Abstract
Pancreatic ductal adenocarcinoma (PDAC) is a highly lethal cancer, with chronic metabolic disorders increasing risk and severity. Prolonged exposure to altered metabolism changes specific metabolite levels, impacting epigenetic landscape contributing neoplastic lesion acquisition. This study examines the interplay between metabolism and epigenetics in dysmetabolic-driven PDAC tumorigenesis, exploiting LSL-KrasG12D;PDX-1-Cre mice (KC mice) exposed to high-fat diet (HFD) and KRAS-mutated human pancreatic ductal epithelial (HPDE) cells. Untargeted metabolomics of HFD-fed KC pancreata reveals altered free fatty acid and elevated S-adenosyl methionine levels during tumorigenesis. Targeted metabolomics shows increased succinate alongside reduced α-ketoglutarate levels. This imbalance suggests an epigenetic derangement, targeting DNA methylation. In KRAS-mutated HPDE cells exposed to altered metabolism, the DNA demethylation complex of ten-to-eleven-translocation methylcytosine 1 and thymine DNA glycosylase (TDG) is disrupted, leading to iterative cytosine modification and apurinic/apyrimidinic (AP) site accumulation. Succinate directly binds TDG at arginine 275, hyperactivating it and increasing AP site formation. This alteration combined with the methylation-prone metabolic environment, impairs the base excision repair pathway by hypermethylating and downmodulating DNA ligases LIG1 and LIG3. This predisposes to genomic instability and pancreatic preneoplastic lesion development. These findings uncover a metabolic-epigenetic axis in dysmetabolic PDAC, highlighting how metabolite-driven epigenetic changes compromise DNA repair and drive tumorigenesis.

Similar content being viewed by others
Introduction
Pancreatic ductal adenocarcinoma (PDAC), the most common form of pancreatic cancer, is a highly aggressive and lethal tumor with a 5-year overall survival rate of 13% [1]. The high mortality is mainly due to late diagnosis, as a consequence of lack of early specific symptoms, and to limited and non-efficient treatment options.
Chronic metabolic alterations, here defined as dysmetabolic condition, include hyperglycemia, type 2 diabetes and obesity [2]. They expose pancreatic tissue to an imbalance of specific metabolites generally involved in cell physiological function and epigenome integrity maintenance, contributing to PDAC development [3,4,5,6,7]. Indeed, dysmetabolic conditions are well-established PDAC risk factors impacting DNA methylation landscape in pancreatic cells [8, 9]. Notably, pancreatic exocrine cells isolated from high fat diet (HFD) fed mice, a common model of chronic metabolic alterations, exhibit different methylated DNA regions, some of which are located within DNA repair genes [8]. Dysmetabolic conditions promote genome instability and impair DNA repair mechanisms [10], in part by impacting expression and/or activity of DNA repair enzymes [10]. One mechanism through which this occurs is the epigenetic silencing of these enzymes. Importantly, epigenetic enzymes require specific metabolites as substrates or cofactors to exert their reactions [11], and for this reason they result extremely sensitive to small fluctuations of metabolite levels.
Among epigenetic reactions, DNA methylation is finely regulated by the activity of DNA methyltransferases (DNMTs), ten-to-eleven-translocation (TETs) methylcytosine dioxygenases and thymine DNA glycosylase (TDG) [12]. Specifically, DNMTs catalyze cytosine methylation (5mC), which in turn can be oxidized by TETs to 5-hydroxymethylcytosine (5hmC), 5-formylcytosine (5fC) and 5-carboxylcytosine (5caC) [13,14,15]. TDG recognizes and removes both 5fC and 5caC, consequently generating an apurinic/apyrimidinic site (AP site) [16, 17]. This event activates the base excision repair (BER) machinery, which recognizes single strand break and restores unmethylated cytosine by activation of DNA ligases 1 (LIG1) and 3 (LIG3), key enzymes of DNA repair [18]. The entire process depends on the availability of specific metabolites [19]. DNMTs are activated by S-adenosyl-methionine (SAM) and inhibited by S-adenosyl-homocysteine (SAH), whereas TETs are positively regulated by α-ketoglutarate (αKG) and inhibited by D-2-hydroxyglutarate (D-2HG), succinate (SA) and fumarate [19, 20]. We recently described that αKG binds TDG at arginine 275 (R275) and positively regulates its activity, supporting its association with TET1 to carry out the DNA demethylation cycle [21]. Interestingly, recent studies correlated TDG overexpression with poor prognosis in hepatocellular carcinoma, melanoma and glioma [22,23,24], shedding light onto its function as epigenome stability keeper. The association between TDG and PDAC tumorigenesis has not been specifically established, as well as the involvement of metabolic derangement in TDG activity regulation supporting PDAC onset and progression.
Hereby, we describe not previously identified metabolic-epigenetic mechanisms supporting pancreatic cancer tumorigenesis, under dysmetabolic conditions. Specifically, metabolomics analyses revealed an imbalance of metabolites derived from the tricarboxylic acid cycle (i.e., decreased αKG and increased SA) with the consequent impairment of the DNA demethylation cycle, due to TET1/TDG complex dissociation. This led to the accumulation of iterative cytosine modifications, including 5fC, and AP sites. AP site accumulation was associated to TDG alteration, as a consequence of SA-dependent TDG hyperactivation and BER impairment due to LIG1 and LIG3 promoter hyper-methylation. This scenario prevents single strand break repair and increases genome instability, supporting pancreatic pre-neoplastic lesion development.
Results
High fat diet fosters PDAC tumorigenesis
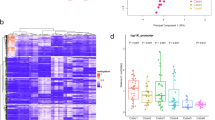
Since PDAC associates with chronic metabolic diseases [3, 7, 25], metabolomics analyses were performed to track metabolic changes during dysmetabolism-dependent PDAC onset. Specifically, to recapitulate metabolic alterations known to support PDAC onset, LSL-KrasG12D/+;PDX-1-Cre (KC) mice were fed either a high-fat diet (HFD) or a low-fat diet (LFD), as a control, for 1.5 (T1) and 5 months (T2). KC mouse model was preferred over LSL-KrasG12D/+;Trp53R172H/+; Pdx-1-Cre (KPC) mice for its much slower progression from PanIN to invasive cancer [26]. This slower advancement allows studies on pancreatic cancer tumorigenesis, particularly enabling the investigation of how metabolic alterations influence this process. Analysis of body weight (Fig. S1A), blood cholesterol (Fig. S1B), and fasting blood glucose levels (Fig. S1C) confirmed dysmetabolism onset in HFD-fed KC mice. Indeed, all metabolic parameters resulted increased already at T1 in HFD-fed KC mice compared to LFD-fed KC mice, indicating a diet-dependent dysmetabolism acquisition, including insulin resistance, as confirmed by the oral glucose tolerance test (OGTT; Fig. S1D). Histological analyses of T1 and T2 HFD-fed KC mice pancreata indicated an acceleration of tissue derangement with clear signs of acinar-to-ductal metaplasia (ADM) and early pancreatic intraepithelial neoplasia (PanIN) at T1 (15.7%), whereas ADM/PanIN (33.2%) and well differentiated PDAC (6.5%) at T2 compared to LFD-fed KC mice (ADM/PanIN: T1 = 6.8%; T2 = 23.6%; Fig. S1E). Nevertheless, no statistically significant differences were found in terms of lesion number at both time points in LFD- and HFD-fed KC mice (Fig. S1F), indicating a similar kras transgene activation. The early HFD-dependent proliferative stimulus at T1 was also confirmed by transcriptomic analysis on whole pancreatic tissue, which well separated transcripts of LFD- and HFD-fed KC mice according to the heatmap (Fig. S1G). Notably, 275 genes were differentially expressed (>1 log2 fold change) in HFD- vs LFD-fed KC mice (Fig. S1H; Supplementary Table 1). Gene-ontology (GO) analysis performed on significantly up- or downregulated transcripts showed an up-regulation of genes mainly grouped in immune system process (GO: 0002376), proliferation (GO: 0090023; GO: 0000165), migration (GO: 2000147), and insulin signaling (GO: 0043568) (Fig. S1H), suggesting the presence of an early proliferative stimulus into pancreata derived from HFD-fed KC mice compared to LFD-ones. Conversely, downregulated genes were associated with glucose metabolism (GO: 0005975; GO: 1904659) and physiological pancreatic function, including pancreatic juice secretion (GO: 0030157) (Fig. S1H). Western blot analyses corroborated these results. Increased levels of ERK1/2 phosphorylation at T202/Y204 and T185/Y186 residues were observed in pancreata from HFD-fed KC mice compared to LFD-fed KC mice at both time points (Fig. S1I). These results imply that different pathways associated with pancreatic cancer onset were induced early at T1 in HFD-fed KC mice with HFD-associated pro-proliferative marker increase sustained till T2, supporting dysmetabolism-dependent pancreatic tumorigenesis acceleration.
Dysmetabolic-dependent PDAC tumorigenesis associates with metabolism reshaping
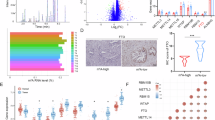
To investigate the metabolic changes during dysmetabolism-dependent PDAC onset, pancreata from LFD- and HFD-fed KC mice were collected at T1 and T2 and analyzed by untargeted metabolomics. 36 statistically significant metabolites (p < 0.05, FDR corrected) resulted differentially modulated in the whole pancreatic tissue at both time points (Fig. 1A; Supplementary Table 2). Interestingly, a selective dysregulation of a specific panel of free fatty acids (FFA) was detected: oleic acid (OA) was upregulated, and linoleic acid was downregulated (Fig. 1B, C), whereas palmitic acid level remained stable (Fig. S2A). In this context, the top three enriched pathways at T2 were glycine and serine metabolism, methylhistidine and methionine metabolism, respectively (Fig. 1D). Besides its known role in fueling one-carbon metabolism in cancer cells, serine is involved in oncogenesis-supportive metabolites as well as in the methionine cycle [27]. Serine metabolism and methionine cycle are correlated through folate cycle and, together, support production of SAM [28]. Interestingly, higher levels of SAM, a crucial metabolite for DNA methylation process representing the methyl-group donor [28], were revealed by untargeted metabolomics analysis (Fig. S2B). This result prompted us to focus on DNA methylation. Specifically, we performed a restricted targeted analysis on a pool of epigenetic-linked metabolites and we found decreased levels of αKG paralleled by increased levels of SA in HFD-fed compared to LFD-fed KC mice, which altered αKG/SA ratio (Fig. 1E–G). Taken together, these data highlight an HFD-dependent metabolic reprogramming responsible for the accumulation of metabolites involved in methylation processes.
A Heatmap showing the top 36 statistically significant differentially regulated metabolites identified by untargeted metabolomics analysis in pancreata derived from LFD and HFD mice at T1 (1.5 month) and T2 (5 months); B MS normalized intensity of OA by untargeted metabolomics analysis in LFD (black circles) and HFD (black squares) mice at T1 and T2; C MS normalized intensity of linoleic acid by untargeted metabolomics analysis in LFD (black circles) and HFD (black squares) at T1 and T2; D Enrichment analysis overview showing the top 25 metabolite pathways enriched in HFD compared to LFD mice at T2. The enrichment ratio is calculated as the number of hits within a particular metabolic pathway divided by the expected number of hits; E Concentration of αKG by targeted metabolomics in LFD (black circles) and HFD (black squares) at T1 and T2; F Concentration of SA by targeted metabolomics in LFD (black circles) and HFD (black squares) at T1 and T2. G Graphs show the αKG/SA ratio quantified by targeted metabolomics analysis in LFD (white bar) and HFD (gray bars) mice at time T1 and T2. B–C–E–F–G. n = 5 mice per group for T1 and n = 6 mice per group for T2. Lines represent median values. Statistical significance was calculated using 2-way ANOVA and is shown as *p < 0.05; **p < 0.01; ***p < 0.001.
High fat diet alters DNA methylation cycle and TET1/TDG complex assembly
TET1/TDG complex formation depends on αKG availability, representing a dysmetabolic sensor directly impacting on the DNA methylation pathway [21]. According to the identified metabolic landscape supporting methylation reactions (see above and Fig. 1), we investigated by immunoprecipitation whether this phenomenon also occurred in pancreata derived from LFD- and HFD-fed KC mice. Interestingly, we found TET1/TDG dissociation early at T1 in pancreata derived from HFD-fed KC mice compared to LFD-fed KC mice (Fig. 2A). These dissociation is sustained also at T2, implying a correlation with dysmetabolic condition over time (Fig. S2C). Consistently, dot blot analyses revealed higher levels of 5mC in pancreata derived from HFD-fed KC mice compared to LFD-fed KC mice (Fig. 2B), suggesting HFD dependent DNA methylation pathway activation. Interestingly, pancreata derived from HFD-fed KC mice displayed global accumulation also of the other iterative cytosine modifications: 5hmC and 5fC (Fig. 2B) [29]. 5fC levels, a direct readout of TET1/TDG complex activity [30], were further investigated in FFPE sections of HFD-fed KC mouse pancreatic tissue to identify the specific cell type accumulating this cytosine modification. Confocal analyses reported prevalent 5fC accumulation in pancreatic E-Cadherin positive epithelial cells compared to αSMA positive fibroblasts (Fig. 2C). These results confirmed TET1/TDG complex sensitivity to the intracellular metabolic status also in pancreas and revealed an accumulation of cytosine modifications, including 5fC, in pre-neoplastic epithelial cells.
A Left, Representative co-IP/WB analysis of the DNA demethylation complex TET1/TDG and quantification of TET1 and TDG levels in LFD (white bars) and HFD (gray bars) mouse pancreata upon TDG IP; Right, Representative co-IP/WB of the DNA demethylation complex TET1/TDG analysis and quantification of TET1 and TDG levels in LFD (white bars) and HFD (gray bars) mouse pancreata upon TET1 IP; B Left, Dot blot analysis of 5mC, 5hmC and 5fC levels performed on genomic DNA extracted from LFD and HFD mouse pancreata. Right, Dot blot densitometry of 5mC, 5hmC and 5fC levels in LFD (white bars) and HFD (gray bars) mouse pancreata. C Left, Representative confocal microscopy images depicting HFD mouse pancreas section probed with an anti-5fC antibody (red) and an anti-E-cadherin (green) in the left panel, and an anti-5fC (red) and an anti-αSMA (green) in the right panel. Nuclei were counterstained with DRAQ5 (blue). Original scale bar, 25 μm. Right, Quantification of 5fC signal intensity in E-cadherin and αSMA cells. A–C. Data expressed as average ± SEM. Statistical significance was calculated using Kolmogorov–Smirnov test (A, B) or Wilcoxon test (C) and is shown as *p < 0.05, **p < 0.01 and ***p < 0.001.
Dysmetabolism promotes TET1/TDG complex dissociation, 5fC accumulation, and a higher proliferation rate in human pancreatic ductal epithelial cells
In vitro validation of our findings was conducted exploiting human pancreatic duct epithelial (HPDE) cells harboring the KRASG12V mutation, as model of pre-neoplastic epithelial cells. Specifically, HPDE cells were exposed to a combination of high glucose (hG; final concentration of 30 mM) and OA (50 µM) to recapitulate dysmetabolic conditions observed in HFD-fed KC mice, as assessed by blood (Fig. S1B) and metabolomics analysis (Fig. 1B), or to solvent control. Dysmetabolic HPDE cells were co-cultured in a 2D culture system together with pancreatic stellate cells (PSCs), stromal cells able to support pancreatic cancer onset and progression once activated [31]. We found a prevalent 5fC accumulation in HPDE cells counterstained by E-Cadherin paralleled by an absence of 5fC signal in PSCs (αSMA positive cells) (Fig. 3A). Moreover, dysmetabolic HPDE cells were analyzed to evaluate SAM, αKG and SA levels by untargeted and targeted metabolomics. Interestingly, we revealed SAM (Fig. 3B) and SA (Fig. 3C) accumulation paralleled by αKG reduction (Fig. 3D), determining αKG/SA ratio imbalance (Fig. S2D) in dysmetabolic HPDE cells, perfectly mirroring what observed in pancreata from HFD-fed KC mice (Fig. 1E–G). Moreover, dysmetabolic HPDE cells were tested for accumulation of iterative cytosine modifications (Fig. 3E). The effect of exposure to the combination of hG and OA can also be mimicked by exposing HPDE cells exclusively to a cell-permeable source of SA, a metabolite supporting methylation reactions [32]. SA was sufficient to induce iterative cytosine modification accumulation (Fig. 3F), TET1/TDG complex dissociation (Fig. 3G), and TET1 nucleus-cytoplasmatic shuttling (Fig. 3H). Specifically, SA was able also to confer a proliferative advantage by increasing both the proliferation rate (Fig. 3I) and pERK levels (Fig. 3J) in SA-treated HPDE cells compared to solvent control. Taken together, these data suggest SA as a crucial metabolite supporting TET1/TDG complex dynamics and global iterative cytosine modification accumulation.
A Representative immunofluorescence (IF) images depicting 2D co-culture system between HPDE cells and PSCs exposed to hG and OA probed by an anti-αSMA (red) and an anti-5fC antibody (green) (upper panel), and an anti-E-cadherin (red) and an anti-5fC (green) (lower panel). Nuclei were counterstained with DAPI (blue). Original scale bar, 50 μm; B MS normalized intensity of SAM by untargeted metabolomics analysis in HPDE cells (white bar) and HPDE cells exposed to hG and OA for 24 h (gray bar); C Concentration of SA by targeted metabolomics in HPDE cells (white bar) and HPDE cells exposed to hG and OA for 24 h (gray bar); D Concentration of αKG by targeted metabolomics in HPDE cells (white bar) and HPDE cells exposed to hG and OA for 24 h (gray bar). E Left, Dot blot analysis of 5mC, 5hmC and 5fC levels performed on genomic DNA extracted from HPDE cells ± hG and OA for 48 h. Right, Dot blot densitometry of 5mC, 5hmC and 5fC levels in HPDE cells (white bars) and HPDE cells exposed to hG and OA for 48 h (gray bars); F Left, Dot blot analysis of 5mC, 5hmC and 5fC levels performed on genomic DNA extracted from HPDE cells ± 50 μM Succinate (SA) for 24 h. Right, Dot blot densitometry of 5mC, 5hmC and 5fC levels in HPDE cells (white bars) and HPDE cells exposed to 50 μM SA for 24 h (black bars); G Representative co-IP/WB analysis of the DNA demethylation complex TET1/TDG and quantification of TET1 and TDG levels in HPDE cells (white bars) and HPDE cells exposed to 50 μM SA for 24 h (black bars); H Left, Representative confocal microscopy images depicting HPDE cells exposed to control solvent (C, upper panels) or SA (lower panels) and probed by an anti-TDG antibody (green, left panels) and an anti-TET1 (red, middle left panels). Nuclei were counterstained with DAPI (blue, middle right panels). Merge signals are depicted in right panels. Original scale bar, 10 μm. Right, Quantification of TET1/TDG colocalization in HPDE cells (C) and HPDE cells exposed to SA (SA). I Left, Representative phase contrast microscopy images of HPDE cells (C) and HPDE cells exposed to 50 μM SA for 24 h (SA). Scale bar, 100 μm. Right, cell number and crystal violet quantification in HPDE cells (C) and and HPDE cells exposed to 50 μM SA for 24 h (SA). J Left, Representative WB analysis of proliferation marker pERK in HPDE cells (C) and HPDE cells exposed to 50 μM SA for 24 h (SA). Loading control: Vinculin; Right, Densitometry of pERK levels in HPDE cells (C, white bar) and HPDE cells exposed to SA (SA, black bar). Data expressed as average ± SEM. Statistical significance was calculated using Kolmogorov–Smirnov test (B–G and I, J) or Wilcoxon test (H) and is shown as *p < 0.05; **p < 0.01; ***p < 0.001.
Succinate binds the thymine DNA glycosylase at arginine 275
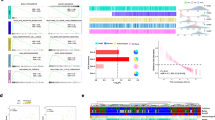
Metabolites are able to bind to several enzymes, affecting their activity [20]. We previously demonstrated that αKG acts as an enzymatic allosteric activator of TDG by direct binding [21]. In this light, the SA role as TDG activity regulator was investigated and possible hotspots for SA binding to TDG were analyzed by molecular dynamics (MD) simulations. Intermolecular recognition was monitored along MD time in three independent MD replicas of 500 ns each, in which the SA ion was initially placed in a random orientation and at a non-binding distance to TDG (i.e., intermolecular distance > 40.0 Å). In all MD replicas, the SA molecule quickly approached TDG due to electrostatic complementarity with the catalytic site. Interestingly, the metabolite interacts with the same arginine 275 (R275) residue (Fig. 4A) crucial for αKG binding [21]. R275 is a key residue of TDG, being responsible for the binding to the phosphate backbone of a DNA AP site, according to previous structural studies [33]. Besides direct H-bond and electrostatic interactions to R275, SA/TDG catalytic site interaction is reinforced by a network of water-bridged H-bonds connecting the molecule to the side chain of Serine 273, and Asparagine 157, as well as to the backbone of Glycine 142 and Alanine 145 (Fig. 4B). The persistence of the direct interaction between SA and R275 was evaluated by monitoring the intermolecular distance along MD time. Results clearly showed that SA recognition of TDG is an early event and that the SA/TDG complex is stable in MD simulation time, except for a few non-significant and short fluctuations of the SA-R275 distance (Fig. 4A). Cellular thermal shift experiments validated specific binding of SA to R275. SA protects wild type TDG (TDGwt) from thermal degradation, whereas TDGR275A mutant is degraded at 54 °C (Fig. 4C). To further characterize SA-TDG binding effect, an active murine recombinant TDG protein (mrTDG) was exploited for surface plasmon resonance (SPR) experiments. (Fig. S3A). All measured dissociation constants (KD) were in the millimolar range, with SA exibhiting the highest affinity for TDG (KD = 2.94 ± 0.03 mM) (Fig. S3A) compared to αKG (KD = 9.01 ± 0.04 mM) (Fig. S3B), demonstrating an effective SA/TDG interaction. To assign a functional role of SA-TDG binding, TDG activity assay was performed on mrTDG in the presence or absence of SA and αKG. Interestingly, SA-dependent TDG activity was higher in comparison to αKG and solvent control (Fig. 4D). This led to an increase of AP sites in the DNA as revealed by dot blot analysis (Fig. 4E). These results demonstrate that TDG turnover is metabolically controlled and sensitive not only to αKG [21], but also to SA fluctuations. SA-dependent TDG higher activity compared to αKG suggests its hyper-activation contributing to pancreatic tumorigenesis upon metabolic derangement.
A Plot of the distance between the mass center of SA and R275 along MD trajectory, showing the recognition and interaction of SA in proximity of R275; B Structural detail of the SA/TDG interaction from the most populated cluster of frames as extrapolated from MD trajectories by cluster analysis. TDG: green cartoon and lines, SA: yellow sticks. H-bond interactions: black dashed lines; TDG residues H-bonded to SA are shown as sticks; C Upper, Cell extract thermal shift assay (CETSA)/WB analysis performed on myc-TDG and myc-TDGR275A overexpressing HEK293T cells evaluated at 54°C ± SA (n = 3); Lower, Densitometry of 3 independent experiments; D TDG activity assay of mrTDG in response to αKG (blue bar) or SA (black bar). Water was used as solvent (white bar); E Left Dot blot analysis performed on genomic DNA extracted from HPDE cells ± 50 μM SA and labeled with aldehyde reactive probe (ARP) to detect AP sites. Right, Dot blot densitometry of AP site levels in HPDE cells (white bar) and HPDE cells exposed to 50 μM SA for 24 h (black bar). Data expressed as average ± SEM. Statistical significance was calculated using Kolmogorov–Smirnov test (C, E) and one-way ANOVA (D) is shown as *p < 0.05; **p < 0.01.
Dysmetabolism-dependent base excision repair machinery impairment and TDG turnover alteration contribute to AP site accumulation in pre-tumoral pancreatic epithelial cells
TDG, a G/T mismatch-specific thymine DNA glycosylase [34], recognizes DNA lesions and catalyzes the first step of the BER pathway, which leads to the generation of an AP site [35]. Thus, we investigated whether dysmetabolism-associated SA-increase could induce TDG hyper-activation, fostering AP site formation and DNA lesion accumulation in HPDE cells. Specifically, HPDE cells were exposed for 48 h to a combination of hG and OA. Thereafter, TDG enzymatic activity and consequent AP site accumulation within the DNA were evaluated. Figure 5A shows a significant increase of TDG activity in dysmetabolic HPDE cells (hG and OA enriched medium). Figure 5B depicts AP site accumulation comparable to that observed with SA treatment (Fig. 4E). Of note, proximity ligation assay (PLA) experiments revealed a specific localization of TDG in AP site enriched regions (Fig. 5C). Similar AP site accumulation was also observed in pancreata of HFD-fed KC mice (Fig. S4A), to further confirm the sensitivity of BER process to metabolic derangement. To test whether this phenomenon involved TDG enzymatic activity, loss-of-function experiments were performed. CRISPR/Cas9-mediated TDG knockdown (Fig. S4B) prevented AP site accumulation in the presence of hG and OA combination (Fig. 5D). When TDGwt or TDGR275A were re-expressed in TDG-deprived HPDE cells (Fig. 5E), AP site accumulation was appreciated only upon TDGwt enzyme re-expression in the presence of hG and OA combination (Fig. 5F). TDGR275A mutant enzyme overexpression, not sensitive to dysmetabolism, does not lead to AP site accumulation upon hG and OA combination (Fig. 5F).
A TDG activity assay in dysmetabolic HPDE cells in response to hG and OA for 48 h (gray bar). Water was used as solvent (white bar); B Left, Dot blot analysis performed on genomic DNA extracted from HPDE cells ± hG and OA for 48 h and labeled with ARP to detect AP sites. Right, Dot blot densitometry of AP site levels in HPDE cells (white bar) and HPDE cells exposed to hG and OA for 48 h (gray bar); C Left, Representative image of a PLA experiment performed on HPDE cells ± hG and OA for 48 h. The IF staining shows TDG level (purple signal), and PLA foci (red signal). DAPI (blue signal) used to normalize the signal. Scale bar: 10 µm. Right, Quantification of PLA foci in HPDE ± hG and OA for 48 h. D Left, Dot blot analysis for AP sites detection performed in dysmetabolic HPDE cells after TDG CRISPR/Cas9 inactivation (LCv2_TDG) compared to control vector (LCv2_NTC); Right, Dot blot densitometry of AP site levels in dysmetabolic HPDE cells + LCv2_NTC (white bar) and dysmetabolic HPDE cells + LCv2_TDG (gray bar); E Left, Representative WB of TDG levels in HPDE cells after CRISPR/Cas9 inactivation (LCv2_TDG compared to control vector LCv2_NTC) and reconstitution by myc-tagged wild-type (myc-TDGWT) or mutant (myc-TDGR275A) TDG; Loading control: Vinculin; Right, Quantification of 3 independent experiments; F Left, Dot blot analysis for AP sites detection performed in HPDE cells ± hG and OA for 48 h after TDG CRISPR/Cas9 inactivation and reconstitution by myc-TDGwt or myc-TDGR275A; Right, Dot blot densitometry of 3 independent experiments; G Left, Representative WB analysis of LIG1 and LIG3, in HPDE cells ± hG and OA for 72 h. Loading control: Tubulin. Right, Densitometry of LIG1 and LIG3 levels in HPDE cells (white bars) and HPDE cells exposed to hG and OA for 72 h (gray bars); H Representative WB analysis of LIG1 and LIG3 in HPDE cells ± SA for 72 h. Loading control: Tubulin. Right, Densitometry of LIG1 and LIG3 levels in HPDE cells (white bars) and HPDE cells exposed to SA for 72 h (black bars); I Left Dot blot analysis performed on genomic DNA extracted from HPDE cells treated with 5 μM, 15 μM, 30 μM L189 for 24 h and labeled with ARP to detect AP sites; Right, Dot blot densitometry of 4 independent experiments; J LIG1 and LIG3 promoters 5mC abundance in dysmetabolic HPDE cells ± αKG (time of treatments: 72 h); K LIG1 and LIG3 promoters 5mC abundance analysis in HPDE cells ± SA for 72 h; L Left, Representative WB analysis of LIG1 and LIG3 in dysmetabolic HPDE cells ± αKG (time of treatments: 72 h). Loading control: Tubulin. Right, Densitometry of 4 independent experiments. M Dot blot analysis performed on genomic DNA extracted from HPDE cells exposed to hG + OA in the presence/absence of αKG for 24 h and labeled with ARP to detect AP sites; Right, Dot blot densitometry of 3 independent experiments. Data expressed as average ± SEM. Statistical significance was calculated using Kolmogorov–Smirnov test (A, B, D, F, G, H), Mann-Whitney test (C), one-way ANOVA (I) and two-way ANOVA (E, J, K, L, M) and is shown as *p < 0.05; **p < 0.01; ***p < 0.001.
To further investigate BER enzymatic machinery integrity under these experimental conditions, the expression of crucial BER enzymes, including APEX1, PARP1, FEN1, XRCC1, LIG1, and LIG3, was assessed by qRT-PCR. Interestingly, we found a significant downregulation of LIG1 and LIG3 at both mRNA (Fig. S4C) and protein levels (Fig. S4D) in HFD-fed KC mice and in dysmetabolic HPDE cells (Fig. 5G). Similar results were obtained in HPDE cells exposed to a cell-permeable source of SA (Fig. 5H), suggesting ligase enzymatic activity impairment in response to metabolic derangement. Notably, LIG1 and LIG3-dependent enzymatic reactions are crucial to accomplish BER pathway and restore DNA integrity [18]. To verify whether dysmetabolism-dependent ligase dysfunction was responsible for TDG-dependent AP site accumulation, L189, a DNA ligase competitive inhibitor, was exploited [36]. A dose-response curve established that L189 treatment was non-toxic for HPDE cells, since proliferation was not affected (Fig. S4E). Moreover, L189 induced a dose-dependent AP site accumulation in HPDE cells (Fig. 5I), suggesting ligase impairment involvement in the DNA lesion propagation of pre-neoplastic epithelial cells. Dysmetabolism-dependent LIG1 and LIG3 level decrease led to a decline of their enzymatic activity supporting DNA damage propagation in proliferating cells. Since our results indicated a link between dysmetabolism and DNA methylation (Fig. 2B, Fig. 3E, F), we wondered whether low expression levels of LIG1 and LIG3 could depend on alteration of their promoter methylation as a consequence of SAM and SA increased levels and αKG decreased levels (Fig. S2B, S2D; Fig. 1E-G). Specifically, quantification of 5mC levels revealed hypermethylation of both LIG1 and LIG3 promoters upon hG and OA exposure or SA supplementation (Fig. 5J, K). Notably, αKG addition, counteracting TET1/TDG complex disassembly [21], was able to reduce the percentage of 5mC in the promoter region of LIG1 and LIG3 (Fig. 5J), ultimately rescuing their expression (Fig. 5L) and reducing AP site levels (Fig. 5M). These data highlight a novel epi-metabolic mechanism harnessing DNA integrity in pre-tumoral pancreatic epithelial cells under dysmetabolic conditions and underscore the sensitivity of ligase enzymes to metabolic derangements as contributor of genome instability and tumorigenesis.
Discussion
Metabolic reprogramming is a core hallmark of cancer, crucial for tumor cell survival and proliferation and supporting biosynthesis and redox balance. This intense metabolic activity, by altering the availability of specific metabolites, might interfere with epigenetic enzyme function, since metabolites usually represent co-factors or substrates essential for epigenetic reactions [11], and with innate immune system function, which can shift from cancer cell surveillance to supporting tumor progression and immune evasion [37]. Several clinical and epidemiological studies identified a specific metabolic dysfunction becoming established prior pancreatic cancer diagnosis [38,39,40]. The dysfunction is the result of a complex cancer-host metabolism interplay affecting glucose and lipid metabolism in the early stages of pancreatic cancer [38,39,40]. Nevertheless, how this intricate interaction translates in specific molecular and/or epigenetic alteration was not described in detail. Metabolic deregulation can promote oncometabolite accumulation, contributing to tumorigenesis by epigenetic landscape alteration. Among oncometabolites, fumarate and SA, usually derived from tumor-associated enzymatic alterations, compete with αKG inhibiting αKG-dependent dioxygenases, including TETs and histone demethylases. This effect supports hypermethylation of DNA and histones, typical of carcinogenesis. Although the effect of hyperglycemia and high caloric intake diets on pancreatic cancer incidence is well known [3,4,5,6,7], how the associated metabolic derangement shapes cancer metabolic reprogramming, fostering pancreatic cancer development, has not been specifically investigated. The present manuscript revealed an alteration of specific metabolite levels prompting to hypothesize a deregulation of the DNA methylation pathway. The observed FFA deregulation was recently defined as an early sign of PDAC onset in type 2 diabetes patients [41]. The involvement of DNA methylation cycle alteration was further supported by αKG and SA level unbalance (Fig. 1). A similar metabolic scenario was reported in p53-deficient PDAC cells. αKG, acting as an effector of p53-mediated tumor suppression, was proposed as an antagonist of malignant progression in a therapeutic perspective [42]. Dysmetabolism, leading to αKG/SA ratio alteration, boosts cell transformation similar to what is observed upon inactivating p53 mutations [42]. Accordingly, a deregulation of αKG metabolism associated to an accumulation of SA during cancer development has been reported in different studies [43,44,45]. In particular, a SA involvement in tumor growth, cell migration, demethylation cycle deregulation, and cancer immune evasion has been described in renal cell carcinoma and in gastric and colon cancer [23, 45, 46], but not clearly addressed in pancreatic cancer. The observed metabolic landscape prompted us to focus on DNA methylation as an essential node of pancreatic tumorigenesis. Consistently, hypermethylation in the promoter regions of DNA repair, tumor suppressor and antitumor response genes has been described in different tumors, paralleled by hypomethylation of genes sustaining cell proliferation, migration, and invasion [47,48,49,50]. We addressed DNA methylation alterations by focusing on the effect of dysmetabolism on the DNA demethylating complex formed by TET1/TDG association, particularly sensitive to metabolic alterations [21]. We demonstrated that TET1/TDG complex is particularly sensitive to metabolic derangement also in pancreas, leading to an accumulation of iterative cytosine modifications (Fig. 2). This observation can partially be explained by the already described TET protein delocalization outside of the nucleus, typical of different tumors [51,52,53], a specific strategy of cancer to alter the DNA methylation landscape. The present manuscript reveals a specific metabolic-epigenetic crosstalk contributing to pancreatic cancer development. Metabolic alteration on one hand drives TDG hyperactivation, which favors AP site accumulation because of increased 5fC levels, promoting genome instability in pre-neoplastic KRAS-mutated epithelial cells. On the other hand, it supports a methylation-prone DNA environment, leading to transcriptional repression of LIG1 and LIG3, which further contributes to genomic instability by limiting BER pathway accomplishment. Remarkably, the above-mentioned metabolic-epigenetic crosstalk occurred in cells with a high proliferative rate induced by a dysmetabolic condition (Fig. S4F). Furthermore, TDG hyperactivation might be detrimental to cell function, sustaining transformation. Indeed, the role of TDG in cancer is controversial: it might both counteract and sustain cancer development and progression depending on the specific pathways in which it is involved [22, 54,55,56]. Here, we analyzed TDG contribution to AP site accumulation, the most common DNA lesion generated during BER pathway activation [16,17,18]. Increased levels of AP sites have been reported in a variety of tumors and associated with DNA damage accumulation [57, 58]. Thus, persistent DNA damage might reflect an impairment of the DNA repair machinery. Compromised BER or aberrant BER protein expression represent two known risk factors of cancer development, and alteration of the associated enzymatic machinery contributes to tumorigenesis supporting genome instability and anti-tumoral response [18, 59]. LIG1 and LIG3 are two ligases acting in the final steps of DNA repair pathway, sealing generated DNA gap, with LIG1 involved in both single-nucleotide BER and long-patch BER and LIG3 only in single-nucleotide BER [18]. Their sensitivity to metabolic derangement has not been investigated until now. Our results show high sensitivity of BER machinery to dysmetabolic conditions, promoting both DNA demethylation cycle impairment due to αKG/SA ratio unbalance. Since LIG1 and LIG3 gene promoters were heavily methylated upon dysmetabolism (Fig. 5J), their activity is downregulated leading to unrepaired DNA damage.
Our results demonstrate that metabolic pressure is associated with an alteration of the DNA damage repair program. Specifically, original results show that dysmetabolism-dependent increased levels of 5mC in the promoter regions of both ligase genes might favor genome instability due to AP site accumulation (Fig. 5). In this light, the metabolic pressure paves the way to 5fC and AP sites as novel pancreatic cancer biomarkers to be exploited for diagnostic and prognostic purposes, as they result from DNA methylation dependent repression of ligases.
Materials and methods
Animals and treatments
FVB PDX-1-Cre; LSL-KrasG12D (KC) mice were generated in the Animal Facility of the Regina Elena National Cancer Institute (Rome, Italy) as previously described [60]. Experiments were performed according with EU Directive 2010/63/EU for animal experiments (protocol permit number: 362/2021-PR). In brief, 4 weeks old male KC mice were randomly divided in 2 groups and fed with a low fat rodent chow (LFD- 10 kJ% fat, 20 kJ% protein, 70 kJ% carbohydrates—D12450J Ssniff—Germany; n = 40) or a high-fat diet (HFD—60 kJ% fat, 20 kJ% protein, 20 kJ% carbohydrates – D12492 Ssniff-Germany; n = 40) to recapitulate metabolic alterations. All groups received drink and food ad libitum. The present study employed male mice due to their higher and faster response to HFD-induced metabolic alterations than female animals [61]. Male mice allowed to observe 3R’s principles for animal studies, since hyperglycemia threshold of 200 mg/dl fixed for the experimental plan was reached faster and more consistently using a less number of animals compared to female mice, usually resistant to HFD-induced metabolic alterations leading to an increased time to achieve hyperglycemia condition. Sample size was determined to reach an 80% power (α error of 0.05), because usually effect below or equal 20% is barely relevant. Pancreata were collected after 1.5 month (T1) and 5 months (T2) of dietary regimen start. Details about mouse monitoring blood analysis and histology provided in Supplementary material.
Untargeted and targeted metabolomics
Pancreatic tissues and cell pellets were lyophilized overnight, and carefully homogenized. Untargeted UHPLC-HRMS/MS analysis was performed on a Thermo Ultimate RS 3000 coupled online to a Q-Exactive hybrid quadrupole Orbitrap mass spectrometer (Thermo Fisher Scientific, Bremen, Germany). Targeted UHPLC-HRMS analyses were performed on a Thermo Vanquish Flex UHPLC system coupled online to a hybrid quadrupole Orbitrap Exploris 120 mass spectrometers (Thermo Fisher Scientific, Bremen, Germany). Details provided in Supplementary material.
Total RNA extraction, sequencing, and bioinformatics analysis
RNA was isolated from mouse pancreata (4 LFD and 4 HFD for each experimental time point) using miRNeasy Mini Kit (Qiagen) combined with DNase digestion (DNase I, Qiagen) to eliminate genomic DNA. Sequencing was performed on rRNA-depleted total RNA on an Illumina HiSeq 2000 platform. Details provided in Supplementary material.
qRT-PCR
RNA was isolated from about 10 mg of mouse pancreatic tissue using miRNeasy Mini Kit (Qiagen) according to manufacturer’s instruction. Details provided in Supplementary material. List of primers provided in Supplementary Table 3.
Cell cultures, treatment, and transfection
HPDE K-RasG12V cells [62], available in the lab [63], were grown in RPMI 1640 medium (Euroclone) containing 10% fetal bovine serum (FBS), 1% L-glutamine and 1% Penicillin/Streptomycin (Euroclone). Human PSCs, purchased by Neo Biotech (Cat# NB-26-01967), were cultured in Stellate Cell Medium (SteCM, ScienCell Research Laboratories) supplemented with 2% FBS (ScienCell Research Laboratories), 1% Stellate Cell Growth Supplement (SteCGS, ScienCell Research Laboratories) and 1% Penicillin/Streptomycin solution (P/S, ScienCell Research Laboratories) at 37 °C in a 5% CO2 environment. All cell lines were routinely tested for mycoplasma contamination. Crystal violet assay was performed in HPDE cells fixed with paraformaldehyde 4% for 20 min at room temperature, washed with PBS and stained with crystal violet solution (0.5% w/v in acetic acid). Stained colonies were solubilized in dimethyl sulfoxide (DMSO) and plate was read at 595 nm at Clariostar microplate reader (BMG, Labtech). HEK293T, purchased by ATCC (Cat# CRL-1573), and HPDE cells were transfected according to Lipofectamine3000 standard procedure (Invitrogen). TDG gain and loss of function experiments were performed as previously described [21]. Details provided in Supplementary material.
Immunoprecipitation and Western blot analysis
Details provided in Supplementary material. Antibody list provided in Supplementary Table 4.
Dot blot
Genomic DNA, from pancreatic mouse tissue or HPDE cells exposed or not to hG (30 mM, Sigma) and OA (50 μM, Sigma) for 48 h or to SA (50 μM Sigma) for 24 h, was extracted with Monarch Genomic DNA purification kit (NEB) according to manufacturer´s instructions. Details provided in Supplementary material. Antibody list provided in Supplementary Table 4.
Immunofluorescence and IHC
HPDE cells were fixed in 4% paraformaldehyde solution for 10 min at room temperature, washed three times for 5 min with PBS, and blocked for 1 h in PBS containing 10% BSA. Cells were incubated overnight at 4 °C with the indicated primary antibodies. After washing, cells were incubated for 1 h at room temperature with a fluorescent secondary antibody (Alexa Fluor 488- or 546-conjugated, Life Technologies) and DAPI (Life Technologies). The immunofluorescence analysis on mouse tissues was performed according to the following protocol. First, 4-μm-thick mouse pancreas FFPE sections were deparaffinized and rehydrated. Antigen-retrieval was performed in the microwave in citrate buffer solution pH 6.0 (ScyTech) for 15 min. After blocking, sections were incubated overnight at 4 °C with following primary antibodies: anti-Ecad (Invitrogen); anti-αSMA (Invitrogen); anti-5-formylcytosine (Active Motif). After washing, cells were incubated for 1 h at room temperature with a fluorescent secondary antibody (Alexa Fluor 488- or 546-conjugated, Life Technologies) and nuclei were counterstained with DRAQ5 (BioStatus). Immunofluorescence was analyzed by Zeiss LSM900 confocal microscope. 5fC levels in E-cad positive cells and in αSMA positive cells were quantified by measuring the Integrated Density (ID) using Fiji ImageJ software on pictures acquired with same settings and % 5fC positive cells was calculated. The Pearson correlation coefficient (PCC) of TET1 and TDG signal in each cell was calculated with a range +1 (perfect correlation) to −1 (perfect exclusion) [64]. Antibody list provided in Supplementary Table 4.
MD simulation
The computational approach follows the procedure already used previously [21, 65]. Briefly, the X-ray crystallographic structure of TDG in complex with DNA (PDB-ID: Z47) was used as receptor in MD simulations, upon removal of the DNA chains [33]. The protein was parametrized with the ff14SB force field, while SA ion with the General Amber Force Field (GAFF) [66, 67]. SA partial charges were computed at the am1-bcc level of theory. TDG and SA were included in a rectilinear box of TIP3P-type water molecules buffering 10 Å from the molecular system, at a non-binding distance >40.0 Å. The total charge was neutralized by the addition of Cl- ions. A 2 fs time-step was used in all MD simulations. The solvent was first energy minimized for 500 steps using the Steepest Descent algorithm (SD) and a further 2500 steps using the Conjugate Gradient algorithm (CG), while keeping the solute as fixed. The solvated solute was then energy minimized for 1000 steps SD and subsequent 9000 steps CG before heating to 300 K at constant volume for 1 ns using the Langevin thermostat. Box density was equilibrated at constant pressure for 1 ns using the Berendsen barostat, then, a preliminary run of 50 ns was carried out at constant pressure, before the final production of trajectories lasting 500 ns. No restraints were used in MD simulations. Three independent MD replicas were run starting from different initial coordinates. MD simulations were run with Ambe18 [68], analysis of MD trajectories was carried out with cpptraj [69].
In vitro surface plasmon resonance experiments
SPR experiments were carried out using a Sartorius Octet SF3 apparatus, to assess the thermodynamic parameters of the interaction between mouse His-tagged TDG (ligand) and the analytes αKG and SA, exploiting an already used approach [70]. Steady-state plateau signal (Req) values and full fittings with 1 and 2 sites were calculated from overall kinetic evaluation of the sensorgrams using the Octet SPR Analysis software. Details provided in Supplementary material.
Cellular Thermal Shift Assay (CETSA)
CETSA was performed as previously described [21]. Briefly, cellular extract was freshly lysed in RIPA buffer (10 mM Tris HCl pH 7.4, 150 mM NaCl, 1% Igepal, 1% sodium deoxycholate (DOC), 0.1% Sodium Dodecyl Sulfate (SDS), 0,1% glycerol, protease and phosphatase inhibitors cocktail, and 2 mM 1,4-dithiothreitol (DTT)). Homogenization was performed by 3 cycles of freeze-thawing and cellular extracts were centrifuged at 20,000 × g for 20 min at 4 °C to separate the soluble fraction from debris. Cellular lysates were divided into two aliquots, with one aliquot being treated with 1 mM SA (Sigma) and the other one exposed to water as solvent (control), and were incubated for 30 min at RT. Subsequently, the lysates were divided into smaller aliquots and heated at the reported temperature for 3 min followed by 3 min cooling at RT. The soluble fraction was separated from debris centrifuging at 20,000 × g for 20 min at 4 °C. All supernatants were analyzed by western blotting and probed with the specific primary antibody.
Enzymatic activity assay
TDG activity was assessed with specific ELISA activity assays following the manufacturer´s instructions (TDG Activity/Inhibition Assay Kit—Epigentek) on HPDE cells exposed to hG and OA for 48 h or using the active full length mrTDG protein (Epigex). αKG disodium salt (Sigma) and SA (Sigma) was used for recombinant TDG protein treatments during assay.
AP site quantification
Genomic DNA, from HPDE cells exposed or not to hG (30 mM, Sigma) and OA (50 μM, Sigma) for 48 h or to SA (50 μM Sigma) for 24 h, was extracted with Monarch Genomic DNA purification kit (NEB) according to manufacturer’s instructions. 500 ng of DNA were incubated with 10 mM aldehyde reactive probe (ARP, Cayman) at 37° for 1 h. After precipitation and quantification, 100 ng of ARP labeled DNA were denatured with 0.4 N NaOH and incubated for 30 min at 80°C. Then, DNA was immobilized on nitrocellulose membrane and probed with Alexa Fluor 790 streptavidin conjugate (Life Technologies) for 1 h at room temperature. Development was performed by ChemiDoc MP Imaging System (Bio-Rad).
Proximity ligation assay
PLA between ARP-biotin and TDG was performed exploiting an approach already used [70]. Briefly HPDE cells were grown on slides, exposed to hG (30 mM, Sigma) and OA (50 μM, Sigma) for 48 h and incubated with 1 mM ARP (Cayman) for the last 3 h of treatment before fixation in 4% paraformaldehyde solution for 10 min at room temperature. Cells were then permeabilised with 0.5% Triton-X 100 for 5 min before blocking in 3% BSA in PBST for 1 h. Slides were incubated with primary antibodies (mouse α-biotin and rabbit α-TDG) for 2 h before processing the samples for PLA following the manufacturer's instructions of the SIGMA Duolink Kit (Sigma). After PLA reactions, cells were incubated with an Alexafluor 647 conjugated anti rabbit antibody for 30 min at room temperature to visualize TDG. Details provided in Supplementary material.
LIG1 and LIG3 promoters 5mC quantification
The level of 5mC in genomic DNA extracted from HPDE cells exposed to SA for 72 h or to hG (30 mM, Sigma) and OA (50 μM, Sigma) ± αKG for 72 h was estimated with the EpiJET 5-mC Analysis Kit (ThermoFisher Scientific). Details provided in Supplementary material. List of primers provided in Supplementary Table 3.
Statistical analyses
Statistical analyses were performed using GraphPad Prism software version 8. Data are shown as mean ± SEM. Statistical significance was determined using 2-way ANOVA or non parametric student’s t test as reported in the corresponding figure legend. A p-value < 0.05 was considered statistically significant and significant differences between experimental samples are represented by asterisks (* = p < 0.05; ** = p < 0.01; *** = p < 0.001). Sample size (N) for each experiment is stated in the related figure legend.
Data availability
The RNA sequencing datasets are publicly available at NCBI’s Gene Expression Omnibus (GEO) repository, under accession number GSE302730 located at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE302730. Any additional information required to reanalyze the data reported in this work paper is available from the lead contact upon request, Dr. Francesco Spallotta (francesco.spallotta@uniroma1.it) and Dr. Chiara Cencioni (chiara.cencioni@cnr.it). This study did not generate new unique reagents.
References
Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA: A Cancer J Clin. 2023;73(1):17–48.
Wood LD, Canto MI, Jaffee EM, Simeone DM. Pancreatic cancer: pathogenesis, screening, diagnosis, and treatment. Gastroenterology. 2022;163(2):386–402.e1.
Carreras-Torres R, Johansson M, Gaborieau V, Haycock PC, Wade KH, Relton CL, et al. The role of obesity, type 2 diabetes, and metabolic factors in pancreatic cancer: a Mendelian randomization study. J Nl Cancer Inst. 2017;109(9).
Jin X, Qiu T, Li L, Yu R, Chen X, Li C, et al. Pathophysiology of obesity and its associated diseases. Acta Pharm Sin B. 2023;13(6):2403–24.
Khandekar MJ, Cohen P, Spiegelman BM. Molecular mechanisms of cancer development in obesity. Nat Rev Cancer. 2011;11(12):886–95.
Klein AP. Pancreatic cancer epidemiology: understanding the role of lifestyle and inherited risk factors. Nat Rev Gastroenterol Hepatol. 2021;18(7):493–502.
Maitra A, Sharma A, Brand RE, Van Den Eeden SK, Fisher WE, Hart PA, et al. A Prospective Study to Establish a New-Onset Diabetes Cohort: From the Consortium for the Study of Chronic Pancreatitis, Diabetes, and Pancreatic Cancer. Pancreas. 2018;47(10):1244–8.
Araki T, Nagashima M, Hirasawa H, Tamalu F, Katagiri Y, Miwa N. Epigenome-wide association analysis of pancreatic exocrine cells from high-fat- and normal diet-fed mice and its potential use for understanding the oncogenesis of human pancreatic cancer. Biochem Biophys Res Commun. 2022;637:50–7.
Cascetta P, Cavaliere A, Piro G, Torroni L, Santoro R, Tortora G, et al. Pancreatic cancer and obesity: molecular mechanisms of cell transformation and chemoresistance. Int J Mol Sci. 2018;19(11).
Wlodarczyk M, Nowicka G. Obesity, DNA damage, and development of obesity-related diseases. Int J Mol Sci. 2019;20(5).
Pavlova NN, Zhu J, Thompson CB. The hallmarks of cancer metabolism: still emerging. Cell Metab. 2022;34(3):355–77.
Kohli RM, Zhang Y. TET enzymes, TDG and the dynamics of DNA demethylation. Nature. 2013;502(7472):472–9.
Jones PA, Baylin SB. The fundamental role of epigenetic events in cancer. Nat Rev Genet. 2002;3(6):415–28.
Patil V, Ward RL, Hesson LB. The evidence for functional non-CpG methylation in mammalian cells. Epigenetics. 2014;9(6):823–8.
Tahiliani M, Koh KP, Shen Y, Pastor WA, Bandukwala H, Brudno Y, et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science. 2009;324(5929):930–5.
Vasovcak P, Krepelova A, Menigatti M, Puchmajerova A, Skapa P, Augustinakova A, et al. Unique mutational profile associated with a loss of TDG expression in the rectal cancer of a patient with a constitutional PMS2 deficiency. DNA Repair. 2012;11(7):616–23.
Wiebauer K, Jiricny J. Mismatch-specific thymine DNA glycosylase and DNA polymerase beta mediate the correction of G.T mispairs in nuclear extracts from human cells. Proc Natl Acad Sci USA. 1990;87(15):5842–5.
Gohil D, Sarker AH, Roy R. Base excision repair: mechanisms and impact in biology, disease, and medicine. Int J Mol Sci. 2023;24(18).
Sun L, Zhang H, Gao P. Metabolic reprogramming and epigenetic modifications on the path to cancer. Protein cell. 2022;13(12):877–919.
Wang P, Chen LL, Xiong Y, Ye D. Metabolite regulation of epigenetics in cancer. Cell Rep. 2024;43(10):114815.
Spallotta F, Cencioni C, Atlante S, Garella D, Cocco M, Mori M, et al. Stable oxidative cytosine modifications accumulate in cardiac mesenchymal cells from type2 diabetes patients: rescue by alpha-ketoglutarate and TET-TDG functional reactivation. Circ Res. 2018;122(1):31–46.
Mancuso P, Tricarico R, Bhattacharjee V, Cosentino L, Kadariya Y, Jelinek J, et al. Correction to: Thymine DNA glycosylase as a novel target for melanoma. Oncogene. 2022;41(23):3300–1.
Wang G, Rao P. Succinate dehydrogenase-deficient renal cell carcinoma: a short review. Arch Pathol Lab Med. 2018;142(10):1284–8.
Xu H, Long S, Xu C, Li Z, Chen J, Yang B, et al. TNC upregulation promotes glioma tumourigenesis through TDG-mediated active DNA demethylation. Cell Death Discov. 2024;10(1):347.
Bracci PM. Obesity and pancreatic cancer: overview of epidemiologic evidence and biologic mechanisms. Mol Carcinog. 2012;51(1):53–63.
Ariston Gabriel AN, Jiao Q, Yvette U, Yang X, Al-Ameri SA, Du L, et al. Differences between KC and KPC pancreatic ductal adenocarcinoma mice models, in terms of their modeling biology and their clinical relevance. Pancreatology. 2020;20(1):79–88.
Geeraerts SL, Heylen E, De Keersmaecker K, Kampen KR. The ins and outs of serine and glycine metabolism in cancer. Nat Metab. 2021;3(2):131–41.
Zeng JD, Wu WKK, Wang HY, Li XX. Serine and one-carbon metabolism, a bridge that links mTOR signaling and DNA methylation in cancer. Pharmacol Res. 2019;149:104352.
Lu X, Han D, Zhao BS, Song CX, Zhang LS, Dore LC, et al. Base-resolution maps of 5-formylcytosine and 5-carboxylcytosine reveal genome-wide DNA demethylation dynamics. Cell Res. 2015;25(3):386–9.
Raiber EA, Murat P, Chirgadze DY, Beraldi D, Luisi BF, Balasubramanian S. 5-Formylcytosine alters the structure of the DNA double helix. Nat Struct Mol Biol. 2015;22(1):44–9.
Apte MV, Wilson JS, Lugea A, Pandol SJ. A starring role for stellate cells in the pancreatic cancer microenvironment. Gastroenterology. 2013;144(6):1210–9.
Letouze E, Martinelli C, Loriot C, Burnichon N, Abermil N, Ottolenghi C, et al. SDH mutations establish a hypermethylator phenotype in paraganglioma. Cancer Cell. 2013;23(6):739–52.
Malik SS, Coey CT, Varney KM, Pozharski E, Drohat AC. Thymine DNA glycosylase exhibits negligible affinity for nucleobases that it removes from DNA. Nucleic Acids Res. 2015;43(19):9541–52.
Chen D, Lucey MJ, Phoenix F, Lopez-Garcia J, Hart SM, Losson R, et al. T:G mismatch-specific thymine-DNA glycosylase potentiates transcription of estrogen-regulated genes through direct interaction with estrogen receptor alpha. J Biol Chem. 2003;278(40):38586–92.
Wilson DM 3rd, Barsky D. The major human abasic endonuclease: formation, consequences and repair of abasic lesions in DNA. Mutat Res. 2001;485(4):283–307.
Tomkinson AE, Howes TR, Wiest NE. DNA ligases as therapeutic targets. Transl Cancer Res. 2013;2(3).
Pandey S, Anang V, Schumacher MM. Mitochondria driven innate immune signaling and inflammation in cancer growth, immune evasion, and therapeutic resistance. Int Rev Cell Mol Biol. 2024;386:223–47.
Chari ST, Wu B, Lopez C, Lustigova E, Chen Q, Van Den Eeden SK, et al. Risk of pancreatic cancer in glycemically defined new-onset diabetes: a prospective cohort study. Gastroenterology. 2025.
Sah RP, Sharma A, Nagpal S, Patlolla SH, Sharma A, Kandlakunta H, et al. Phases of metabolic and soft tissue changes in months preceding a diagnosis of pancreatic ductal adenocarcinoma. Gastroenterology. 2019;156(6):1742–52.
Sharma A, Smyrk TC, Levy MJ, Topazian MA, Chari ST. Fasting blood glucose levels provide estimate of duration and progression of pancreatic cancer before diagnosis. Gastroenterology. 2018;155(2):490–500.e2.
Gumpper-Fedus K, Hart PA, Belury MA, Crowe O, Cole RM, Pita Grisanti V, et al. Altered plasma fatty acid abundance is associated with cachexia in treatment-naive pancreatic cancer. Cells. 2022;11(5).
Morris, JPt, Yashinskie JJ, Koche R, Chandwani R, Tian S, Chen CC, et al. alpha-Ketoglutarate links p53 to cell fate during tumour suppression. Nature. 2019;573(7775):595–9.
Atallah R, Olschewski A, Heinemann A. Succinate at the crossroad of metabolism and angiogenesis: roles of SDH, HIF1alpha and SUCNR1. Biomedicines. 2022;10(12).
Laukka T, Mariani CJ, Ihantola T, Cao JZ, Hokkanen J, Kaelin WG Jr, et al. Fumarate and succinate regulate expression of hypoxia-inducible genes via TET enzymes. J Biol Chem. 2016;291(8):4256–65.
Selak MA, Armour SM, MacKenzie ED, Boulahbel H, Watson DG, Mansfield KD, et al. Succinate links TCA cycle dysfunction to oncogenesis by inhibiting HIF-alpha prolyl hydroxylase. Cancer Cell. 2005;7(1):77–85.
Zhang W, Lang R. Succinate metabolism: a promising therapeutic target for inflammation, ischemia/reperfusion injury and cancer. Front Cell Dev Biol. 2023;11:1266973.
Das PM, Singal R. DNA methylation and cancer. J Clin Oncol. 2004;22(22):4632–42.
Tan AC, Jimeno A, Lin SH, Wheelhouse J, Chan F, Solomon A, et al. Characterizing DNA methylation patterns in pancreatic cancer genome. Mol Oncol. 2009;3(5-6):425–38.
Ueki T, Toyota M, Sohn T, Yeo CJ, Issa JP, Hruban RH, et al. Hypermethylation of multiple genes in pancreatic adenocarcinoma. Cancer Res. 2000;60(7):1835–9.
Luo L, Fu S, Du W, He LN, Zhang X, Wang Y, et al. LRRC3B and its promoter hypomethylation status predicts response to anti-PD-1 based immunotherapy. Front Immunol. 2023;14:959868.
Atlante S, Visintin A, Marini E, Savoia M, Dianzani C, Giorgis M, et al. alpha-ketoglutarate dehydrogenase inhibition counteracts breast cancer-associated lung metastasis. Cell Death Dis. 2018;9(7):756.
Huang Y, Wang G, Liang Z, Yang Y, Cui L, Liu CY. Loss of nuclear localization of TET2 in colorectal cancer. Clin Epigenet. 2016;8:9.
Muller T, Gessi M, Waha A, Isselstein LJ, Luxen D, Freihoff D, et al. Nuclear exclusion of TET1 is associated with loss of 5-hydroxymethylcytosine in IDH1 wild-type gliomas. Am J Pathol. 2012;181(2):675–83.
Xu X, Yu T, Shi J, Chen X, Zhang W, Lin T, et al. Thymine DNA glycosylase is a positive regulator of Wnt signaling in colorectal cancer. J Biol Chem. 2014;289(13):8881–90.
Xu X, Watt DS, Liu C. Multifaceted roles for thymine DNA glycosylase in embryonic development and human carcinogenesis. Acta Biochim Biophys Sin. 2016;48(1):82–9.
Zhou W, Zhang L, Chen P, Li S, Cheng Y. Thymine DNA glycosylase-regulated TAZ promotes radioresistance by targeting nonhomologous end joining and tumor progression in esophageal cancer. Cancer Sci. 2020;111(10):3613–25.
Nowsheen S, Wukovich RL, Aziz K, Kalogerinis PT, Richardson CC, Panayiotidis MI, et al. Accumulation of oxidatively induced clustered DNA lesions in human tumor tissues. Mutat Res. 2009;674(1-2):131–6.
Wei S, Perera MLW, Sakhtemani R, Bhagwat AS. A novel class of chemicals that react with abasic sites in DNA and specifically kill B cell cancers. PloS one. 2017;12(9):e0185010.
Tong J, Song J, Zhang W, Zhai J, Guan Q, Wang H, et al. When DNA-damage responses meet innate and adaptive immunity. Cell Mol Life Sci. 2024;81(1):185.
de Latouliere L, Manni I, Iacobini C, Pugliese G, Grazi GL, Perri P, et al. A bioluminescent mouse model of proliferation to highlight early stages of pancreatic cancer: a suitable tool for preclinical studies. Ann Anat = Anatomischer Anz. 2016;207:2–8.
Pettersson US, Walden TB, Carlsson PO, Jansson L, Phillipson M. Female mice are protected against high-fat diet induced metabolic syndrome and increase the regulatory T cell population in adipose tissue. PloS One. 2012;7(9):e46057.
Siddiqui I, Erreni M, Kamal MA, Porta C, Marchesi F, Pesce S, et al. Differential role of Interleukin-1 and Interleukin-6 in K-Ras-driven pancreatic carcinoma undergoing mesenchymal transition. Oncoimmunology. 2018;7(2):e1388485.
Middonti E, Astanina E, Vallariello E, Hoza RM, Metovic J, Spadi R, et al. A neuroligin-2-YAP axis regulates progression of pancreatic intraepithelial neoplasia. EMBO Rep. 2024;25(4):1886–908.
Adler J, Parmryd I. Quantifying colocalization by correlation: the Pearson correlation coefficient is superior to the Mander’s overlap coefficient. Cytom Part A. 2010;77(8):733–42.
Humbert N, Kovalenko L, Saladini F, Giannini A, Pires M, Botzanowski T, et al. Thia)calixarenephosphonic acids as potent inhibitors of the nucleic acid chaperone activity of the HIV-1 nucleocapsid protein with a new binding mode and multitarget antiviral activity. ACS Infect Dis. 2020;6(4):687–702.
Tian C, Kasavajhala K, Belfon KAA, Raguette L, Huang H, Migues AN, et al. ff19SB: amino-acid-specific protein backbone parameters trained against quantum mechanics energy surfaces in solution. J Chem Theory Comput. 2020;16(1):528–52.
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA. Development and testing of a general amber force field. J Comput Chem. 2004;25(9):1157–74.
Rubenstein AB, Blacklock K, Nguyen H, Case DA, Khare SD. Systematic comparison of amber and rosetta energy functions for protein structure evaluation. J Chem Theory Comput. 2018;14(11):6015–25.
Roe DR, Cheatham TE 3rd. PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J Chem Theory Comput. 2013;9(7):3084–95.
Cipolla L, Bertoletti F, Maffia A, Liang CC, Lehmann AR, Cohn MA, et al. UBR5 interacts with the replication fork and protects DNA replication from DNA polymerase eta toxicity. Nucleic Acids Res. 2019;47(21):11268–83.
Funding
This research was funded by the AIRC, Associazione Italiana per la Ricerca sul Cancro (AIRC), My First AIRC “Giorgio e Adriana Squinzi” MFAG number 23099 to Francesco Spallotta, MFAG number 28858 to Livia Perfetto, Start Up number 30656 to Gian Luca Rampioni Vinciguerra and IG number 22910 to Federico Bussolino; Sapienza University of Rome, “Progetto Ateneo 2023” to Francesco Spallotta; funded by European Union-Next Generation EU, Missione 4 C2 Investimento 1.1 PRIN-PNRR number P2022R7WRC; CUP B53D23025120001 to Chiara Cencioni and Eduardo Maria Sommella; PRIN-PNRR number P2022E3BTH; CUP B53D23024970001 to Francesco Spallotta; funded by PNRR M4C2—Dalla ricerca all’impresa—3.1: Fondo per la realizzazione di un sistema integrato di infrastrutture di ricerca e innovazione “Potentiating the Italian Capacity for Structural Biology Services in Instruct-ERIC (ITACA.SB)” CUP: B53C22001790006; PNRR PE8 Age-IT., cofounding from Next Generation EU [DM 1557 11.10.2022], in the context of the National Recovery and Resilience Plan, Investment PE8—Project Age-It: “Ageing Well in an Ageing Society”; Project PRIN MIUR 2022HYF8KS to Gianni Colotti; MUR (PNRR D3 4 Health) and FPRC 5xmille Ministero Salute 2022 – CARESS and Ricerca Corrente 2025 to Federico Bussolino; Project "Pathogen Readiness Platform for CERIC ERIC upgrade" - PRP@CERIC CUP J97G22000400006 to Pietro Campiglia.
Author information
Authors and Affiliations
Contributions
Each author significantly contributed to the conceptualization of the study, the acquisition, analysis, or interpretation of data, as well as the drafting of the paper. All authors approved the final version of the manuscript. CC and FS designed the research and carried out experiments; SM, VVB, ES, BI, IM, and EM performed the experiments; VL, LP, EMS analyzed data and performed bioinformatics; GLRV performed histological evaluation; EMS supervised metabolomics analyses; MM performed molecular dynamics; LC and SS performed PLA experiments; FT and GC performed SPR experiments; GP, FB, FDN, PC gave conceptual advice; CC and FS wrote the manuscript and supervised the study. All authors discussed the results and implications of the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
All methods were performed in accordance with the relevant guidelines and regulations. All animal studies were approved by the Institutional Animal Care of Regina Elena National Cancer Institute (Rome, Italy) and by the Government Committee of National Minister of Health (protocol permit number: 362/2021-PR) and conducted according to EU Directive 2010/63/EU and Italian D.L. 2614/2014 for animal experiments following the Institutional Guidelines for Animal Care and Welfare. The present study does not include human subjects.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Dr Gerry Melino
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Malatesta, S., Vigiano Benedetti, V., Salviati, E. et al. α-ketoglutarate/succinate ratio imbalance impairs thymine DNA glycosylase function and base excision repair process increasing susceptibility to pancreatic cancer. Cell Death Dis 17, 242 (2026). https://doi.org/10.1038/s41419-026-08475-w
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41419-026-08475-w