Abstract
Background
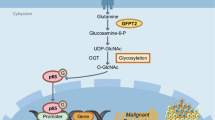
Mesothelioma is an aggressive malignancy with limited therapeutic options. Genetic alterations involving the Hippo pathway are commonly observed. O-GlcNAcylation is frequently elevated in cancer and drives tumour progression. However, its relationship with Hippo pathway dysfunction in mesothelioma remains unclear.
Methods
O-GlcNAcylation levels were examined in mesothelioma samples and cell lines, and O-GlcNAcylated proteins were detected by mass spectrometry. The functional impact of O-GlcNAcylation was determined by quantifying nuclear transport dynamics using light-induced live-cell imaging. Genetic and pharmacological inhibition of O-GlcNAcylation was evaluated in vitro. Treatment with the nuclear export inhibitor KPT-330 (Selinexor) was assessed in vitro and in a mouse xenograft model.
Results
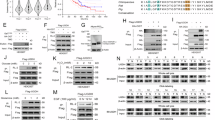
O-GlcNAcylation was markedly increased in mesothelioma cells with Hippo pathway inactivation. This modification primarily targeted nuclear pore complex proteins, including NUP214 and NUP62, and significantly accelerated nuclear export rates. Suppression of O-GlcNAcylation diminished nuclear export and inhibited cell proliferation. Importantly, pharmacological blockade of nuclear export using KPT-330 suppressed cell growth in vitro and produced significant antitumour effects in vivo.
Conclusions
These findings demonstrate O-GlcNAcylation-driven enhancement of nuclear export as a therapeutically actionable vulnerability in mesothelioma with inactivation of the Hippo pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The mass spectrometry proteomics data generated in this study have been deposited to the jPOST repository [48] and are accessible via the ProteomeXchange accession number PXD070876.
References
Carbone M, Adusumilli PS, Alexander HR Jr., Baas P, Bardelli F, Bononi A, et al. Mesothelioma: scientific clues for prevention, diagnosis, and therapy. CA Cancer J Clin. 2019;69:402–29.
Asciak R, George V, Rahman NM. Update on biology and management of mesothelioma. Eur Respir Rev. 2021;30:200226.
Baas P, Scherpereel A, Nowak AK, Fujimoto N, Peters S, Tsao AS, et al. First-line nivolumab plus ipilimumab in unresectable malignant pleural mesothelioma (CheckMate 743): a multicentre, randomised, open-label, phase 3 trial. Lancet. 2021;397:375–86.
Sekido Y, Pass HI, Bader S, Mew DJ, Christman MF, Gazdar AF, et al. Neurofibromatosis type 2 (NF2) gene is somatically mutated in mesothelioma but not in lung cancer. Cancer Res. 1995;55:1227–31.
Murakami H, Mizuno T, Taniguchi T, Fujii M, Ishiguro F, Fukui T, et al. LATS2 is a tumor suppressor gene of malignant mesothelioma. Cancer Res. 2011;71:873–83.
Tanaka I, Osada H, Fujii M, Fukatsu A, Hida T, Horio Y, et al. LIM-domain protein AJUBA suppresses malignant mesothelioma cell proliferation via Hippo signaling cascade. Oncogene. 2015;34:73–83.
Tranchant R, Quetel L, Tallet A, Meiller C, Renier A, de Koning L, et al. Co-occurring mutations of tumor suppressor genes, LATS2 and NF2, in malignant pleural mesothelioma. Clin Cancer Res. 2017;23:3191–202.
Bueno R, Stawiski EW, Goldstein LD, Durinck S, De Rienzo A, Modrusan Z, et al. Comprehensive genomic analysis of malignant pleural mesothelioma identifies recurrent mutations, gene fusions and splicing alterations. Nat Genet. 2016;48:407–16.
Hmeljak J, Sanchez-Vega F, Hoadley KA, Shih J, Stewart C, Heiman D, et al. Integrative molecular characterization of malignant pleural mesothelioma. Cancer Discov. 2018;8:1548–65.
Mizuno T, Murakami H, Fujii M, Ishiguro F, Tanaka I, Kondo Y, et al. YAP induces malignant mesothelioma cell proliferation by upregulating transcription of cell cycle-promoting genes. Oncogene. 2012;31:5117–22.
Matsushita A, Sato T, Mukai S, Fujishita T, Mishiro-Sato E, Okuda M, et al. TAZ activation by Hippo pathway dysregulation induces cytokine gene expression and promotes mesothelial cell transformation. Oncogene. 2019;38:1966–78.
Pobbati AV, Kumar R, Rubin BP, Hong W. Therapeutic targeting of TEAD transcription factors in cancer. Trends Biochem Sci. 2023;48:450–62.
Kakiuchi-Kiyota S, Schutten MM, Zhong Y, Crawford JJ, Dey A. Safety considerations in the development of Hippo pathway inhibitors in cancers. Front Cell Dev Biol. 2019;7:156.
Akao K, Sato T, Mishiro-Sato E, Mukai S, Ghani FI, Kondo-Ida L, et al. TEAD-independent cell growth of Hippo-inactive mesothelioma cells: Unveiling resistance to TEAD inhibitor K-975 through MYC signaling activation. Mol Cancer Ther. 2024;24:709–19.
Torres CR, Hart GW. Topography and polypeptide distribution of terminal N-acetylglucosamine residues on the surfaces of intact lymphocytes. Evidence for O-linked GlcNAc. J Biol Chem. 1984;259:3308–17.
Holt GD, Hart GW. The subcellular distribution of terminal N-acetylglucosamine moieties. Localization of a novel protein-saccharide linkage, O-linked GlcNAc. J Biol Chem. 1986;261:8049–57.
Yang X, Qian K. Protein O-GlcNAcylation: emerging mechanisms and functions. Nat Rev Mol Cell Biol. 2017;18:452–65.
Lu Q, Zhang X, Liang T, Bai X. O-GlcNAcylation: an important post-translational modification and a potential therapeutic target for cancer therapy. Mol Med. 2022;28:115.
Slawson C, Hart GW. O-GlcNAc signalling: implications for cancer cell biology. Nat Rev Cancer. 2011;11:678–84.
Zhang X, Qiao Y, Wu Q, Chen Y, Zou S, Liu X, et al. The essential role of YAP O-GlcNAcylation in high-glucose-stimulated liver tumorigenesis. Nat Commun. 2017;8:15280.
Peng C, Zhu Y, Zhang W, Liao Q, Chen Y, Zhao X, et al. Regulation of the Hippo-YAP pathway by glucose sensor O-GlcNAcylation. Mol Cell. 2017;68:591–604.e595.
Kim E, Kang JG, Kang MJ, Park JH, Kim YJ, Kweon TH, et al. O-GlcNAcylation on LATS2 disrupts the Hippo pathway by inhibiting its activity. Proc Natl Acad Sci USA. 2020;117:14259–69.
Hakiri S, Osada H, Ishiguro F, Murakami H, Murakami-Tonami Y, Yokoi K, et al. Functional differences between wild-type and mutant-type BRCA1-associated protein 1 tumor suppressor against malignant mesothelioma cells. Cancer Sci. 2015;106:990–9.
Sanjaya A, Goenawan H, Setiawan I, Gunadi JW, Limyati Y, Lesmana R. Elaborating the physiological role of YAP as a glucose metabolism regulator: a systematic review. Cell Physiol Biochem. 2021;55:193–205.
Chatham JC, Zhang J, Wende AR. Role of O-linked N-acetylglucosamine protein modification in cellular (patho) physiology. Physiol Rev. 2021;101:427–93.
Zhao P, Viner R, Teo CF, Boons GJ, Horn D, Wells L. Combining high-energy C-trap dissociation and electron transfer dissociation for protein O-GlcNAc modification site assignment. J Proteome Res. 2011;10:4088–104.
Yoo TY, Mitchison TJ. O-GlcNAc modification of nuclear pore complexes accelerates bidirectional transport. J Cell Biol. 2021;220:e202010141.
Niopek D, Wehler P, Roensch J, Eils R, Di Ventura B. Optogenetic control of nuclear protein export. Nat Commun. 2016;7:10624.
Chen YF, Adams DJ. Therapeutic targeting of exportin-1 beyond nuclear export. Trends Pharm Sci. 2025;46:20–31.
Ali A, Kim SH, Kim MJ, Choi MY, Kang SS, Cho GJ, et al. O-GlcNAcylation of NF-κB promotes lung metastasis of cervical cancer cells via upregulation of CXCR4 expression. Mol Cells. 2017;40:476–84.
Kim M, Kim YS, Kim H, Kang MY, Park J, Lee DH, et al. O-linked N-acetylglucosamine transferase promotes cervical cancer tumorigenesis through human papillomaviruses E6 and E7 oncogenes. Oncotarget. 2016;7:44596–607.
Zhu G, Qian M, Lu L, Chen Y, Zhang X, Wu Q, et al. O-GlcNAcylation of YY1 stimulates tumorigenesis in colorectal cancer cells by targeting SLC22A15 and AANAT. Carcinogenesis. 2019;40:1121–31.
Wu N, Jiang M, Han Y, Liu H, Chu Y, Liu H, et al. O-GlcNAcylation promotes colorectal cancer progression by regulating protein stability and potential catcinogenic function of DDX5. J Cell Mol Med. 2019;23:1354–62.
Wulff-Fuentes E, Berendt RR, Massman L, Danner L, Malard F, Vora J, et al. The human O-GlcNAcome database and meta-analysis. Sci Data. 2021;8:25.
Lin DH, Hoelz A. The structure of the nuclear pore complex (an update). Annu Rev Biochem. 2019;88:725–83.
Nofrini V, Di Giacomo D, Mecucci C. Nucleoporin genes in human diseases. Eur J Hum Genet. 2016;24:1388–95.
Hutten S, Kehlenbach RH. CRM1-mediated nuclear export: to the pore and beyond. Trends Cell Biol. 2007;17:193–201.
Roloff S, Spillner C, Kehlenbach RH. Several phenylalanine-glycine motives in the nucleoporin Nup214 are essential for binding of the nuclear export receptor CRM1. J Biol Chem. 2013;288:3952–63.
Port SA, Monecke T, Dickmanns A, Spillner C, Hofele R, Urlaub H, et al. Structural and functional characterization of CRM1-Nup214 interactions reveals multiple FG-binding sites involved in nuclear export. Cell Rep. 2015;13:690–702.
Tarbet HJ, Toleman CA, Boyce M. A sweet embrace: control of protein-protein interactions by O-linked β-N-acetylglucosamine. Biochemistry. 2018;57:13–21.
Kırlı K, Karaca S, Dehne HJ, Samwer M, Pan KT, Lenz C, et al. A deep proteomics perspective on CRM1-mediated nuclear export and nucleocytoplasmic partitioning. Elife. 2015;4:e11466.
Azmi AS, Uddin MH, Mohammad RM. The nuclear export protein XPO1—from biology to targeted therapy. Nat Rev Clin Oncol. 2021;18:152–69.
Lai C, Xu L, Dai S. The nuclear export protein exportin-1 in solid malignant tumours: From biology to clinical trials. Clin Transl Med. 2024;14:e1684.
Bogani G, Monk BJ, Coleman RL, Vergote I, Oakin A, Ray-Coquard I, et al. Selinexor in patients with advanced and recurrent endometrial cancer. Curr Probl Cancer. 2023;47:100963.
De Cesare M, Cominetti D, Doldi V, Lopergolo A, Deraco M, Gandellini P, et al. Anti-tumor activity of selective inhibitors of XPO1/CRM1-mediated nuclear export in diffuse malignant peritoneal mesothelioma: the role of survivin. Oncotarget. 2015;6:13119–32.
Fu SC, Fung HYJ, Cağatay T, Baumhardt J, Chook YM. Correlation of CRM1-NES affinity with nuclear export activity. Mol Biol Cell. 2018;29:2037–44.
Sun Q, Carrasco YP, Hu Y, Guo X, Mirzaei H, Macmillan J, et al. Nuclear export inhibition through covalent conjugation and hydrolysis of Leptomycin B by CRM1. Proc Natl Acad Sci USA. 2013;110:1303–8.
Okuda S, Yoshizawa AC, Kobayashi D, Takahashi Y, Watanabe Y, Moriya Y, et al. jPOST environment accelerates the reuse and reanalysis of public proteome mass spectrometry data. Nucleic Acids Res. 2025;53:D462–67.
Acknowledgements
We thank Teruaki Fujishita and Masahiro Aoki for technical support and comments on the manuscript. We also thank the Division for Medical Research Engineering, Nagoya University Graduate School of Medicine, for using Q Exactive. We thank Editage for English language editing. This work was supported by JSPS KAKENHI Grant Number JP22H04926, Grant-in-Aid for Transformative Research Areas—Platforms for Advanced Technologies and Research Resources ‘Advanced Bioimaging Support’.
Funding
This work was supported by JSPS KAKENHI (22K07182 (SM), 25K10503 (SM), 24K02336 (YS), and 24K22079 (YS)), Aichi Cancer Research Foundation (SM), Daiko Foundation (SM), Takeda Science Foundation (SM), and Research Grant of the Princess Takamatsu Cancer Research Fund (YS).
Author information
Authors and Affiliations
Contributions
SM: Conceptualisation; Formal analysis; Founding acquisition; Investigation; Project acquisition; Validation; Visualisation; Writing—original draft; Writing—review and editing. TS: Resources; Supervision; Writing—review and editing. YK and KK: Methodology; Resources. ES and LI: Investigation. NY: Supervision; Writing—review and editing. KH: Investigation; Resources. YS: Resources; Supervision; Funding acquisition; Writing—review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the ethics committee of Aichi Cancer Center (approval number IR041047; approval date, September 2022) and performed in accordance with the Declaration of Helsinki. The requirement for informed consent was waived by the ethics committee because of the minimal risk involved. All animal experiments were performed according to the protocols approved by the Institutional Animal Care and Use Committee of the Aichi Cancer Center.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mukai, S., Sato, T., Kamei, Y. et al. Enhanced nuclear export caused by O-GlcNAcylation of nucleoporins is a potential therapeutic target in mesothelioma. Br J Cancer (2026). https://doi.org/10.1038/s41416-026-03369-2
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41416-026-03369-2