Abstract
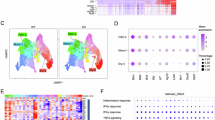
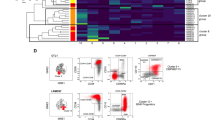
Histone methyltransferase SETD2 is recurrently mutated in hematopoietic malignancies. Our previous study showed that Setd2 deficiency impairs the self-renewal potential of murine hematopoietic stem cells (HSCs) and drives myelodysplastic syndrome (MDS)-like disorders. However, the precise oncogenic advantages conferred upon HSCs by Setd2 loss remain unclear. In this study, we found that Setd2 deficiency disrupted the fidelity of HSC lineage differentiation with preferential erythroid commitment and excessive macrophage priming, leading to ineffective erythropoiesis and the production of inflammatory embryonic-derived Kupffer cell (EmKC)-like cells. Notably, these EmKC-like cells exhibited HSC-independent self-renewal capability and remotely perturbed intramedullary hematopoiesis by inducing systemic inflammation. Furthermore, macrophage depletion effectively alleviated the inflammatory state and relieved MDS-like symptoms. Mechanistically, Setd2 loss leads to significant changes in DNA methylation and chromatin accessibility, resulting in the activation of Irf8. These findings suggest that the long-lived inflammatory cells may compensate for the HSC self-renewal defects, triggering systemic inflammation and driving hematopoietic malignant transformation. This paradigm provides a new understanding of hematopoietic malignancies with functional defects and exhaustion of HSCs.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All raw and processed data from scRNA-seq, bulk RNA-seq, ATAC-seq, and WGBS are available in the National Omics Data Encyclopedia (NODE) under Access number: OEP004811. The RNA-seq of BM LSKs from old WT and KO-MDS mice have been previously published (GEO: GSE108617). Source data are provided with this paper. The detailed experimental procedures and analytical methodologies are provided in the Supplementary Information.
References
Issa JP. The myelodysplastic syndrome as a prototypical epigenetic disease. Blood. 2013;121:3811–7.
Sperling AS, Gibson CJ, Ebert BL. The genetics of myelodysplastic syndrome: from clonal haematopoiesis to secondary leukaemia. Nat Rev Cancer. 2017;17:5–19.
Kristinsson SY, Bjorkholm M, Hultcrantz M, Derolf AR, Landgren O, Goldin LR. Chronic immune stimulation might act as a trigger for the development of acute myeloid leukemia or myelodysplastic syndromes. J Clin Oncol. 2011;29:2897–903.
Sallman DA, List A. The central role of inflammatory signaling in the pathogenesis of myelodysplastic syndromes. Blood. 2019;133:1039–48.
Barreyro L, Chlon TM, Starczynowski DT. Chronic immune response dysregulation in MDS pathogenesis. Blood. 2018;132:1553–60.
Ganan-Gomez I, Wei Y, Starczynowski DT, Colla S, Yang H, Cabrero-Calvo M, et al. Deregulation of innate immune and inflammatory signaling in myelodysplastic syndromes. Leukemia. 2015;29:1458–69.
Varney ME, Melgar K, Niederkorn M, Smith M, Barreyro L, Starczynowski DT. Deconstructing innate immune signaling in myelodysplastic syndromes. Exp Hematol. 2015;43:587–98.
Basiorka AA, McGraw KL, Eksioglu EA, Chen X, Johnson J, Zhang L, et al. The NLRP3 inflammasome functions as a driver of the myelodysplastic syndrome phenotype. Blood. 2016;128:2960–75.
Raaijmakers MH, Mukherjee S, Guo S, Zhang S, Kobayashi T, Schoonmaker JA, et al. Bone progenitor dysfunction induces myelodysplasia and secondary leukaemia. Nature. 2010;464:852–7.
Zambetti NA, Ping Z, Chen S, Kenswil KJG, Mylona MA, Sanders MA, et al. Mesenchymal inflammation drives genotoxic stress in hematopoietic stem cells and predicts disease evolution in human pre-leukemia. Cell Stem Cell. 2016;19:613–27.
Chen X, Eksioglu EA, Zhou J, Zhang L, Djeu J, Fortenbery N, et al. Induction of myelodysplasia by myeloid-derived suppressor cells. J Clin Invest. 2013;123:4595–611.
Schneider RK, Schenone M, Ferreira MV, Kramann R, Joyce CE, Hartigan C, et al. Rps14 haploinsufficiency causes a block in erythroid differentiation mediated by S100A8 and S100A9. Nat Med. 2016;22:288–97.
Ribezzo F, Snoeren IAM, Ziegler S, Stoelben J, Olofsen PA, Henic A, et al. Rps14, Csnk1a1 and miRNA145/miRNA146a deficiency cooperate in the clinical phenotype and activation of the innate immune system in the 5q- syndrome. Leukemia. 2019;33:1759–72.
Gomez Perdiguero E, Klapproth K, Schulz C, Busch K, Azzoni E, Crozet L, et al. Tissue-resident macrophages originate from yolk-sac-derived erythro-myeloid progenitors. Nature. 2015;518:547–51.
Hoeffel G, Chen J, Lavin Y, Low D, Almeida FF, See P, et al. C-Myb(+) erythro-myeloid progenitor-derived fetal monocytes give rise to adult tissue-resident macrophages. Immunity. 2015;42:665–78.
Guilliams M, Scott CL. Liver macrophages in health and disease. Immunity. 2022;55:1515–29.
Scott CL, Zheng F, De Baetselier P, Martens L, Saeys Y, De Prijck S, et al. Bone marrow-derived monocytes give rise to self-renewing and fully differentiated Kupffer cells. Nat Commun. 2016;7:10321.
Li W, Yang Y, Yang L, Chang N, Li L. Monocyte-derived Kupffer cells dominate in the Kupffer cell pool during liver injury. Cell Rep. 2023;42:113164.
Tran S, Baba I, Poupel L, Dussaud S, Moreau M, Gelineau A, et al. Impaired Kupffer cell self-renewal alters the liver response to lipid overload during non-alcoholic steatohepatitis. Immunity. 2020;53:627–40.e5.
Seidman JS, Troutman TD, Sakai M, Gola A, Spann NJ, Bennett H, et al. Niche-specific reprogramming of epigenetic landscapes drives myeloid cell diversity in nonalcoholic steatohepatitis. Immunity. 2020;52:1057–74.e7.
Remmerie A, Martens L, Thone T, Castoldi A, Seurinck R, Pavie B, et al. Osteopontin expression identifies a subset of recruited macrophages distinct from Kupffer cells in the fatty liver. Immunity. 2020;53:641–57.e14.
Fan N, Lavu S, Hanson CA, Tefferi A. Extramedullary hematopoiesis in the absence of myeloproliferative neoplasm: Mayo Clinic case series of 309 patients. Blood Cancer J. 2018;8:119.
Sun XJ, Wei J, Wu XY, Hu M, Wang L, Wang HH, et al. Identification and characterization of a novel human histone H3 lysine 36-specific methyltransferase. J Biol Chem. 2005;280:35261–71.
Hu M, Sun XJ, Zhang YL, Kuang Y, Hu CQ, Wu WL, et al. Histone H3 lysine 36 methyltransferase Hypb/Setd2 is required for embryonic vascular remodeling. Proc Natl Acad Sci USA. 2010;107:2956–61.
Kizer KO, Phatnani HP, Shibata Y, Hall H, Greenleaf AL, Strahl BD. A novel domain in Set2 mediates RNA polymerase II interaction and couples histone H3 K36 methylation with transcript elongation. Mol Cell Biol. 2005;25:3305–16.
Baubec T, Colombo DF, Wirbelauer C, Schmidt J, Burger L, Krebs AR, et al. Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature. 2015;520:243–7.
Huang H, Weng H, Zhou K, Wu T, Zhao BS, Sun M, et al. Histone H3 trimethylation at lysine 36 guides m(6)A RNA modification co-transcriptionally. Nature. 2019;567:414–9.
Luco RF, Pan Q, Tominaga K, Blencowe BJ, Pereira-Smith OM, Misteli T. Regulation of alternative splicing by histone modifications. Science. 2010;327:996–1000.
Wen H, Li Y, Xi Y, Jiang S, Stratton S, Peng D, et al. ZMYND11 links histone H3.3K36me3 to transcription elongation and tumour suppression. Nature. 2014;508:263–8.
Li F, Mao G, Tong D, Huang J, Gu L, Yang W, et al. The histone mark H3K36me3 regulates human DNA mismatch repair through its interaction with MutSalpha. Cell. 2013;153:590–600.
Pfister SX, Ahrabi S, Zalmas LP, Sarkar S, Aymard F, Bachrati CZ, et al. SETD2-dependent histone H3K36 trimethylation is required for homologous recombination repair and genome stability. Cell Rep. 2014;7:2006–18.
Chen K, Liu J, Liu S, Xia M, Zhang X, Han D, et al. Methyltransferase SETD2-mediated methylation of STAT1 is critical for interferon antiviral activity. Cell. 2017;170:492–506.e14.
Park IY, Powell RT, Tripathi DN, Dere R, Ho TH, Blasius TL, et al. Dual chromatin and cytoskeletal remodeling by SETD2. Cell. 2016;166:950–62.
Yuan H, Han Y, Wang X, Li N, Liu Q, Yin Y, et al. SETD2 restricts prostate cancer metastasis by integrating EZH2 and AMPK signaling pathways. Cancer Cell. 2020;38:350–65.e7.
Lu M, Zhao B, Liu M, Wu L, Li Y, Zhai Y, et al. Pan-cancer analysis of SETD2 mutation and its association with the efficacy of immunotherapy. NPJ Precis Oncol. 2021;5:51.
McKinney M, Moffitt AB, Gaulard P, Travert M, De Leval L, Nicolae A, et al. The genetic basis of hepatosplenic T-cell lymphoma. Cancer Discov. 2017;7:369–79.
Mar BG, Bullinger LB, McLean KM, Grauman PV, Harris MH, Stevenson K, et al. Mutations in epigenetic regulators including SETD2 are gained during relapse in paediatric acute lymphoblastic leukaemia. Nat Commun. 2014;5:3469.
Zhu X, He F, Zeng H, Ling S, Chen A, Wang Y, et al. Identification of functional cooperative mutations of SETD2 in human acute leukemia. Nat Genet. 2014;46:287–93.
Leung W, Teater M, Durmaz C, Meydan C, Chivu AG, Chadburn A, et al. SETD2 haploinsufficiency enhances germinal center-associated AICDA somatic hypermutation to drive B-cell lymphomagenesis. Cancer Discov. 2022;12:1782–803.
Parker H, Rose-Zerilli MJ, Larrayoz M, Clifford R, Edelmann J, Blakemore S, et al. Genomic disruption of the histone methyltransferase SETD2 in chronic lymphocytic leukaemia. Leukemia. 2016;30:2179–86.
Li J, Peng Z, Luo F, Chen Y. SET domain containing 2 deficiency in myelodysplastic syndrome. Front Genet. 2020;11:794.
Chen BY, Song J, Hu CL, Chen SB, Zhang Q, Xu CH, et al. SETD2 deficiency accelerates MDS-associated leukemogenesis via S100a9 in NHD13 mice and predicts poor prognosis in MDS. Blood. 2020;135:2271–85.
Zhang YL, Sun JW, Xie YY, Zhou Y, Liu P, Song JC, et al. Setd2 deficiency impairs hematopoietic stem cell self-renewal and causes malignant transformation. Cell Res. 2018;28:476–90.
Zhou Y, Yan X, Feng X, Bu J, Dong Y, Lin P, et al. Setd2 regulates quiescence and differentiation of adult hematopoietic stem cells by restricting RNA polymerase II elongation. Haematologica. 2018;103:1110–23.
Chu SH, Chabon JR, Matovina CN, Minehart JC, Chen BR, Zhang J, et al. Loss of H3K36 methyltransferase SETD2 impairs V(D)J recombination during lymphoid development. iScience. 2020;23:100941.
Ji Z, Sheng Y, Miao J, Li X, Zhao H, Wang J, et al. The histone methyltransferase Setd2 is indispensable for V(D)J recombination. Nat Commun. 2019;10:3353.
Ding Z, Cai T, Tang J, Sun H, Qi X, Zhang Y, et al. Setd2 supports GATA3(+)ST2(+) thymic-derived Treg cells and suppresses intestinal inflammation. Nat Commun. 2022;13:7468.
Liu Y, Pop R, Sadegh C, Brugnara C, Haase VH, Socolovsky M. Suppression of Fas-FasL coexpression by erythropoietin mediates erythroblast expansion during the erythropoietic stress response in vivo. Blood. 2006;108:123–33.
Shim YA, Campbell T, Weliwitigoda A, Dosanjh M, Johnson P. Regulation of CD71(+)TER119(+) erythroid progenitor cells by CD45. Exp Hematol. 2020;86:53–66.e1.
Mass E, Nimmerjahn F, Kierdorf K, Schlitzer A. Tissue-specific macrophages: how they develop and choreograph tissue biology. Nat Rev Immunol. 2023;23:563–79.
Barisas DAG, Choi K. Extramedullary hematopoiesis in cancer. Exp Mol Med. 2024;56:549–58.
Jiang K, Tian K, Yu Y, Wu E, Yang M, Pan F, et al. Kupffer cells determine intrahepatic traffic of PEGylated liposomal doxorubicin. Nat Commun. 2024;15:6136.
Liu J, Zhu Z, Leung GK. Erythrophagocytosis by microglia/macrophage in intracerebral hemorrhage: from mechanisms to translation. Front Cell Neurosci. 2022;16:818602.
Soyfer EM, Fleischman AG. Myeloproliferative neoplasms - blurring the lines between cancer and chronic inflammatory disorder. Front Oncol. 2023;13:1208089.
Rodriguez-Sevilla JJ, Colla S. Inflammation in myelodysplastic syndrome pathogenesis. Semin Hematol. 2024;61:385–96.
Stubbins RJ, Platzbecker U, Karsan A. Inflammation and myeloid malignancy: quenching the flame. Blood. 2022;140:1067–74.
Kesharwani P, Dash D, Koiri RK. Deciphering the role of hepcidin in iron metabolism and anemia management. J Trace Elem Med Biol. 2025;87:127591.
Sun Y, Tong H, Chu X, Li Y, Zhang J, Ding Y, et al. Notch1 regulates hepatic thrombopoietin production. Blood. 2024;143:2778–90.
Merz AMA, Platzbecker U. Treatment of lower-risk myelodysplastic syndromes. Haematologica. 2025;110:330–8.
Gonzalez-Menendez P, Phadke I, Olive ME, Joly A, Papoin J, Yan H, et al. Arginine metabolism regulates human erythroid differentiation through hypusination of eIF5A. Blood. 2023;141:2520–36.
Mass, E, I Ballesteros, M Farlik, F Halbritter, P Gunther, L Crozet, et al. Specification of tissue-resident macrophages during organogenesis. Science. 2016;353:aaf4238.
Simon JM, Hacker KE, Singh D, Brannon AR, Parker JS, Weiser M, et al. Variation in chromatin accessibility in human kidney cancer links H3K36 methyltransferase loss with widespread RNA processing defects. Genome Res. 2014;24:241–50.
Xie Y, Sahin M, Sinha S, Wang Y, Nargund AM, Lyu Y, et al. SETD2 loss perturbs the kidney cancer epigenetic landscape to promote metastasis and engenders actionable dependencies on histone chaperone complexes. Nat Cancer. 2022;3:188–202.
Domcke S, Bardet AF, Adrian Ginno P, Hartl D, Burger L, Schubeler D. Competition between DNA methylation and transcription factors determines binding of NRF1. Nature. 2015;528:575–9.
Yin, Y, E Morgunova, A Jolma, E Kaasinen, B Sahu, S Khund-Sayeed, et al. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science. 2017;356:eaaj2239.
Izzo F, Lee SC, Poran A, Chaligne R, Gaiti F, Gross B, et al. DNA methylation disruption reshapes the hematopoietic differentiation landscape. Nat Genet. 2020;52:378–87.
Guo Y, Xue Z, Yuan R, Li JJ, Pastor WA, Liu W. RAD: a web application to identify region associated differentially expressed genes. Bioinformatics. 2021;37:2741–3.
Wang L, You X, Ruan D, Shao R, Dai HQ, Shen W, et al. TET enzymes regulate skeletal development through increasing chromatin accessibility of RUNX2 target genes. Nat Commun. 2022;13:4709.
Friedrich, C and O Kosmider. The mesenchymal niche in myelodysplastic syndromes. Diagnostics. 2022;12:1639.
Kouroukli O, Symeonidis A, Foukas P, Maragkou MK, Kourea EP. Bone marrow immune microenvironment in myelodysplastic syndromes. Cancers. 2022;14:5656.
Tanaka TN, Bejar R. MDS overlap disorders and diagnostic boundaries. Blood. 2019;133:1086–95.
Guryanova OA, Lieu YK, Garrett-Bakelman FE, Spitzer B, Glass JL, Shank K, et al. Dnmt3a regulates myeloproliferation and liver-specific expansion of hematopoietic stem and progenitor cells. Leukemia. 2016;30:1133–42.
Kuhn R, Schwenk F, Aguet M, Rajewsky K. Inducible gene targeting in mice. Science. 1995;269:1427–9.
Georgiades P, Ogilvy S, Duval H, Licence DR, Charnock-Jones DS, Smith SK, et al. VavCre transgenic mice: a tool for mutagenesis in hematopoietic and endothelial lineages. Genesis. 2002;34:251–6.
Funding
This work was supported by National Natural Science Foundation of China (81870102, 82070150), the Young Scientists Fund of the National Natural Science Foundation of China (82400168), Natural Science Foundation of Shanghai Science and Technology Innovation Plan (23ZR1439700), Postdoctoral Fellowship Program of CPSF (GZC20241054), and Samuel Waxman Cancer Research Foundation.
Author information
Authors and Affiliations
Contributions
YLZ, QHH and XJS designed the research. YLZ, JCS, FHW, YYX, XYL, and HT performed experiments. YLZ, JCS and HFZ performed bioinformatics analysis. YLZ, JCS, FHW, YYX, XYL, HT, RF, YZ, JW, TH, HFZ, XJS, and QHH analyzed and discussed data. YLZ, JCS, and QHH wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Song, J., Wang, F., Xie, Y. et al. Aberrant Kupffer-like differentiation of hematopoietic stem cell is critical for the MDS pathogenesis in Setd2-deficient mice. Cell Death Differ (2026). https://doi.org/10.1038/s41418-026-01715-8
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41418-026-01715-8