Abstract
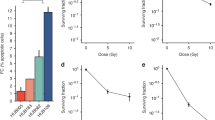
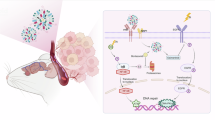
Resistance to neoadjuvant chemoradiotherapy in rectal cancer diminishes survival benefits, potentially due to dysregulated lipid metabolism, though the mechanisms are unclear. Using the MSigDB database and GSE68204 cohort, we identified lipid metabolism genes linked to radiotherapy resistance. We developed resistant cell lines and xenograft models, and through multi-algorithm analysis (SVM-RFE, RF, LASSO), pinpointed key genes. Molecular mechanisms were explored via Western blotting, co-immunoprecipitation, molecular docking, and functional assays, validated in patient-derived organoids. Our study found that radiotherapy-resistant rectal cancer shows a lipid accumulation phenotype, with an inverse relationship between lipid droplet deposition and radiosensitivity in resistant cell models. The multi-algorithm screening identified GDPD5 as a key regulator. Silencing GDPD5 reduced lipid accumulation and increased radiosensitivity. Mechanistically, GDPD5 competes with CD55, disrupting its interaction with EGFR and promoting EGFR nuclear translocation, which suppresses p53 and leads to lipid buildup and radiotherapy resistance in tumors. Clinical samples showed high GDPD5 and low CD55 levels correlate with EGFR nuclear localization. Patient-derived organoids with high GDPD5 also showed increased radiotherapy resistance. Our findings indicate that GDPD5 facilitates EGFR nuclear translocation by binding to CD55, suppressing p53, and causing lipid accumulation and radiotherapy resistance in tumors. Targeting the GDPD5-CD55-EGFR interaction may enhance radiosensitivity.
Similar content being viewed by others
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request. RNA-seq data for HCT116 (GSE319208) were downloaded from the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/).
References
Benson AB, Venook AP, Al-Hawary MM, Azad N, Chen YJ, Ciombor KK, et al. Rectal cancer, version 2.2022, NCCN clinical practice guidelines in oncology. J Natl Compr Canc Netw. 2022;20:1139–67.
Kagawa Y, Smith JJ, Fokas E, Watanabe J, Cercek A, Greten FR, et al. Future direction of total neoadjuvant therapy for locally advanced rectal cancer. Nat Rev Gastroenterol Hepatol. 2024;21:444–55.
Scott AJ, Kennedy EB, Berlin J, Brown G, Chalabi M, Cho MT, et al. Management of locally advanced rectal cancer: ASCO guideline. J Clin Oncol. 2024;42:3355–75.
Coelho D, Estêvão D, Oliveira MJ, Sarmento B. Radioresistance in rectal cancer: can nanoparticles turn the tide?. Mol Cancer. 2025;24:35.
Buckley AM, Lynam-Lennon N, O’Neill H, O’Sullivan J. Targeting hallmarks of cancer to enhance radiosensitivity in gastrointestinal cancers. Nat Rev Gastroenterol Hepatol. 2020;17:298–313.
Yu Y, Yu J, Ge S, Su Y, Fan X. Novel insight into metabolic reprogrammming in cancer radioresistance: a promising therapeutic target in radiotherapy. Int J Biol Sci. 2023;19:811–28.
An D, Zhai D, Wan C, Yang K. The role of lipid metabolism in cancer radioresistance. Clin Transl Oncol. 2023;25:2332–49.
Liang J, Liao L, Xie L, Tang W, Yu X, Lu Y, et al. PITPNC1 Suppress CD8(+) T cell immune function and promote radioresistance in rectal cancer by modulating FASN/CD155. J Transl Med. 2024;22:117.
Borza R, Matas-Rico E, Perrakis A, Moolenaar WH. Unlocking the signaling potential of GPI-anchored proteins through lipolytic cleavage. Trends Cell Biol. 2025;35:732–44.
Feng C, Zhang L, Sun Y, Li X, Zhan L, Lou Y, et al. GDPD5, a target of miR-195-5p, is associated with metastasis and chemoresistance in colorectal cancer. Biomed Pharmacother. 2018;101:945–52.
Cao MD, Döpkens M, Krishnamachary B, Vesuna F, Gadiya MM, Lønning PE, et al. Glycerophosphodiester phosphodiesterase domain containing 5 (GDPD5) expression correlates with malignant choline phospholipid metabolite profiles in human breast cancer. NMR Biomed. 2012;25:1033–42.
Wijnen JP, Jiang L, Greenwood TR, Cheng M, Döpkens M, Cao MD, et al. Silencing of the glycerophosphocholine phosphodiesterase GDPD5 alters the phospholipid metabolite profile in a breast cancer model in vivo as monitored by (31) P MRS. NMR Biomed. 2014;27:692–9.
Luo T, Peng J, Li Q, Zhang Y, Huang Y, Xu L, et al. GDPD5 related to lipid metabolism is a potential prognostic biomarker in neuroblastoma. Int J Mol Sci. 2022;23:13740.
Busato F, Khouzai BE, Mognato M. Biological mechanisms to reduce radioresistance and increase the efficacy of radiotherapy: state of the art. Int J Mol Sci. 2022;23:10211.
Shen A, Chen Y, Liu L, Huang Y, Chen H, Qi F, et al. EBF1-mediated upregulation of ribosome assembly factor PNO1 contributes to cancer progression by negatively regulating the p53 signaling pathway. Cancer Res. 2019;79:2257–70.
Wang X, Xu M, Peng Y, Naren Q, Xu Y, Wang X, et al. Triptolide enhances lipolysis of adipocytes by enhancing ATGL transcription via upregulation of p53. Phytother Res. 2020;34:3298–310.
Han Z, Liu M, Xie Y, Zeng K, Zhan Z, Chen Y, et al. Derepression of the USP22-FASN axis by p53 loss under oxidative stress drives lipogenesis and tumorigenesis. Cell Death Discov. 2022;8:445.
Ding J, Li X, Khan S, Zhang C, Gao F, Sen S, et al. EGFR suppresses p53 function by promoting p53 binding to DNA-PKcs: a noncanonical regulatory axis between EGFR and wild-type p53 in glioblastoma. Neuro Oncol. 2022;24:1712–25.
Zhao K, Zhou Y, Qiao C, Ni T, Li Z, Wang X, et al. Oroxylin A promotes PTEN-mediated negative regulation of MDM2 transcription via SIRT3-mediated deacetylation to stabilize p53 and inhibit glycolysis in wt-p53 cancer cells. J Hematol Oncol. 2015;8:41.
Liu Y, Bodmer WF. Analysis of P53 mutations and their expression in 56 colorectal cancer cell lines. Proc Natl Acad Sci USA. 2006;103:976–81.
Tan Z, Xiao L, Tang M, Bai F, Li J, Li L, et al. Targeting CPT1A-mediated fatty acid oxidation sensitizes nasopharyngeal carcinoma to radiation therapy. Theranostics. 2018;8:2329–47.
Kery M, Papandreou I. Emerging strategies to target cancer metabolism and improve radiation therapy outcomes. Br J Radiol. 2020;93:20200067.
Zhang Y, Yang Z, Liu Y, Pei J, Li R, Yang Y. Targeting lipid metabolism: novel insights and therapeutic advances in pancreatic cancer treatment. Lipids Health Dis. 2025;24:12.
Wang CJ, Li D, Danielson JA, Zhang EH, Dong Z, Miller KD, et al. Proton pump inhibitors suppress DNA damage repair and sensitize treatment resistance in breast cancer by targeting fatty acid synthase. Cancer Lett. 2021;509:1–12.
Marchan R. GDPD5, a choline-generating enzyme and its novel role in tumor cell migration. Arch Toxicol. 2016;90:3143–4.
Matas-Rico E, van Veen M, Leyton-Puig D, van den Berg J, Koster J, Kedziora KM, et al. Glycerophosphodiesterase GDE2 promotes neuroblastoma differentiation through glypican release and is a marker of clinical outcome. Cancer Cell. 2016;30:548–62.
Yu EY, Cheung NV, Lue NF. Connecting telomere maintenance and regulation to the developmental origin and differentiation states of neuroblastoma tumor cells. J Hematol Oncol. 2022;15:117.
Westerhaus A, Joseph T, Meyers AJ, Jang Y, Na CH, Cave C, et al. The distribution and function of GDE2, a regulator of spinal motor neuron survival, are disrupted in amyotrophic lateral sclerosis. Acta Neuropathol Commun. 2022;10:73.
Liu Y, Gu W. The complexity of p53-mediated metabolic regulation in tumor suppression. Semin Cancer Biol. 2022;85:4–32.
Kang JG, Lago CU, Lee JE, Park JH, Donnelly MP, Starost MF, et al. A mouse homolog of a human TP53 germline mutation reveals a lipolytic activity of p53. Cell Rep. 2020;30:783–92.
Koo KY, Moon K, Song HS, Lee MS. Metabolic regulation by p53: implications for cancer therapy. Mol Cells. 2025;48:100198.
Song B, Yang P, Zhang S. Cell fate regulation governed by p53: friends or reversible foes in cancer therapy. Cancer Commun. 2024;44:297–360.
Huang RX, Zhou PK. DNA damage response signaling pathways and targets for radiotherapy sensitization in cancer. Signal Transduct Target Ther. 2020;5:60.
Chen X, Zhang T, Su W, Dou Z, Zhao D, Jin X, et al. Mutant p53 in cancer: from molecular mechanism to therapeutic modulation. Cell Death Dis. 2022;13:974.
Huang R, Zhou PK. DNA damage repair: historical perspectives, mechanistic pathways and clinical translation for targeted cancer therapy. Signal Transduct Target Ther. 2021;6:254.
Funk JS, Klimovich M, Drangenstein D, Pielhoop O, Hunold P, Borowek A, et al. Deep CRISPR mutagenesis characterizes the functional diversity of TP53 mutations. Nat Genet. 2025;57:140–53.
Yan S, Zhan F, He Y, Zhu Y, Ma Z. p53 in colorectal cancer: from a master player to a privileged therapy target. J Transl Med. 2025;23:684.
Lane DP. Mutant p53 gain-of-function in the spotlight: are we suffering a GOF delusion? Cancer Discov. 2024;14:211–3.
Kennedy MC, Lowe SW. Mutant p53: it’s not all one and the same. Cell Death Differ. 2022;29:983–7.
Hu J, Cao J, Topatana W, Juengpanich S, Li S, Zhang B, et al. Targeting mutant p53 for cancer therapy: direct and indirect strategies. J Hematol Oncol. 2021;14:157.
Kastenhuber ER, Lowe SW. Putting p53 in context. Cell. 2017;170:1062–78.
Levine AJ. p53: 800 million years of evolution and 40 years of discovery. Nat Rev Cancer. 2020;20:471–80.
Zhu J, Wu Z, Shan G, Huang Y, Liang J, Zhan C. Nuclear epidermal growth factor receptor (nEGFR) in clinical treatment. Heliyon. 2024;10:e40150.
Yang CC, Lin LC, Lin YW, Tian YF, Lin CY, Sheu MJ, et al. Higher nuclear EGFR expression is a better predictor of survival in rectal cancer patients following neoadjuvant chemoradiotherapy than cytoplasmic EGFR expression. Oncol Lett. 2019;17:1551–8.
Ewen-Campen B, Perrimon N. Wnt signaling modulates the response to DNA damage in the Drosophila wing imaginal disc by regulating the EGFR pathway. PLoS Biol. 2024;22:e3002547.
Bharti R, Dey G, Lin F, Lathia J, Reizes O. CD55 in cancer: complementing functions in a non-canonical manner. Cancer Lett. 2022;551:215935.
Lala T, Hall RA. Adhesion G protein-coupled receptors: structure, signaling, physiology, and pathophysiology. Physiol Rev. 2022;102:1587–624.
Saygin C, Wiechert A, Rao VS, Alluri R, Connor E, Thiagarajan PS, et al. CD55 regulates self-renewal and cisplatin resistance in endometrioid tumors. J Exp Med. 2017;214:2715–32.
Leung TH, Tang HW, Siu MK, Chan DW, Chan KK, Cheung AN, et al. Human papillomavirus E6 protein enriches the CD55(+) population in cervical cancer cells, promoting radioresistance and cancer aggressiveness. J Pathol. 2018;244:151–63.
Olcina MM, Balanis NG, Kim RK, Aksoy BA, Kodysh J, Thompson MJ, et al. Mutations in an innate immunity pathway are associated with poor overall survival outcomes and hypoxic signaling in cancer. Cell Rep. 2018;25:3721–32.
Li G, Yin Q, Ji H, Wang Y, Liu H, Jiang L, et al. A study on screening and antitumor effect of CD55-specific ligand peptide in cervical cancer cells. Drug Des Devel Ther. 2018;12:3899–912.
Yang L, Wei W, Yuan X, Guo E, Peng P, Wang J, et al. Targeting DNA damage repair to enhance antitumor immunity in radiotherapy: mechanisms and opportunities. Int J Mol Sci. 2025;26:3743.
Khwairakpam AD, Banik K, Girisa S, Shabnam B, Shakibaei M, Fan L, et al. The vital role of ATP citrate lyase in chronic diseases. J Mol Med. 2020;98:71–95.
Fu Y, Zou T, Shen X, Nelson PJ, Li J, Wu C, et al. Lipid metabolism in cancer progression and therapeutic strategies. MedComm. 2021;2:27–59.
Chen J, Zhang F, Ren X, Wang Y, Huang W, Zhang J, et al. Targeting fatty acid synthase sensitizes human nasopharyngeal carcinoma cells to radiation via downregulating frizzled class receptor 10. Cancer Biol Med. 2020;17:740–52.
Kubik J, Humeniuk E, Adamczuk G, Madej-Czerwonka B, Korga-Plewko A. Targeting energy metabolism in cancer treatment. Int J Mol Sci. 2022;23:5572.
Kwon YS, Lee MG, Baek J, Kim NY, Jang H, Kim S. Acyl-CoA synthetase-4 mediates radioresistance of breast cancer cells by regulating FOXM1. Biochem Pharmacol. 2021;192:114718.
Nisticò C, Pagliari F, Chiarella E, Fernandes Guerreiro J, Marafioti MG, Aversa I, et al. Lipid droplet biosynthesis impairment through DGAT2 inhibition sensitizes MCF7 breast cancer cells to radiation. Int J Mol Sci. 2021;22:10102.
Zhang C, Liu P. The new face of the lipid droplet: lipid droplet proteins. Proteomics. 2019;19:e1700223.
Liu H, Du J, Chao S, Li S, Cai H, Zhang H, et al. Fusobacterium nucleatum promotes colorectal cancer cell to acquire stem cell-like features by manipulating lipid droplet-mediated numb degradation. Adv Sci. 2022;9:e2105222.
Dörr D, Obermayer B, Weiner JM, Zimmermann K, Anania C, Wagner LK, et al. C/EBPβ regulates lipid metabolism and Pparg isoform 2 expression in alveolar macrophages. Sci Immunol. 2022;7:eabj0140.
Liu H, Jing L, Li Y, Zhou J, Cui X, Li S, et al. Lipid droplet-organized MDM2-mediated P53 degradation: a metabolic switch governing diet-driven tumor progression. Adv Sci. 2025;12:e03473.
Calabria A, Cipriani C, Spinozzi G, Rudilosso L, Esposito S, Benedicenti F, et al. Intrathymic AAV delivery results in therapeutic site-specific integration at TCR loci in mice. Blood. 2023;141:2316–29.
Nuñez JK, Chen J, Pommier GC, Cogan JZ, Replogle JM, Adriaens C, et al. Genome-wide programmable transcriptional memory by CRISPR-based epigenome editing. Cell. 2021;184:2503–19.
Jiang H, Chen HC, Lafata KJ, Bashir MR. Week 4 liver fat reduction on MRI as an early predictor of treatment response in participants with nonalcoholic steatohepatitis. Radiology. 2021;300:361–8.
Kupczyk PA, Kurt D, Endler C, Luetkens JA, Kukuk GM, Fronhoffs F, et al. MRI proton density fat fraction for estimation of tumor grade in steatotic hepatocellular carcinoma. Eur Radiol. 2023;33:8974–85.
Acknowledgements
The mechanistic scheme of this study was drawn by Biorender (www.biorender.com).
Funding
This work was supported by Scientific Research Program for Young Talents of China National Nuclear Corporation, Gusu Health Talent Research Program (GSWS2023043), Suzhou Science and Technology Development Program (SYW2025102), Suzhou Radiotherapy Clinical Medical Center (Szlcyxzx202103), and Young Talent Support Project of the Second Affiliated Hospital of Soochow University (XKTJ-RC202408).
Author information
Authors and Affiliations
Contributions
RQZ, MYL, and YS initiated and designed the research. Experimental work wasconducted by RQZ, MYL, and YS, with data analysis performed by LZ, YTS, LMJ, and YQZ. QLP wrote the manuscript and supervised the study, with all authors approving the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
In vivo experiments met relevant ethical guidelines for animal research, followed AstraZeneca’s global bioethics policy, and had the approval of the ethical committee of Soochow University (Approval Number: SUDA202411A1189). All human tissue research in this study had the approval of the ethics committees of the Second Affiliated Hospital of Soochow University (Approval Number: JD-LK2022167-IR02). Informed consent was obtained from all participants involved in this study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Zhi-Xiong Xiao
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhu, R., Li, M., Shen, Y. et al. GDPD5-CD55-EGFR competitive binding axis regulates radioresistance and lipid accumulation in rectal cancer. Cell Death Dis (2026). https://doi.org/10.1038/s41419-026-08711-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-026-08711-3