Abstract
Ovarian cancer (OC) is a highly aggressive malignancy in women, and platinum resistance remains a major clinical obstacle. p53 mutations are prevalent in OC and exhibit functional heterogeneity that is associated with therapeutic response and disease progression. However, the roles and mechanisms underlying the functional heterogeneity of p53 mutations in platinum-resistant OC remain elusive. This investigation delineated that p53 mutations within the Loop 2, Loop 3, and β-strand S10 regions were closely linked to platinum resistance. In particular, functional assays unveiled that p53R175H and p53R175G mutations at Arg175 revealed distinct roles in tumor cell migration and drug resistance, with p53R175G conferring resistance to agents targeting p53R175H. Through multi-omics sequencing analysis, it was discerned that p53R175H and p53R175G promoted tumor progression through distinct cofactors and regulatory networks. p53R175H mediated upregulation of extracellular matrix-related genes, whereas p53R175G activated pathways associated with cytokine receptor interaction and membrane trafficking. Notably, the chromatin remodeling protein CHD1 selectively interacted with p53R175G, but not p53R175H, and regulated the transcriptional activity of p53R175G, including target genes such as IL7R. Moreover, CHD1 knockdown or pharmacological inhibition of IL7R synergistically enhanced platinum sensitivity, suggesting promising combination therapies specifically targeting the R175G mutation. The findings revealed that p53 mutations at the same residue exhibited distinct functional properties and relied on unique cofactors, offering valuable insights for precision therapy in OC.

Similar content being viewed by others
Introduction
Ovarian cancer (OC) is among the most lethal malignancies affecting the female reproductive system globally [1, 2]. Cytoreductive surgery combined with platinum-based chemotherapy remains the first-line standard of care [3]. Despite initial therapeutic responses, over 80% of patients experience disease recurrence [4], with nearly all eventually developing platinum resistance. For patients with platinum-resistant recurrence, effective treatment options are extremely limited, and the median progression-free survival is less than 1 year [5], underscoring the urgent need for innovative precision therapies.
High-throughput sequencing technologies have characterized the mutational landscape of OC. TP53 is the most frequently mutated gene in OC, with a mutation rate exceeding 80% [6, 7]. The majority of TP53 mutations are missense, with over 2000 identified, primarily concentrated in the DNA-binding domain (DBD) [8,9,10,11]. Emerging evidence has demonstrated substantial functional and prognostic heterogeneity among different p53 hotspot mutations, suggesting mutation-specific influences on therapeutic response and drug selection [12, 13]. However, the roles and mechanisms underlying the functional heterogeneity of p53 mutations in platinum-resistant recurrent OC require further elucidation, which is essential for advancing the development of personalized precision medicine strategies.
The transcription factor p53 plays a crucial role in suppressing tumor proliferation by inducing cell cycle arrest, senescence, and apoptosis [14, 15]. TP53 mutations result in a loss of p53’s tumor-suppressor function (LOF), thereby contributing to tumorigenesis in both humans and mice [16, 17]. Furthermore, mutant p53 exhibits oncogenic gain-of-function (GOF) properties, reprogramming the tumor microenvironment (TME) to promote immunosuppression, immune evasion, and therapeutic resistance, thereby accelerating tumor progression [18, 19]. Platinum resistance is a multifactorial process involving enhanced DNA repair, drug efflux pump activation, apoptotic suppression, cancer stem cell maintenance, and TME reprogramming [20,21,22]. An increasing body of evidence has demonstrated that mutant p53 influences tumor metastasis, epithelial-mesenchymal transition, and other biological processes [23,24,25]. Mutant p53 can alter genomic functions through interactions with transcription factors, mediation of chromatin remodeling, and acquiring novel DNA-binding activities [26,27,28,29,30,31]. Moreover, different p53 mutations may exhibit distinct GOF activities, and even an identical mutation may function through diverse mechanisms depending on the cellular context [13, 32,33,34], further increasing the complexity of the TP53 mutation spectrum.
Funk et al. systematically mapped the functional spectrum of TP53 missense mutations, revealing unique LOF and GOF effects among different mutations, even within the same residue. In addition, targeted therapies developed for the R175H mutation, such as APR-246 and ZMC1, have demonstrated limited efficacy in restoring the tumor-suppressive function in other R175 variants [11]. These limitations underscore the urgent need for alternative therapeutic strategies targeting non-R175H variants at Arg175. Recent studies have increasingly demonstrated the pronounced functional heterogeneity among mutation variants at the same TP53 residue [11, 35, 36]. Given the significant prognostic value of p53 mutations and their demonstrated impact on therapeutic response and drug selection [25], targeting p53 mutations represents a promising strategy to overcome platinum resistance in OC. However, the functional heterogeneity of p53 mutations necessitates further elucidation of their roles and underlying mechanisms in platinum-resistant recurrence of OC.
In this study, we investigated the association between TP53 missense mutations and platinum resistance in OC, with particular emphasis on the functional heterogeneity and molecular mechanisms distinguishing the Arg175 variants (R175H and R175G) in driving platinum resistance. This work aimed to elucidate the therapeutic implications of p53 mutation heterogeneity for individualized treatment, thereby providing new perspectives for advancing personalized precision medicine in OC.
Results
TP53 missense mutations contribute to platinum resistance in ovarian cancer
To investigate the association between TP53 mutations and platinum resistance in OC, we profiled TP53 mutations in 96 high-grade serous ovarian cancer (HGSOC) tumor samples, stratified into four groups: primary sensitive (PS), primary resistant (PR), relapsed sensitive (RS), and relapsed resistant (RR) (Fig. 1A; Details in “Methods”). Somatic TP53 mutations were identified in 83.3% of patients, which mirrors the mutation prevalence reported in Asian populations (Fig. 1B) [6, 7]. Among these, missense mutations were predominant (57%) and were localized within the DBD (Supplementary Fig. S1A, B). The identified TP53 hotspot mutations, R175, Y220, R248, R273, and G245, were consistent with the most frequently mutated TP53 residues reported in both The Cancer Genome Atlas (TCGA) and Chinese patient cohorts (Fig. 1C–E).
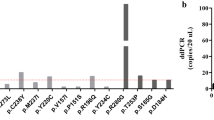
A 96 HGSOC patients were categorized into four groups: primary-sensitive, primary-resistant, relapse-sensitive, and relapse-resistant. B Somatic mutations of TP53. C–E TP53 hotspot mutations of ovarian cancer identified in this study, TCGA, and the Chinese cohort. F The frequency of TP53 missense mutations within the DBD. G The proportion of missense mutations in each secondary domain across the four groups. The Loop 3 (L3), β-strand S10 (S10), and Loop 2 (L2) domains were highlighted in color, while other regions were represented in gray. H, I CCK-8 assay validating the cisplatin sensitivity of the mutations in the primary-resistant and relapse-resistant groups. The data are presented as a nonlinear fit. J, K 50% inhibitory concentration (IC50, with 95% CI) for CDDP with NC (red) and WT (yellow) as reference.
Previous studies have predominantly focused on DBD hotspot mutations; however, the contribution of missense mutations in secondary or tertiary domains to platinum resistance remains poorly defined. Here, we comprehensively analyzed the distribution of missense mutations within the DBD across the four groups (Fig. 1F) and quantified the proportion of mutations within each secondary domain (Fig. 1G). Notably, mutations in Loop 3 (particularly amino acids 240–250) were significantly more prevalent in the PR group. Likewise, mutations in the β-strand S10 and Loop 2 were more frequent in the RR group by Fisher’s exact test (Tables S1–S3). Additionally, three-dimensional structural analyses revealed that mutations in the PR group predominantly clustered at the direct DNA-contact interface of the p53 protein (Supplementary Fig. S1C).
To validate the findings, representative mutations from the PR and RR groups were introduced into p53-null SKOV3 cells (Supplementary Fig. S1D–F; Table S4). Cell Counting Kit-8 (CCK-8) assay showed that these mutations significantly enhanced cellular tolerance to cisplatin (Fig. 1H, I), as reflected by elevated cisplatin IC50 values compared to the negative control (NC: 1.54 µM) and wild-type control (WT: 2.62 µM) (Fig. 1J, K; Table S5–S8). These statistical and functional assays indicated a close association between missense mutations in the Loop 2, Loop 3, and β-strand S10 regions of TP53 and platinum resistance in SKOV3 cells.
Notably, two relapse-resistant variants at Arg175 (R175H and R175G) exhibited distinct levels of cisplatin resistance, with R175G conferring nearly double the IC50 of R175H. Analysis of the National Cancer Institute (NCI) database revealed that R175H was more prevalent in colorectal cancer (16%), while R175G predominated in lung cancer (33%) (Supplementary Fig. S1G, H). Although both variants are linked to drug resistance, whether they share regulatory networks or possess unique functional roles warrants further investigation.
p53R175G confers enhanced cisplatin resistance and promotes tumor cell migration compared to p53R175H
To further elucidate the functional differences between p53R175G and p53R175H, we stably expressed p53NC (negative control), p53R175H, and p53R175G in p53-null SKOV3 and H1299 cells as well as in OVCAR8 cells that express endogenous p53 (Fig. 2A; Supplementary Figs. S2A and S3A). CCK-8 assay showed that both p53R175G and p53R175H significantly increased the resistance of tumor cells to cisplatin compared to the negative control, with p53R175G conferring a markedly higher level of resistance than did p53R175H (Fig. 2B; Supplementary Figs. S2B and S3B). Apoptosis assay further demonstrated that both variants significantly reduced cisplatin-induced apoptosis, with p53R175G exhibiting the strongest anti-apoptotic effect (Fig. 2C, D; Supplementary Figs. S2C, D and S3C, D). Furthermore, cell migration assay revealed that both p53R175G and p53R175H substantially promoted the migratory potential of tumor cells relative to the negative control, with the pro-migratory effect of p53R175G being particularly pronounced (Fig. 2E, F; Supplementary Figs. S2E, F and S3E, F). Wound healing assay corroborated these observations, further highlighting the superior migration-promoting effect of p53R175G (Fig. 2G, H; Supplementary Figs. S2G, H and S3G, H). Notably, p53R175G did not enhance the proliferation capacity of tumor cells; in fact, its proliferation rate was lower than that observed with p53R175H (Fig. 2I; Supplementary Figs. S2I and S3I). To further compare the phenotypic characteristics of p53R175G and p53R175H, we measured the expression of proliferation markers (cyclin E1, cyclin D1) and epithelial–mesenchymal transition markers (E-cadherin, N-cadherin, vimentin). Cells expressing p53R175G exhibited elevated N-cadherin and vimentin levels relative to those expressing p53R175H, consistent with enhanced migratory capacity, whereas cyclin E1 expression was lower, indicating reduced proliferative potential (Fig. 2J; Supplementary Figs. S2J and S3J). Additionally, we assessed the sensitivity of p53R175G to p53R175H-targeting drugs (APR-246, COTI-2) via CCK-8 assay. At concentrations that did not significantly affect p53NC or p53shTP53cell viability, inhibition of p53R175G cell viability was markedly less than that observed in p53R175H cells (Fig. 2K, L; Supplementary Figs. S2K, L and S3K, L).
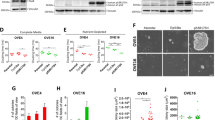
A Western blot (WB) assay validating the successful establishment of p53-null (p53NC), p53R175H, and p53R175G mutations in TP53-null SKOV3 cells. B CCK-8 assay validating the cisplatin resistance of p53NC, p53R175G and p53R175H in SKOV3 cells. The data are presented as a nonlinear fit. C, D Apoptosis assay validating the cisplatin-induced apoptosis of p53NC, p53R175G and p53R175H in SKOV3 cells. E, F Cell migration assay validating the cell migration capacity of p53R175G and p53R175H in SKOV3 cells. G, H Wound healing assay validating the wound healing rate of p53NC, p53R175G and p53R175H in SKOV3 cells. I CCK-8 assay validating the proliferation of p53NC, p53R175G and p53R175H in SKOV3 cells. J WB assay validating the expression levels of E-Cadherin, N-Cadherin, Vimentin, cyclin E1 (CCNE1) and cyclin D1 (CCND1) of p53NC, p53R175G and p53R175H in SKOV3 cells. K, L CCK-8 assay validating the sensitivity of p53NC, p53R175G and p53R175H to p53R175H-targeting drugs APR-246 and COTI-2. The data are presented as mean ± standard deviation: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, significant difference; ns, no significant difference.
Taken together, these findings suggest that p53R175G and p53R175H possess distinct functional characteristics, with p53R175G exhibiting stronger cisplatin resistance and migration-promoting properties than p53R175H. These findings indicate that the two variants may function through different molecular mechanisms.
Distinct transcriptomic signatures underlying cisplatin resistance in p53R175G and p53R175H
To further delineate the cofactors and regulatory networks associated with p53R175H and p53R175G in promoting tumor progression, RNA sequencing (RNA-seq) was conducted on SKOV3 and H1299 cell lines stably expressing either p53R175H or p53R175G, before and after cisplatin treatment. Differentially expressed genes (DEGs) were identified, and enrichment analysis prioritized genes and pathways that were consistently regulated across both cell lines, enhancing the robustness of the findings. Volcano plots illustrated genes associated with p53R175H that were significantly upregulated, downregulated, or unchanged following cisplatin treatment (Fig. 3A, B). Significantly upregulated genes were further analyzed using Metascape, revealing enrichment of pathways such as the NABA CORE MATRISOME, mitotic cell cycle, and signaling by Rho GTPases in both SKOV3 and H1299 cells treated with cisplatin (R175HCDDP group, Fig. 3E). Additionally, gene set enrichment analysis (GSEA) demonstrated a positive correlation between the ECM-receptor interaction pathway and the R175HCDDP group, involving core regulatory molecules such as COL4A6, COL6A1, FN1, ITGB3, and THBS1 (Fig. 3G, I). A parallel approach was used to examine the transcriptomic response of p53R175G cells to cisplatin (Fig. 3C, D). Functional enrichment analyses indicated significant activation of the cellular response to cytokine stimulus pathway in both cell lines (Fig. 3F). Moreover, GSEA revealed a positive correlation between the cytokine-cytokine receptor interaction pathway and the R175GCDDP group, with key molecules including IL7R, IL32, TNFRSF9, CXCL11, CXCR4, and INHBB (Fig. 3H, J).
A, B Volcano plots for DEGs in p53R175H cells treated with cisplatin, analyzed in SKOV3 and H1299 cells. Colored dots represented DEGs with p < 0.05 and fold change ≥1.5. C, D Volcano plots for DEGs in p53R175G cells treated with cisplatin, analyzed in SKOV3 and H1299 cells. Colored dots represented DEGs with p < 0.05 and fold change ≥1.5. E Enriched pathways of upregulated DEGs in R175HCDDP compared to R175H were identified using Metascape, and p values calculated via a one-sided Fisher’s exact test. F Enriched pathways of upregulated DEGs in R175GCDDP compared to R175G were identified using Metascape, and p values calculated via a one-sided Fisher’s exact test. G, H GSEA enrichment plot for DEGs in p53R175H and p53R175G cells treated with cisplatin. Nominal p < 0.05. I Key factors involved in the ECM receptor interaction pathway. J Key factors involved in the cytokine-cytokine receptor interaction pathway.
To further dissect the molecular distinctions between p53R175G and p53R175H in driving cancer progression, genes that were significantly upregulated in p53R175G compared with p53R175H (Fig. 4A, B) were subjected to functional enrichment analysis. The analysis revealed abnormal activation of VEGFA-VEGFR2 signaling and vasculature development pathways in the p53R175G group, which may underlie its stronger pro-migration capacity (Fig. 4E). Membrane trafficking, cytokine signaling in immune system, and cell morphogenesis pathways were also enriched in the p53R175GCDDP group, further suggesting an association with cisplatin resistance (Fig. 4C, D, F). To identify key drivers of p53R175G-mediated cisplatin resistance, we identified DEGs between R175GCDDP and R175HCDDP following cisplatin treatment (fold change ≥ 2, padj < 0.05) and filtered for those shared by both SKOV3 and H1299 cells, resulting in 24 candidate genes (Supplementary Fig. S4A–C). Among these, interleukin-7 receptor (IL7R) was markedly overexpressed in p53R175G (Fig. 4G, H). Functional enrichment revealed that IL7R was involved in cytokine-cytokine receptor interactions, membrane trafficking, cytokine signaling in immune system, and cell morphogenesis pathways, suggesting that IL7R may drive the oncogenic phenotype of p53R175G through multidimensional regulation (Fig. 4I). In summary, these analyses demonstrate that p53R175G and p53R175H mediate cisplatin resistance and cell migration via distinct molecular mechanisms: p53R175G-driven resistance is associated with upregulation of IL7R and activation of cytokine-related pathways, whereas p53R175H relies more on ECM-related and cell cycle regulatory networks.
A–D Volcano plots for the DEGs between p53R175G and p53R175H in SKOV3 and H1299 cells before and after cisplatin treatment. Colored dots indicated DEGs with p < 0.05 and fold change ≥1.5. E Enriched pathways of upregulated DEGs in R175G compared to R175H were identified using Metascape, with q values calculated using a one-sided Fisher’s exact test. F Enriched pathways of upregulated DEGs in R175GCDDP compared to R175HCDDP were also identified using Metascape, with q values calculated using a one-sided Fisher’s exact test. G, H Significant upregulated DEGs in R175GCDDP compared to R175HCDDP in SKOV3 and H1299 cells (fold change ≥ 2, padj < 0.05). I IL7R is implicated in cytokine-cytokine receptor interaction, membrane trafficking, cytokine signaling in immune system, and cell morphogenesis pathways.
Pharmacological targeting of IL7R overcomes p53R175G-driven tumor progression and enhances cisplatin efficacy
Previous studies have shown that IL7R promotes tumor cell survival by activating the JAK-STAT pathway [37]. To further explore the biological functions of IL7R in p53R175G cells, we first assessed IL7R expression using WB assay. The results revealed that IL7R protein levels were significantly higher in p53R175G cells compared to p53R175H cells, and this elevation was accompanied by a marked increase in STAT1 phosphorylation (p-STAT1) (Fig. 5A, B). To further substantiate the synergistic effects of OSE-127 and cisplatin, we evaluated the pharmacological inhibition of IL7R by OSE-127 in combination with cisplatin in p53R175G cells by CCK-8 assay. The combination exhibited synergistic effects, as demonstrated by the Chou–Talalay method in both cell lines (Supplementary Fig. S4D–G), whereas no significant effect was observed in p53R175H cells (Fig. 5C, D). Similarly, cell migration and wound healing assays indicated that OSE-127 treatment markedly inhibited migration capacity of p53R175G cells but had no significant effect on p53R175H-mediated migration (Fig. 5E–I). WB assay further showed that OSE-127 treatment alone significantly reduced p-STAT1 in p53R175G cells, while cisplatin treatment alone increased p-STAT1. Moreover, combined OSE-127 and cisplatin treatment reversed p-STAT1 activation and upregulated the expression of the pro-apoptotic protein Caspase-1. Under cisplatin stress, OSE-127 effectively inhibited the p-STAT1 downstream of IL7R in p53R175G cells, even when IL7R expression was increased (Fig. 5J). These findings indicate that high IL7R expression promotes p-STAT1 activation, driving cisplatin resistance and migration in p53R175G cells, while IL7R pharmacological inhibition markedly suppresses tumor progression and enhances the efficacy of cisplatin treatment.
A WB assay validating the expression levels of IL7R, STAT1, and phosphorylated STAT1 (p-STAT1) in p53R175G and p53R175H cells. B Expression validation of IL7R, STAT1, and p-STAT1 in SKOV3 and H1299 cells accompanied by the protein band intensities relative to β-Actin in scrambled cells. C, D CCK-8 assay validating that OSE-127 (10 ng/ml) significantly improved the cisplatin sensitivity of p53R175G cells. E–I Cell migration and wound healing assays validating that OSE-127 (10 ng/ml) significantly suppressed the migration of p53R175G in both SKOV3 and H1299 cell lines. J WB assay validating the expression levels of IL7R, STAT1, p-STAT1, and Caspase-1 in p53R175G cells treated with cisplatin alone, OSE-127 (10 ng/ml) alone, or the combination of both. The data are presented as mean ± standard deviation: ****P < 0.0001, significant difference; ns, no significant difference.
Distinct and shared transcriptional regulation of p53R175G and p53R175H
p53 is a key transcription factor, and its mutant forms can regulate the genome by interacting with other transcription factors, mediating chromatin remodeling and acquiring specific DNA-binding capabilities [38, 39]. To explore the transcriptional regulatory roles of p53R175H and p53R175G, we performed chromatin immunoprecipitation sequencing (ChIP-seq) on SKOV3 and H1299 cell lines. p53R175G and p53R175H displayed both shared and distinct genome-binding profiles, with SKOV3 cells showing 6092 overlapping peaks and H1299 cells 902. Furthermore, read distribution and peak localization differed between the variants (Supplementary Fig. S5A–D). These results indicated that while p53R175G and p53R175H co-regulate some targets, each mutant also binds unique genomic regions.
In SKOV3 and H1299 cells, 241 peak-associated genes were co-bound by p53R175G and p53R175H, while 1712 and 855 genes were uniquely associated with p53R175G and p53R175H, respectively (Fig. 6A–C). Functional enrichment analysis revealed that p53R175G-bound genes were mainly involved in signaling, phosphorylation, morphogenesis, and angiogenesis (Fig. 6D), whereas p53R175H-bound genes were enriched in extracellular matrix organization, chromosome organization, positive regulation of angiogenesis, and related pathways (Fig. 6E). Co-bound genes were linked to actin filament processes, cell junction organization, and cell-cell adhesion (Fig. 6F). Integration of RNA-seq and ChIP-seq analyses further demonstrated that while p53R175H and p53R175G co-regulate angiogenesis-related genes, they promote tumor progression via distinct pathways: p53R175H upregulates extracellular matrix genes, while p53R175G activates membrane trafficking and cell morphogenesis pathways.
A Wayne diagram showing p53R175G binding genes in SKOV3 and H1299 cells. B Wayne diagram showing p53R175H binding genes in SKOV3 and H1299 cells. C Wayne diagram showing peak-associated genes targeted by p53R175G and p53R175H. D Pathway enrichment plot for genes in p53R175G-target (1 712 genes). E Pathway enrichment plot for genes in p53R175H-target (855 genes). F Pathway enrichment plot for genes jointly bound by p53R175G and p53R175H (241 genes). G Wayne diagram showing p53R175G-target genes (ChIP-seq) and DEGs (fold change ≥ 2, padj < 0.05) from RNA-seq (R175GCDDP vs R175HCDDP). H ChIP-seq signals for p53R175G target genes, including DGKI, PPP2R2C, BTNL9 and KALRN. I ChIP-qPCR assay showing p53R175G enrichment at the promoter region of IL7R. J ChIP-seq signal showing p53R175G binding to the IL7R gene.
To further identify the direct transcriptional targets of p53R175G, we overlapped the p53R175G-bound genes with the 24 DEGs identified in the RNA-seq comparison between R175GCDDP and R175HCDDP. 13 direct transcriptional targets were identified, including DGKI, PPP2R2C, KALRN, and BTNL9, which were bound by p53R175G in both SKOV3 and H1299 cells (Fig. 6G). ChIP-seq data confirmed that p53R175G bound to the transcription start site (TSS) regions of these four genes (Fig. 6H). Chromatin immunoprecipitation-quantitative (ChIP)-qPCR assay was conducted and revealed that p53R175G was enriched approximately sevenfold at the IL7R promoter compared to the IgG negative control (Fig. 6I), consistent with ChIP-seq signal showing a binding peak in the IL7R promoter region (Fig. 6J). Together, these results indicated that while both p53R175G and p53R175H co-regulate angiogenesis-related genes, p53R175G upregulates IL7R and other oncogenic targets, whereas p53R175H upregulates extracellular matrix-related genes.
CHD1 specifically interacts with p53R175G
To investigate the underlying mechanism by which p53R175H and p53R175G promote tumor progression, co-immunoprecipitation-mass spectrometry (IP-MS) was applied to characterize their interacting protein profiles. Several proteins previously reported as p53R175H interactors, such as TP53BP1, USP28, and SP1, were identified. Both variants were found to interact with proteins involved in cell cycle regulation and DNA damage repair (Fig. 7A, B). Notably, pathway analysis revealed that p53R175G, compared to p53R175H, exhibited significant activation of pathways related to mRNA processing, cell cycle progression, and chromatin remodeling, particularly following cisplatin exposure (Fig. 7C, D).
A Wayne diagram showing the differentially expressed proteins (DEPs) of p53R175H or p53R175G before and after cisplatin treatment in SKOV3 and H1299 cells. DEPs were defined as proteins with a p < 0.05 and a fold change ≥1.5. B Pathway enrichment plot for the overlapping DEPs (in A) was conducted using Metascape, with q values calculated via a one-sided Fisher’s exact test. C Wayne diagram showing the DEPs (R175G vs R175H) before and after cisplatin treatment in SKOV3 and H1299 cells. DEPs were defined as proteins with a p < 0.05 and a fold change ≥1.5. D Pathway enrichment plot for the overlapping DEPs (in C) was conducted using Metascape, with q values calculated via a one-sided Fisher’s exact test. E CHD1 was identified as a key interacting protein specific to p53R175G in SKOV3 and H1299 cells. F, G Reciprocal Co-IP of CHD1 with p53NC, p53R175G and p53R175H in SKOV3 and H1299 cells. H Co-IP of CHD1 with p53WT, p53R273H and p53R175G in SKOV3 and H1299 cells.
Among the chromatin remodeling proteins, Chromodomain-helicase-DNA-binding protein 1 (CHD1) exhibited a notably stronger interaction with p53R175G than with p53R175H (Fig. 7E). This preferential interaction was further validated by reciprocal co-immunoprecipitation (Co-IP) assays. In these experiments, CHD1 was immunoprecipitated and probed for p53 to confirm the bidirectional nature of the interaction (Fig. 7F, G). To assess the specificity of this interaction, we examined the binding of CHD1 to p53WT and p53R273H in SKOV3 and H1299 cells. Neither p53WT nor p53R273H showed detectable interaction with CHD1, in contrast to the strong association observed with p53R175G (Fig. 7H), thereby highlighting the specificity of the R175G–CHD1 axis. Furthermore, survival analysis demonstrated that high CHD1 expression correlated with poorer prognosis in patients with p53-mutant OC (log-rank P < 0.05) (Supplementary Fig. S5E).
To further explore the functional significance of CHD1, we knocked down CHD1 by siRNA and observed a concomitant reduction in its association with p53R175G cells (Fig. 8A, B). CHD1 silencing also significantly decreased the expression of multiple p53R175G target genes, including IL7R, DGKI, SIRPA, MYEOV, and INHBB (Fig. 8C, D and Supplementary Fig. S5F), using RT-qPCR analyses. A luciferase assay targeting the IL7R promoter demonstrated that p53R175G overexpression increased IL7R activity in HEK293T cells, and this effect was significantly suppressed by CHD1 depletion (Supplementary Fig. S5G). Moreover, ablation of CHD1 by siRNA dramatically prevented the upregulation of IL7R and p-STAT1 in p53R175G cells (Fig. 8E, F). Functional assays revealed that CHD1 knockdown, in combination with cisplatin treatment, significantly enhanced tumor suppression and reduced the cisplatin IC50 by ~50% compared to cisplatin alone in p53R175G cells (Fig. 8G, H). Additionally, cell migration and wound healing assays showed that CHD1 silencing markedly impaired the migratory capacity of p53R175G cells (Fig. 8I–L). Collectively, these findings demonstrate that CHD1 supports p53R175G-driven transcriptional activation of oncogenic genes such as IL7R, promoting tumor progression. Conversely, depletion of CHD1 synergistically sensitizes p53R175G cells to cisplatin and suppresses migratory potential, providing a rationale for combination therapies specifically targeting the R175G mutation.
A, B WB assay demonstrated that CHD1 knockdown using siRNA disrupted its interaction with p53R175G. C RT-PCR assay showed that CHD1 knockdown significantly downregulated the mRNA levels of p53R175G-driven genes, including IL7R, DGKI, SIRPA, SORCS2, AP1M2, and MYEOV in SKOV3 cells. D RT-PCR assay showed that CHD1 knockdown significantly decreased the mRNA levels of p53R175G-driven genes, including IL7R, DGKI, DCAF4L1, SIRPA, SORCS2, AP1M2, MFAP5, and INHBB in H1299 cells. E, F WB assay showed that CHD1 knockdown decreased IL7R protein levels and reduced p-STAT1 expression in SKOV3 and H1299 cells. G, H CCK-8 assay showed that CHD1 knockdown significantly reduced cisplatin resistance of p53R175G in SKOV3 and H1299 cells. I, J Cell migration assay showed that CHD1 knockdown inhibited the migration ability of p53R175G in SKOV3 and H1299 cells. K, L Wound healing assay showed that CHD1 knockdown significantly reduced the wound healing rate of p53R175G in SKOV3 and H1299 cells. The data are presented as mean ± standard deviation: *P < 0.05, ***P < 0.001, and ****P < 0.0001, significant differences.
Discussion
Although the functional heterogeneity of TP53 mutations is recognized [26,27,28,29,30,31, 40], the functional differences and mechanistic underpinnings of TP53 mutations at the same amino acid remain poorly understood. In this study, we have demonstrated that p53R175H and p53R175G exerted distinct effects on cell migration and platinum resistance. Remarkably, even substitutions at the same codon could exhibit unique cofactor interactions. For instance, p53R175G, but not p53R175H, specifically interacted with CHD1. This unique interaction regulated the transcriptional activity of p53R175G and modulated the IL7R-STAT1 signaling pathway, thereby promoting tumor progression. Furthermore, inhibition of either CHD1 or IL7R sensitized p53R175G cells to platinum, suggesting a potential therapeutic strategy for tumors harboring the R175G mutation.
Growing evidence demonstrates significant functional divergence among TP53 mutations at the same residue. For example, in non-small-cell lung cancer, R248Q mutation enhanced tumor invasiveness, while R248W did not [35]. Similarly, R273H and R273C mutations promoted proliferation, invasion, and drug resistance, while R273G mutation lacked these effects [36]. Recent studies on R175 variants indicated that R175H mutation primarily drove migration, invasion, and metastasis, whereas R175T and R175S mutations promoted apoptosis and cell cycle arrest, respectively [11]. In our study, we found that R175G mutation exerted a stronger effect on apoptosis compared to proliferation and exhibited superior platinum resistance and pro-migratory capacity compared to R175H mutation. Moreover, R175G mutation was unresponsive to targeted agents such as APR-246 and COTI-2, corroborating previous reports of resistance in R175 mutations [11]. These results underscore the importance of considering the functional heterogeneity and cofactor dependence of TP53 mutations in precision medicine, as both the molecular mechanisms and clinical intervention strategies may vary among mutations at the same residue, thus providing critical insights for personalized treatment approaches.
Our finding showed that CHD1 specifically interacted with p53R175G to regulate a distinct transcriptional program, including the upregulation of IL7R, DGKI, SIRPA, MYEOV, and INHBB, which provided new insight into the epigenetic mechanisms of platinum resistance. As an ATP-dependent chromatin remodeling enzyme, CHD1 plays a crucial role in regulating chromatin structure, gene expression, DNA repair, and replication [41, 42]. Previous studies have indicated that CHD1 supports tumor growth by modulating metabolic genes such as HK2 and LDHA, with its inhibition suppressing proliferation and inducing apoptosis, particularly in PTEN-deficient models [43]. Extending these insights, our results suggested that CHD1 may regulate genes associated with platinum sensitivity and cell migration, making the development of selective CHD1 inhibitors or degraders a promising precision therapeutic strategy for R175G mutation cancers.
Similarly, cytokine signaling in promoting tumor progression and therapy resistance has been increasingly recognized [44,45,46,47,48,49]. IL7R activation is well documented in hematological malignancies such as acute lymphoblastic leukemia, where it drives pro-survival and proliferation pathways (notably via JAK/STAT, MAPK/ERK, and PI3K/AKT/mTOR signaling) [37, 50,51,52,53,54]. Our findings showed that IL7R was especially upregulated in p53R175G cells, leading to enhanced STAT1 activation and extending the significance of IL7R signaling to mutant p53-driven solid tumors. While IL7/IL7R signaling was primarily associated with STAT5 activation, emerging reports and our findings indicated that STAT1 could also be engaged [55], imparting a dual function: although classically a tumor suppressor, STAT1 has been implicated in chemoresistance [56, 57] and immune evasion through induction of factors such as PD-L1 [58, 59] and ROS-scavenging proteins [60]. This aligns with our observation that STAT1 phosphorylation sustained platinum resistance and cell motility, while its inhibition sensitized cells to cisplatin-induced apoptosis. Taken together with the existing literature, these findings suggested that targeting CHD1 or IL7R may represent promising therapeutic strategies for overcoming platinum resistance and cell migration in p53R175G-mutant cancers.
Our findings on the distinct roles of p53R175H and p53R175G in cancer progression inspire promising research avenues. While structural biochemical analyses were not included, integrating crystallography and biophysical methods offers a prime opportunity to clarify how these mutations alter p53 structure/function and tumorigenic networks. Our discovery that CHD1/IL7R inhibition enhances platinum sensitivity in p53R175G cells identifies novel therapeutic targets; future in vivo and clinical validation will strengthen translational potential and enable personalized strategies for p53-mutant cancers. Exploring other Arg175 mutations is critical, as these studies, combined with our results, will construct a comprehensive functional map of p53 mutations at Arg175.
In conclusion, our study elucidates the distinct functional roles and molecular mechanisms of p53R175H and p53R175G mutations in platinum-resistant OC. We demonstrate that mutations at the same arginine residue can drive tumor progression through divergent transcriptional programs and unique cofactor interactions, notably involving CHD1 in the case of p53R175G. These findings underscore the necessity of characterizing mutation-specific pathways and cofactor dependencies for the development of effective, individualized therapeutic strategies.
Materials and methods
Ethics approval and consent to participate
Secondary analyses of de-identified, publicly available data only. Original ethics approvals and informed consent for the HGSOC cohort were obtained by Qian et al. (Zhejiang Cancer Hospital IRB-2020-155; Westlake University 20190401GTN0009) in accordance with the Declaration of Helsinki. No new participants were enrolled and no identifiable information was used.
Public data
TP53-targeted sequencing data from 96 HGSOC patients were obtained from the Genome Sequence Archive, accession number HRA007126 [2]. Patients were classified into four groups based on recurrence-free survival (RFS) following platinum chemotherapy: primary sensitive (PS, n = 27, RFS > 6 months), primary resistant (PR, n = 26, RFS ≤ 6 months), relapsed sensitive (RS, n = 26, RFS > 6 months), and relapsed resistant (RR, n = 17, RFS ≤ 6 months). Additional data on TP53 missense mutations were retrieved from TCGA, cBio Cancer Genomics Portal (cBioPortal), and NCI. All datasets are publicly available, and all analyses complied with respective data access and publication policies.
Cell culture
All cell lines, including human OC cell lines (SKOV3, COV362, Kuramochi, OVCAR4, OVCAR8, A2780), human non-small cell lung cancer (H1299), and human embryonic kidney cells (293T), were all obtained from Wuhan Pricella Biotechnology Co., Ltd (Wuhan, China). Cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM; Gibco, 11965092, Waltham, MA, USA) supplemented with 10% (v/v) fetal bovine serum (FBS; Excell, FSD500, Jiangyin, China) at 37 °C in a humidified incubator with 5% CO2.
Lentiviral infection
Full-length TP53 was synthesized (Hangzhou Loche Biomedical Science and Technology Co., Ltd., Hangzhou, China), and mutations were introduced by site-directed mutagenesis using overlapping primers (Supplemental primers), then cloned into the pCDH-flagN-eGFP plasmid. The pLKO.1 plasmid was used for TP53 and CHD1 knockdown via siRNA.
Lentiviruses were produced in 293T cells by co-transfecting psPAX2, pMD2.G, and either pCDH-flagN-eGFP or pLKO.1 plasmids with TG transfection reagent. Viral supernatants were collected, filtered, and stored at −80 °C. For transduction, target cells were seeded in 24-well plates and infected with lentivirus and Polybrene (6 μg/ml; Beyotime, Shanghai, China). Medium was replaced after 24 h, and cells were cultured for an additional 48 h. Transfection efficiency was assessed by WB assay.
Cell viability assay
The Cell Counting Kit-8 (CCK-8) assay was employed to evaluate cell viability. Cells were seeded into 96-well plates at a density of ~5000 cells per well and exposed to varying concentrations of cisplatin (Selleck, S1166, Houston, TX, USA), APR-246 (Selleck, S7724, Houston, TX, USA), COTI-2 (Selleck, S8580, Houston, TX, USA), or OSE-127 (MCE, HY-P99412, New Jersey, USA) for 48 h. Subsequently, 10 μl of CCK-8 reagent (Beyotime, Shanghai, China) was added to each well, followed by incubation at 37 °C for 1 h. The optical density at 450 nm was measured using a SpectraMax M5 Automated Microplate Reader (Molecular Devices, USA).
Dose–response combination assays and calculation of drug combination index
For dose–response combination assays, cells were treated for 48 h with varying concentrations of two drugs, either alone or in combination. Cell viability was measured using a CCK-8 kit (Beyotime, Shanghai, China) according to the manufacturer’s protocol. Drug interactions were evaluated using CompuSyn software based on the Chou–Talalay method, which calculates the combination index (CI) via the isobologram equation, where CI < 1, =1, and >1 indicate synergism, additive effect, and antagonism, respectively [61].
Cell migration assay
3 × 10⁴ cells were resuspended in 400 μl of serum-free medium and seeded into the upper chamber of a 24-well culture plate containing an 8.0 μm pore-size polycarbonate membrane (Corning, 3422, New York, USA). 1 ml of medium supplemented with 10% FBS was added to the lower chamber. After 24 h, the cells were fixed with 4% paraformaldehyde and stained with crystal violet (Beyotime, C0121, Shanghai, China) for 30 min. The migrated cells were imaged using an optical microscope.
Wound healing assay
Cells were seeded into a 6-well plate and cultured until reaching 90% confluence the following day. A 200 μl pipette tip was used to create a scratch in the cell monolayer, simulating a wound. The medium was replaced with serum-free medium, and the cells were incubated for 24 h. Wound closure was imaged using a microscope (Olympus, Japan) and quantified with ImageJ software.
Apoptosis assay
Due to the spontaneous green fluorescence of the cells, the Annexin V-PE apoptosis detection kit (Beyotime, C1065L, Shanghai, China), which emits red fluorescence, was selected to detect apoptosis signals. The procedure was conducted in accordance with the manufacturer’s instructions. Following cisplatin treatment for 48 h, both the supernatant and cells were collected. The cells were resuspended in 195 μl of Annexin V-PE binding solution, and 5 μl of Annexin V-PE was added. The mixture was incubated in the dark for 20 min and analyzed using flow cytometry (FACSCALBUR, BD Biosciences, USA).
Co-immunoprecipitation (Co-IP) and Western blot assays
Cells were lysed in RIPA lysis buffer (Beyotime, P0013C) supplemented with 1% protease inhibitors. Anti-FLAG M2 beads (Sigma, M8823, Saint Louis, USA) were incubated with cell lysates overnight at 4 °C to immunoprecipitate FLAG-tagged proteins. The beads were washed, and the eluted proteins were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). The proteins were transferred onto a polyvinylidene fluoride membrane (Millipore, IPFL00010, Massachusetts, USA). The membrane was blocked with 5% skim milk at room temperature for 1 h, followed by overnight incubation at 4 °C with primary antibodies. The following day, the membrane was incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies (Abclonal, 1:10,000, Wuhan, China) at room temperature for 1 h. Protein bands were detected using enhanced chemiluminescence (ECL) reagents (Solarbio, Beijing, China).
The primary antibodies used were as follows: anti-FLAG (AE004, 1:4000; ABclonal, Wuhan, China), anti-β-actin (66009-1-Ig, 1:5000; Proteintech, Wuhan, China), IL7R (A11678, 1:2000; ABclonal, Wuhan, China), STAT1 (A19563, 1:2000; ABclonal, Wuhan, China), Phospho-STAT1-Y701 (AP0135, 1:2000; ABclonal, Wuhan, China), CHD1 (1:1000, Santa Cruz, Texas, USA), p53 (10442-1-AP, 1:10,000; Proteintech, Wuhan, China), HRP Goat Anti-Rabbit IgG (AS014, 1:10,000; ABclonal, Wuhan, China), HRP Goat Anti-Mouse IgG (AS003, 1:10,000; ABclonal, Wuhan, China), Caspase-1 (P79884R2S, 1:1000, Abmart, Shanghai, China). E-Cadherin (A20798, 1:1000; ABclonal, Wuhan, China), N-Cadherin (A19083, 1:1000; ABclonal, Wuhan, China), Vimentin (A19607, 1:1000; ABclonal, Wuhan, China), Cyclin E1 (11554-1-AP, 1:1000, Proteintech, Wuhan, China), and Cyclin D1 (60186-1-Ig, 1:1000, Proteintech, Wuhan, China).
RNA extraction, RT-PCR and qRT-PCR analysis
Total RNA was extracted from treated cells using the NcmSpin Cell/Tissue Total RNA kit (NCM, M5105) following the manufacturer’s protocol. Complementary DNA (cDNA) was synthesized via reverse transcription (RT-PCR) using the PrimeScript RT reagent kit (Takara, RR037Q, Japan). Quantitative real-time PCR (qRT-PCR) was performed using the TB Green Premix Ex Taq II kit (Takara, RR820Q, Japan) on a Bio-Rad CFX Opus 96 Real-Time PCR System in accordance with the manufacturer’s instructions. The primers used are listed in the Supplemental Materials.
Dual-luciferase reporter assay
To evaluate the effect of p53R175G and CHD1 on IL7R promoter activity, the IL7R promoter, TP53 R175G, and siCHD1 sequences were cloned into the pGL4-Luc and pCDNA3.1-HA vectors, respectively, and co-transfected into HEK293T cells for 48 h. Luciferase activity was measured using the Dual-Luciferase Reporter Assay System (Promega, E1910), with firefly signals normalized to Renilla luciferase activity.
RNA sequencing (RNA-seq)
RNA-seq was performed on SKOV3 and H1299 cell lines treated with or without 1 µM cisplatin (Selleck, S1166, Houston, TX, USA), with three biological replicates per group. Total RNA was extracted and assessed for quality using an Agilent 2100 Bioanalyzer. Libraries were prepared using the NEB general protocol, quantified with a Qubit 2.0 Fluorometer, and evaluated for insert size before sequencing on an Illumina platform. Library preparation, sequencing, and data analysis were conducted by Novogene Co., Ltd. (Beijing, China).
Immunopurification–mass spectrometry (IP-MS)
SKOV3 and H1299 cells expressing FLAG-p53 were lysed in IP buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 1% NP-40, and 1 mM EDTA) supplemented with a protease inhibitor cocktail (Selleck, B14001, Houston, TX, USA). The lysates were incubated overnight with Anti-FLAG M2 beads (Sigma, M8823, Saint Louis, USA) at 4 °C. The beads were washed, enzymatically digested with mass spectrometry-grade trypsin (Promega, V5280), desalted using Pierce C18 Tips (Thermo Scientific™, 87784, USA), vacuum-dried, and frozen. The resulting peptides were analyzed using Orbitrap Exploris™ 480 (Thermo Fisher Scientific, USA), and raw data were processed with MaxQuant software for protein identification and label-free quantification.
Chromatin immunoprecipitation (ChIP) assay
Cells were crosslinked with 1% formaldehyde for 10 min at room temperature and quenched with 2.5 M glycine. After washing with cold TBS, cells were lysed in cell lysis buffer (10 mM Tris-HCl, pH 7.5, 10 mM NaCl, 0.5% NP-40) and incubated on ice for 10 min. For 3–4 × 10⁶ cells, chromatin was digested in 0.5 ml MNase digestion buffer with 5 µl of 1:10 diluted MNase (NEB, M0247S) at 37 °C with shaking for 20 min. Chromatin was sheared by sonication. 1% Input was retained at −20 °C. Immunoprecipitation was performed overnight at 4 °C with indicated antibodies, followed by incubation with protein A/G beads at 4 °C for 3 h. Beads were washed sequentially with 1× Stop/ChIP buffer, high-salt buffer, Tris/LiCl buffer, and TE buffer twice. Chromatin was eluted in elution buffer (10 mM Tris-HCl, pH 8.0, 10 mM EDTA, 150 mM NaCl, 5 mM DTT, 1% SDS) at 65 °C, and crosslinks were reversed by overnight incubation at 65 °C. RNA and proteins were removed by RNase A (TRAN, GE101-01, Beijing, China) and Proteinase K digestion (TRAN, GE201-01, Beijing, China), respectively. DNA was purified using a PCR Purification Kit (Qiagen, 28106, Hilden, Germany) and analyzed by qRT-PCR for target enrichment; primers are listed in the supplemental materials. ChIP-seq libraries were prepared and sequenced on an Illumina platform (Novogene Co., Ltd., Beijing, China).
Statistical analysis
Each experiment in this study was conducted at least three times. Data analysis was performed using GraphPad Prism 10.0 and R 4.3.2. Statistical comparisons were evaluated using Fisher’s exact test, Student’s t-test, and one-way ANOVA. Statistical significance was indicated as follows: *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
Data availability
The raw RNA-seq and ChIP-seq data are deposited in the Sequence Read Archive (SRA) under accession ID PRJNA1233431 and PRJNA1245600. Additional data are available within the Article, and Supplementary Information.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Qian L, Zhu J, Xue Z, Zhou Y, Xiang N, Xu H, et al. Proteomic landscape of epithelial ovarian cancer. Nat Commun. 2024;15:6462.
Ledermann JA. First-line treatment of ovarian cancer: questions and controversies to address. Ther Adv Med Oncol. 2018;10:1758835918768232.
Lheureux S, Gourley C, Vergote I, Oza AM. Epithelial ovarian cancer. Lancet. 2019;393:1240–53.
Pujade-Lauraine E, Fujiwara K, Ledermann JA, Oza AM, Kristeleit R, Ray-Coquard IL, et al. Avelumab alone or in combination with chemotherapy versus chemotherapy alone in platinum-resistant or platinum-refractory ovarian cancer (JAVELIN Ovarian 200): an open-label, three-arm, randomised, phase 3 study. Lancet Oncol. 2021;22:1034–46.
Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474:609–15.
Ahmed AA, Etemadmoghadam D, Temple J, Lynch AG, Riad M, Sharma R, et al. Driver mutations in TP53 are ubiquitous in high grade serous carcinoma of the ovary. J Pathol. 2010;221:49–56.
Attardi LD, Boutelle AM. Targeting p53 gain-of-function activity in cancer therapy: a cautionary tale. Cell Death Differ. 2024;31:133–5.
Hoyos D, Greenbaum B, Levine AJ. The genotypes and phenotypes of missense mutations in the proline domain of the p53 protein. Cell Death Differ. 2022;29:938–45.
Donehower LA, Soussi T, Korkut A, Liu Y, Schultz A, Cardenas M, et al. Integrated analysis of TP53 gene and pathway alterations in the Cancer Genome Atlas. Cell Rep. 2019;28:1370–.e5.
Funk JS, Klimovich M, Drangenstein D, Pielhoop O, Hunold P, Borowek A, et al. Deep CRISPR mutagenesis characterizes the functional diversity of TP53 mutations. Nature Genet. 2025;57:140–53.
Tuna M, Ju Z, Yoshihara K, Amos CI, Tanyi JL, Mills GB. Clinical relevance of TP53 hotspot mutations in high-grade serous ovarian cancers. Br J Cancer. 2020;122:405–12.
Hanel W, Marchenko N, Xu S, Yu SX, Weng W, Moll U. Two hot spot mutant p53 mouse models display differential gain of function in tumorigenesis. Cell Death Differ. 2013;20:898–909.
Janic A, Abad E, Amelio I. Decoding p53 tumor suppression: a crosstalk between genomic stability and epigenetic control?. Cell Death Differ. 2025;32:1–8.
Kaiser AM, Attardi LD. Deconstructing networks of p53-mediated tumor suppression in vivo. Cell Death Differ. 2018;25:93–103.
Harvey M, McArthur MJ, Montgomery CA Jr, Bradley A, Donehower LA. Mice deficient for p53 are developmentally normal but susceptible to spontaneous tumours. Nature. 1992;356:215–21.
Malkin D, Li FP, Strong LC, Fraumeni JF Jr, Nelson CE, Kim DH, et al. Germ line p53 mutations in a familial syndrome of breast cancer, sarcomas, and other neoplasms. Science. 1990;250:1233–8.
Kennedy MC, Lowe SW. Mutant p53: it’s not all one and the same. Cell Death Differ. 2022;29:983–7.
Mantovani F, Collavin L, Del Sal G. Mutant p53 as a guardian of the cancer cell. Cell Death Differ. 2019;26:199–212.
Rottenberg S, Disler C, Perego P. The rediscovery of platinum-based cancer therapy. Nat Rev Cancer. 2021;21:37–50.
Richardson DL, Eskander RN, O’Malley DM. Advances in ovarian cancer care and unmet treatment needs for patients with platinum resistance: a narrative review. JAMA Oncol. 2023;9:851–9.
Damia G, Broggini M. Platinum resistance in ovarian cancer: role of DNA repair. Cancers. 11.2019;11:119.
Aschauer L, Muller PAJ. Novel targets and interaction partners of mutant p53 Gain-Of-Function. Biochem Soc Trans. 2016;44:460–6.
Bargonetti J, Prives C. Gain-of-function mutant p53: history and speculation. J Mol Cell Biol. 2019;11:605–9.
Muller PAJ, Vousden KH. Mutant p53 in cancer: new functions and therapeutic opportunities. Cancer Cell. 2014;25:304–17.
Di Agostino S, Sorrentino G, Ingallina E, Valenti F, Ferraiuolo M, Bicciato S, et al. YAP enhances the pro-proliferative transcriptional activity of mutant p53 proteins. EMBO Rep. 2016;17:188–201.
Kim E, Deppert W. Interactions of mutant p53 with DNA: guilt by association. Oncogene. 2007;26:2185–90.
Will K, Warnecke G, Wiesmüller L, Deppert W. Specific interaction of mutant p53 with regions of matrix attachment region DNA elements (MARs) with a high potential for base-unpairing. Proc Natl Acad Sci USA. 1998;95:13681–6.
Zhu J, Sammons MA, Donahue G, Dou Z, Vedadi M, Getlik M, et al. Gain-of-function p53 mutants co-opt chromatin pathways to drive cancer growth. Nature. 2015;525:206–11.
Monteith JA, Mellert H, Sammons MA, Kuswanto LA, Sykes SM, Resnick-Silverman L, et al. A rare DNA contact mutation in cancer confers p53 gain-of-function and tumor cell survival via TNFAIP8 induction. Mol Oncol. 2016;10:1207–20.
Stiewe T, Haran TE. How mutations shape p53 interactions with the genome to promote tumorigenesis and drug resistance. Drug Resist Updat. 2018;38:27–43.
Freed-Pastor WA, Mizuno H, Zhao X, Langerød A, Moon SH, Rodriguez-Barrueco R, et al. Mutant p53 disrupts mammary tissue architecture via the mevalonate pathway. Cell. 2012;148:244–58.
Weissmueller S, Manchado E, Saborowski M, Morris JP, Wagenblast E, Davis CA, et al. Mutant p53 drives pancreatic cancer metastasis through cell-autonomous PDGF receptor β signaling. Cell. 2014;157:382–94.
Muller PA, Trinidad AG, Timpson P, Morton JP, Zanivan S, van den Berghe PV, et al. Mutant p53 enhances MET trafficking and signalling to drive cell scattering and invasion. Oncogene. 2013;32:1252–65.
Yoshikawa K, Hamada J, Tada M, Kameyama T, Nakagawa K, Suzuki Y, et al. Mutant p53 R248Q but not R248W enhances in vitro invasiveness of human lung cancer NCI-H1299 cells. Biomed Res. 2010;31:401–11.
Li J, Yang L, Gaur S, Zhang K, Wu X, Yuan YC, et al. Mutants TP53 p.R273H and p.R273C but not p.R273G enhance cancer cell malignancy. Hum Mutat. 2014;35:575–84.
Lodewijckx I, Mentens N, Jacobs K, Cools J. Oncogenic cooperation between IL7R-JAK-STAT pathway mutations. Hemasphere. 2021;5:e637.
Di Agostino S, Strano S, Emiliozzi V, Zerbini V, Mottolese M, Sacchi A, et al. Gain of function of mutant p53: the mutant p53/NF-Y protein complex reveals an aberrant transcriptional mechanism of cell cycle regulation. Cancer Cell. 2006;10:191–202.
Zhou X, Hao Q, Lu H. Mutant p53 in cancer therapy-the barrier or the path. J Mol Cell Biol. 2019;11:293–305.
Sabapathy K, Lane DP. Therapeutic targeting of p53: all mutants are equal, but some mutants are more equal than others. Nat Rev Clin Oncol. 2018;15:13–30.
Iyer J, Gentry LK, Bergwell M, Smith A, Guagliardo S, Kropp PA, et al. The chromatin remodeling protein CHD-1 and the EFL-1/DPL-1 transcription factor cooperatively down regulate CDK-2 to control SAS-6 levels and centriole number. PLoS Genet. 2022;18:e1009799.
Kari V, Mansour WY, Raul SK, Baumgart SJ, Mund A, Grade M, et al. Loss of CHD1 causes DNA repair defects and enhances prostate cancer therapeutic responsiveness. EMBO Rep. 2018;17:1609–23.
Zhao D, Lu X, Wang G, Lan Z, Liao W, Li J, et al. Synthetic essentiality of chromatin remodelling factor CHD1 in PTEN-deficient cancer. Nature. 2017;542:484–8.
Chen Y, Guo Y, Li S, Xu J, Zhao C, Wang J, et al. Tumor-derived IL-6 promotes chordoma invasion by stimulating tumor-associated macrophages M2 polarization and TNFα secretion. Int Immunopharmacol. 2024;143:113315.
Zheng P, He J, Fu Y, Yang Y, Li S, Duan B, et al. Engineered bacterial biomimetic vesicles reprogram tumor-associated macrophages and remodel tumor microenvironment to promote innate and adaptive antitumor immune responses. ACS Nano. 2024;18:6863–86.
Qiao X, Zhu L, Song R, Shang C, Guo Y. CD44 occurring alternative splicing promotes cisplatin resistance and evokes tumor immune response in oral squamous cell carcinoma cells. Transl Oncol. 2023;31:101644.
Mao X, Xu J, Wang W, Liang C, Hua J, Liu J, et al. Crosstalk between cancer-associated fibroblasts and immune cells in the tumor microenvironment: new findings and future perspectives. Mol Cancer. 2021;20:131.
Tang B, Zhu J, Shi Y, Wang Y, Zhang X, Chen B, et al. Tumor cell-intrinsic MELK enhanced CCL2-dependent immunosuppression to exacerbate hepatocarcinogenesis and confer resistance of HCC to radiotherapy. Mol Cancer. 2024;23:137.
Kumari N, Choi SH. Tumor-associated macrophages in cancer: recent advancements in cancer nanoimmunotherapies. J Exp Clin Cancer Res. 2022;41:68.
Wang X, Chang S, Wang T, Wu R, Huang Z, Sun J, et al. IL7R is correlated with immune cell infiltration in the tumor microenvironment of lung adenocarcinoma. Front Pharm. 2022;13:857289.
Barata JT, Durum SK, Seddon B. Flip the coin: IL-7 and IL-7R in health and disease. Nat Immunol. 2019;20:1584–93.
Oliveira ML, Veloso A, Garcia EG, Iyer S, Pereira C, Barreto VM, et al. Mutant IL7R collaborates with MYC to induce T-cell acute lymphoblastic leukemia. Leukemia. 2022;36:1533–40.
Silva A, Almeida A, Cachucho A, Neto JL, Demeyer S, de Matos M, et al. Overexpression of wild-type IL-7Rα promotes T-cell acute lymphoblastic leukemia/lymphoma. Blood. 2021;138:1040–52.
González-García S, Mosquera M, Fuentes P, Palumbo T, Escudero A, Pérez-Martínez A, et al. IL-7R is essential for leukemia-initiating cell activity of T-cell acute lymphoblastic leukemia. Blood. 2019;134:2171–82.
Huber A, Allam AH, Dijkstra C, Thiem S, Huynh J, Poh AR, et al. Mutant TP53 switches therapeutic vulnerability during gastric cancer progression within interleukin-6 family cytokines. Cell Rep. 2024;43:114616.
Chang W-M, Chang YC, Yang YC, Lin SK, Chang PM, Hsiao M. AKR1C1 controls cisplatin-resistance in head and neck squamous cell carcinoma through cross-talk with the STAT1/3 signaling pathway. J Exp Clin Cancer Res. 2019;38:245.
Bharadwaj U, Eckols TK, Xu X, Kasembeli MM, Chen Y, Adachi M, et al. Small-molecule inhibition of STAT3 in radioresistant head and neck squamous cell carcinoma. Oncotarget. 2016;7:26307–30.
Ma H, Yang W, Zhang L, Liu S, Zhao M, Zhou G, et al. Interferon-alpha promotes immunosuppression through IFNAR1/STAT1 signalling in head and neck squamous cell carcinoma. Br J Cancer. 2019;120:317–30.
Chen S, Crabill GA, Pritchard TS, McMiller TL, Wei P, Pardoll DM, et al. Mechanisms regulating PD-L1 expression on tumor and immune cells. J Immunother Cancer. 2019;7:305.
Li Y-J, Zhang C, Martincuks A, Herrmann A, Yu H. STAT proteins in cancer: orchestration of metabolism. Nat Rev Cancer. 2023;23:115–34.
Chou T-C. Theoretical basis, experimental design, and computerized simulation of synergism and antagonism in drug combination studies. Pharmacol Rev. 2006;58:621–81.
Acknowledgements
We are grateful to Beijing Novogene Co., Ltd. for assisting in sequencing and/or bioinformatics analysis. Graphical Abstract was created with BioRender.com. This work was supported by grants from the National Key Research and Development Program of China (2017YFA0503900), the Zhejiang Provincial Natural Science Foundation of China (LDQ23C050002), the Fundamental Research Funds for the Central Universities (No. 20220228), and Zhejiang Province Basic Public Welfare Research Program (LGC22H160009).
Author information
Authors and Affiliations
Contributions
YH, HX, and ZZ designed and supervised the project; YL, MS, and XY performed experiments. YL, YZ, BT, and LW interpreted results of experiments. YL, ZZ, and MN interpreted the proteomic data. YL and YZ prepared figures. YL wrote the manuscript with inputs from co-authors. YH and HX supported and supervised the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Giovanni Blandino
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, Y., Zheng, Z., Ni, M. et al. Distinct functional heterogeneity of TP53 R175 mutations in platinum-resistant ovarian cancer: unveiling molecular mechanisms and therapeutic targets. Cell Death Dis 16, 837 (2025). https://doi.org/10.1038/s41419-025-08172-0
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41419-025-08172-0