Abstract
Background
Because breast cancer (BC) is molecularly heterogeneous, diagnosis and treatment will likely benefit from comprehensive genetic profiling. However, routine, high-resolution sequencing is not feasible yet, due to implementation challenges associated with whole-genome sequencing of formalin-fixed paraffin embedded (FFPE) BC samples. Therefore, we explored the potential of an alternative low-resolution, genome-wide testing approach that is able to capture the copy number (CN) landscape, including actionable alterations, in FFPE derived DNA.
Methods
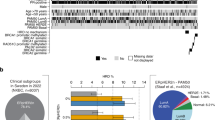
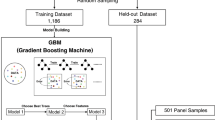
The performance of the genome-wide CN testing approach, including CN signatures/focal CN alterations, was evaluated in two phases: (i) exploration and (ii) feasibility phase. First, high-resolution sequencing data of a previously published triple-negative BC cohort (n = 237) was leveraged to benchmark the homologous recombination deficiency (HRD)-related CN signature using a comprehensive, multimodal approach incorporating both genetic and functional HRD tests. Secondly, the low-resolution testing strategy’s feasibility was prospectively evaluated in a BC cohort of patients referred to clinical genetic services (n = 147).
Results
Applying the HRD threshold that was established using both genomic and functional HRD data, we identified a 100% sensitivity for BC with BRCA1/BRCA2/PALB2 pathogenic variants in the prospective cohort. Moreover, the success rate of the low-resolution testing approach proved high, regardless of input material. Finally, additional CN alterations were enriched in the HR-proficient BC population, indicating potential actionable CN-alterations beyond HRD.
Conclusions
In conclusion, low-resolution, genome-wide sequencing has shown high potential in capturing the CN landscape, including features associated with HRD, in BC patients. This preselection testing approach is likely to maximize potential for personalized medicine and genetic counseling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The WGS data within the exploration phase (SCAN-B cohort) were derived from the following public domain resource: https://data.mendeley.com/datasets/2mn4ctdpxp/1, version 3. The RAD51-FFPE data of the SCAN-B cohort were generated and are available upon reasonable request from the corresponding author (MV). For the exploration phase, the data generated are available upon reasonable request from the corresponding author (MV).
References
Rennert G, Bisland-Naggan S, Barnett-Griness O, Bar-Joseph N, Zhang S, Rennert HS, et al. Clinical outcomes of breast cancer in carriers of BRCA1 and BRCA2 mutations. N Engl J Med. 2007;357:115–23.
Parise CA, Caggiano V. Breast Cancer Survival Defined by the ER/PR/HER2 Subtypes and a Surrogate Classification according to Tumor Grade and Immunohistochemical Biomarkers. J Cancer Epidemiol. 2014;2014:469251.
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, et al. Molecular portraits of human breast tumours. Nature. 2000;406:747–52.
van ‘t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature. 2002;415:530–6.
Angus L, Smid M, Wilting SM, van Riet J, Van Hoeck A, Nguyen L, et al. The genomic landscape of metastatic breast cancer highlights changes in mutation and signature frequencies. Nat Genet. 2019;51:1450–8.
Nathanson KL, Wooster R, Weber BL. Breast cancer genetics: what we know and what we need. Nat Med. 2001;7:552–6.
Breast Cancer Association C, Dorling L, Carvalho S, Allen J, Gonzalez-Neira A, Luccarini C, et al. Breast Cancer Risk Genes - Association Analysis in More than 113,000 Women. N. Engl J Med. 2021;384:428–39.
Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA Cancer J Clin. 2023;73:17–48.
Piccart-Gebhart MJ, Procter M, Leyland-Jones B, Goldhirsch A, Untch M, Smith I, et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N Engl J Med. 2005;353:1659–72.
Davies H, Glodzik D, Morganella S, Yates LR, Staaf J, Zou X, et al. HRDetect is a predictor of BRCA1 and BRCA2 deficiency based on mutational signatures. Nat Med. 2017;23:517–25.
Nguyen L, Martens JWM, Van Hoeck A, Cuppen E. Pan-cancer landscape of homologous recombination deficiency. Nat Commun. 2020;11:5584.
Staaf J, Glodzik D, Bosch A, Vallon-Christersson J, Reutersward C, Hakkinen J, et al. Whole-genome sequencing of triple-negative breast cancers in a population-based clinical study. Nat Med. 2019;25:1526–33.
Telli ML, Timms KM, Reid J, Hennessy B, Mills GB, Jensen KC, et al. Homologous Recombination Deficiency (HRD) Score Predicts Response to Platinum-Containing Neoadjuvant Chemotherapy in Patients with Triple-Negative Breast Cancer. Clin Cancer Res. 2016;22:3764–73.
Robson M, Im SA, Senkus E, Xu B, Domchek SM, Masuda N, et al. Olaparib for Metastatic Breast Cancer in Patients with a Germline BRCA Mutation. N Engl J Med. 2017;377:523–33.
Litton JK, Rugo HS, Ettl J, Hurvitz SA, Goncalves A, Lee KH, et al. Talazoparib in Patients with Advanced Breast Cancer and a Germline BRCA Mutation. N Engl J Med. 2018;379:753–63.
van Wijk LM, Kramer CJH, Vermeulen S, Ter Haar NT, de Jonge MM, Kroep JR, et al. The RAD51-FFPE Test; Calibration of a functional homologous recombination deficiency test on diagnostic endometrial and ovarian tumor blocks. Cancers. 2021;13:2994.
Kramer CJ, Llop-Guevara A, Yaniz-Galende E, Pellegrino B, Ter Haar NT, Herencia-Ropero A, et al. RAD51 as a biomarker for homologous recombination deficiency in high-grade serous ovarian carcinoma: robustness and interobserver variability of the RAD51 test. J Pathol Clin Res. 2023;9:442–8.
Castroviejo-Bermejo M, Cruz C, Llop-Guevara A, Gutierrez-Enriquez S, Ducy M, Ibrahim YH, et al. A RAD51 assay feasible in routine tumor samples calls PARP inhibitor response beyond BRCA mutation. EMBO Mol Med. 2018;10:e9172.
Cruz C, Castroviejo-Bermejo M, Gutierrez-Enriquez S, Llop-Guevara A, Ibrahim YH, Gris-Oliver A, et al. RAD51 foci as a functional biomarker of homologous recombination repair and PARP inhibitor resistance in germline BRCA-mutated breast cancer. Ann Oncol. 2018;29:1203–10.
Eikesdal HP, Yndestad S, Elzawahry A, Llop-Guevara A, Gilje B, Blix ES, et al. Olaparib monotherapy as primary treatment in unselected triple negative breast cancer. Ann Oncol. 2021;32:240–9.
Llop-Guevara A, Loibl S, Villacampa G, Vladimirova V, Schneeweiss A, Karn T, et al. Association of RAD51 with homologous recombination deficiency (HRD) and clinical outcomes in untreated triple-negative breast cancer (TNBC): analysis of the GeparSixto randomized clinical trial. Ann Oncol. 2021;32:1590–6.
van Wijk LM, Vermeulen S, Ter Haar NT, Kramer CJH, Terlouw D, Vrieling H, et al. Performance of a RAD51-based functional HRD test on paraffin-embedded breast cancer tissue. Breast Cancer Res Treat. 2023;202:607–16.
Meijer TG, Verkaik NS, Sieuwerts AM, van Riet J, Naipal KAT, van Deurzen CHM, et al. Functional ex vivo assay reveals homologous recombination deficiency in breast cancer beyond BRCA gene defects. Clin Cancer Res. 2018;24:6277–87.
Fu S, Yao S, Yuan Y, Previs RA, Elias AD, Carvajal RD, et al. Multicenter Phase II Trial of the WEE1 inhibitor adavosertib in refractory solid tumors harboring CCNE1 amplification. J Clin Oncol. 2023;41:1725–34.
Drews RM, Hernando B, Tarabichi M, Haase K, Lesluyes T, Smith PS, et al. A pan-cancer compendium of chromosomal instability. Nature. 2022;606:976–83.
Saal LH, Vallon-Christersson J, Hakkinen J, Hegardt C, Grabau D, Winter C, et al. The Sweden Cancerome Analysis Network - Breast (SCAN-B) Initiative: a large-scale multicenter infrastructure towards implementation of breast cancer genomic analyses in the clinical routine. Genome Med. 2015;7:20.
Ryden L, Loman N, Larsson C, Hegardt C, Vallon-Christersson J, Malmberg M, et al. Minimizing inequality in access to precision medicine in breast cancer by real-time population-based molecular analysis in the SCAN-B initiative. Br J Surg. 2018;105:e158–e68.
Robbe P, Popitsch N, Knight SJL, Antoniou P, Becq J, He M, et al. Clinical whole-genome sequencing from routine formalin-fixed, paraffin-embedded specimens: pilot study for the 100,000 Genomes Project. Genet Med. 2018;20:1196–205.
Guo Q, Lakatos E, Bakir IA, Curtius K, Graham TA, Mustonen V. The mutational signatures of formalin fixation on the human genome. Nat Commun. 2022;13:4487.
Lam SY, Ioannou A, Konstanti P, Visseren T, Doukas M, Peppelenbosch MP, et al. Technical challenges regarding the use of formalin-fixed paraffin embedded (FFPE) tissue specimens for the detection of bacterial alterations in colorectal cancer. BMC Microbiol. 2021;21:297.
Spencer DH, Sehn JK, Abel HJ, Watson MA, Pfeifer JD, Duncavage EJ. Comparison of clinical targeted next-generation sequence data from formalin-fixed and fresh-frozen tissue specimens. J Mol Diagn. 2013;15:623–33.
Akhoundova D, Rubin MA. The grand challenge of moving cancer whole-genome sequencing into the clinic. Nat Med. 2024;30:39–40.
Specialisten FMSFM. Breast cancer - Criteria for referral Clinical Genetics (Borstkanker - Criteria voor verwijzing KG) [Available from: https://richtlijnendatabase.nl/richtlijn/borstkanker/screening/screening_buiten_het_bob/criteria_voor_verwijzing_kg.html.
Daly MB, Pal T, Berry MP, Buys SS, Dickson P, Domchek SM, et al. Genetic/Familial High-Risk Assessment: Breast, Ovarian, and Pancreatic, Version 2.2021, NCCN Clinical Practice Guidelines in Oncology. J Natl Compr Canc Netw. 2021;19:77–102.
Beitsch PD, Whitworth PW, Hughes K, Patel R, Rosen B, Compagnoni G, et al. Underdiagnosis of hereditary breast cancer: are genetic testing guidelines a tool or an obstacle? J Clin Oncol. 2019;37:453–60.
Loibl S, Andre F, Bachelot T, Barrios CH, Bergh J, Burstein HJ, et al. Early breast cancer: ESMO Clinical Practice Guideline for diagnosis, treatment and follow-up. Ann Oncol. 2024;35:159–82.
Aine M, Boyaci C, Hartman J, Hakkinen J, Mitra S, Campos AB, et al. Molecular analyses of triple-negative breast cancer in the young and elderly. Breast Cancer Res. 2021;23:20.
Danecek P, Bonfield JK, Liddle J, Marshall J, Ohan V, Pollard MO, et al. Twelve years of SAMtools and BCFtools. Gigascience. 2021;10.
Breast cancer, Dutch Guideline, version 2.0. In: NABON IKN-, editor. 2012.
Tan PH, Ellis I, Allison K, Brogi E, Fox SB, Lakhani S, et al. The 2019 World Health Organization classification of tumours of the breast. Histopathology. 2020;77:181–5.
Acknowledgements
The authors would like to acknowledge patients and clinicians participating in the SCAN-B study, personnel at the central SCAN-B laboratory at the Division of Oncology, Lund University, the Swedish national breast cancer quality registry (NKBC), Regional Cancer Center South, RBC Syd, and the South Sweden Breast Cancer Group (SSBCG). Moreover, we thank all contributing clinical geneticists, clinical laboratory geneticists, genetic counselors and clinical molecular biologists in Pathology in the Leiden University Medical Center, specifically A.M. Cleton, D. Terlouw, R. van Eijk, E.M.P. Steeghs, R.B. van der Luijt, C.M. Tops, A.V.E. Harder, N. Hofland, P.A.M. de Koning, T.P. Potjer, S. Moghadasi. Moreover, the authors thank N.T. ter Haar, T.A. Rutten, R. van Rijnsoever, and E.J.J. Leijnse (all from Leiden University Medical Center), as well as J. Häkkinen (Lund University Cancer Centre), for their excellent technical support. We also thank K. Yost for support in figure illustrations.
Funding
This work was supported by the Dutch Cancer Society (grant number 12995 to TB and MV). Next-generation sequencing of breast cancer specimens (n = 150) as well as assessing BRCA1 promotor hypermethylation was financially supported by AstraZeneca (ESR-21-21497 to MV).
Author information
Authors and Affiliations
Contributions
Conceptualization: CK, KT, TB, CA, MV; Methodology: CK, TB, JS, HV, CA, MV; Formal analysis: CK, MV; Investigation: CK, LW, DR, SGV; Resources: KT, DC, TB, VS, NS, CA, JS, JVC; Data curation: CK, LW, DR, SV; Writing – original draft: CK, MV; Writing – review & editing: LW, DR, SGV, KT, NS, TW, JW, VS, DC, HV, JS, TB, CA, JVC; Visualization: CK, MV; Supervision: TB, CA, MV; Project administration: CK, CA; Funding acquisition: TB, CA, MV.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The SCAN-B study (exploration phase) (ClinicalTrials.gov ID NCT02306096) was approved by the Regional Ethical Review Board in Lund, Sweden (applicable registration numbers 2009/658, 2015/277, 2016/742, 2018/267 and 2019/01252) [26, 27]. All patients provided written informed consent prior to enrollment. For the feasibility cohort, the study design was approved by the medical ethics committee of the Leiden University Medical Center (N20.183). The study was performed in accordance with the Declaration of Helsinki.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kramer, C.J.H., van Wijk, L.M., Ruano, D. et al. Capturing breast cancers’ copy-number landscape in routine pathology: Exploiting low-resolution, genome-wide sequencing to identify HRD and beyond. Br J Cancer 133, 1199–1207 (2025). https://doi.org/10.1038/s41416-025-03134-x
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41416-025-03134-x