Abstract
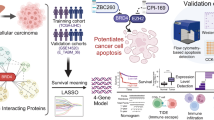
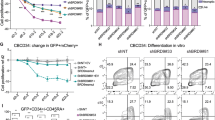
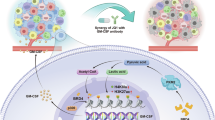
Histone lactylation couples glycolytic metabolism to oncogenic transcription, but its mechanistic readers remain poorly defined. Here, we identify bromodomain-containing protein 9 (BRD9) as a lactyl-lysine reader that links lactate-driven H3K18 lactylation (H3K18la) to chromatin remodeling in hepatocellular carcinoma (HCC). Clinically, elevated H3K18la levels correlate with poor HCC prognosis. Structural (NMR) and biophysical analyses demonstrate that BRD9’s bromodomain engages H3K18la with weak, transient affinity through its conserved acetyl-lysine pocket, distinct from its stable H3K18ac binding. This enables BRD9 to function as a metabolic-epigenetic sensor, dynamically recruited to chromatin in response to glycolytic flux. Multi-omics profiling reveals that H3K18la recruits BRD9 and the non-canonical BRG1-associated factor (ncBAF) chromatin remodeling complex to active enhancers and promoters, promoting chromatin accessibility and driving oncogenic transcription (SPARC, TMEM64, ANGEL1, SCARB1). Glycolytic inhibition or BRD9 targeting displaces BRD9 from chromatin, suppresses oncogenes, and impairs HCC proliferation. Modulating the lactylation vis p300 or HDAC inhibition attenuates transcription and reduces tumor viability. In vivo, glycolytic inhibition suppresses tumor growth. Our findings establish a feedforward loop wherein glycolytic flux promotes H3K18la-dependent BRD9-ncBAF recruitment to remodel chromatin and sustain oncogenic transcription, defining BRD9 as a critical metabolic-epigenetic mediator and a promising therapeutic target in HCC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support this study are available from the corresponding authors upon reasonable request. All protocols used in this study are available in the Methods section. RNA-seq, ChIP-seq, and ATAC-seq data generated in this study have been deposited in the NCBI SRA database under BioProject accession code (GSE314155). The structure coordinates for the BRD9 bromodomain in complex with the H3K18la peptide have been deposited in the Protein Data Bank under PDB accession codes (21GE, 21HJ). NMR data are available in the Biological Magnetic Resonance Data Bank (BMRB code: 36816, 36817). Source data are provided with this paper. Supplementary Information is available in the online version of the paper.
References
Li Y, Mo H, Wu S, Liu X, Tu K. A novel lactate metabolism-related gene signature for predicting clinical outcome and tumor microenvironment in hepatocellular carcinoma. Front Cell Dev Biol. 2021;9:801959.
Ganapathy-Kanniappan S, Kunjithapatham R, Geschwind JF. Glyceraldehyde-3-phosphate dehydrogenase: a promising target for molecular therapy in hepatocellular carcinoma. Oncotarget. 2012;3:940–53.
Vogel A, Cervantes A, Chau I, Daniele B, Llovet JM, Meyer T, et al. Hepatocellular carcinoma: ESMO clinical practice guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2018;29:iv238–iv55.
Shiani A, Narayanan S, Pena L, Friedman M. The role of diagnosis and treatment of underlying liver disease for the prognosis of primary liver cancer. Cancer Control. 2017;24:1073274817729240.
Forner A, Reig M, Bruix J. Hepatocellular carcinoma. Lancet. 2018;391:1301–14.
Dai X, Lv X, Thompson EW, Ostrikov KK. Histone lactylation: epigenetic mark of glycolytic switch. Trends Genet. 2022;38:124–7.
Vander Heiden MG, Cantley LC, Thompson CB. Understanding the warburg effect: the metabolic requirements of cell proliferation. Science. 2009;324:1029–33.
Koppenol WH, Bounds PL, Dang CV. Otto Warburg’s contributions to current concepts of cancer metabolism. Nat Rev Cancer. 2011;11:325–37.
Liberti MV, Locasale JW. The Warburg effect: how does it benefit cancer cells? Trends Biochem Sci. 2016;41:211–8.
Zhang D, Tang Z, Huang H, Zhou G, Cui C, Weng Y, et al. Metabolic regulation of gene expression by histone lactylation. Nature. 2019;574:575–80.
Yang J, Luo L, Zhao C, Li X, Wang Z, Zeng Z, et al. A positive feedback loop between inactive vhl-triggered histone lactylation and PDGFRβ signaling drives clear cell renal cell carcinoma progression. Int J Biol Sci. 2022;18:3470–83.
Yu J, Chai P, Xie M, Ge S, Ruan J, Fan X, et al. Histone lactylation drives oncogenesis by facilitating m(6)A reader protein YTHDF2 expression in ocular melanoma. Genome Biol. 2021;22:85.
Wang N, Wang W, Wang X, Mang G, Chen J, Yan X, et al. Histone lactylation boosts reparative gene activation post-myocardial infarction. Circ Res. 2022;131:893–908.
Pan RY, He L, Zhang J, Liu X, Liao Y, Gao J, et al. Positive feedback regulation of microglial glucose metabolism by histone H4 lysine 12 lactylation in Alzheimer’s disease. Cell Metab. 2022;34:634–48.e6.
Xie B, Lin J, Chen X, Zhou X, Zhang Y, Fan M, et al. CircXRN2 suppresses tumor progression driven by histone lactylation through activating the Hippo pathway in human bladder cancer. Mol Cancer. 2023;22:151.
Li F, Si W, Xia L, Yin D, Wei T, Tao M, et al. Positive feedback regulation between glycolysis and histone lactylation drives oncogenesis in pancreatic ductal adenocarcinoma. Mol Cancer. 2024;23:90.
Klein DC, Lardo SM, Hainer SJ. The ncBAF complex regulates transcription in AML through H3K27ac sensing by BRD9. Cancer Res Commun. 2024;4:237–52.
Mathupala SP, Ko YH, Pedersen PL. Hexokinase-2 bound to mitochondria: cancer’s stygian link to the “Warburg Effect” and a pivotal target for effective therapy. Semin Cancer Biol. 2009;19:17–24.
Yu D, Liang Y, Kim C, Jaganathan A, Ji D, Han X, et al. Structural mechanism of BRD4-NUT and p300 bipartite interaction in propagating aberrant gene transcription in chromatin in NUT carcinoma. Nat Commun. 2023;14:378.
Remillard D, Buckley DL, Paulk J, Brien GL, Sonnett M, Seo HS, et al. Degradation of the BAF complex factor BRD9 by heterobifunctional ligands. Angew Chem Int Ed Engl. 2017;56:5738–43.
Shi J, Wang Y, Zeng L, Wu Y, Deng J, Zhang Q, et al. Disrupting the interaction of BRD4 with diacetylated twist suppresses tumorigenesis in basal-like breast cancer. Cancer Cell. 2014;25:210–25.
Liu Z, Ye Y, Liu Y, Liu Y, Chen H, Shen M, et al. RNA helicase DHX37 facilitates liver cancer progression by cooperating with PLRG1 to drive superenhancer-mediated transcription of cyclin D1. Cancer Res. 2022;82:1937–52.
Michel BC, D’Avino AR, Cassel SH, Mashtalir N, McKenzie ZM, McBride MJ, et al. A non-canonical SWI/SNF complex is a synthetic lethal target in cancers driven by BAF complex perturbation. Nat Cell Biol. 2018;20:1410–20.
Wang X, Wang S, Troisi EC, Howard TP, Haswell JR, Wolf BK, et al. BRD9 defines a SWI/SNF sub-complex and constitutes a specific vulnerability in malignant rhabdoid tumors. Nat Commun. 2019;10:1881.
Moreno-Yruela C, Zhang D, Wei W, Bæk M, Liu W, Gao J, et al. Class I histone deacetylases (HDAC1-3) are histone lysine delactylases. Sci Adv. 2022;8:eabi6696.
Hu X, Huang X, Yang Y, Sun Y, Zhao Y, Zhang Z, et al. Dux activates metabolism-lactylation-MET network during early iPSC reprogramming with Brg1 as the histone lactylation reader. Nucleic Acids Res. 2024;52:5529–48.
Zhai G, Niu Z, Jiang Z, Zhao F, Wang S, Chen C, et al. DPF2 reads histone lactylation to drive transcription and tumorigenesis. Proc Natl Acad Sci USA. 2024;121:e2421496121.
Carey MF, Peterson CL, Smale ST. Dignam and Roeder's nuclear extract preparation. Cold Spring Harb Protoc. 2009;2009:pdb.prot5330.
Han X, Yu D, Gu R, Jia Y, Wang Q, Jaganathan A, et al. Roles of the BRD4 short isoform in phase separation and active gene transcription. Nat Struct Mol Biol. 2020;27:333–41.
Acknowledgements
We thank the biobank at the First Hospital of Jilin University (Changchun, China) for liver samples from HCC patients. We acknowledge the BGI Group for generating RNA-seq data and Seqhealth Technology Co., Ltd. for ChIP-seq data and ATAC-seq data. We thank Di Liu for the assistance with ITC and NMR experiments, and Naicui Zhai for Mass spectrometric analysis, both conducted in the core facility of the First Hospital of Jilin University. This work was supported in part by the research fund from the First Hospital of Jilin University (Changchun, China), International Center of Future Science of Jilin University, and National Natural Science Foundation of China (82372736; LZ).
Author information
Authors and Affiliations
Contributions
LZ, M-MZ and YL conceived the project. EW designed and performed experiments. EW, WZ, YL, YL and LX performed transfection in cells and qRT-PCR, EW, YJ and DJ for cellular experiments. EW, CG and BJ worked on HCC patient tissue experiments, and EW, MY and GS performed ChIP experiments. CW and CZ worked on NMR experiments. ZS and YY analyzed sequencing data. LZ and M-MZ wrote the manuscript with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics declarations
Written informed consent was obtained from all patients, and all procedures were approved by the Institute Research Ethics Committee of the First Hospital of Jilin University (No.: 2025-591). All animal experiments were approved by the Animal Ethics Committee of the College of Basic Medical Sciences of Jilin University (No.: 2025-591) and were conducted in compliance with the Guiding Principles established by the Laboratory Animal Management Committee of the College of Basic Medical Sciences of Jilin University.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wei, E., Ji, D., Jia, Y. et al. BRD9 recognizes lactate-induced H3K18 lactylation to drive oncogenic chromatin remodeling in hepatocellular carcinoma. Cell Death Differ (2026). https://doi.org/10.1038/s41418-026-01698-6
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41418-026-01698-6