Abstract
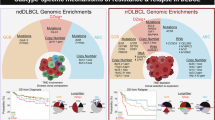
With tumor genomic and gene-expression profiling (GEP), this study investigated the immune-molecular signatures of a unique cohort of diffuse large B-cell lymphoma of the bone (bone-DLBCL), including primary bone (PB-DLBCL, n = 52) and polyostotic-DLBCL (n = 20), in comparison to nodal DLBCLs with germinal center B-cell (GCB) phenotype (nodal-DLBCL-GCB, n = 34). PB-DLBCL and polyostotic-DLBCL shared similar genomic profiles and transcriptomic signatures, justifying their collective analysis as bone-DLBCL. Differential incidences of EZH2, HIST1H1E, and MYC aberrations (p < 0.05) confirmed the distinct oncogenic evolution between bone-DLBCL and nodal-DLBCL-GCB. Differentially expressed genes were identified between bone-DLBCL and nodal-DLBCL-GCB (p < 0.001), substantiated by distinct gene-set enrichment analysis (GSEA). In contrast to a more ‘depleted’ phenotype for nodal-DLBCL-GCB, bone-DLBCL primarily exhibited an ‘intermediate/rich’ tumor microenvironment (TME) signature (p = 0.001), as determined by a previously published gene set. Unsupervised clustering defined two distinct groups that aligned with previously reported immune-enriched TME clusters: an ‘immune-rich’ cluster largely consisting of bone-DLBCLs (75%, p = 0.002) with superior survival (p = 0.030), and a poor-prognostic ‘immune-low’ cluster, including mostly nodal-DLBCL-GCB (61%). Single-sample (ss)GSEA showed higher scores for regulatory T cells, immunosuppressive/prolymphoma cytokines, and vascular endothelial cells in immune-rich samples (p < 0.001). Additionally, CIBERSORTx revealed a higher abundance of regulatory T cells and activated mast cells in the immune-rich cluster (p < 0.001). These findings were confirmed at protein level, where CD3 and FOXP3 immunochemistry showed significant overlap with the gene-expression data (p < 0.001). Conclusively, PB-DLBCL and polyostotic-DLBCL share immune-molecular TME characteristics, supporting their classification as a unified bone-DLBCL entity. The distinct immune-rich TME profile of bone-DLBCL associated with superior survival potentially shapes emerging immunomodulatory strategies

Similar content being viewed by others
Introduction
Extranodal diffuse large B-cell lymphoma of the bone (bone-DLBCL) is a rare subtype of DLBCL that exhibits relatively favorable survival outcomes [1,2,3,4,5,6]. Bone-DLBCL is classified into three subtypes based on the International Extranodal Lymphoma Study Group (IELSG)-14 study: primary bone DLBCL (PB-DLBCL; single bone localization with or without regional lymphadenopathy), polyostotic-DLBCL (multiple bone localizations in one or several bones), and disseminated-bone-DLBCL with concomitant bone involvement along with other (extra)nodal localizations [3, 6, 7]. Recent studies have demonstrated that bone-DLBCL are primarily of the germinal center B-cell (GCB) phenotype, according to either the immunohistochemistry-based Hans’s or nanoString-based Lymph2Cx algorithm [2, 6, 8,9,10,11].
While PB-DLBCL is a rare disease, polyostotic-DLBCL is even more uncommon [5, 9, 12, 13]. Several small case-cohort studies have grouped polyostotic-DLBCL and PB-DLBCL together without performing direct comparative analyses [9, 14]. Only two studies have clearly distinguished PB-DLBCL and polyostotic-DLBCL, limiting our understanding of immune-molecular features, and survival outcomes [3, 15]. By definition, polyostotic-DLBCL, with more than one extranodal localization, is categorized as Ann Arbor stage IV. Though, compared to disseminated-bone-DLBCL, also categorized as Ann Arbor stage IV, polyostotic-DLBCL demonstrates markedly superior survival outcomes [5, 12, 13].
With the advent of targeted and immunomodulatory therapies, treatment could potentially be tailored based on the immune-molecular aspects of DLBCLs. There is a pressing need to elucidate the molecular profile along with the tumor microenvironment (TME) of homogeneous cohorts of anatomical DLBCL entities [16]. As previously reported, our group utilized custom-developed targeted next-generation sequencing (tNGS) BLYMFv2 panel (including 128 lymphoma-relevant genes) and gene-expression profiling (GEP) to identify immune-molecular profiles of DLBCLs originating within particular anatomical localizations, such as intravascular-DLBCL, DLBCL-leg type, ocular-adnexal-DLBCL and sinonasal-DLBCL [17,18,19,20].
To understand the importance of the TME and comprehensively explore this aspect of lymphomagenesis, a custom knowledge-based, biology-driven GEP panel called BLYM-777 (nanoString nCounter platform) was developed [21]. The BLYM-777 probe-set captures several crucial features of lymphomagenesis such as the COO classification (Lymph2Cx) [10] and different TME compositions, such as specific immune clusters delineated by two independent studies of Ciavarella et al. and Kotlov et al. [22, 23], enabling an in-depth investigation into lymphoma biology and its tumor microenvironment. This study intends to identify distinct immune-molecular profiles within a unique, well-annotated and homogenous bone-DLBCL cohort, comprising polyostotic-DLBCL and PB-DLBCL. By integrating BLYM-777 GEP signatures with genotyping via BLYMFv2 tNGS, these profiles were compared to DLBCLs with exclusively nodal localizations and GCB-phenotype, to investigate the biology of DLBCLs originating in these specific sites.
Methods
Patient characteristics
This retrospective study analyzed 72 adult bone-DLBCL patients, including 20 polyostotic-DLBCLs and 52 PB-DLBCLs (Supplementary Table 1). With the predominant GCB-phenotype for bone-DLBCL, a comparator cohort of GCB-type DLBCL (nodal-DLBCL-GCB, n = 34) with limited Ann Arbor stage I/II and strictly nodal localizations was included. Patient selection was based on pathology surveys of DLBCL with bone or nodal tissue. Disseminated-bone-DLBCLs were excluded due to difficulties in accurately determining its anatomic origin. T-cell rich DLBCLs were not included. As high-grade B-cell lymphomas with MYC and BCL2 translocations are exceedingly rare in bone-DLBCL, these were excluded for the nodal-DLBCL-GCB cohort. All cases (n = 106) were diagnosed (2002–2024) at the Leiden University Medical Center (LUMC) or collaborating hospitals (Supplementary Table 2). Imaging was performed using computed tomography, magnetic resonance imaging, or positron emission tomography (-computed tomography), depending on the local standard procedure at time of diagnosis. Patient characteristics and survival outcomes were collected from the medical records. All cases were classified according to the Ann Arbor staging system and the International Prognostic Index (IPI). The study was performed in accordance with the Dutch Code for Proper Secondary Use of Human Tissue, the local institutional board requirements, and the revised Declaration of Helsinki (2008) and approved with a waiver of informed consent by the LUMC’s medical ethics committee (B16.048 and RP23.065) and University Hospital Leuven (S66625).
Histopathological evaluation
Immunohistochemistry (IHC) and fluorescence in situ hybridization (FISH) analyses were performed according to the 4th and 5th edition of the WHO classification of lymphoid neoplasms and reviewed by expert hematopathologists [24]. Antibodies for CD10, BCL6, and MUM1 were used for the Hans’s algorithm-based COO classification and scored as positive with ≥30% staining of tumor cells. General and regulatory T-cell infiltration was assessed using CD3 and FOXP3 immunohistochemistry, with positivity defined as ≥30% for CD3 and ≥20% for FOXP3. FISH using break-apart probes for MYC, BCL2 and BCL6 rearrangement status was performed according to standard procedures. Epstein-Barr virus (EBV) status was determined by EBV-encoded RNA in situ hybridization. Supplementary methods describe further details.
Nucleic acid isolation
FFPE samples were cut into 10-µm sections (median tumor cells, 70%; range, 30–90%). Microdissection was performed to assure high tumor cell percentage. Subsequently, DNA and RNA were extracted using the fully automated Tissue Preparation System (Siemens Healthcare Diagnostics) [25]. DNA and RNA concentrations were quantified with the QubitTM dsDNA HS Assay Kit and the QubitTM RNA HS Assay Kit with the QubitTM 2.0 Fluorometer (Invitrogen) with minimum concentrations of 0.11 ng/µL DNA and 4 ng/µL RNA.
Tumor genomic profiling
The used custom BLYMFv2 tNGS panel contains 128 lymphoma-relevant genes (Supplementary Table 3), with an overlap of 97% with the proposed consensus tNGS panel for mature lymphoid malignancies [26]. As described previously and detailed in the Supplementary Methods, the BLYMFv2 panel was validated using samples previously analyzed with the routine diagnostic tNGS LYMFv1 panel, including 52 lymphoma-relevant genes, or with whole-exome sequencing (WES) [17, 18]. BLYMFv2 libraries were generated using the Ion AmpliSeqTM Library Kit 2.0 (ThermoFisher Scientific) and sequenced with an Ion Torrent S5-system (ThermoFisher Scientific). Sequence reads were aligned to the human reference genome (GRCh37/hg19) using TMAP 5.07 software, with default parameters (https://github.com/iontorrent/TS) [27]. The average read count was 1284 (Range, 201–7464) (Supplementary Table 1). Variants were called by Torrent Variant Caller. The minimum thresholds were ≥100 on-target reads and a variant allele frequency ≥10% (Supplementary Table 4). Samples were excluded if insufficient reads or the transition to transversion rate was >5, indicating potential presence of excess formalin fixation artefacts. All variants were annotated in the Geneticist Assistant NGS Interpretive Workbench (SoftGenetics), into class 1 benign, class 2 likely benign, class 3 unknown significance, class 4 likely pathogenic, or class 5 pathogenic [28]. Class 4 and 5 variants were designated as pathogenic mutations, followed by class 3 variants with a high CADD-phred score ( > 25) and/or a pathogenic prediction from more than two of four selected prediction scores (LRT, MutationTaster, Polyphen, Sift). For genotyping, LymphGen, 2-step LymphGen, LymphPlex, and Guidance-20 were performed. For the LymphGen classification, both the pathogenic and non-pathogenic variants were uploaded to the online LymphGen 2.0 tool (https://llmpp.nih.gov/lymphgen/index.php) [29]. For the 2-step LymphGen, LymphPlex, and Guidance-20, the provided scripts on the git repositories were used (described in Supplementary Methods) [27, 30, 31]. Because some samples failed sequencing with BLYMFv2, a minority of tumor samples (n = 20) were analyzed with LYMFv1.
Gene-expression profiling
The custom ‘BLYM-777’ GEP panel uses the nanoString nCounter platform and includes 797 probes for genes of interest involved in lymphomagenesis and TME [21]. The raw counts generated with the Lymph2Cx probes were uploaded at the Lymphoma/Leukemia Molecular Profiling Project website for COO categorization (https://llmpp.nih.gov/LYMPHCX/) [10]. For the remaining analyses, the data were standardized by the geometric mean of inherent positive controls to reduce potential technical variation. Subsequently, standardized data was normalized with the housekeeping genes and log2 and transformed to z-scores. Next, a principal component analysis identified the possible presence of outliers (deviance from median >6 times the median absolute deviation) or ‘batch effects’. Unsupervised clustering analysis on GEP-data was performed using k-means clustering. Using the ‘glmnet’ package, a penalized generalized linear model with LASSO regularization was fitted to define gene-expression features distinctive for sub-categories (e.g., Polyostotic-DLBCL, PB-DLBCL, or nodal-DLBCL-GCB). DESeq2 (R-package) was used to calculate the log2-fold change and adjusted p-value of each gene between groups, for gene-set enrichment analysis (GSEA) with the clusterProfiler package. The raw nanoString nCounter data reported in this article have been deposited in the Gene Expression Omnibus database (GSE273595) and in Supplementary Table 5.
Tumor microenvironment phenotyping
The BLYM-777 probe encompasses all 45 genes that define the TME clusters identified by Ciavarella et al. This clustering method focuses on the expression of myofibroblast, dendritic and CD4 positive T cells ultimately splitting the data into three clusters with a high, mid, or low immune expression profile [22]. To accurately cluster these TME signatures, GEP data from an in-house reference cohort of 629 samples of (extra)nodal DLBCLs was utilized. The three distinct clusters were initially defined using k-means clustering on the z-scores of the normalized data. The Kotlov signature described four lymphoma microenvironment clusters based on so-called functional gene-enrichment signatures (FGES) and three progeny scores [23]. The BLYM-777 contains 77% of the genes delineating these FGES (n = 225). However, five of the 22 FGES specifically those associated with lymphatic endothelial cells, cancer-associated fibroblasts, fibroblastic reticular cells, granulocyte traffic, and activated macrophages—had less than 80% gene overlap and were therefore excluded from the analysis. The FGES were computed using single-sample gene-set enrichment analysis (ssGSEA), while progeny scores were determined through matrix multiplication with progeny coefficients, as described in the authors’ published Python script. The z-scores of the FGES and progeny data were then subjected to k-means clustering. The identified clusters were assigned to their respective lymphoma microenvironments based on published reference data. CIBERSORTx was applied to the normalized GEP data using the online CIBERSORTx tool (cibersortx.stanford.edu) and the LM22 reference matrix.
Statistical analysis
All analyses were performed using RStudio (R-4.4.0, clusterProfiler-4.12.0, ComplexHeatmap-2.20.0, DESeq2-1.44.0, fgsea-1.30.0, glmnet-4.1-8, msigdbr-7.5.1, survival-3.6-4). Fisher exact or Student t test were applied for analyzing categorical or continuous variables between groups. Progression-free (PFS), overall (OS), or disease-specific survival (DSS) were defined as the date from initial diagnosis to date of event. Event was determined by progression/relapse or death by any cause for PFS, death by any cause for OS, and disease-related death for DSS. For the survival analysis, patients were censored after 5 years of follow-up or censored at last follow-up if there was no event. The Kaplan-Meier method determined median follow-up time and constructed survival curves that were compared with a log-rank test. If statistically significant (alpha-level of 0.05), corresponding hazard ratio and 95% confidence intervals (CIs) were calculated with a Cox proportional hazard model.
Results
Patient cohorts
Clinical characteristics of polyostotic-DLBCL (n = 20), PB-DLBCL (n = 52) and nodal-DLBCL-GCB patients (n = 34) are shown in Fig. 1A, Table 1 and Supplementary Table 1. The median ages of polyostotic-DLBCL, PB-DLBCL and nodal-DLBCL-GCB were 60, 60.0 and 64 years, respectively (p = 0.142). By definition, all polyostotic-DLBCL cases, characterized by ≥2 extranodal sites, were classified as Ann Arbor stage IV, leading to a significantly higher IPI score (p < 0.001). Patients were predominantly treated with R-CHOP(-like) chemoimmunotherapy (n = 103), typically receiving either three to four or six to eight cycles. Polyostotic cases were more frequently treated with six to eight cycles (n = 12) than three to four cycles (n = 2). PB-DLBCL cases were evenly distributed between both regimens (n = 21 and n = 20, respectively), while nodal DLBCL-GCB cases were more often treated with fewer cycles (n = 16 vs. n = 13). Additionally, PB-DLBCL received more often additive radiotherapy (27 out of 49 cases), than polyostotic-DLBCL (6/20) and nodal-DLBCL-GCB (12/34). Nonetheless, both discrepancies in treatment modalities were not significantly different. No differences were observed in survival outcomes for polyostotic-DLBCL, PB-DLBCL and stage I/II nodal-DLBCL-GCB (Fig. 1B, C).
A Schematic overview of polyostotic-DLBCL, PB-DLBCL and nodal-DLBCL-GCB patients and successfully performed targeted NGS and GEP panels. B Clinical course and median survival times per DLBCL subtype. C Comparison of the overall, progression-free and disease-specific survival of the three cohorts.
Using the Hans’s algorithm, 65% of polyostotic-DLBCL and 68% of PB-DLBCL were designated as GCB-phenotype, similar to prior studies [2, 6, 8, 9]. Regarding Lymph2Cx, predominantly GCB phenotypes were found (95%, 91/96) (Supplementary Table 1). Three PB-DLBCL (5%, 3/62) were classified as ABC-phenotype, and one polyostotic case was categorized as intermediate/unclassifiable (2%). As anticipated, 97% (33/34) of nodal-DLBCL-GCBs, were annotated as GCB-phenotype and one case (3%) as intermediate phenotype.
Genomic profiles: similar in PB-DLBCL and polyostotic-DLBCL, distinct from nodal-DLBCL
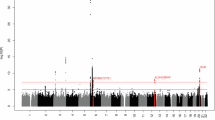
Figure 2A provides an overview of the mutations identified using BLYMFv2 (n = 82) and LYMFv1 (n = 21) panels. The most prevalent identified mutations are typically associated with GCB-phenotypes, such as B2M, KLHL6, SOCS1, and TNFRSF14. Except for a notable absence of IRF8 mutations in polyostotic-DLBCL (p < 0.050), PB-DLBCL and polyostotic-DLBCL displayed a similar mutational profile with frequent mutations in BCL6, EZH2, KLHL6, and TNFRSF14 (Fig. 2A, B). These molecular similarities justified grouping PB-DLBCL and polyostotic-DLBCL together as a relatively homogenous group of bone-DLBCL for subsequent analysis. Significant differences in mutational profiles were observed between bone-DLBCL and nodal-DLBCL-GCB (Fig. 2C), including frequent EZH2 mutations in bone-DLBCL and HIST1H1E and MYC mutations or translocations in nodal-DLBCL-GCB (Supplementary Fig. 1A, C).
A Oncoprintplot visualizing the mutational profile per patient, each mutation type (e.g., nonsynonymous or stopgain) is showed per gene per sample. Furthermore, annotations show the metadata of each sample: tNGS panel, LymphGen, LymphPlex (with or without TP53 status), two-step LymphGen, Guidance-20 algorithms, and the cell-of-origin (COO) using the Hans’s algorithm or Lymph2Cx gene-expression profiling. B Differential variant frequency plots of polyostotic-DLBCL against PB-DLBCL and C bone-DLBCL against nodal-DLBCL-GCB.
Correlating mutational status with clinical data revealed no significant overlap between most clinical characteristics and the most frequently mutated genes, including the bone- or nodal-DLBCL specific mutations. However, TNFRSF14 mutations were more commonly found in younger patients compared to older patients (p = 0.017). Additionally, mutations in B2M were significantly associated with inferior survival (p = 0.013), while KLHL6 mutations were associated with superior survival (p = 0.049). Of these two mutations, only B2M was identified as an independent risk factor (hazard ratio: 4.5, 95% confidence interval: 1.50–13.62, p = 0.007), when adjusted for age and disease type (bone vs nodal). However, it was borderline significant when adjusted for the IPI score (age, LDH, Ann Arbor, WHO status), with a hazard ratio of 2.9 (95% confidence interval: 0.93–8.82, p = 0.067).
Without significant differences between cohorts, LymphGen genotyping primarily categorized bone-DLBCL and nodal-DLBCL-GCB as either EZB (EZH2 & BCL2 aberrations, n = 22 and n = 8) or ST2 (SGK1 & TET2 mutations, n = 8 and n = 8) (Fig. 2A, Supplementary Table 6). Forty-four cases (43%) were unclassified following LymphGen classification. The LymphPlex classification successfully categorized 70% of the cases, revealing a relatively higher prevalence of ST2 subtypes for nodal-DLBCL-GCB compared to the bone-DLBCL (6 and 2 patients, respectively, p = 0.023). As opposed to the LymphGen and LymphPlex algorithm, the ‘two-step LymphGen’ algorithm is less convoluted and only takes into account if the major mutational markers were mutated for classification. Therefore, this method is capable of classifying almost any sample, and was able to assign 77% of the cases. Of note, is the increase in MCD classification in the bone-DLBCL group (19%, n = 13) that was not significantly different from nodal-DLBCL-GCB (6%, n = 2, p = 0.212).
Lastly, our findings were compared to NGS data from four independent molecular studies of large DLBCL cohorts (Supplementary Fig. 1D) [32,33,34,35]. The bone-DLBCL specific mutations (EZH2 and IRF8) were significantly different compared to the DLBCL derived from the literature cohort when examined irrespective of COO subtype (p < 0.001). Nonetheless, no statistically significant differences were observed when comparing bone-DLBCL to the GCB-DLBCL subtype in the literature overview.
Supervised GEP clustering: An immune-rich TME signature for bone-DLBCL
GEP data was effectively collected for 96 cases (91%), comprising 62 bone-DLBCLs (including 19 polyostotic-DLBCLs and 43 PB-DLBCLs) as well as 34 nodal-DLBCL-GCB cases. No significant batch effects were detected with principal component analysis (Supplementary Fig. 2). Direct comparison between PB-DLBCL and polyostotic-DLBCL revealed no significant differences (Fig. 3A), further supporting the rationale for merging these groups into a unified ‘bone-DLBCL’ entity. Comparing bone-DLBCL with nodal-DLBCL-GCB identified several significantly (adjusted p value < 0.001) differentially expressed genes (n = 30, Fig. 3B). Subsequent elastic net regression using a generalized linear model (alpha=0.5) distinguished a total of 44 genes that discriminated between bone-DLBCL and nodal-DLBCL-GCB cohorts (Fig. 3C), emphasizing significantly distinct GEP signatures for both DLBCL entities.
A Volcano plot of differentially expressed genes between polyostotic-DLBCL and PB-DLBCL. B Volcano plot of differentially expressed genes between bone-DLBCL (polyostotic-DLBCL and PB-DLBCL) and nodal-DLBCL-GCB. C Heatmap with gene-selection based on elastic net regression analysis from a generalized linear model of the difference between bone-DLBCL and nodal-DLBCL-GCB. D Gene-set enrichment analysis on differentially expressed genes between bone-DLBCL and nodal-DLBCL-GCB with Gene Ontology (GO) datasets. The normalized enrichment score (color) and p-value (size) are plotted against the gene ratio (enriched genes of the panel divided by the total GO dataset genelist). E Matrix visualizing the distribution of the bone-DLBCL and nodal-DLBCL-GCB cohort over tumor microenvironment clusters by Ciavarella et al. and Kotlov et al.
To further explore the TME differences between the GEP signatures of bone-DLBCL and nodal-DLBCL-GCB, gene-set enrichment analysis (GSEA) was applied to the differentially expressed genes. GSEA revealed that bone-DLBCL was notably enriched for gene ontology datasets related to the ‘extracellular matrix’ and bone-associated pathways (Fig. 3D). Conversely, nodal-DLBCL-GCB exhibited enrichment for datasets associated with the ‘cell cycle process’, various cellular components, and the ‘immune response-regulating signaling pathway’. Unsupervised clustering as delineated by Ciavarella et al. and using the in-house reference cohort, allowed for categorization of our dataset. Bone-DLBCLs were predominantly classified into the ‘intermediate’ or ‘rich’ categories (95%) with abundant expression of genes associated with CD4 T cells, dendritic cells, and myofibroblasts, whereas nodal-DLBCL-GCB cases were more commonly associated with the ‘rich/depleted’ signatures (p = 0.001, Fig. 3E, Supplementary Fig. 3). Applying ssGSEA for lymphoma microenvironment cluster allocation as outlined by Kotlov et al., both bone-DLBCL and nodal-DLBCL-GCB demonstrated enrichment in the mesenchymal/inflammatory clusters (respectively 79% and 68%, Fig. 3E, Supplementary Fig. 3). Although not statistically significant (p = 0.067), 24% (n = 8) of the nodal-DLBCL-GCB cases were assigned to the depleted cluster, compared to 10% (n = 6) of the bone-DLBCL cases. Altogether, these supervised clustering analyses indicated a relatively uniform ‘immune rich’ TME in bone-DLBCL, while the nodal-DLBCL-GCB cases exhibited greater diversity in TME richness and subtypes.
Unsupervised clustering: immune-rich cluster in bone-DLBCL with superior survival
Unsupervised clustering on GEP data using all 784 endogenous mRNA genes of the BLYM-777 panel revealed two distinct clusters (Fig. 4A). This clustering analysis revealed a marked separation for bone-DLBCL cases, with 75% (n = 51/68, p = 0.002) clustering into a single group (Fig. 4B). In contrast, nodal-DLBCL-GCB cases were evenly distributed across both clusters. The cluster dominated by the bone-DLBCL cases demonstrated superior survival outcomes (Fig. 4C) compared to the cluster with a higher proportion of nodal-DLBCL-GCBs, with improved OS (p = 0.027), PFS (p = 0.020), and DSS (p = 0.022), respectively. Multivariable analysis using a Cox regression model demonstrated immune clustering as an independent risk factor when adjusted for age, disease subtype, or IPI score (0–1 vs. 2–5) (hazard ratio: 3.7/3.37, 95% CI: 1.17–11.82/1.07–10.63, p = 0.026/0.038).
A Unsupervised clustering on the entire dataset with bone-DLBCL and nodal-DLBCL-GCB. B Matrix of the distribution of bone-DLBCL and nodal-DLBCL-GCB cohorts over the immune-rich and immune-low clusters. C Comparison of the overall, progression-free and disease-specific survival between the immune-rich and immune-low cluster.
Additionally, unsupervised clustering was performed on the bone-DLBCL cases alone, clustering the data into two groups (Supplementary Fig. 4). Comparison of the unsupervised clustering from the total cohort and the bone cohort, showed highly significant overlap (p < 0.001). Although a survival difference was observed between the clusters within the bone-DLBCL cohort, it did not reach statistical significance, likely due to the limited number of events.
To further investigate the difference between these two clusters, supervised analyses were conducted comparing the previously identified TME signatures with the newly defined clusters. This comparison revealed a significant overlap within and between the immune enriched TME clusters from both signatures (identified by Ciavarella et al. and Kotlov et al.) and the bone-DLBCL-dominated cluster (p < 0.001 and p = 0.003, respectively) (Fig. 5A, Supplementary Fig. 3). Consequently, our identified cluster was designated as an ‘immune-rich’ cluster. Conversely, the remaining cluster characterized by a relatively immune-depleted TME, was termed an ‘immune-low’ cluster.
A Matrix visualizing the distribution of the immune-rich and immune-low clusters over tumor microenvironment clusters by Ciavarella et al. and Kotlov et al. B Gene-set enrichment analysis on differentially expressed genes between immune-rich and immune-low with Gene Ontology (GO) datasets. The normalized enrichment score (color) and p value (size) are plotted against the gene ratio (enriched genes of the panel divided by the total GO dataset genelist). C Deconvolution analysis with CIBERSORTx and ssGSEA with FGES.
Immune-rich tumor microenvironment characterized by regulatory T-cell abundance
GSEA (Fig. 5B) demonstrated significant downregulation in several gene ontology (GO) datasets within the immune-rich cluster compared to the immune-low cluster, including ‘B cell receptor signaling pathway’, ‘cell cycle’ and related pathways. Conversely, the immune-rich cluster exhibited positive enrichment for ‘growth factor receptor binding’ ‘signaling receptor activator activity’, and channel-associated datasets. These findings indicate that the immune-rich cluster’s tumor microenvironment is characterized by less extensive cellular proliferation and greater cell interaction as opposed to the immune-low cluster with more pronounced B-cell receptor activity.
To further differentiate these clusters, ssGSEA was applied using the functional gene expression signatures (FGES) established by Kotlov et al. (Fig. 5C, Supplementary Fig. 5A). The immune-rich samples had significantly higher FGES-scores for regulatory T cells, immunosuppressive/prolymphoma cytokines, and vascular endothelial cells. Furthermore, these immune-rich cases demonstrated enrichment for tumor infiltrating lymphocytes, follicular helper T cells, T-cell trafficking, natural killer cells, and follicular dendritic cells. In contrast, the immune-low cases were characterized by higher FGES-scores for MHC-I/II, B cells, and cellular proliferation. Altogether these findings underscore the distinct differences in immune richness between the immune-rich and immune-low tumor microenvironment clusters. Additionally, using CIBERSORTx, bulk GEP data was processed to estimate cell fractions within the tumor microenvironment. This analysis revealed a higher abundance of regulatory T cells and activated mast cells in the immune-rich cluster, while immune-low cases had more CD4 memory-activated T cells (Fig. 5C, Supplementary Fig. 5B). Conclusively, both deconvolution techniques confirmed significantly higher levels of regulatory T cells and other immune cell types, reinforcing the immune cell enrichment observed in the immune-rich cluster compared to the immune-low cluster.
Increased CD3 protein expression for immune-rich tumor microenvironment
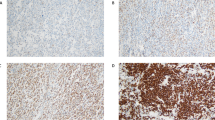
Using CD3 and FOXP3 immunohistochemistry for T-cell and regulatory T-cell infiltration, a subset of cases (n = 76 for CD3 and n = 74 for FOXP3) was evaluated by two expert hematopathologists. The threshold for CD3 was set at 30% and 20% for FOXP3 (Fig. 6A). Comparing these ‘low’ and ‘high’ T-cell infiltration groups at the CD3 protein level with the ‘low’ and ‘high’ regulatory T-cell infiltration at the FOXP3 protein level showed significant overlap (p < 0.001). Of the cases, 29% (n = 21/73) were double positive, 38% (n = 28/73) were double negative, 32% were only CD3 positive (n = 23/73), and 1% were only FOXP3 positive (1/73). Furthermore, CD3 protein expression showed significant overlap with the average gene-expression levels of CD3D, CD3E, and CD3G (p < 0.001, Fig. 6B). Likewise, FOXP3 scoring was significantly associated with the average CD3 expression (p < 0.001), though not with FOXP3 expression (p = 0.22). Subsequently, ‘high’ protein T-cell infiltration was correlated to elevated FGES-scores for T-cell trafficking (p = 0.039), follicular helper T cells (p = 0.014), regulatory T cells (p < 0.001), and tumor infiltrating lymphocytes (p < 0.001), as determined by ssGSEA (Fig. 6B). The same holds true for FOXP3 (p = 0.03, p = 0.012, p = 0.031, p < 0.001). Altogether, these significant associations independently confirm the ‘immune-high/low’ TME classification, particularly highlighting the abundance of regulatory T cells, as identified through unsupervised GEP analysis at the protein dimension level.
A Immunohistochemistry examples of low ( < 30%) and high ( ≥ 30%) CD3 infiltration B and low ( < 20%) and high ( ≥ 20%) FOXP3 expression. C Comparison of CD3 and FOXP3 protein levels with the average gene-expression levels of CD3D, CD3E, and CD3G and correlation with FGES-scores for T-cell trafficking, T follicular helper cells, regulatory T cells, and tumor infiltrating lymphocytes.
Discussion
This study describes an in-depth analysis of the immune-molecular characteristics of a large and distinctive bone-DLBCL cohort, including PB-DLBCLs and polyostotic-DLBCLs. Using previously established, knowledge-driven targeted gene panels (BLYMFv2 and BLYM777), mutational profiles and GEP signatures of bone-DLBCLs were compared to nodal-DLBCL-GCB cases with similar clinical features [32,33,34,35]. Except for IRF8 mutations, PB-DLBCL and polyostotic-DLBCL shared similar mutational profiles and GEP signatures, supporting the rationale for their classification as a unified bone-DLBCL entity. Frequent mutations in EZH2 and HIST1H1E along with MYC aberrations, defined significant differences between bone-DLBCL and nodal-DLBCL-GCB, suggesting distinct oncogenic development. Furthermore, unsupervised GEP clustering identified two different clusters that were attributed to TME differences. An immune-rich cluster, characterized by extensive presence of specific T cells (f.e. regulatory T cells), was predominantly composed of bone-DLBCLs. Conversely, an immune-low cluster marked by increased cellular proliferation, contained a higher proportion of nodal-DLBCL-GCBs. The immune-rich cluster was associated with superior survival outcomes compared to the immune-low group. Finally, the ‘immune-high/low’ classification determined through unsupervised GEP analysis was validated at the protein dimension level using CD3 and FOXP3 immunohistochemistry on tumor tissue.
Based on tNGS, polyostotic-DLBCL and PB-DLBCL demonstrated nearly indistinguishable mutational profiles. Although polyostotic-DLBCL lacked IRF8 mutations, both cohorts displayed frequently occurring mutations in BCL6, EZH2, KLHL6, and TNFRSF14. Moreover, mutations in TNFRSF14 were predominantly found in younger patients, and B2M was independently associated with inferior survival. B2M mutations were also significantly more common in the inferior immune-low group (p = 0.033). Most bone-DLBCLs were classified into the EZB genetic subtype (characterized by EZH2 and BCL2 alterations) according to the LymphGen classification (55% of classified cases), followed by the ST2 (20%) genetic subtype (defined by SGK1 and TET2 mutations). On average, 58% of our mutational data was classified using the LymphGen 2.0 tool, despite the absence of CNA data and, in some cases, missing FISH data. This classification rate is comparable to the 63.1% achieved in the original study [29]. Mutations in bone-DLBCL primarily affected epigenetic and immune-regulating genes, whereas nodal-DLBCL-GCB contained frequent alterations in HIST1H1E, MYC, SGK1, SOCS1, and STAT3, resulting in a more even distribution across the LymphGen clusters EZB and ST2 ( ~ 45% each). While the LymphPlex algorithm showed a higher prevalence of ST2 in nodal-DLBCL-GCB, the two-step LymphGen method was more frequently associated with MCD classification in bone-DLBCL.
The GEP signatures of polyostotic-DLBCL and PB-DLBCL were identical. In contrast, significant GEP differences were found between bone-DLBCL and nodal-DLBCL-GCB. GSEA revealed that nodal-DLBCL-GCB was associated with active ‘cell cycle process’ pathways, while bone-DLBCL exhibited enrichment in pathways related to the extracellular matrix and bone tissue. Bone-DLBCL primarily exhibited an intermediate/rich TME profile according to classification by Ciavarella et al. and Kotlov et al. [22, 23]. Specifically, bone-DLBCL showed a higher abundance of myofibroblasts, dendritic, and CD4 positive T cells compared to nodal-DLBCL-GCB. Unsupervised GEP clustering identified two distinct clusters: one primarily consisting of bone-DLBCLs and the other predominantly nodal-DLBCL-GCB cases. Furthermore, unsupervised clustering on the bone cohort, without nodal-DLBCL-GCB, had significant overlap with the cluster-distribution from the total cohort. Subsequent supervised analyses comparing the previously identified TME signatures with the newly defined clusters revealed a significant overlap between the immune-enriched TME clusters from both signatures and the bone-DLBCL-dominated cluster. As a result, our identified cluster with a high proportion of bone-DLBCL was annotated as immune-rich, while the other cluster was considered immune-low. The immune-rich cluster was independently associated with superior survival. GSEA further indicated that immune-rich DLBCLs were associated with increased activity in receptor-related datasets, while immune-low DLBCLs was enriched for cell proliferation, DNA damage response, and B-cell receptor signaling pathway. Lastly, deconvolution techniques like CIBERSORTx and ssGSEA with FGES both showed significant higher contribution of regulatory T cells in the immune-rich group. CIBERSORTx marked among others increased levels of CD4 memory activated T cells in the immune-low group. The ssGSEA with FGES identified elevated presence of genes associated with vascular endothelial cells and immune-suppressive/prolymphoma cytokines in the immune-rich group. Conversely, the immune-low cluster exhibited increased expression of genes related to cell proliferation, aligning with the enriched GSEA GO datasets for ‘cell cycle process’. The higher levels of T cells, particularly regulatory T cells, in the immune-rich cluster, as indicated by GEP clustering, were corroborated and correlated with protein expression through immunohistochemistry of CD3 and FOXP3.
Overall, the combination of supervised, unsupervised, GSEA, deconvolution analysis and protein dimension level consistently indicated that bone-DLBCL is characterized by an immune-rich TME. Our cohort also confirmed the association between an immune-rich environment and a favorable prognosis, as described in the literature [22, 23, 36]. Specifically, the presence of regulatory T cells was enriched in the superior-prognostic immune-rich as opposed to the poor-prognostic immune-low cluster. The presence of specific T-cell subsets could therefore hypothetically play a crucial role in orchestrating an effective immune response in bone-DLBCLs, potentially triggered by therapeutic interventions, facilitating the destruction of lymphoma cells. Besides, the substantial differences in survival outcomes among TME clusters highlight the urgent need to explore which immune context and interactions provide optimal benefit from innovative immunomodulatory therapies.
Our findings suggest that bone-DLBCLs generally have a favorable prognosis and could potentially benefit from de-escalating immuno-chemotherapy regimens, with treatments tailored to their immune-molecular subtype. The Guidance trial implemented a customized LymphGen genotyping assay (Guidance-20) to assign DLBCL patients to targeted treatment regimens, leading to improved survival outcomes [31]. The EZB cluster, prevalent in bone-DLBCL, was treated with R-CHOP combined with tucidinostat (histone deacetylase inhibitor). Furthermore, the immune-rich nature of this cluster might indicate sensitivity to emerging immunomodulatory therapies such as immunomodulatory drug (IMIDs), bispecific antibodies, or chimeric antigen receptor (CAR) T-cell therapy. In an exploratory analysis of the phase 3 ZUMA-7 trial data, Locke et al. showed that stromal immune-suppressive TME lacking CD19 expression and B-cell signature correlated with inferior outcomes after axicabtagene ciloleucel CAR-T-cell treatment [37]. Extrapolating these findings to our study suggests that the immune-rich phenotype of the bone-DLBCL might be more responsive to CAR T-cell treatment.
This study has several limitations, primarily due to its retrospective nature and reliance of targeted gene panels rather than comprehensive genome/exome-wide approaches, which may introduce biases. The challenge of retrieving adequate DNA/RNA from decalcified bone biopsies resulted in the loss of 15% of NGS and 10% of GEP data. Furthermore, fifteen PB-DLBCL and three polyostotic-DLBCL were excluded due to insufficient material. Our study would also benefit from additional large-scale spatial analyses to further elucidate the vulnerabilities of the immune-rich TME for bone-DLBCL at the level of individual immune cells and their interactions. In addition, multivariable Cox regression analysis was not conducted due to the limited number of events in our cohort. Despite these limitations, our results underline the impact of in-depth immune-molecular investigations in a homogeneous DLBCL cohort with exclusive anatomical localizations [16].
In conclusion, this study is the first to demonstrate that PB-DLBCL and polyostotic-DLBCL share similar mutational and gene-expression profiles, distinct from those of nodal-DLBCL-GCB, supporting the rationale for their classification as a unified bone-DLBCL entity. Furthermore, bone-DLBCL is linked to an immune-rich tumor microenvironment, characterized by an abundance of specific T-cell subsets and correlated with superior survival outcomes. Consequently, these patients may benefit from less intensive treatment strategies. Our findings emphasize the potential to tailor novel immunomodulatory strategies, like IMIDs, bispecific antibodies or CAR T-cell therapy, according to specific mutational profiles and TME signatures. This targeted approach could enhance treatment efficacy, reduce immuno-chemotherapy toxicity, and improve survival outcomes for patients with bone-DLBCL.
Data availability
The raw data from the targeted next-generation sequencing with the Ion Torrent S5 and gene-expression profiling with the nanoString nCounter are publicly available in the NCBI Sequence Read Archive (SRA) (RRID:SCR_004891) at PRJNA1152334 and Gene Expression Omnibus (GEO) (RRID:SCR_005012) at GSE273595, respectively. Furthermore, the raw nanoString data was added to Supplementary Table-5.
Code availability
Code used for all analysis of this study and the creation of the figures can be found at https://git.lumc.nl/raldegroen/bonedlbclanalysis.
References
Bruno Ventre M, Ferreri AJ, Gospodarowicz M, Govi S, Messina C, Porter D, et al. Clinical features, management, and prognosis of an international series of 161 patients with limited-stage diffuse large B-cell lymphoma of the bone (the IELSG-14 study). Oncologist. 2014;19:291–8. https://doi.org/10.1634/theoncologist.2013-0249
Li X, Xu-Monette ZY, Yi S, Dabaja BS, Manyam GC, Westin J, et al. Primary Bone Lymphoma Exhibits a Favorable Prognosis and Distinct Gene Expression Signatures Resembling Diffuse Large B-Cell Lymphoma Derived From Centrocytes in the Germinal Center. Am J Surg Pathol. 2017;41:1309–21. https://doi.org/10.1097/pas.0000000000000923
Messina C, Christie D, Zucca E, Gospodarowicz M, Ferreri AJ. Primary and secondary bone lymphomas. Cancer Treat Rev. 2015;41:235–46. https://doi.org/10.1016/j.ctrv.2015.02.001
Tao R, Allen PK, Rodriguez A, Shihadeh F, Pinnix CC, Arzu I, et al. Benefit of consolidative radiation therapy for primary bone diffuse large B-cell lymphoma. Int J Radiat Oncol, Biol, Phys. 2015;92:122–9. https://doi.org/10.1016/j.ijrobp.2015.01.014
Wu H, Bui MM, Leston DG, Shao H, Sokol L, Sotomayor EM, et al. Clinical characteristics and prognostic factors of bone lymphomas: focus on the clinical significance of multifocal bone involvement by primary bone large B-cell lymphomas. BMC cancer. 2014;14:900. https://doi.org/10.1186/1471-2407-14-900
de Groen RAL, van Eijk R, Bohringer S, van Wezel T, Raghoo R, Ruano D, et al. Frequent mutated B2M, EZH2, IRF8, and TNFRSF14 in primary bone diffuse large B-cell lymphoma reflect a GCB phenotype. Blood Adv. 2021;5:3760–75. https://doi.org/10.1182/bloodadvances.2021005215
Board WCoTE. Soft Tissue and Bone Tumours. WHO Classification of Tumours. 5th ed. IARC; 2020:489:chap 3.
Koens L, Heyning FH, Szepesi A, Matolcsy A, Hogendoorn PC, Jansen PM. Nuclear factor-kappaB activation in primary lymphoma of bone. Virchows Arch: Int J Pathol 2013;462:349–54. https://doi.org/10.1007/s00428-013-1372-x
Xu Y, Li J, Ouyang J, Li J, Xu J, Zhang Q, et al. Prognostic relevance of protein expression, clinical factors, and MYD88 mutation in primary bone lymphoma. Oncotarget. 2017;8:65609–19. https://doi.org/10.18632/oncotarget.19936
Scott DW, Wright GW, Williams PM, Lih CJ, Walsh W, Jaffe ES, et al. Determining cell-of-origin subtypes of diffuse large B-cell lymphoma using gene expression in formalin-fixed paraffin-embedded tissue. Blood. 2014;123:1214–7. https://doi.org/10.1182/blood-2013-11-536433
Hans CP, Weisenburger DD, Greiner TC, Gascoyne RD, Delabie J, Ott G, et al. Confirmation of the molecular classification of diffuse large B-cell lymphoma by immunohistochemistry using a tissue microarray. Blood. 2004;103:275–82. https://doi.org/10.1182/blood-2003-05-1545
Messina C, Ferreri AJ, Govi S, Bruno-Ventre M, Gracia Medina EA, Porter D, et al. Clinical features, management and prognosis of multifocal primary bone lymphoma: a retrospective study of the international extranodal lymphoma study group (the IELSG 14 study). Br J Haematol. 2014;164:834–40. https://doi.org/10.1111/bjh.12714
Wu H, Zhang L, Shao H, Sokol L, Sotomayor E, Letson D, et al. Prognostic significance of soft tissue extension, international prognostic index, and multifocality in primary bone lymphoma: a single institutional experience. Br J Haematol. 2014;166:60–8. https://doi.org/10.1111/bjh.12841
Xiao Y, Zhu F, Liu T, Li Q, Li X, Wu G, et al. Beneficial effect of consolidative radiotherapy for patients with lymphoma and skeletal involvement. Medicine. 2019;98:e16688 https://doi.org/10.1097/md.0000000000016688
de Leval L, Braaten KM, Ancukiewicz M, Kiggundu E, Delaney T, Mankin HJ, et al. Diffuse large B-cell lymphoma of bone: an analysis of differentiation-associated antigens with clinical correlation. Am J Surgl Pathol. 2003;27:1269–77.
Vermaat JS, Pals ST, Younes A, Dreyling M, Federico M, Aurer I, et al. Precision medicine in diffuse large B-cell lymphoma: hitting the target. Haematologica. 2015;100:989–93. https://doi.org/10.3324/haematol.2015.128371
Eriksen PRG, de Groot F, Clasen-Linde E, de Nully Brown P, de Groen R, Melchior LC, et al. Sinonasal DLBCL: molecular profiling identifies subtypes with distinctive prognosis and targetable genetic features. Blood Adv. 2024;8:1946–57. https://doi.org/10.1182/bloodadvances.2023011517
Vest SD, Eriksen PRG, de Groot FA, de Groen RAL, Kleij AHR, Kirkegaard MK et al. The Genetic Profile of Large B-Cell Lymphomas Presenting in the Ocular Adnexa. Int J Mol Sci. 2024;25. https://doi.org/10.3390/ijms25063094
Schrader AMR, Jansen PM, Willemze R, Vermeer MH, Cleton-Jansen AM, Somers SF, et al. High prevalence of MYD88 and CD79B mutations in intravascular large B-cell lymphoma. Blood. 2018;131:2086–9. https://doi.org/10.1182/blood-2017-12-822817
Schrader AMR, de Groen RAL, Willemze R, Jansen PM, Quint KD, Cleven AHG et al. Genetic Stability of Driver Alterations in Primary Cutaneous Diffuse Large B-Cell Lymphoma, Leg Type and Their Relapses: A Rationale for the Use of Molecular-Based Methods for More Effective Disease Monitoring. Cancers (Basel). 2022;14. https://doi.org/10.3390/cancers14205152
de Groot FA, de Groen RAL, van den Berg A, Jansen PM, Lam KH, Mutsaers P et al. Biological and Clinical Implications of Gene-Expression Profiling in Diffuse Large B-Cell Lymphoma: A Proposal for a Targeted BLYM-777 Consortium Panel as Part of a Multilayered Analytical Approach. Cancers (Basel). 2022;14. https://doi.org/10.3390/cancers14081857
Ciavarella S, Vegliante MC, Fabbri M, De Summa S, Melle F, Motta G, et al. Dissection of DLBCL microenvironment provides a gene expression-based predictor of survival applicable to formalin-fixed paraffin-embedded tissue. Ann Oncol. 2018;29:2363–70. https://doi.org/10.1093/annonc/mdy450
Kotlov N, Bagaev A, Revuelta MV, Phillip JM, Cacciapuoti MT, Antysheva Z, et al. Clinical and Biological Subtypes of B-cell Lymphoma Revealed by Microenvironmental Signatures. Cancer Discov. 2021;11:1468–89. https://doi.org/10.1158/2159-8290.CD-20-0839
Swerdlow SH, Campo E, Harris NL, Jaffe ES, Pileri SA, Stein H et al. WHO Classification of Tumour of Haematopoietic and Lymphoid Tissues, Revised 4th edition. IARC Press, Lyon, 2017.
van Eijk R, Stevens L, Morreau H, van Wezel T. Assessment of a fully automated high-throughput DNA extraction method from formalin-fixed, paraffin-embedded tissue for KRAS, and BRAF somatic mutation analysis. Exp Mol Pathol. 2013;94:121–5. https://doi.org/10.1016/j.yexmp.2012.06.004
Sujobert P, Le Bris Y, de Leval L, Gros A, Merlio JP, Pastoret C, et al. The Need for a Consensus Next-generation Sequencing Panel for Mature Lymphoid Malignancies. Hemasphere. 2019;3:e169. https://doi.org/10.1097/HS9.0000000000000169
Mishina T, Oshima-Hasegawa N, Tsukamoto S, Fukuyo M, Kageyama H, Muto T, et al. Genetic subtype classification using a simplified algorithm and mutational characteristics of diffuse large B-cell lymphoma in a Japanese cohort. Br J Haematol. 2021;195:731–42. https://doi.org/10.1111/bjh.17765
Thompson BA, Spurdle AB, Plazzer JP, Greenblatt MS, Akagi K, Al-Mulla F, et al. Application of a 5-tiered scheme for standardized classification of 2,360 unique mismatch repair gene variants in the InSiGHT locus-specific database. Nat Genet. 2014;46:107–15. https://doi.org/10.1038/ng.2854
Wright GW, Huang DW, Phelan JD, Coulibaly ZA, Roulland S, Young RM, et al. A Probabilistic Classification Tool for Genetic Subtypes of Diffuse Large B Cell Lymphoma with Therapeutic Implications. Cancer Cell. 2020;37:551–568.e14. https://doi.org/10.1016/j.ccell.2020.03.015
Pedrosa L, Fernandez-Miranda I, Perez-Callejo D, Quero C, Rodriguez M, Martin-Acosta P, et al. Proposal and validation of a method to classify genetic subtypes of diffuse large B cell lymphoma. Sci Rep. 2021;11:1886 https://doi.org/10.1038/s41598-020-80376-0
Zhang MC, Tian S, Fu D, Wang L, Cheng S, Yi HM, et al. Genetic subtype-guided immunochemotherapy in diffuse large B cell lymphoma: The randomized GUIDANCE-01 trial. Cancer Cell. 2023;41:1705–1716.e5. https://doi.org/10.1016/j.ccell.2023.09.004
Schmitz R, Wright GW, Huang DW, Johnson CA, Phelan JD, Wang JQ, et al. Genetics and Pathogenesis of Diffuse Large B-Cell Lymphoma. N. Engl J Med. 2018;378:1396–407. https://doi.org/10.1056/NEJMoa1801445
Reddy A, Zhang J, Davis NS, Moffitt AB, Love CL, Waldrop A, et al. Genetic and Functional Drivers of Diffuse Large B Cell Lymphoma. Cell. 2017;171:481–494 e15. https://doi.org/10.1016/j.cell.2017.09.027
Lacy SE, Barrans SL, Beer PA, Painter D, Smith AG, Roman E, et al. Targeted sequencing in DLBCL, molecular subtypes, and outcomes: a Haematological Malignancy Research Network report. Blood. 2020;135:1759–71. https://doi.org/10.1182/blood.2019003535
Chapuy B, Stewart C, Dunford AJ, Kim J, Kamburov A, Redd RA, et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat Med. 2018;24:679–90. https://doi.org/10.1038/s41591-018-0016-8
Autio M, Leivonen SK, Bruck O, Karjalainen-Lindsberg ML, Pellinen T, Leppa S. Clinical Impact of Immune Cells and Their Spatial Interactions in Diffuse Large B-Cell Lymphoma Microenvironment. Clin Cancer Res. 2022;28:781–92. https://doi.org/10.1158/1078-0432.CCR-21-3140
Locke FL, Filosto S, Chou J, Vardhanabhuti S, Perbost R, Dreger P, et al. Impact of tumor microenvironment on efficacy of anti-CD19 CAR T cell therapy or chemotherapy and transplant in large B cell lymphoma. Nat Med. 2024;30:507–18. https://doi.org/10.1038/s41591-023-02754-1
Acknowledgements
The authors would like to thank Lotte Vissers, Annika Seitz, and Mirjam Mastik for their valuable technical assistance. We acknowledge the support provided by Louis M. Staudt’s Laboratory at the National Cancer Institute of the National Institutes of Health for the online analysis of Lymph2Cx raw data for COO characterization and LymphGen clustering.
Funding
This study is funded by the Stichting Fonds Oncologie Holland.
Author information
Authors and Affiliations
Contributions
Conceptualization was performed by R.A.L.D.G, A.H.G.C., H.V., and J.S.P.V.; Data Curation was established by R.A.L.D.G. and E.J.K.; Formal Analysis was done by R.A.L.D.G. and S.B.; Funding Acquisition was established by J.S.P.V; Investigation was performed by R.A.L.D.G., F.A.D.G., S.B., E.J.K., L.M.D.H, T.N., S.B., R.E.W.J., R.R., L.T.B., V.T., H.L., E.A., E.F.M.P., I.F.S., L.H., W.C.E.D.H., A.V.D.B., P.M., K.H.L., M.W.M.V.D.P., M.A.H., F.J.S.H.W.-A-J., A.J., T.T., J.V.M.G.B., L.K., A.D., A.H.G.C., M.J.K., P.M.J., and M.N.; The Methodology was supported by R.A.L.D.G., L.M.H., T.V.W., R.V.E., D.R., A.V.D.B., and P.M.J.; Project Administration was done by H.V. and J.S.P.V.; Resources were provided by S.B., T.V.W., R.V.E., D.R., L.T.B., V.T., H.L., E.A., E.F.M.P., I.F.S., L.H., W.C.E.D.H., A.V.D.B., P.M., K.H.L., M.W.M.V.D.P., M.A.H., F.J.S.H.W.-A-J., A.J., T.T., J.V.M.G.B., L.K., A.D., A.H.G.C., M.J.K., P.M.J., and M.N.; Software was coded by R.A.L.D.G., S.B., and D.R.; H.V. and J.S.P.V. performed Supervision. Validation was done by R.A.L.D.G., S.B., and E.J.K.; Visualization was performed by R.A.L.D.G.; R.A.L.D.G., F.A.D.G., A.V.D.B., A.D., A.H.G.C., H.V., M.N., T.J.A., and J.S.P.V., aided with writing of the original draft. Everyone assisted with the review & editing of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
A.D. received research funding from and participated in advisory board of Takeda. F.J.S.H.W.-A-J. participated in educational event from Takeda and participated in advisory board of Recordati. M.J.K. has received honoraria from and performed a consulting/advisory role for Celgene subsidiary of Bristol Myers Squibb, Kite subsidiary of Gilead, Miltenyi Biotec, Novartis, Adicet Bio, and Roche; research funding from Kite (Gilead), Roche, Takeda and Celgene (BMS); and travel support from Kite (Gilead), Miltenyi Biotec, Novartis, Abbvie, and Roche. P.J.L. has received honoraria/research funding from Takeda, Servier, Genmab, Roche, Genentech, Celgene (BMS), Incyte, and Regeneron. The remaining authors declare no competing (financial) interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
de Groen, R.A.L., de Groot, F.A., Böhringer, S. et al. Superior survival in diffuse large B cell lymphoma of the bone with immune rich tumor microenvironment. Blood Cancer J. 15, 82 (2025). https://doi.org/10.1038/s41408-025-01291-z
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41408-025-01291-z
This article is cited by
-
Anatomical patterns at first confirmed relapse or refractory disease predict outcomes in diffuse large B-cell lymphoma
International Journal of Hematology (2025)