, Schaum N, Hahn O, Kahraman M, Li Y, et al. Common diseases alter the physiological age-related blood microRNA profile. Nat Commun. 2020;11:5958.
Ziats MN, Rennert OM. Identification of differentially expressed microRNAs across the developing human brain. Mol Psychiatry. 2014;19:848–52.
Benesova S, Kubista M, Valihrach L. Small RNA-Sequencing: approaches and considerations for miRNA analysis. Diagnostics (Basel). 2021;11:964.
Nersisyan S, Shkurnikov M, Poloznikov A, Turchinovich A, Burwinkel B, Anisimov N, et al. A post-processing algorithm for miRNA microarray data. Int J Mol Sci. 2020;21:1228.
Evans C, Hardin J, Stoebel DM. Selecting between-sample RNA-Seq normalization methods from the perspective of their assumptions. Brief Bioinform. 2018;19:776–92.
Morin RD, O’Connor MD, Griffith M, Kuchenbauer F, Delaney A, Prabhu AL, et al. Application of massively parallel sequencing to microRNA profiling and discovery in human embryonic stem cells. Genome Res. 2008;18:610–21.
Kozomara A, Birgaoanu M, Griffiths-Jones S. miRBase: from microRNA sequences to function. Nucleic Acids Res. 2019;47:D155–D162.
Tomasello L, Distefano R, Nigita G, Croce CM. The MicroRNA family gets wider: the isomirs classification and role. Front Cell Dev Biol. 2021;9:668648.
Yang A, Bofill-De Ros X, Stanton R, Shao TJ, Villanueva P, Gu S. TENT2, TUT4, and TUT7 selectively regulate miRNA sequence and abundance. Nat Commun. 2022;13:5260.
Telonis AG, Loher P, Jing Y, Londin E, Rigoutsos I. Beyond the one-locus-one-miRNA paradigm: microRNA isoforms enable deeper insights into breast cancer heterogeneity. Nucleic Acids Res. 2015;43:9158–75.
Yu F, Pillman KA, Neilsen CT, Toubia J, Lawrence DM, Tsykin A, et al. Naturally existing isoforms of miR-222 have distinct functions. Nucleic Acids Res. 2017;45:11371–85.
Cherlin T, Jing Y, Shah S, Kennedy A, Telonis AG, Pliatsika V, et al. The subcellular distribution of miRNA isoforms, tRNA-derived fragments, and rRNA-derived fragments depends on nucleotide sequence and cell type. BMC Biol. 2024;22:205.
Pliatsika V, Loher P, Magee R, Telonis AG, Londin E, Shigematsu M, et al. MINTbase v2.0: a comprehensive database for tRNA-derived fragments that includes nuclear and mitochondrial fragments from all the cancer genome atlas projects. Nucleic Acids Res. 2018;46:D152–D159.
Pliatsika V, Cherlin T, Loher P, Vlantis P, Nagarkar P, Nersisyan S, et al. MINRbase: a comprehensive database of nuclear- and mitochondrial-ribosomal-RNA-derived fragments (rRFs). Nucleic Acids Res. 2024;52:D229–D238.
Nersisyan S, Montenont E, Loher P, Middleton EA, Campbell R, Bray P, et al. Characterization of all small RNAs in and comparisons across cultured megakaryocytes and platelets of healthy individuals and COVID-19 patients. J Thromb Haemost. 2023;21:3252–67.
Telonis AG, Magee R, Loher P, Chervoneva I, Londin E, Rigoutsos I. Knowledge about the presence or absence of miRNA isoforms (isomiRs) can successfully discriminate amongst 32 TCGA cancer types. Nucleic Acids Res. 2017;45:2973–85.
Magee R, Londin E, Rigoutsos I. TRNA-derived fragments as sex-dependent circulating candidate biomarkers for Parkinson’s disease. Parkinsonism Relat Disord. 2019;65:203–9.
Telonis AG, Loher P, Honda S, Jing Y, Palazzo J, Kirino Y, et al. Dissecting tRNA-derived fragment complexities using personalized transcriptomes reveals novel fragment classes and unexpected dependencies. Oncotarget. 2015;6:24797–822.
Cherlin T, Magee R, Jing Y, Pliatsika V, Loher P, Rigoutsos I. Ribosomal RNA fragmentation into short RNAs (rRFs) is modulated in a sex- and population of origin-specific manner. BMC Biol. 2020;18:38.
Loher P, Karathanasis N, Londin E, Paul FB, Pliatsika V, Telonis AG, et al. IsoMiRmap: fast, deterministic and exhaustive mining of isomiRs from short RNA-seq datasets. Bioinformatics. 2021;37:1828–38.
Loher P, Telonis AG, Rigoutsos I. MINTmap: fast and exhaustive profiling of nuclear and mitochondrial tRNA fragments from short RNA-seq data. Sci Rep. 2017;7:41184.
Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10:R25.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics. 2012;28:882–3.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Nersisyan S, Loher P, Rigoutsos I. CorrAdjust unveils biologically relevant transcriptomic correlations by efficiently eliminating hidden confounders. Nucleic Acids Res. 2025;53:gkaf444.
Akins RB, Ostberg K, Cherlin T, Tsiouplis NJ, Loher P, Rigoutsos I. The Typical tRNA Co-Expresses Multiple 5’ tRNA halves whose sequences and abundances depend on isodecoder and isoacceptor and change with tissue type, cell type, and disease. Noncoding RNA. 2023;9:96.
Zhiyanov A, Nersisyan S, Tonevitsky A. Hairpin sequence and structure is associated with features of isomiR biogenesis. RNA Biol. 2021;18:430–8.
Fromer M, Roussos P, Sieberts SK, Johnson JS, Kavanagh DH, Perumal TM, et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat Neurosci. 2016;19:1442–53.
Shaham L, Binder V, Gefen N, Borkhardt A, Izraeli S. MiR-125 in normal and malignant hematopoiesis. Leukemia. 2012;26:2011–8.
Yin J, Lin J, Luo X, Chen Y, Li Z, Ma G, et al. miR-137: a new player in schizophrenia. Int J Mol Sci. 2014;15:3262–71.
Khudayberdiev SA, Zampa F, Rajman M, Schratt G. A comprehensive characterization of the nuclear microRNA repertoire of post-mitotic neurons. Front Mol Neurosci. 2013;6:43.
Evangelista JE, Clarke DJB, Xie Z, Lachmann A, Jeon M, Chen K, et al. SigCom LINCS: data and metadata search engine for a million gene expression signatures. Nucleic Acids Res. 2022;50:W697–W709.
Miranda KC, Huynh T, Tay Y, Ang YS, Tam WL, Thomson AM, et al. A pattern-based method for the identification of MicroRNA binding sites and their corresponding heteroduplexes. Cell. 2006;126:1203–17.
Skoufos G, Kakoulidis P, Tastsoglou S, Zacharopoulou E, Kotsira V, Miliotis M, et al. TarBase-v9.0 extends experimentally supported miRNA-gene interactions to cell-types and virally encoded miRNAs. Nucleic Acids Res. 2024;52:D304–D310.
Hu Z, Gao S, Lindberg D, Panja D, Wakabayashi Y, Li K, et al. Temporal dynamics of miRNAs in human DLPFC and its association with miRNA dysregulation in schizophrenia. Transl Psychiatry. 2019;9:196.
Ragan C, Patel K, Edson J, Zhang ZH, Gratten J, Mowry B. Small non-coding RNA expression from anterior cingulate cortex in schizophrenia shows sex specific regulation. Schizophr Res. 2017;183:82–87.
Liu Y, Chang X, Hahn CG, Gur RE, Sleiman PAM, Hakonarson H. Non-coding RNA dysregulation in the amygdala region of schizophrenia patients contributes to the pathogenesis of the disease. Transl Psychiatry. 2018;8:44.
Yang A, Bofill-De Ros X, Shao TJ, Jiang M, Li K, Villanueva P, et al. 3’ Uridylation Confers miRNAs with Non-canonical target repertoires. Mol Cell. 2019;75:511–522 e514.
Shamsuzzama, Kumar L, Haque R, Nazir A. Role of MicroRNA Let-7 in modulating multifactorial aspect of neurodegenerative diseases: an overview. Mol Neurobiol. 2016;53:2787–93.
Lehmann SM, Kruger C, Park B, Derkow K, Rosenberger K, Baumgart J, et al. An unconventional role for miRNA: let-7 activates Toll-like receptor 7 and causes neurodegeneration. Nat Neurosci. 2012;15:827–35.
Bian S, Xu TL, Sun T. Tuning the cell fate of neurons and glia by microRNAs. Curr Opin Neurobiol. 2013;23:928–34.
McGowan H, Mirabella VR, Hamod A, Karakhanyan A, Mlynaryk N, Moore JC, et al. hsa-let-7c miRNA regulates synaptic and neuronal function in human neurons. Front Synaptic Neurosci. 2018;10:19.
Swahari V, Nakamura A, Hollville E, Stroud H, Simon JM, Ptacek TS, et al. MicroRNA-29 is an essential regulator of brain maturation through regulation of CH methylation. Cell Rep. 2021;35:108946.
Blaze J, Akbarian S. The tRNA regulome in neurodevelopmental and neuropsychiatric disease. Mol Psychiatry. 2022;27:3204–13.
Cooke WR, Jiang P, Ji L, Bai J, Jones GD, Lo YMD, et al. Differential 5’-tRNA fragment expression in circulating preeclampsia syncytiotrophoblast vesicles drives macrophage inflammation. Hypertension. 2024;81:876–86.
Gandal MJ, Haney JR, Parikshak NN, Leppa V, Ramaswami G, Hartl C, et al. Shared molecular neuropathology across major psychiatric disorders parallels polygenic overlap. Science. 2018;359:693–7.
Nguyen TT, Eyler LT, Jeste DV. Systemic biomarkers of accelerated aging in schizophrenia: a critical review and future directions. Schizophr Bull. 2018;44:398–408.
Sabunciyan S. Gene expression profiles associated with brain aging are altered in schizophrenia. Sci Rep. 2019;9:5896.
Arakelyan A, Avagyan S, Kurnosov A, Mkrtchyan T, Mkrtchyan G, Zakharyan R, et al. Temporal changes of gene expression in health, schizophrenia, bipolar disorder, and major depressive disorder. Schizophrenia (Heidelb). 2024;10:19.
Lin CW, Chang LC, Ma T, Oh H, French B, Puralewski R, et al. Older molecular brain age in severe mental illness. Mol Psychiatry. 2021;26:3646–56.
Ling E, Nemesh J, Goldman M, Kamitaki N, Reed N, Handsaker RE, et al. A concerted neuron-astrocyte program declines in ageing and schizophrenia. Nature. 2024;627:604–11.
Magee R, Telonis AG, Cherlin T, Rigoutsos I, Londin E. Assessment of isomiR discrimination using commercial qPCR Methods. Noncoding RNA. 2017;3:18.
Honda S, Kirino Y. Dumbbell-PCR: a method to quantify specific small RNA variants with a single nucleotide resolution at terminal sequences. Nucleic Acids Res. 2015;43:e77.
Acknowledgements
The work of the Thomas Jefferson University team was supported by University funds (IR) and NIH grant R01HG012784 (IR). The work of the Mount Sinai School of Medicine team was supported by NIH grants R01MH109677, U01MH116442, R01MH110921, R01MH125246, R01MH109897, R01AG067025, R01AG065582, R01AG050986 (PR). We also acknowledge the use of the Cancer Genomics Shared Resource at the Sidney Kimmel Cancer Center (SKCC) of Thomas Jefferson University: SKCC is supported by an NIH Cancer Center Support Grant P30CA056036. We also thank Dr. Gerhard Schratt for providing us with the raw small RNA sequencing data of rat neurons from his earlier study [50].
Author information
Authors and Affiliations
Contributions
IR and PR designed and supervised the study. IN, PL, and SN developed the small RNA-seq mapping pipeline. ZS, JFF, and GV participated in the selection of samples and isolation of RNA. SN, PL, and IR analyzed the data and interpreted the results with contributions from PR and KG. SN and IR prepared the figures and tables. SN and IR wrote the manuscript with contributions from PR and KG. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nersisyan, S., Loher, P., Nazeraj, I. et al. Several novel classes of small regulatory RNAs show widespread changes in schizophrenia and bipolar disorder and extensive linkages to critical brain processes. Transl Psychiatry 16, 72 (2026). https://doi.org/10.1038/s41398-026-03808-x
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41398-026-03808-x