Abstract
TP53-mutant acute lymphoblastic leukemia (ALL) in adults is a high-risk subtype with poor outcomes, yet its molecular landscape and clinical implications remain incompletely defined. In this multi-institutional study of 830 adult ALL patients treated at eight academic centers between 2010 and 2024, we demonstrated that TP53 mutations are independent predictors of inferior overall survival in both B-ALL (median, 1.9 vs 5 years) and T-ALL (1.6 vs 9.5 years), irrespective of age, biologic disease subtype, or therapy. Genomic profiling revealed that >90% of TP53 mutations were DNA-binding domain missense variants, frequently co-occurring with hypodiploidy in B-ALL and NOTCH1/FBXW7 mutations in T-ALL. Unlike myeloid malignancies, biallelic TP53 mutations did not worsen outcomes, and variant type (missense vs truncating) did not influence survival. TP53-mutant B-ALL exhibited higher CD20 positivity than TP53-wild type B-ALL (65% vs 31%) but had inferior responses to conventional chemotherapy. Novel immunotherapies (e.g., inotuzumab/blinatumomab) or venetoclax-containing combination regimens improved remission rates, yet relapses were common, often with CD19/CD20/CD22 loss (triple-negative) or acquisition of new mutations. Allogeneic transplantation in first remission trended toward survival benefit (median, 3.3 vs 2.2 years). These findings underscore TP53-mutant ALL as a distinct, chemo-resistant entity necessitating tailored approaches, with antigen escape highlighting challenges of immunotherapy durability.
Similar content being viewed by others
Introduction
Acute lymphoblastic leukemia (ALL) is a curable disease in children and young adults [1]. However, outcomes of B- and T-lineage ALL are inferior in older adults due to the higher frequency of adverse-risk molecular characteristics and more limited ability to utilize intensive pediatric-inspired chemotherapy regimens [1, 2]. Biologically, the majority of adult ALL cases harbor aneuploidy (e.g., low hypodiploidy), high-risk fusions (e.g., BCR::ABL1-like signature), or mutations in TP53 or non-TP53 myeloid genes [3,4,5]. We and others have shown that adult TP53-mutant ALL may arise from pre-existing TP53-mutant clonal hematopoiesis (CH) [4, 6, 7]. In adults, TP53 mutations are somatic in >90% of patients, which contrasts the high incidence of Li Fraumeni syndrome seen in 45% of pediatric low hypodiploid TP53-mutant ALL cases [8]. TP53 mutations are found in 15–20% of adults with ALL and are associated with poor chemotherapy response and overall survival (OS) [4, 9, 10]. In this large multi-institution cohort study, we aim to decipher the unique molecular landscape of adult TP53-mutant ALL, as well as the clinical implications of TP53 mutations in the context of established molecular disease subtypes.
TP53 encodes the transcription factor p53, which is a tumor suppressor that can be induced in response to DNA damage, cellular stress or oncogenic hyperproliferation [11]. TP53 is the most frequently mutated gene across all cancers, and TP53-mutant myeloid neoplasms have an extremely poor prognosis [12, 13]. In acute myeloid leukemia (AML) and myelodysplastic syndromes (MDS), biallelic TP53 mutations predict higher risk for relapse and death when compared to monoallelic mutations [12, 14]. In addition, 80% of TP53 mutations across all cancer types are protein-altering missense variants in the DNA-binding domain [15]. While some studies in solid tumors and leukemias suggest an oncogenic gain-of-function (GoF) phenotype associated with these missense variants [16, 17], other mechanistic studies found no evidence to support GoF role for mutant p53 in AML [18]. Instead, mutations reduced p53 tumor-suppressive activity in a dominant-negative manner. Biological and clinical significance of TP53 allelic state and variant type is not established for ALL.
We studied a large cohort of adult patients with ALL treated at eight academic institutions in the US. Through comprehensive genomic profiling and correlative analyses with real-world outcomes data, we found that TP53 mutations are independent predictors of poor outcomes in adult ALL when adjusted for age, molecular subtype, allelic state or variant type. The adverse impact of TP53 mutations is driven by inferior chemotherapy response and high rates of CD19, CD20, or CD22 negative relapses upon treatment with newer antibody-based therapies.
Methods
Patient cohort
A total of 830 adult patients with ALL (age ≥18 years) treated at 8 academic centers between the years 2010 and 2024 were included in this study. Diagnosis, relapse, and disease status were confirmed and assigned according to the World Health Organization (WHO) criteria [19]. Patients with mixed phenotype acute leukemia were excluded. Complete remission (CR) was defined as no circulating blasts or extramedullary disease, <5% bone marrow blasts, adequate neutrophil and platelet count recovery. Measurable residual disease (MRD) was assessed with multiparameter flow cytometry (sensitivity of 0.01%) assay in Clinical Laboratory Improvement Amendments (CLIA)-certified laboratories of collaborating institutions [20]. The study was approved by the Institutional Review Boards of participating sites and conducted in accordance with the Declaration of Helsinki. Informed consent has been obtained from participants.
Molecular profiling
Molecular subtype classification was based on the use of multiple assays, including cytogenetics (karyotype), fluorescence in situ hybridization (FISH), polymerase chain reaction for known fusions, DNA and RNA sequencing, and chromosome microarrays for copy-number abnormalities. These assays were performed as part of routine clinical care in CLIA-certified laboratories. Karyotyping of diagnostic bone marrow aspirate specimens was performed by counting 20 metaphase cells. Ph-like gene signature was determined by FISH for common fusions (CRLF2, JAK2, ABL1, ABL2, CSF1R, PDGFR) and RNA-sequencing. Patient samples from the University of Chicago underwent high-throughput targeted next-generation sequencing (NGS) with the OncoPlus assay (178 genes) [21]. Similarly, diagnostic NGS testing was performed with pan-heme NGS panel at the Northwestern University (204 genes) [22], Hematologic Neoplasm Mutation Panel at The Ohio State University [23], Leukemia NGS panel at Johns Hopkins University (94 genes) [24], Myeloid NGS test at Duke University (75 genes) [25], Comprehensive OncoHeme NGS assay at Mayo Clinic [26], FocusHeme NGS panel at the Medical University of South Carolina (49 genes) and Foundation One Heme Assay at the Moffitt Cancer Center (455 genes) [4]. Genes that are shared between the NGS panels of collaborating sites were used for further analysis. For pathogenic and likely pathogenic variants, an allelic frequency (AF) cutoff of 5% was used.
Statistical analysis
Clinical data for patients were available from medical records. The associations between clinical variables and different classes of mutations were investigated by calculating the odds ratios (OR) with the Fisher exact test for categorical variables and the Wilcoxon rank-sum test for continuous variables. The impact of covariates on survival outcomes was investigated with univariable and multivariable Cox regression analyses. OS was defined as the time from diagnosis until death or last follow-up. Recurrence-free survival (RFS) was defined as the time from first CR until disease relapse, death, or last follow-up. OS and RFS estimates were calculated with the Kaplan–Meier method, and differences were compared using the log-rank test. All statistical analyses were performed using R v.4.4.2. The figures were generated with the ggplot2 package in R and GraphPad Prism version 9 (GraphPad Software).
Results
TP53 mutations are independent predictors of adverse outcomes in ALL
We retrospectively studied molecular characteristics and survival outcomes of 830 adult patients with ALL (709 with B-ALL, 121 with T-lineage ALL) treated at the collaborating institutions within the recent era of new ALL therapeutics (2010–2024). Demographic characteristics of the cohort are summarized in Table 1. Patients with B-ALL were classified into six genetic subtypes based on the molecular data available from all participating sites. ALL cases harboring BCL2 or MYC fusions, KMT2A rearrangements, low hypodiploidy (≤40 chromosomes), or Ph-like signature (i.e., BCR::ABL1-like) had inferior OS and RFS outcomes when compared to B-ALL patients with hyperdiploidy, BCR::ABL1, and other genetic alterations (Fig. 1A).
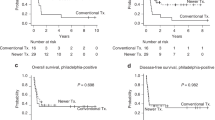
A Overall and relapse-free survival curves for B-ALL patients with different molecular subtypes of disease. Multivariable analysis of overall survival for patients with B-ALL (B) and T-lineage ALL (C). Survival outcomes of B-ALL (D) and T-lineage ALL (E) patients stratified based on their TP53 status. CI confidence interval, HR, hazard ratio, OS overall survival, WT wild type.
TP53 mutations can be seen across different WHO-defined genetic subtypes of ALL [9]. The frequency of TP53 mutations was 17% in our adult ALL cohort. We investigated the prognostic impact of TP53 mutation in the context of previously established predictors of adverse outcomes in ALL. Using multivariable Cox regression analysis, TP53 mutation was an independent predictor of poor OS in B-ALL when adjusted for age, sex, year of ALL diagnosis and molecular subtype (Fig. 1B). Similarly, TP53 mutation was associated with inferior OS in T-lineage ALL when adjusted for age, sex, year of ALL diagnosis and ETP-status (Fig. 1C). Comparing TP53-mutant vs wild-type (WT) ALL, median OS was significantly shorter in TP53-mutant B-ALL (1.9 vs 5 years, p < 0.001) and T-lineage ALL (1.6 vs 9.5 years, p = 0.03) (Fig. 1D, E). The adverse prognostic impact of TP53 mutations was seen in both younger (<40 years) and older (≥40 years) patients with B- and T-lineage ALL (Supplementary Fig. 1). Altogether, these data suggest that TP53 mutations confer adverse prognostic risk for B- and T-ALL patients, independent from traditional high-risk features.
Molecular landscape of TP53-mutant ALL
To understand the determinants of poor outcomes in TP53-mutant ALL, we first studied the genomic landscape associated with TP53 mutations. Similar to the distributions observed in AML [18] and MDS [14], pathogenic or likely pathogenic TP53 variants were concentrated in the DNA-binding domain in >90% of patients with ALL (Fig. 2A). TP53 mutations were most frequent in patients with hypodiploid B-ALL, but rare in patients with BCL/MYC, BCR::ABL1, BCR::ABL1-like or hyperdiploid B-ALL (Fig. 2B, C). We found that 14% of B-ALL and 9% of T-ALL patients had biallelic TP53 mutations, defined by WHO criteria as multiple mutations or mutation with concurrent deletion of the other allele (Fig. 2D) [27]. As expected, biallelic TP53 alterations were more frequent in B-ALL patients with hypodiploidy (Fig. 2E). We also investigated the landscape of co-occurring gene mutations and found differences between B- and T-lineage ALL (Fig. 2F). Mutations in RB1 (13%), IKZF1 (10%), NF1 (10%), TET2 (9%) and NRAS (7%) were more frequent in TP53-mutant B-ALL (Fig. 2G), while NOTCH1 (62%), FBXW7 (31%), PHF6 (31%), ASXL1 (23%) and WT1 (23%) mutations were more frequent in TP53-mutant T-ALL (Fig. 2H). CDKN2A mutations were common in both B- and T-ALL. Collectively, these data suggest that the majority of TP53 mutations are missense variants affecting the DNA-binding domain, and TP53-mutant B- vs T-lineage ALL may harbor co-mutations in different genes associated with lineage.
A Pathogenic and likely pathogenic TP53 variants spanning different functional domains of TP53 gene. Pie charts demonstrate WHO-defined genetic subtypes in TP53-mutant (B) and wild-type (C) B-ALL. Distribution of TP53 mutations based on allelic state in B- vs T-ALL (D) and across genetic subtypes of B-ALL (E). F Bar graphs comparing the frequencies of mutations that frequently co-occur with TP53 in B- and T-ALL. Oncoprints summarizing molecular landscape of TP53-mutant B-ALL (G) and T-ALL (H).
Clinical outcomes of TP53-mutant B-ALL
To determine the biological and clinical factors associated with TP53-mutant B-ALL, we studied the associations between TP53 status and other demographic and clinicopathological variables. Notably, TP53 mutations were more common with older age (OR 7.53, 95% confidence interval [CI]: 3.3–18, p < 0.001) and therapy-related B-ALL (OR 2.6, 95% CI: 1.6–4.1, p < 0.001), but less common in individuals of self-reported Hispanic ancestry (OR 0.4, 95% CI: 0.2–0.9) (Fig. 3A). CD20 positivity was more common in TP53-mutant B-ALL blasts when compared to TP53 WT blasts (65% vs 31%, p = 0.001) (Fig. 3B). Other therapeutically targetable surface markers, CD19 and CD22 were expressed in >90% of B-ALL cases at diagnosis. Over-expression of the missense-mutant p53 protein was detectable with immunohistochemistry in diagnostic bone marrow slides, co-localizing with CD20 positive blasts (Fig. 3C).
A Forest plot demonstrates the odds ratios for the associations between clinical variables and TP53 mutation status in B-ALL. B CD19, CD20, and CD22 surface marker expression in TP53 wild-type vs mutant B-ALL. C Representative immunohistochemistry slides showing CD20 and TP53 co-expression in B-ALL.
Next, we investigated the predictors of worse OS in patients with TP53-mutant B-ALL. In univariable Cox regression analysis, older age was associated with inferior outcomes, while patients treated between 2020 and 2024 had better OS than those treated in earlier years (Fig. 4A). TP53 missense mutations in DNA binding domain may have a novel oncogenic GoF phenotype in different cancers. However, we observed similar OS outcomes between patients harboring missense-mutant TP53 vs other (e.g., frameshift, splice site mutations) types of loss-of-function TP53 variants (Fig. 4B). Biallelic TP53 mutations are associated with worse outcomes than monoallelic TP53 mutations in some subtypes of myeloid neoplasms. However, TP53 allelic state did not predict OS or RFS in patients with B-ALL (Fig. 4C, D). To study the roles of different first-line therapies in TP53-mutant B-ALL outcomes, we compared CR with flow MRD-negativity rates in patients treated with hyper-CVAD (cyclophosphamide, vincristine, doxorubicin, and dexamethasone alternating with high-dose methotrexate and cytarabine), pediatric protocols (i.e., CALGB 10403 regimen), hyper-CVD (cyclophosphamide, vincristine, dexamethasone) + venetoclax and novel antibody-based approaches (i.e., A41703 regimen of sequential inotuzumab and blinatumomab) (Fig. 4E). Patients with TP53-mutant B-ALL had inferior responses to chemotherapy approaches when compared to patients with TP53-WT B-ALL, while responses were similar when newer therapies (venetoclax or antibodies) are introduced into frontline regimens. Finally, we compared outcomes of TP53-mutant B-ALL patients stratified based on receipt of allogeneic hematopoietic cell transplantation (HCT) (Fig. 4F). We used a 3-month landmark analysis to include patients who achieved CR after first-line therapy. Allogeneic HCT was associated with longer OS, but the difference did not reach statistical significance (median OS, 3.3 vs 2.2 years, p = 0.07).
A Univariable Cox regression analysis for predictors of adverse overall survival. Kaplan-Meier survival curves comparing outcomes of TP53-WT vs TP53-mutant patients stratified based on TP53 mutation type (B) or allelic state (C, D). E Bar graphs showing complete remission (CR) with flow cytometric measurable residual disease (MRD)-negativity rates between patients with TP53-WT vs TP53-mutant B-ALL treated with different first-line approaches. F Kaplan-Meier overall survival curves comparing outcomes of TP53-mutant B-ALL patients stratified based on performance of allogeneic hematopoietic cell transplantation (HCT).
Altogether, these data indicate higher rates of CD20 positivity in TP53-mutant B-ALL, in which the type of TP53 mutation or its allelic state did not predict OS. These patients may benefit from novel antibody-based therapies or BH3 mimetic combinations in first-line therapy, and a subset may benefit from allogeneic HCT in CR1.
Clonal evolution in relapsed TP53-mutant B-ALL
To gain insights into the clonal evolution and acquired mechanisms of resistance, we performed genomic and immunophenotypic analysis of serial diagnosis and relapse samples in TP53-mutant B-ALL. TP53 mutations were present in relapse samples of all patients, further confirming the stability of these variants in founder leukemic clones (Fig. 5A). Next, we studied clonal dynamics in patients who developed new acquired mutations at the time of relapse. In B-ALL1, pre-leukemic TP53 mutation was detectable at low AF (3%) when the patient was in remission, supporting previous observations that TP53-mutant CH is a precursor for B-ALL in adults (Fig. 5B) [4, 6]. CDKN2A and PTPN11 mutations re-emerged at the time of relapse, which was also characterized by the loss of CD19 expression after blinatumomab therapy and new somatic deletion in the STK11 gene. We observed loss of surface marker expression as a mechanism of immunotherapy resistance in other TP53-mutant B-ALL samples. In B-ALL2, first-line treatment with inotuzumab followed by blinatumomab effectively treated TP53/RB1-mutant B-ALL, but the disease re-emerged with CD19/CD22-negative relapse (Fig. 5C). CR2 was achieved with rituximab, hyper-CVD, venetoclax combination therapy, but the patient relapsed again with triple-negative (CD19, CD20, CD22) disease and newly acquired MSH2 mutation resulting in new-onset microsatellite instability (MSI) and high tumor mutation burden in lymphoblasts. Finally, we investigated patterns of relapse after allogeneic HCT. In B-ALL3, relapse post-transplant was associated with an acquired CCND3 mutation in TP53, CDKN2A, NF1-mutant clone (Fig. 5D). Patient achieved CR2 with inotuzumab and blinatumomab combination therapy. In summary, these data indicate that TP53 mutations are pre-leukemic events in B-ALL and clonal evolution is characterized by antigenic loss and emergence of new mutations in genes that are less frequently altered in treatment-naive disease.
A Line graph showing evolution of TP53 clone size between diagnosis vs relapse in TP53-mutant B-ALL. B–D Fish plots showing evolution of B-ALL clones at the time of relapse, characterized by the acquisition of new mutations (STK11 loss in B-ALL1, MSH2 and microsatellite instability [MSI] in B-ALL2, CCND3 in B-ALL3) and loss of CD19, CD20, or CD22 surface markers under selective pressure of antibody-based targeted therapies.
Discussion
We established clinical and molecular characteristics of adult TP53-mutant ALL by studying a large multi-center cohort of real-world patients. Traditionally, pediatric ALL is more comprehensively studied than adult ALL, and insights obtained from large cohorts of children and young adults with ALL formed the basis for the genetic subtypes established by the WHO and International Consensus Classification [19, 28]. Older adults (age >40 years) were often excluded from adult ALL trials, further exacerbating the gaps in our knowledge on ALL biology, which is very different between children and adults. Several WHO-defined ALL subtypes are either not seen in adults (e.g., ETV6::RUNX1) or not routinely assessed in clinical practice with available commercial or in-house assays (e.g., MEF2D-rearrangement, NUTM1-rearrangement, ETV6::RUNX1-like). There exists a discordance between prognostic models proposed by the classification guidelines and what can be feasibly implemented in clinical testing to inform therapeutic decision making for adults with ALL. In this study, we established adverse prognostic impact of high-risk genetic features (i.e., BCL/MYC rearrangement, hypodiploidy, KMT2A-rearrangement, BCR::ABL1-like signature) in the modern era of ALL therapeutics. These subtypes can be readily identified with available sequencing platforms. Given the independent prognostic role of TP53 mutations in adult ALL, we examined the previously unanswered questions of the significance of TP53 allelic state, mutation type and allogeneic HCT in predicting outcomes for TP53-mutant ALL. While biallelic TP53 mutations did not portend worse outcomes than monoallelic mutations, a subgroup of patients with TP53-mutant ALL may benefit from allogeneic HCT in CR1. This question should be further investigated in future studies examining larger cohorts derived from transplant registries.
TP53-mutant leukemias have a poor prognosis due to their refractoriness to traditional cytotoxic chemotherapy. Importantly, we observed high MRD-negative CR rates when these patients are treated with newer immunotherapy approaches (inotuzumab and blinatumomab) or venetoclax-based combination regimens (hyper-CVD plus venetoclax). However, triple-negative relapses (CD19/CD20/CD22 negative) are common in patients who receive antibody-based therapies. We also observed unique genotypes at the time of relapse, such as acquisition of MSH2 mutation leading to deficient mismatch repair. MSI-high status has not been observed in large cohorts of AML [29]. Emergence of this phenotype in late stages of TP53-mutant lymphoblastic leukemogenesis may indicate a lineage-specific phenomenon with genomic instability.
This study has a few limitations. We performed molecular subclassification based on the genes and fusions that are mutually covered by participating institutions. Some of the recently annotated B-ALL genetic subtypes and most T-ALL genetic subtypes are not covered by these panels, which limited our analyses in T-ALL. However, the clinical and therapeutic significance of these unprofiled genetic alterations (e.g., NUTM1, MEF2D, DUX4, ZEB2, CDX2/UBTF) is not fully established in large cohorts of older adults with ALL. Therefore, we focused on the biological and clinical significance of TP53 mutations, which have an independent but underappreciated prognostic impact in ALL. As is the case for any retrospective analysis, conclusions related to the role of allogeneic HCT should be interpreted with caution and validated further in larger cohorts.
In conclusion, TP53-mutant ALL is a high-risk disease characterized by inferior response to cytotoxic chemotherapy, high-rates of antigen-negative relapse after immunotherapies, and distinct patterns of clonal evolution. Biallelic mutations or missense mutations, which are thought to confer GoF phenotype in other cancers, do not confer higher risk when compared to monoallelic or truncating mutations, respectively. Nevertheless, given the worse outcomes overall, there is an unmet need for new therapeutics in TP53-mutant ALL, akin to the significant therapeutic challenge that exists for TP53-mutant myeloid neoplasms.
Data availability
Access to anonymized clinical data might be granted upon request to the corresponding author.
References
Pagliaro L, Chen SJ, Herranz D, Mecucci C, Harrison CJ, Mullighan CG, et al. Acute lymphoblastic leukaemia. Nat Rev Dis Prim. 2024;10:41.
Johnston H, Youshanlouei HR, Osei C, Patel AA, DuVall A, Wang P, et al. Socioeconomic determinants of the biology and outcomes of acute lymphoblastic leukemia in adults. Blood Adv. 2024;8:164–71.
Creasey T, Barretta E, Ryan SL, Butler E, Kirkwood AA, Leongamornlert D, et al. Genetic and genomic analysis of acute lymphoblastic leukemia in older adults reveals a distinct profile of abnormalities: analysis of 210 patients from the UKALL14 and UKALL60+ clinical trials. Haematologica. 2022;107:2051–63.
Saygin C, Zhang P, Stauber J, Aldoss I, Sperling AS, Weeks LD, et al. Acute lymphoblastic leukemia with myeloid mutations is a high-risk disease associated with clonal hematopoiesis. Blood Cancer Discov. 2024;5:164–79.
Paietta E, Roberts KG, Wang V, Gu Z, Buck GAN, Pei D, et al. Molecular classification improves risk assessment in adult BCR-ABL1-negative B-ALL. Blood. 2021;138:948–58.
Kim R, Bergugnat H, Larcher L, Duchmann M, Passet M, Gachet S, et al. Adult low-hypodiploid acute lymphoblastic leukemia emerges from preleukemic TP53-mutant clonal hematopoiesis. Blood Cancer Discov. 2023;4:134–49.
Chitadze G, Stengel A, John-Klaua C, Bruckmuller J, Trautmann H, Kotrova M, et al. Somatic TP53 mutations are pre-leukemic events in acute lymphoblastic leukemia. Blood. 2023;141:1640–4.
Comeaux EQ, Mullighan CG. TP53 mutations in hypodiploid acute lymphoblastic leukemia. Cold Spring Harb Perspect Med. 2017;7:a026286.
Aldoss I, Li S, Zhang J, Clark MC, Agrawal V, Pourhassan H, et al. TP53 mutations are associated with CD19- relapse and inferior outcomes after blinatumomab in adults with ALL. Blood Adv. 2025;9:2159–72.
Patel M, Zabor EC, Mohamed A, Zureigat H, Chen MJ, Nakitandwe J, et al. Impact of TP53 mutation on survival outcomes in acute lymphoblastic leukemia at a tertiary center. Leuk Lymphoma. 2025:66:1509–14.
Chen X, Zhang T, Su W, Dou Z, Zhao D, Jin X, et al. Mutant p53 in cancer: from molecular mechanism to therapeutic modulation. Cell Death Dis. 2022;13:974.
Kaur A, Rojek AE, Symes E, Nawas MT, Patel AA, Patel JL, et al. Real world predictors of response and 24-month survival in high-grade TP53-mutated myeloid neoplasms. Blood Cancer J. 2024;14:99.
Badar T, Foran JM, Bewersdorf JP, Wang YH, Coltoff A, El Kettani M, et al. Heterogeneity in outcomes of TP53-mutated myeloproliferative neoplasms based on disease phenotype and mutational status. Br J Haematol. 2025. https://doi.org/10.1111/bjh.20187.
Bernard E, Nannya Y, Hasserjian RP, Devlin SM, Tuechler H, Medina-Martinez JS, et al. Implications of TP53 allelic state for genome stability, clinical presentation and outcomes in myelodysplastic syndromes. Nat Med. 2020;26:1549–56.
Kastenhuber ER, Lowe SW. Putting p53 in context. Cell. 2017;170:1062–78.
Baugh EH, Ke H, Levine AJ, Bonneau RA, Chan CS. Why are there hotspot mutations in the TP53 gene in human cancers? Cell Death Differ. 2018;25:154–60.
Wong TN, Link DC. Are TP53 mutations all alike? Hematol Am Soc Hematol Educ Program. 2024;2024:321–5.
Boettcher S, Miller PG, Sharma R, McConkey M, Leventhal M, Krivtsov AV, et al. A dominant-negative effect drives selection of TP53 missense mutations in myeloid malignancies. Science. 2019;365:599–604.
Alaggio R, Amador C, Anagnostopoulos I, Attygalle AD, Araujo IBO, Berti E, et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Lymphoid Neoplasms. Leukemia. 2022;36:1720–48.
Saygin C, Cannova J, Stock W, Muffly L. Measurable residual disease in acute lymphoblastic leukemia: methods and clinical context in adult patients. Haematologica. 2022;107:2783–93.
Kadri S, Long BC, Mujacic I, Zhen CJ, Wurst MN, Sharma S, et al. Clinical validation of a next-generation sequencing genomic oncology panel via cross-platform benchmarking against established amplicon sequencing assays. J Mol Diagn. 2017;19:43–56.
Symes EO, Wang P, Sojitra P, Menon MP, Patel AA, Hasan F, et al. Somatic co-alteration signatures are prognostic in high-grade TP53-mutated myeloid neoplasms. Br J Haematol. 2025;206:1103–8.
Borate U, Yang F, Press R, Ruppert AS, Jones D, Caruthers S, et al. Samples from patients with AML show high concordance in detection of mutations by NGS at local institutions vs central laboratories. Blood Adv. 2023;7:6048–54.
Pasca S, Haldar SD, Ambinder A, Webster JA, Jain T, Dalton WB, et al. Outcome heterogeneity of TP53-mutated myeloid neoplasms and the role of allogeneic hematopoietic cell transplantation. Haematologica. 2024;109:948–52.
Zhao Y, Siddiqi I, Wildes TJ, Charles D, Deak K, Wang E. Blast phase of myeloproliferative neoplasm resembles acute myeloid leukemia, myelodysplasia-related, in clinical presentation, cytogenetic pattern, and genomic profile, and often undergoes reversion to second chronic phase status after induction chemotherapy. Arch Pathol Lab Med. 2024;148:1310–9.
Shah MV, Hung K, Baranwal A, Wechalekar G, Al-Kali A, Toop CR, et al. Validation of the 5th edition of the World Health Organization and International Consensus Classification guidelines for TP53-mutated myeloid neoplasm in an independent international cohort. Blood Cancer J. 2025;15:88.
Khoury JD, Solary E, Abla O, Akkari Y, Alaggio R, Apperley JF, et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Myeloid and Histiocytic/Dendritic Neoplasms. Leukemia. 2022;36:1703–19.
Arber DA, Orazi A, Hasserjian RP, Borowitz MJ, Calvo KR, Kvasnicka HM, et al. International Consensus Classification of Myeloid Neoplasms and Acute Leukemias: integrating morphologic, clinical, and genomic data. Blood. 2022;140:1200–28.
Walker CJ, Eisfeld AK, Genutis LK, Bainazar M, Kohlschmidt J, Mrozek K, et al. No evidence for microsatellite instability in acute myeloid leukemia. Leukemia. 2017;31:1474–6.
Funding
CS is supported by the Department of Defense Peer Reviewed Cancer Research Program (HT94252510221), American Society of Hematology Scholar Award and Leukemia Lymphoma Society Special Fellow Award. A-KE is supported by R01 CA262496, R01CA284595-01, R01CA283574-01, R01 LM013879, Leukemia & Lymphoma Society, and the American Cancer Society.
Author information
Authors and Affiliations
Contributions
EH, DAK, and CS analyzed and interpreted the data and prepared the manuscript. MS, YA, TK, A-KE, CA, CL, YAMV, TB, AC, TCK, NO, HRY, SC, AAP, ASD, MWD, PW, MT, JPS, GV, SG, JXC, DA, RAL, OO, JW, BS, and WS contributed patients and data, interpreted data, and wrote and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
A-KE’s spouse is employed by Karyopharm Therapeutics, she serves on the scientific advisory boards of Syndax, Daiichi Sankyo, and Servier, as well as the DEI AB of Astra Zeneca. CL is on the Advisory Board of Autolus, ADC therapeutics, and consults for Rigel. AC received honoraria from Sobi, PharmaEssentia, and research funding from Merck, Kartos, Oncoverity. AAP received honoraria from AbbVie, Amgen, Astellas, Jazz, Sobi, Syndax; research funding from Pfizer, Servier, Incyte, Sumitomo. MWD has consulting or advisory roles (Argenx, Blueprint Medicines); honoraria for educational writing (American Society of Hematology Self-Assessment Program); honoraria (Novartis). RAL has acted as a consultant or advisor to Ariad/Takeda, CVS/Caremark, Daiichi Sankyo, Epizyme/Ipsen, and Novartis, and has received clinical research support to his institution from Ascentage, Astellas, Biomea Fusion, Cellectis, Daiichi Sankyo, and Novartis, and royalties from UpToDate.
Ethics approval and consent to participate
The study was approved by the institutional review boards and ethical committee clearance was obtained (IRB09-130-B and IRB16-0892). All clinical investigations were conducted according to the principles of the Declaration of Helsinki. Informed consent was obtained from all study participants or their legal guardians.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Harris, E.J., Karaoglu, D.A., Sukhanova, M. et al. Clinical and molecular characterization of TP53-mutant acute lymphoblastic leukemia in adults. Blood Cancer J. 15, 138 (2025). https://doi.org/10.1038/s41408-025-01350-5
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41408-025-01350-5