Abstract
Heterogeneity is widely recognised across different cell types. Human oligodendrocyte progenitor cells (hOPCs), essential for myelination, exhibit considerable heterogeneity, which has not been fully characterised. In the current study, by examining the transcriptome of hOPCs at the single-cell level, three distinct subclusters were identified: PRE-OPCs, OPCs, and PRE-OLs. Single-cell RNA-sequencing and RNA-Scope detected high platelet-derived growth factor receptor alpha (PDGFRA) expression. PDGFR-α+ hOPCs exhibited greater myelination, migration, and proliferation capabilities compared to both unsorted hOPCs and PDGFR-α– hOPCs. These enhanced functions may be associated with the activation of the PI3K-AKT-mTOR and TGF-β signalling pathways, which support oligodendrocyte differentiation.

hOPCs were induced by hNSCs, their characteristics were identified. RNA-Scope and single-cell RNA Seq sequencing showed PDGFRA were highly expressed at mRNA and protein level. hOPCs were sorted by MACS using PDGFR-α beads. The myelination, migration, and proliferation abilities of PDGFR-α+ hOPCs were higher than that of un-sorting hOPCs and PDGFR-α– hOPCs, possibly being associated with the activation of PI3K–AKT–mTOR and TGF-β signalling pathways, which support oligodendrocyte differentiation (Partly created with Scientific Image and Illustration Software BioRender).
Similar content being viewed by others
Facts
-
The myelination, migration, and proliferation abilities of PDGFR-α+ hOPCs are predominant.
-
scRNA-sequencing and RNA-Scope are combined to characterise hOPCs heterogeneity.
-
This study provides a theoretical basis for selecting appropriate hOPCs for myelination.
-
Activation of TGF-β signalling pathways and stem cell transplantation may synergistically promote myelination and represent therapeutic targets for myelin regeneration.
Introduction
Recent studies have identified partial heterogeneity among oligodendrocyte progenitor cells (OPCs). However, most of the current understanding of this heterogeneity is based on differences in gene or surface receptors, developmental origin, and anatomical location, (e.g., the grey versus white matter) [1], with functional differences between OPC subpopulations often overlooked [2]. Currently, the terms ‘heterogeneity’ more commonly scenarios in which human OPCs (hOPCs) exhibit variability in gene expression, morphology, phenotypic profiles, and functions, sush as proliferation capacity, migration and myelination abilities [2]. Obtaining a sufficient number of purified hOPCs remains a significant challenge for single-cell investigations of genotype/phenotype, morphology, and function.
Primary hOPCs are scarce, difficult to isolate, and demonstrate limited expansion after isolation [3]. Human pluripotent stem cells (hPSCs), including both human embryonic stem cells (hESCs) and induced pluripotent stem cells (hiPSCs), have become valuable resources for differentiating in vitro into hOPCs, facilitating the evaluation of myelination and serving as a potential source for cell-based therapies. Nonetheless, hPSCs-derived hOPCs present several inherent challenges: incomplete differentiation may increase tumorigenic risks [4]; differentiation protocols are lengthy and complex, typically requiring weeks to several months, and may result in varied efficiency and purity [5,6,7]. In contrast, hNSCs derived from the human foetal brain tissue, as progenitors already committed to neural lineages, possess relatively limited self-renewal and differentiation capacities. Theoretically, this translates to lower tumorigenicity, which is critical for safe cell therapy [8,9,10].
Zhou et al. investigated functional differences when sorting hOPCs and found the myelination, migration, and proliferation abilities of NG2−/A2B5− hOPCs to be superior to those of NG2+/A2B5+ hOPCs. They attributed these enhanced capabilities to an enrichment of platelet-derived growth factor receptor (PDGFR)-α+ cells within the NG2−/A2B5− group [11]. We have captured the transcriptomes of hOPCs derived from hNSCs at the single-cell level. Based on the cluster assignment probabilities and label predictions, three sub-clusters of PRE-OPCs, OPCs, and PRE-OLs were predicted. Furthermore, the gene and protein expression of hOPCs were examined using RNA-Scope. Integrating Zhou et al.’s findings with our RNA-scope and single-cell RNA sequencing (scRNA-seq) results identified PDGFRA as a key regulator of hOPC for proliferation, migration, and myelination. hOPCs were immunopurified using anti-PDGFR-α microbeads and magnetic-activated cell sorting (MACS) for in vivo and in vitro experiments. Downstream validations are driven by marker-based hypotheses; alongside validating these hypotheses at the single-cell level using hOPCs mRNAs and proteins, emphasis is placed on selecting cells with superior proliferation, migration, and myelination capabilities.
Results
Identification of hNSCs and their induction into hOPCs
hNSCs were cultured as ‘neurospheres’. Throughout days 1–3 of the neurospheres culture, the spheres appeared well-defined with visible protrusions surrounding them. Immunofluorescence staining confirmed that hNSCs expressed the specific markers, Nestin and SOX2, were positive (Fig. 1A). Days 5–7 of the neurospheres (labelled) culture, cells began to migrate from the sphere and spread, forming bipolar, bead-like cells. At this stage, immunofluorescence revealed that one of the specific markers of oligodendrocyte line, OLIG2 was positive, and one of the specific markers of the early hOPCs, NG2 was positive (Fig. 1B). Flow cytometry results of hNSCs (n = 3, from three different biological replicates) demonstrated the following expression rates for key markers: Nestin (95.32 ± 0.50%), Musashi (64.4 ± 0.66%), SOX2 (76.5%, median), Vimentin (98.43 ± 0.76%), CD133 (92.7 ± 0.26%), and PAX6 (73.9 ± 3.73%; Fig. 1C, D).
A Morphology of NSCs observed via phase-contrast microscopy, scale bars = 50, or 20 μm; Nestin and SOX2 in NSCs confirmed by immunofluorescence assay; scale bars = 250 μm. B Morphologies of differentiation from hNSCs into hPOCs, scale bar = 50 μm; OLIG2 and NG2 confirmed by immunofluorescence assay; scale bar = 250 μm. C, D Proportions of Nestin+, Musashi+, SOX2+, vimentin+, CD133+, and PAX6+ hNSCs analysed as by flow cytometry, blue and orange curves mean without Ab (negative control) and with Ab (sample), respectively. The positive rates (Nestin, Musashi, vimentin, CD133, and PAX6) that conformed to normality were presented as the average value, otherwise it was presented as the median, such as SOX2.
Identification of hOPCs induced by hNSCs
The hOPCs derived from hNSCs displayed cell bodies that were round, elliptical, or spindle-shaped, with protrusions appearing as elongated, beaded-like bipolar, or multipolar extensions (Fig. 2A, B). Immunofluorescence staining showed positive results for markers associated with the oligodendrocyte lineage (OLIG2), early oligodendrocyte precursors (NG2 and A2B5), the typical marker PDGFR-α, and late oligodendrocyte precursor cell markers O4 and CNPase (Fig. 2C). Flow cytometry analysis of hOPCs indicated average positivity rates for PDGFR-α at 80.8 ± 1.44%, A2B5 at 46.83 ± 2.37%, OLIG2 at 95.13 ± 0.42%, and SOX10 at 93.2 ± 1.27% (Fig. 2E, F). Bulk RNA-seq revealed a heatmap showing high levels of VIM and NES in hNSCs, while early OPC-related genes like ST8SIA1 and CSPG4, the typical hOPC gene PDGFRA, and late-stage hOPC genes APC and GALC were predominantly expressed in hOPCs (Fig. 2D). Subsequent qPCR validation (n ≥ 3 per group), confirmed these results (Fig. 2G).
A, B Morphologies of hOPCs as observed; scale bar = 50 or 20 μm. C Protein expressions of OLIG2, NG2, A2B5, PDGFRA, O4, and CNPase in hOPCs as confirmed by immunofluorescence assay; scale bar = 20 μm. D Heatmap of genes of VIM, NES, OLIG2, OLIG1, ST8SIA1, CSPG4, APC, PDGFRA, and GALC expressed in hNSCs and hOPCs. E, F Proportions of PDGFR-α+, A2B5+, OLIG2+, and SOX10+ hOPCs as analysed by flow cytometry; The positive rates of PDGFR-α, A2B5, OLIG2, and SOX10 conformed to normality were presented as the average value (each n ≥ 3). G mRNAs levels of NSCs and OPC markers as detected by qPCR (n ≥ 3 each group). In hNSCs vs hOPCs, the gene expression levels of hNSCs and hOPCs conformed to both normality and homogeneity of variance, and the independent sample t-test was used in NESTIN (p = 0.025), VIM (p = 0.052), PDGFRA (p <0.01), ST8SIA1(A2B5) (p <0.01), OLIG1 (p < 0.01) and CSPG4 (p <0.01). However, gene expression levels in hNSCs and hOPCs conformed to normality but not homogeneity of variance, and Welch’s t-test was performed in OLIG2 (p = 0.008) and GALC (p < 0.01), *p < 0.05, **p < 0.01.
Single-cell RNA sequencing (scRNA-seq) of hOPCs
An RNA-seq strategy was used to examine the mRNAs profiles of hOPCs-1 at the single-cell level. After quality control, a total of 7885 cells selected for subsequent investigation, revealing three distinct subpopulations (Fig. 3A). Furthermore, a scRNA-seq data were collected from the hOPCs -1, hOPCs -2, and hOPCs -3 (hOPCs -1, 2, and 3 derived from three different biological replicates; Fig. 3B). A total of 28 801 cells were scored and categorised into three populations: PRE-OPCs, OPCs and PRE-OLs (Fig. 3B, C). The OPC population in our study shared the expressions of 12 887 genes, including PDGFRA, OLIG1, OLIG2, ST8SIA1, NKX2-2, and CNP, with those previously reported in human OPCs [12] (Fig. 3D). Similarly, the PRE-OL population in our study shared expression of 13,199 genes, including PDGFRA, OLIG1, OLIG2, CNP, PLLP, PLP1, APC, and PTEN, with those previously reported for human immune oligodendroglia (IMOIGs) [12] (Fig. 3D). The PRE-OPCs, OPCs, and PRE-OLs populations in our study shared similarities with the OPCs sampled on Days 77, 89, 85, and 104 in a prior study[13] (Fig. 3E). Volcano plots depicting DE mRNAs in pairwise comparisons among OPCs versus PRE-OLs, OPCs versus PRE-OPCs, and PRE-OPCs versus PRE-OLs (Figure S2A), highlighted PDGFRA as highly expressed across all stages. Finally, upregulated and downregulated DE mRNAs underwent GO enrichment analysis and were visualised using Cytoscape (Fig. S2B–G).
A Single-cell transcriptional profiles of 7885 cells visualised with t-SNE embedding. B Co-imbedding of our hOPCs from hOPCs -1, hOPCs -2, and hOPCs-3 were predicted into PRE-OPCs, OPCs, and PRE-OLs. C Distribution of genetic markers among the different clusters presented as an enrichment heatmap. Green colour represents PRE-OPCs, pink colour represents OPCs, and blue colour represents PRE-OLs. D Venn diagram depicting the overlap of genes in OPCs enriched in our system vs human OPCs previously reported, in PRE-OLs (our system) vs IMOLGs previously reported, with examples of overlapping and different genes. E The correlations between the PRE-OPCs, OPCs and PRE-OLs populations in our study and the previously reported OPCs at 77, 89, 85 and 104 days were analysed. Average cell-type label probability cell-type label probability assignment combined reference. Rows: cells from reported data set, column: cell types from the data set of our system. Red colour represents higher probability and blue is used for lower probability.
RNA-Scope demonstrates mRNA and protein expression across three developmental stages
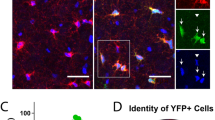
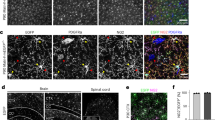
During the initial differentiation stage, i.e., PRE-OPCs, cells exhibited a unipolar morphology. In the OPCs stage, their morphology shifted to biopolar or multipolar forms, while cells in the PRE-OL stage predominantly showed a multipolar structurre (Fig. 4A) [3]. Using RNA-Scope technology, PDGFRA mRNAs was fluorescently labelled with purple, CNP with orange, and SOX2 with green pseudo-colours (Fig. 4B, C), respectively, with the corresponding proteins highlighted in white pseudo-colours (Fig. 4D). The abundance of mRNAs represented as average dots /μm2, and protein levels were quantified by mean intensity. Results from experiments performed on three culture slides, analysing 11 862 hOPCs, indicated that PDGFRA mRNAs had the highest average number of dots per μm2, followed by SOX2 with intermediate levels and CNP with the lowest (Fig. 4E). Protein expression patterns paralleled those observed for mRNAs (Fig. 4F). Overall, RNA-Scope analysis showed that PDGFRA, CNP, and SOX2 mRNAs and proteins were expressed during the PRE-OPC, OPC, and PRE-OL stages, with PDGFRA consistently expressed at higher levels than the other two markers (Fig. S3A–D).
A Bright-field images of PRE-OPCs with a unipolar morphology, OPCs with a biopolar (or multipolar) morphology, and PRE-OLs with a multipolar regraded morphology; scale bar = 50 or 10 μm. B Representative Tissue FAXS Viewer images of RNA Scope®-PDGFRA, CNP, and SOX2 mRNAs expression in hOPCs, scale bar = 5 μm. C, D FAXS Viewer images showed (C) average dots /μm2 of mRNAs and D the intensities of proteins of PDGFRA, CNP and SOX2. Purple pseudo-colour (Opal™ 520) repressents the number of mRNA of average dots/μm2 of PDGFRA, orange pseudo-colour (Opal™ 570) represents the number of mRNA of average dots/μm2 of CNP, green pseudo-colour (Opal™ 620) represents the number of mRNA of average dots/μm2 of SOX2, and white pseudo-colour (Opal™ 690) represents the corresponding fluorescence intensities of each protein, scale bar = 5 μm. Statistical analyses of (E) the average dots /μm2 of mRNAs of PDGFRA (9.815), CNP (5.015), and SOX2 (5.743) and F the mean intensities of proteins of PDGFR-α (94.664), CNPase (76.120), and SOX2 (73.209). For the data of PDGFRA, CNP, and SOX2 mRNAs in RNA-SCOPE conformed to normality but not to homogeneity of variance. Therefore, pairwise comparisons were performed using Welch’s t-test, with Šidák correction applied to control the family-wise error rate. The results were as follows: SOX2 vs. CNP (mean difference = 0.728 ± 0.083, Šidák-adjusted p < 0.0001, 95% CI [0.531, 0.925]); SOX2 vs. PDGFRA (mean difference = −4.072 ± 0.083, Šidák-adjusted p < .0001, 95% CI [−4.269, −3.875]); and CNP vs. PDGFRA (mean difference = −4.800 ± 0.083, Šidák-adjusted p < 0.0001, 95% CI [−4.997, −4.603]). The protein expression data for SOX2, CNP, and PDGFRA were non-normally distributed. We performed the three-group comparisons using the Kruskal–Wallis test, followed by Dunn’s multiple comparisons test for post hoc analysis. The Kruskal–Wallis test was significant (p < 0.0001). Dunn’s test results showed that PDGFRA vs. SOX2 (adjusted p < 0.0001), and PDGFRA vs. CNP (adjusted p < .0001) were significant, while SOX2 vs. CNP was not significant (adjusted p = 0.112).
Functional verification of PDGFR-α+, PDGFR-α–, and unsorted hOPCs
The PDGFRA mRNA positivity rates determined by RNA-Scope and scRNA-seq were 93.47% and 68.58%, respectively. Both mRNAs and proteins levels in PDDFRA were highly expressed in three stages. Building upon the previous work of Zhou et al. [11], hOPCs were immunopurified using anti-PDGFR-α microbeads and magnetic-activated cell sorting (MACS). Before MACS, the positive rate of PDGFR-α+ hOPCs was71.56 ± 15.22%; which increased to 90.33% ± 12.15% after MACS (Fig. 5A). Post-sorting identification confirmed that PDGFR-α+ hOPCs displayed positive immunofluorescence staining for the PDGFR-α antibody, while PDGFR-α– hOPCs showed no detectable staining (Fig. S4).
A Gating strategy to define PDGFR-α+ cell populations: the positive rate of PDGFR-α+ cell populations before and after MASCs, the value was expressed as the x ± STDEV.P. B Representative TEM images of myelin sheath in the corpus callosum in PDGFR-α+, PDGFR-α–, and unsorted hOPCs transplanted and control groups. The scale bar represents 2 µm and 200 nm for the insert pictures. C Alexa 488-labelled, donor-derived MBP (green) of four groups. Scale bar = 500 μm or 20 μm. Statistical analyses of the (D) G-ratio and E the remyelination rate of four groups. When comparing the G-ratio and remyelination rate in PDGFR-α+ hOPCs vs hOPCs, for the data of G-ratio, data conformed to normality but not homogeneity of variance, we used Welch’s t-test, ****p < 0.0001. for the data of remyelination, Welch’s t-test were also used, ****p < 0.0001. F Migration assay: microscopic images show DAPI staining of migrating cell nuclei. Scale bar = 20 μm. G Statistical analyses of the cell migration rate of PDGFR-α+ hOPCs, PDGFR-α– hOPCs, and unsorted hOPCs groups; The pairwise comparisons were performed using Šidák’s multiple comparisons test following a significant ordinary one-way ANOVA. All adjusted p-values were below the 0.05 threshold: hOPCs vs. PDGFR-α+ hOPCs (adjusted p = 0.038), 95% CI [−8.73, −0.25]; hOPCs vs. PDGFR-α– hOPCs (adjusted p = 0.0019), 95% CI [3.16, 11.64]; and PDGFR-α+ hOPCs vs. PDGFR-α– hOPCs (adjusted p < 0.0001), 95% CI [7.65, 16.13]. H Statistical analyses of OD value (450 nm) of PDGFR-α+ hOPCs, PDGFR-α– hOPCs, and unsored hOPCs groups.
Remyelination capacity of PDGFR-α+, PDGFR-α–, and unsorted hOPCs was assessed via electron microscopy three weeks post-transplantation into shiverer–/– mice (Fig. 5B). Myelin sheaths in the PDGFR-α+ and unsorted groups appeared denser and more compact than those of the PDGFR-α– group (Fig. 5B, red Box). Immunofluorescence staining demonstrated prominent filamentous MBP expression in the PDGFR-α+ and unsored groups (Fig. 5C, red Box). The remyelination was highest in the PDGFR-α+ hOPC group when compared to the unsorted hOPC, PDGFR-α– hOPC, and untransplanted (control) groups (Fig. 5E). Conversely, G-ratio values were lower in the PDGFR-α+ hOPC group relative to the unsorted hOPC, PDGFR-α– hOPC, and control groups (Fig. 5D).
Migratory capabilities of the PDGFR-α+, PDGFR-α–, and unsorted hOPCs were evaluated by measuring the distance traversed from the upper to the lower chamber after a 16–18 h attachment period (Fig. 5F). After 18 h, the migration rate of the PDGFR-α+ hOPCs group exceeded those the PDGFR-α– group (p < 0.0001) and unsorted groups (p = 0.0379; Fig. 5G). Additionally, proliferation of PDGFR-α+ and unsorted hOPCs was greater than that of PDGFR-α– hOPCs from day 1 to day 11 (Fig. 5H).
GO and KEGG terms enriched from DE mRNAs of in PDGFR-α+ and PDGFR-α– populations
Volcano plots illustrating the expression of DE mRNAs expression between PDGFR-α+ versus PDGFR-α– hOPCs were generated (Fig. 6A). Go analysis identified enrichment in glial cell differentiation, gliogenesis, compact myelin, and myelin sheath processes within PDGFR-α+ compared to PDGFR-α– hOPCs (Fig. 6B). KEGG pathway revealed that the DE mRNAs were associated with TGF-β signalling, Notch signalling, glioma, mammalian target of rapamycin (mTOR) signalling, vascular endothelial growth factor (VEGF) signalling, gap junctions, and Wnt signalling pathways in PDGFR-α+ versus PDGFR-α– hOPCs (Fig. 6C). Integration of GO enrichments and related DE genes in Cytoscape indicated that upregulated genes (PDGFRA, CNP, OLIG1, OLIG2, VIM, CD9, PTN, and PLP1) were predominantly enriched in glial cell differentiation, myelin sheath and gliogenesis (Fig. 6D, threshold: -0.82 <avg_logFC <0.49). Further integration of KEGG pathway data and associated DE genes demonstrated that upregulated genes (PDGFRA, APC, EGFR, ID4, and GLUL) were enriched in TGF-β signalling, neurotrophin signalling, and gap junction pathways (Fig. 6E, threshold: −0.10 <avg_logFC <0.10).
A DE mRNAs associated with differentiation and maturation of OLs drawn in volcano plots in comparisons of PDGFR-α+ and PDGFR-α– cells. B GO and C KEGG pathway enrichment terms for DE mRNAs related to differentiation and maturation of OLs in comparisons of PDGFR-α+ and PDGFR-α– populations. Interaction network for DE mRNAs and (D) GO and E KEGG pathway enrichment terms in PDGFR-α+ and PDGFR-α– populations comparisons. Yellow ovals represent GO enrichment and KEGG pathway terms; red ovals represent upregulated mRNAs (avg_logFCå 0), and green ovals represent downregulated mRNAs (avg_logFC <0). Functions that may influence the differentiation and maturation of OLs are shown in bold black font. D Threshold: −0.82 <avg_logFC <0.49; E threshold: −0.10 <avg_logFC <0.10.
Interactions between the PI3K–AKT–mTOR and TGF-β signalling pathways in regulating hOPCs proliferation
Analysis of KEGG pathways enriched by the DE mRNAs in PDGFR-α+ and PDGFR-α– hOPCs revealed that the TGF-β signalling pathway exhibited the highest -log10 (padj) value (Fig. 6C). Moreover, PDGFR-α+ hOPCs activated the PI3K–AKT–mTOR signalling pathway (Fig. 7A), which promotes myelination [14,15,16]. The binding between PDGFR-α and PDGFRAA and their ligands initiates PI3K-AKT-mTOR signalling, resulting in upregulation of SMAD3 expression and subsequent activation of the TGF-β signalling pathway [17] (Fig. 7B). These findings suggest that crosstalk between the PI3K–AKT–mTOR and TGF-β signalling pathways may influence the proliferation, migration, and myelination capacities of PDGFR-α+ hOPCs.
A PDGFR-α+ hOPCs activated PI3K–AKT–mTOR signalling pathways in our study. B The diagram of the interaction between the PI3K–AKT–mTOR signalling pathway and the TGF-β signalling pathway (Created with Scientific Image and Illustration Software | BioRender).
Role of TGF-β pathway activation in enhancing hOPCs myelination
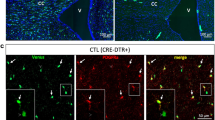
SRI-011381, an activator of the TGF-β pathway[18] was applied to in vitro cultures of PDGFR-α+ hOPCs. Fluorescent labelling of PDGFRA-, CNP-, and SOX2- mRNAs was conducted in untreated PDGFR-α+ hOPCs and PDGFR-α+ hOPCs treated with SRI-011381 (Fig. 8A). The average dots /μm2 for PDGFRA, CNP, and SOX2 was higher in the SRI-011381-treated group compared to the untreated controls (Fig. 8B). Quantitative PCR analysis further revealed elevated expression levels of PDGFRA, OLIG2, CSPG4, NKX2-2, CNP, and PLP1 mRNAs in the SRI-011381-treated population relative to the untreated cells (Fig. 8C). Electron microscopy conducted three weeks post-transplantation into shiverer–/– mice assessed myelin sheath formation (Fig. 8D). Immunofluorescence staining demonstrated filamentous MBP expression in mice transplanted with PDGFR-α+ hOPCs or SRI-011381-treated PDGFR-α+ hOPCs (Fig. 8E). Notably, the remyelination rate was higher in the group transplanted with SRI-011381-treated PDGFR-α+ hOPCs than in those transplanted with untreated PDGFR-α+ hOPCs (Fig. 8F).
A Tissue FAXS Viewer images of RNA Scope® showed the mRNAs of PDGFRA, CNP, and SOX2 in PDGFR-α+ hOPCs and PDGFR-α+ hOPCs + SRI-011381, scale bar=5 µm. B Statistical analyses of the average dots /μm2 of mRNAs of PDGFRA, CNP, and SOX2 in PDGFR-α+ hOPCs and PDGFR-α+ hOPCs+SRI-011381, respectively. The data of two groups conformed to both normality and homogeneity of variance, and the Independent Samples t-test was used, P-values are displayed in graphs. C mRNA levels of PDGFRA, OLIG2, CSPG4, NKX2.2, CNP, MBP, and PLP1 in PDGFR-α+ hOPCs and PDGFR-α+ hOPCs + SRI-011381 as detected by qPCR. In PDGFR-α+ hOPCs vs. PDGFR-α+ hOPCs + SRI-011381, the data conformed to neither normality and to homogeneous variance, and the Mann-Whitney U Tests were used in PDGFRA (p = 0.030), OLIG2 (p = 0.037), CSPG4 (p = 0.0189), NKX2.2 (p <0.0001), CNP (p = 0.0190), MBP (p = 0.087), and PLP1(p = 0.024). D Representative TEM images of myelin sheath in the corpus callosum in PDGFR-α+ hOPCs, and PDGFR-α+ hOPCs. The scale bar represents 2 µm and 200 nm for the insert pictures. E Alexa 488-labelled, donor-derived MBP (green) of two groups. Scale bar = 10 μm or 20 μm. F Statistical analyses of the remyelination rate of the two groups. The data of two groups conformed to both normality and to homogeneous variance, the Independent Samples t-test was used. p < 0.0001.
Discussion
The optimal cell type for myelination of stem cell transplantation may not necessarily be mature hOLs, which are fragile and lack the ability to migrate or proliferate. These limitations restrict their utility in stem cell therapy, particularly in terms of migration and mechanical tolerance—factors crucial for efficient transmission of electrical signals in neurons [19,20,21]. This underscores the importance of considering the transcriptional diversity, developmental stage, heterogeneity, and primary functional populations of hOPCs [22]. Key questions arise regarding the nature of transcriptional heterogeneity and the methods used to study it in hOPCs. Evidence supporting heterogeneity in both OLs and OPCs has increased substantially over the past decade [23]. scRNA-seq of OLs has facilitated a more detailed exploration of their diversity. For instance, Marques et al. identified 12 distinct mouse oligodendrocyte lineage subsets spanning various OPC stages from differentiated to mature OLs across ten regions of the central nervous system using scRNA-seq [24]. Although a few studies have described the heterogeneity of hOPCs, several technical challenges remain for in vitro-cultured hOPCs in RNA-seq, including limited RNA quantities, hOPC purity and yield, and their vulnerabilities [25]. Nevertheless, hOPCs induced from hNSCs obtained from the human brain are of sufficient yield and quality to meet scRNA-seq technical requirements[26, 27].
DNA in situ hybridisation and immunohistochemistry are commonly used to assess DNA and proteins expression. However, the in situ analysis of RNA is rarely reported. RNA-Scope enables the detection of mRNAs biomarkers labelled with multiple fluorescent chromogenic dyes, which can be visualised using bright-field at the single-cell level. In the current study, RNA-Scope (93.47%) and scRNA-seq (68.58%) showed that high PDGFRA mRNAs expression, albeit with certain discrepancies. The phenomenon of ‘gene loss’ in scRNA-seq refers to transcripts of medium and low abundance that are not effectively captured or detected [28]. Single-cell suspensions may bias capture towards particular cell types, affecting the representativeness of sequencing data [29]. In contrast, RNA-Scope’s high sensitivity facilitates the reliable detection of low-abundance mRNA in tissue sections. The novel probe design strategy in RNA in situ hybridisation can amplify signals while suppressing background, striking a balance between cell morphology and single-molecule visualisation [30]. Thus, the observed consistency in expression patterns across techniques in the current study indicates PDGFRA as the predominant marker of hOPCs.
Zhou et al. previously reported that NG2−/A2B5− hOPCs, which contain a higher proportion of PDGFR-α+ cells, exhibit superior myelination, migration, and proliferation compared to NG2+/A2B5+ hOPCs[11]. Building on this, in the current study, MACS with anti-PDGFR-α microbeads was utilised to separate PDGFR-α+ and PDGFR-α− hOPCs, enabling investigation of their functional differences. Shiverer–/– mice lacks myelin basic protein (MBP), which is essential for the compaction of multilayered myelin. As a result, these mice are unable to form compact myelin sheaths. In shiverer–/– mice, axons are typically either completely unmyelinated or surrounded by a single, loose, and uncompacted myelin layer that appears thin and fragmented [31]. The PDGFR-α+ hOPCs demonstrated greater myelination, migration, and proliferation than their PDGFR-α− counterparts, highlighting the functional significance of PDGFRA in linking the transcriptome to cellular behaviour. Future downstream validation will be based on direct functional comparison of original transcriptomic clusters. Advanced approaches such as combining barcode lineage tracing with scRNA-seq [32, 33], or directly performing scRNA-seq on transplanted cells and computationally aligning them with initial reference maps [34] will be implemented in subsequent research. The PDGFR-α+ cell population expressed a number of genes (PDGFRA, CNP, OLIG1, OLIG2, VIM, and PLP1)—at higher levels than that of PDGFR-α− cells. These genes are associated with which involved in oligodendrocyte-genesis, glial cell differentiation, gliogenesis, compact myelin, and myelin sheath. Moreover, the KEGG pathway which related to oligodendrocyte revealed involvement in Notch signalling [35, 36], transforming growth factor TGF-β signalling [37], mammalian target of rapamycin (mTOR) signalling [38, 39], vascular endothelial growth factor (VEGF) signalling [40], gap junctions [41], and the Wnt signalling pathway [42, 43]. In particular, upregulated genes such as PDGFRA, APC, EGFR, ID4, and GLUL were enriched in TGF-β, neurotrophin, and Wnt signalling pathways. Collectively, these results support the potential of PDGFR-α+ cells for oligodendrocyte differentiation.
The value of -log10 (padj) in the TGF-β signalling pathways was notably high. For regulating TGF-β signalling, SMAD families such as inhibitory SMADs (I-SMADS) play a key role [44]. After bone morphogenetic proteins (BMPs), one of the ligands of the TGF-β superfamily, bind to the receptor, SMAD1/5/8 binds to SMAD4 and enters the nucleus [44]. After binding of TGF-β superfamily ligand to type II receptors, type II receptors recruit the phosphorylated type I receptors that regulate R-SMADs (SMAD2/3) and bind coSMAD-SMAD4. A complex of R-SMAD/coSMAD as a transcription factor accumulates in the nucleus is involved in regulating the expression of target genes[45]. Compared with PDGFR-α− hOPCs, upregulation of ID2/ID3 of PDGFR-α+ hOPCs is linked with neuronal differentiation and central nervous system development, while downregulation of ID4 is associated with negative regulation of neuron and astrocyte differentiation (Fig. S5).
Changes in these target genes driven by TGF-β signalling may promote oligodendrocyte differentiation. PDGFR-α+ hOPCs also activated the PI3K–AKT–mTOR signalling pathway, which reportedly promotes myelination[14,15,16]. Meanwhile, it has been reported that the interaction between PDGFR-α and PDGFRAA ligands activates PI3K–AKT–mTOR signalling, which regulates SMAD3 expression and subsequently triggers the TGF-β signalling pathway [17]. SRI-011381, a TGF-β pathway activator, exerts neuroprotective effects [18]; when added to PDGFR-α+ hOPCs cultures, SRI-011381 upregulated PDGFRA, CNP, and SOX2 mRNA expression. Additionally, the expression of PDGFRA, OLIG2, CSPG4, NKX2-2, CNP, and PLP1 mRNAs was higher in SRI-011381-treated PDGFR-α+ hOPCs than in untreated PDGFR-α+ hOPCs. The remyelination rate was also higher in SRI-011381-treated PDGFR-α+ hOPCs. It is speculated that the PI3K–AKT–mTOR pathway may interacts with the TGF-β signalling pathway, contributing to the proliferation, migration, and myelination functions of PDGFR-α+ hOPCs. However, the conclusions of this study are based on transcriptomic differential expression and KEGG enrichment analysis; thus, direct functional verification of pathway inhibition or activation remains to be performed. Future research will include the use of pathway inhibitors or gene silencing to evaluate proliferation, migration, and myelination functions in these cells.
Conclusions
In summary, hOPCs display heterogeneity and can be classified into PRE-OPCs, OPCs and PRE-OLs stages using scRNA-seq analysis. Both scRNA-Seq and RNA-Scope show that PDGFR mRNA and protein are highly expressed across all stages. PDGFR-α+ hOPCs excel in myelination, proliferation, and migration, which may be linked to activation of the PI3K–AKT–mTOR and TGF-β pathways. Collectively, these findings offer a new perspective for selecting stem cells for transplantation to treat myelin disorders.
Materials and methods
Culture of hNSCs and hOPCs
hNSCs are derived from neural stem cell lines isolated from human fetal brain in the paediatric laboratory of the Sixth Medical Centre of Chinese PLA General Hospital. When the ‘neurospheres’ in hNSC medium [46] exhibited strong refraction and displayed protrusions at the edges, the hNSCs were ready for passage (typically after 14–15 days of culture). hNSCs were incubated for 10 d for induction of hOPCs. We dissociated neurospheres into single cells using Accutase (Cat.#A1110501, Gibco, USA) and resuspended them in hOPC medium [26]. The single cells were inoculated on a 6-well plate (Cat.#3516, Coring, USA) pre-coated with fibronectin human plasma (FN, 1.0 mg/mL, Cat.#F1056, Sigma-Aldrich, MO, USA) and laminin (LN, 1.06 mg/mL, Cat.#2399234, Gibco, USA). Half of the hOPCs medium (1.5 L) was replenished every 3 days. When hOPCs stretched and sticked to the bottom of well, with a bipolar protrusion, sometimes accompanied by a multipolar or a multipolar and graded protrusion, hOPCs were ready for passage. hOPCs were digested into a single cell by Accutase and incubated on pre-coated 6-well plates at a 200,000 cells/well density. hOPCs were maintained in hOPCs medium at 37 °C in a humidified atmosphere with 5% CO2.
Single-cell RNA sequencing (scRNA‑seq) of hOPCs
The adherent hOPCs were digested into single-cell suspension. The total number of cells in each suspension sample should exceed 1 × 106. The viability and cell counted were measured by Countess II automatic cell counter (Novogene, Beijing, China), requiring a final concentration of 700–1200 cells/µL. The counted cell suspensions were analysed for mRNA expression using Chromo4 Real-Time PCR detection system (Cat.#3591590 G, Bio-Rad, USA). After mixing the samples with enzymes and Gel beads containing Barcode sequence, the mixture binds to the oil surfactant to form the GEMs (Gel Bead-in-emulsion). We collect the GEMs streams into the reservoir, dissolve the Gel beads and release the Barcode, and label cells with identical sequences using the same barcode. The oil droplets were purified to construct reverse transcription and standard sequencing libraries using reagents from the Chromium Single Cell 3’v2 kit (10×genomics, USA). Illumina Hi seq 2000 platform (HiSeq2000, Illumina, USA) was used to generate 50,000 to 100,000 reads per cell. After Illumina Hi Seq high-throughput sequencing, raw image files were converted into sequence read segments using CASAVA base-calling and stored in FASTQ format. Violin plots showed the distributions of mcount_RNA, nFeature_RNA, and percent_mito across samples before and after filtration (Fig. S1). Original data quality filtration employed the following criteria: nFeature_RNA < 200 or nFeature_RNA > 6000; cells with a mitochondrial content greater than 30% were also removed (%mito > 30%). Secondary cell and feature filtration was performed in Seurat v4.4.0 package[47] (https://github.com/satijalab/seurat), retaining genes detected in at least three cells expressing a minimum of 200 genes; potential doublets—two or more cells captured within a single droplet—were removed using the Doublet Finder R package v2.0.2 (https://github.com/chris-mcginnis-ucsf/DoubletFinder). Downstream analyses, encompassing reanalysis with Cell Ranger and further processing in Seurat, utilised the filtered gene expression matrix. Subsequently, the cell count decreased from the original 400,017 to 28,801 (the number of cells removed at each step is provided in Fig. S1). Cell Ranger v6.0.2 (https://support.10xgenomics.com/single-cell-gene expression/software/pipelines/latest/installation) performed barcode counting on FASTQ files. Seurat v4.4.0 package[47] (https://github.com/satijalab/seurat) was used for data normalisation, dimension reduction, clustering, analysis of difference expressions. Based on the gene expression matrix, the edgeR v3.34.1 package[48] (https://github.com/Bioconductor-mirror/edgeR) was used for differential expression analysis among samples to obtain region-specific marker genes. The parameter of RunUMAP/RunTSNE was dim 20. For PCA, the first 30 principal components were used for subsequent analysis. The resolution was 0.6. We used Loupe cell browser (https://www.10xgenomics.com/products/loupe-browser/) software based on the expression of oligodendrocyte lineage specific genes on Maker Cell (http://xteam.xbio.top/CellMarker/) to integrate 28 801 cells from three samples of hOPCs-1/2/3 (from three different biological replicates) to obtain differential gene which were finally displayed in t-SNE and predicted three sub-clusters of PRE-OPCs, OPCs, and PRE-OLs by combining Maker Cell (http://xteam.xbio.top/CellMarker/).
Bulk RNA sequencing of hNSCs and hOPCs
hNSCs after being passaged to 10–12 generations (P10-P12) and hOPCs (P3-P5) were cleaved by Trizol (Cat.# NP80, 15596026, Thermo Fisher Scientific, USA) for Bulk RNA-seq at the Beijing Novo-gene Bioinformatics Technology Co. (Beijing, China). The Bioanalyzer 2100 system (Agilent Technologies, Santa Clara) and RNA Nano 6000 assay kit were used to assess the RNA integrity of each sample. RNA libraries were obtained by purifying the RNA products using a Nanoophotometer® (Cat.#NP80, Implen, Germany). The qualified libraries were sequenced on an Illumina NovaSeq 6000 platform. Image data were transformed into sequence data (reads) through recognition by CASAVA. High-quality data (clean reads) were obtained for downstream analysis after sequencing, followed by filtering to control the false discovery rate (FDR), removal of low-quality reads, and assessment of GC content distribution. After correcting for depth and gene length, the fragments per kilobase of transcript per million (FPKM) values were calculated based on each gene’s length and the number of reads.
MACS® cell sorting of hOPCs
The input samples (n = 3, from three different biological replicates) contained 1 × 107 hOPCs in 60 μL buffer (0.5% BSA, Cat #BP1600-100, Sigma-Aldrich, St. Louis, MO, USA), FcR Blocking Reagent (Fc, Cat. #130-059-901, Miltenyi Biotec, Germany) were blocked for 10 min at 2–8 °C. A volume of 20 μL of anti-PDGFR-α MicroBeads (Cat.#130-101-547, Miltenyi Biotec, Germany) was added to each 1 × 10⁷ cells and incubated at 2–8 °C for 15 minutes. Before entering the MiniMACS™ separator (Cat.#130-042-102, Miltenyi Biotec, Germany), hOPCs were filtered using a 30 μM nylon mesh (MACS Smart Strainers, Cat.#130-098-458, Miltenyi Biotec, Germany) to remove cell clumps that may block the column. hOPCs labelled with anti-PDGFR-α antibody-conjugated MicroBeads were subsequently isolated using a MACS® LS column (Cat. # 130-042-401, Miltenyi Biotec, Germany). By applying a magnetic field to the matrix, PDGFR-α+ hOPCs bound to magnetic beads were retained in the LS column and prevented from passing through, and unlabelled hOPCs can pass through the magnetic field. After removing the column from the magnetic field, the sorting column was washed with 1 mL buffer and quickly pushed down with a piston to avoid bubbles to collect PDGFR-α+ hOPCs.
Flow cytometry of hNSCs and hOPCs
To analyse protein expression, following digestion and centrifugation, hNSCs and hOPCs were resuspended in TF Perm/Wash buffer (Cat.#0114775, BD Biosciences, NJ, USA) centrifuged at 2000 × g for 5 min. Human TruStain Fc X™Fc Receptor Blocking Solution (5 μL/test, Cat. #422302, Biolegend) was added to the cell suspension and incubated for 10 minutes at 26 °C. hNSCs (n = 3, from three different biological replicates) were stained with PE Mouse IgG1, κ Isotype Control (5 μL/Test, Cat.# 554680, BD Biosciences), PE Nestin mouse-anti-human (5 μL/Test, Cat.#561230, BD Biosciences), Alexa Fluor® 488 Mouse IgG1, κ Isotype Control (5 μL/Test, Cat.#557721, BD Biosciences), Alexa Fluor®AF488 Vimentin mouse anti-human (8 μL/Test, Cat.#562338, BD Biosciences), PE Mouse IgG1 κ Isotype Control, PE Musashi mouse anti-human (5 μL/Test, Cat.#561468, BD Biosciences), Alexa Fluor® 647 Mouse IgG1, κ Isotype Control (5 μL/Test, Cat.#566011, BD Biosciences) and Alexa Fluor® 647 PAX6 mouse anti-human (5 μL/Test, Cat.#562249, BD Biosciences), respectively; hOPCs (n = 3) were stained with BV421 Mouse IgG2a, k Isotype Control (5 μL/Test, Cat.# 562439, BD Biosciences), BV421 PDGFR-α mouse-anti-human (5 μL/Test,Cat.#562799,BD Biosciences), Mouse IgG2b Kappa Light Chain Isotype Control (MG2b) [FITC] (5 μL/Test, Cat.#NBP1-43954, NOVUS Biologicals), FITC SOX10 mouse-anti-human (10 μL/Test, Cat.#NBP2-47709F, NOVUS Biologicals), PE Mouse IgG1, κ Isotype Control, PE A2B5 mouse anti human (10 μL/Test, Cat.#130-093-581, Miltenyi Biotec), Mouse IgG2b Kappa Light Chain Isotype Control (MG2b) [FITC] and FITC OLIG2 mouse-anti human (10 μL/Test, Cat.#AB9610, Millipore) antibodies, respectively incubated for 30 minutes at 26 °C. After staining, hNSCs and hOPCs were washed and centrifuged at 2000 × g for 5 min. hNSCs and hOPCs were filtered through a 30 μM nylon mesh prior to analysis, and were identified using the FlowSight® imaging flow cytometer (Amnis®, part of EMD Millipore). The data were analysed using Flow Jo software v10 (Flow Jo, Ashland, OR, USA).
Immunofluorescence staining
hNSCs (‘neurospheres’) exhibiting good refractive properties, appearing white or beige in colour with a distinct edge protrusion, and having an average diameter of approximately 200 μM, as well as hOPCs displaying bipolar or multipolar processes cultured for 6–7 days, were plated onto Cell Culture 8-Slide pre-coated with LN (1.06 mg/mL) and FN (1.0 mg/mL) for immunofluorescence staining. After 24–48 h, hNSCs and hOPCs were fixed with 4% tissue fixative. hNSCs were identified using mouse anti-Nestin (1:300, Cat.#ab6320, Abcam, UK), rabbit anti-SOX2 (1:150, Cat. #MAB1416, R&D Systems, USA), rabbit anti-OLIG2 (1:300, Cat.#AB9610, Millipore, USA) and rabbit anti-NG2 (1:100, Cat.#ab83178, Abcam, UK), respectively. We used rabbit anti-OLIG2 (1:300), rabbit anti-NG2 (1:100), mouse anti-A2B5 (1:50, Cat. #MAB1416, R&D Systems, USA), rabbit anti-PDGFRA (PDGFR-α, 1:800, Cat.#C2318, CST, USA), mouse anti-O4 (1:300, Cat.#MAB1326, R&D Systems, USA), and mouse anti-CNP (1:100, Cat.#ab6319, Abcam, UK), respectively to identify hOPCs. Secondary antibodies, Alexa 488-conjugated donkey anti-mouse (1:500, Cat.#ab150105, Abcam, UK) and Alexa 488-conjugated donkey anti-rabbit (1:500, Cat.#ab150153, Abcam, UK) were used to fluorescently label cells, respectively. The nuclei were stained with 300 μL DAPI (1:20, Cat.#28718-90-3, Sigma-Aldrich, USA) for 10-15 min. Images were captured via a digital confocal laser scanning microscope (20× and 40× objective lens, 26 °C, FV1000-D).
RT-qPCR
After digestion with Accutase, the reaction was terminated by adding DMEM/F-12 medium (Cat.#C11330500BT, Gibco, USA) at five times the volume of Accutase. Total RNA was extracted using an RNA prep Pure Cell/Bacteria Kit (Cat. #DP430, Tiangen, China). The quality and quantity of RNA solutions were assessed using a NanoPhotometer® ultra-microspectrophotometer. RNA samples with an OD260/OD280 ratio close to 2.0 were considered to be of high purity. Only RNA samples free from genomic DNA contamination and exhibiting clear ribosomal RNA bands on gel electrophoresis were selected for subsequent experiments. After adding 5×Prime Script RT Master Mix (Cat.#RR036A, TAKARA, Japan) into RNA, a reverse transcription reaction was performed using the C1000TMThermal Cycler (Bio-Rad, Hercules, CA, USA) PCR apparatus. Reverse transcription reaction: 37°C for 5 min; reverse transcriptase inactivation: 85°C for 5 s; cDNA was held at 4 °C for 3–5 min. A qPCR reaction solution containing TB Green® Fast qPCR Mix (Cat.#RR430A, TAKARA, Japan), PCR Forward and Reverse Primer, DNA template, and RNase Free dH2O were prepared on ice in a dark environment. The qPCR reaction mixture was then loaded into the Chromo4 Real-Time PCR Detection System (Cat.#3591590 G, Bio-Rad Laboratories, USA) for amplification and analysis. The housekeeping gene used for normalisation was β-actin, and relative gene expression levels were calculated using the 2 − ΔΔCt method. For primer details, please refer to Table S1.
GO and KEGG
The DESeq2 package[48] (v1.34.0) (https://github.com/al-mcintyre/DEQ) in R-Studio (the R Foundation, Austria) was used to perform differential expression (DE) analysis between hNSCs and hOPCs (3 biological replicates per group). The comparison of OPCs vs PRE-OPCs, OPCs vs PRE-OLs, and PRE-OPCs vs PRE-OLs were as well as conducted to analyse DE mRNAs. The threshold for DE mRNAs was defined as adjusted P-value ≤ 0.05 and |log2[fold change] ≥ 1. GO analysis was conducted using a comprehensive database of gene function annotations, encompassing Molecular Function (MF), Biological Process (BP), and Cellular Component (CC). The KEGG pathway is a molecular database (http://www.genome.jp/kegg/) that integrates chemical, genomic, and functional information. The ClusterProfiler R package [49] (v4.2.2) (https://github.com/YuLab-SMU/clusterProfiler) was used to perform GO and KEGG pathway enrichment analysis of DE mRNAs. GO and KEGG terms with adjusted P-value < 0.05 and log2[foldchange] ≥ 1 were considered significant. The interaction networks were performed to predict the interaction between target genes and their associated functions or pathways and were visualised via Cytoscape software v3.10.0 [50] (https://cytoscape.org/). For details of Software and algorithms, please refer to Table S2.
Cell proliferation assay
PDGFR-α+ hOPCs, PDGFR-α– hOPCs, and unsorted hOPCs were seeded into a 96-well-plate (Cat.#CLS3340, Corning, USA) at a density of 6000 cells per well and cultured under 5% CO2 and 37 °C for 24 h. On days 1, 3, 5, 7, 9, and 11, 10 μL Cell Counting Kit-8 solution (CCK8, Cat.#CK04, Dojindo, Japan) was added to each well, respectively. The group that received only the same volume of medium without any cells served as the control group. Six parallel wells were established in each experimental group and control group. After 3.5 h of incubation, the OD value was measured at 450 nm using Spectrophotometer (Thermo Fisher Scientific, Waltham, Massachusetts, USA). We measured the OD values on Days 1, 3, 5, 7, 9, and 11, with the experiment repeated three times at each time point. The cell proliferation rate at each time point was calculated using the following formula: proliferation rate = (ODn-OD0)/OD0 × 100%, where ODn represented the OD value at each time point, and OD0 represented the OD value of the medium after inoculation.
Cell transplantation in shiverer− / − mice
Shiverer–/– mice (The Jackson Laboratory, USA) were raised in a specific pathogen-free (SPF) laboratory located on the 4th floor of Building B, Incubation Building, No. 7, Fengxian Middle Road, Haidian District, Beijing, China. All animal procedures were approved by the Experimental Animal Welfare and Ethics Committee of Beijing Physical and Chemical Analysis and Testing Center (Ethics number: 210520-SWDWF-003). The mouse brain stereotaxic apparatus (Cat.#51730, Stoelting, USA) enabled precise targeting of the transplantation site (AP = –1.0 mm; L = 1.0 mm; V= –1.0 mm) in shiverer–/– mice. The impact of the stereoscope in combination with the 33 G microsyringe needle (Stoelting, USA) on brain tissue was minimal or even negligible. Newborn pups one day after birth were randomly assigned into the control (n = 6), PDGFR-α+ hOPCs (n = 11), PDGFR-α– hOPCs (n = 6, under the assumptions of a significance level (α) of 0.05 and a statistical power of 80%, approximately 3 animals per group would be required to achieve adequate statistical power), and unsorted hOPCs (n = 12) transplantation groups. PDGFR-α+ hOPCs, PDGFR-α- hOPCs, and unsorted hOPCs populations of 1 × 10⁵ cells in 1.5 µL was injected into the t corpus callosum (AP = –1.0 mm, L = 1.0 mm, V= –1.0 mm). Meanwhile, shiverer–/– mice without cell transplantation were randomly selected and assigned to the control group. After surgery, the pups were returned to their mothers and housed separately after weaning. Three months after transplantation, the shiverer–/– mice were euthanized via an overdose chloral hydrate (Cat. #82128-69-6, Sigma-Aldrich, MO, USA) and perfused with 4% paraformaldehyde (PFA, Cat. #15827, Sigma-Aldrich, USA). The brain tissue (n ≥ 3 in each group) was cut coronally along the sagittal suture, and one half of them were used for immunofluorescence staining with a rat anti-MBP antibody (Cat. #ab62631, Abcam, UK), and the stained sections were examined under a fluorescence microscope (IX-70, Olympus Corporation, Japan) followed by blinding rules. The others (n ≥ 3 in each group) were processed for transmission electron microscopy also followed by blinding rules (TEM, H7650-B, HITACHI, Japan). For microscopy, the fields were randomly selected form the corpus callosum—three fields per animal (at 20 000× magnification)—with 3–4 myelin sheaths analysed per field. Automated image analysis was performed with pre-defined parameters using ImageJ software, ensuring consistency across all experimental groups. The G-ratio and remyelination rates were calculated as follows: G-ratio value = axon diameter/total myelin fibre diameter; myelination rate = Q1/Q2, where Q1 represents the number of axons wrapped in the myelin sheath, and Q2 represents the total quantity of axons.
RNA-SCOPE
Cell preparation for RNA-scope
hOPCs were seeded onto 8-well culture slides (Cat.#072108, BIOLOGIX, China) pre-coated with FN and LN in hOPCs medium, cultured for 24–48 h, and then fixed with 4% PFA for 15 min prior to performing RNAscope® Multiplex Fluorescent v2 Assay combined with Immunofluorescence-Integrated Co-Detection. The medium and chambers were then removed, and the slides were immersed in 10% Neutral Buffered Formalin (NBF, Cat.#G2161, Solarbio, China) for 30 min. The slides were sequentially immersed in 50% ethanol for 5 min, 70% ethanol for 5 min, 100% ethanol for 5 min, and 100% ethanol for 10 min. Water was removed from the cells for preservation by storing them in 100% ethanol at −20 °C for 6 months. To rehydrate cells, the slides were immersed in 70% ethanol for 2 min, followed by 50% ethanol for 2 minutes, and finally in PBS for 10 min. The hOPCs were then processed for probe hybridisation and simultaneous protein detection.
RNA-scope® multiple fluorescence labelled mRNAs combined with immunofluorescence labelled proteins
The slides were washed in 1 × PBS-T (Cat. #P1031, Solarbio, China), and were then drawn 2–4 times around the edges of adherent hOPCs via the ImmEdge™ hydrophobic barrier pen (Cat. #310018, Advanced Cell Diagnostics, USA). Rabbit anti- PDGFR-α (1:120, Cat. #C5241, CST, USA), mouse mouse anti-CNP (1:120, Cat. # ab6319, Abcam, UK), and rabbit anti-SOX2 (1:150, Cat. #MAB1416, R&D Systems, USA) were diluted in Co-detection Antibody Diluent and applied to each slide to ensure complete coverage. The slides were then incubated overnight at 4 °C in a humidified oven overnight. Afterward, the slides were submerged in PBS-T solution, followed by fixation in 10% neutral buffered formalin (NBF) for 30 minutes. Then, 2–4 drops of RNAscope® Protease Ⅲ were applied to the tissue sections which were subsequently placed in the HybEZ™ oven at 40 °C for 30 minutes. The slides were washed in fresh distilled water. During the incubation process, the probes were pre-warmed for 10 min in a water bath at 40 °C, then allowed to cool to room temperature (RT, 26 °C). One volume of C2-CNPase probe, one volume of C3-SOX2 probes, and 50 volumes of C1 PDGFRA probe were pipetted into a tube. We applied 4–6 drops of Negative Control Probe, target RNA probes, and Positive Control Probe on slides followed by incubation for 2 h in a HybEZ™ oven at 40 °C. Subsequently, slides were incubated sequentially with Amp 1 for 30 min, Amp 2 for 30 min, Amp 3 for 15 min. This was followed by HRP-C1 (15 min) and diluted Opal™ 520 (30 min), then HRP blocker (15 min); next, HRP-C2 (15 min) and diluted Opal™ 570 (30 min), followed again by HRP blocker (15 min); and finally, HRP-C3 (15 min) and diluted Opal™ 620 (30 min), with HRP blocker (15 min) concluding the staining steps—all performed at 40 °C in the HybEZ™ oven. The slides were washed twice with fresh buffer for 2 minutes at room temperature (RT) at each interval. Following probe incubation, Goat anti-rabbit IgG H&L (HRP) (1:100, Cat. #ab205718, Abcam, UK) and Goat anti-mouse IgG H&L (HRP) (1:1000, Cat. #ab205719, Abcam, UK) were added and incubated for 30 minutes at RT (26 °C), diluted in Co-Detection Antibody Diluent. Subsequently, Opal™ 690 reagent was applied and incubated for 10 minutes at RT. The slides were then treated with Co-Detection Blocker and incubated for 30 minutes in a HybEZ™ Oven (Cat. #310013, Advanced Cell Diagnostics, Boston, MA, USA) at 40 °C. Due to the limitations of instrument channel constraints and spectral overlap of RNA-Scope technology, each sample could be labelled with multiple (three) mRNAs, but only one protein could be simultaneously detected. At the end, DAPI was added to each section for 30 s at RT. For detailed information on matierials and regeants of RNAscope, please refer to Table S3.
Data collection and analysis
RNA and proteins signals were acquired using Tissue Gnostics Tissue FAXS Spectra panoramic tissue scanning system (Tissue Gnostics, Beijing, China), which provided a standardised, fully automated, and tightly integrated hardware–software platform. Sequential multi-channel scanning was performed, progressing through channels such as DAPI, Opal ™ 520, Opal ™ 570, Opal ™ 620, and Opal ™ 690, with each scan involving filter switching and exposure adjustment. Before scanning the entire slide, representative tissue areas were pre-scanned to optimise the exposure time and LED light intensity for each channel. This step ensured that the signal strength for each channel was maintained within the linear range and avoided saturation. Once optimal settings were determined, these parameters were applied consistently across the entire slide during the scanning process. The system automatically adjusted focus at nine to sixteen points to generate a focusing plane map. During scanning, real-time Z-axis compensation was implemented using this map, ensuring consistently clear imaging throughout the tissue, even when surface irregularities were present. After scanning, the software automatically stitched all fields of view to create a comprehensive panoramic digital slide. This slide retained all channel information and could be exported as a multi-layer TIFF. Morphological analysis, cellular identification, and quantification were performed using Tissue Quest software (Tissue Gnostics, Beijing, China). Nuclear segmentation was achieved via DAPI staining, after which cytoplasmic regions were delineated. Quantitative measurements of fluorescence intensity and cell area were performed across the various fluorescence channels. For signal recognition, a threshold was established at 3 standard deviations (SDs) above the background fluorescence intensity. Cells with low DAPI signal intensity, overflowing cells, tissue debris, or overexposed signals were excluded from the analysis. The scanning sample cohort was organised with positive and negative control slides randomly distributed among the experimental samples. This arrangement facilitated minimising technical variability throughout the collection and analysis processes.
Replication, statistical tests, and reporting of sample sizes
Replication
Based on the expected effect size derived from Zhou et al.‘s published paper[11] in remyelination rate, under the assumptions of a significance level (α) of 0.05 and a statistical power of 80%, approximately 3 animals per group would be required to achieve adequate statistical power. For cell transplantation, the pups being numbered, and after shuffling the numbers, each numbered individual should have an equal chance of being included in the control (n = 6), PDGFR-α+ hOPCs (n = 11), PDGFR-α– hOPCs (n = 6), and unsoring hOPCs (n = 12) transplantation groups. The low positive rate and low activity rate [51, 52] after MACS of the PDGFR-α– hOPCs result in a smaller number of cells available for transplantation of 6 pups. As for the replication of scRNA-seq, a total of 28,801 cells (hOPCs-1/2/3, derived from three different biological replicates) were captured and the mRNA expression levels in each single cell were studied.
Statistical tests
The data in this study were analysed using the GraphPad Prism software v10.1 (GraphPad Software, San Diego, CA, USA, www.graphpad.com). All measurement data are expressed as \(\bar{{\rm{x}}}\) ± STDEV.P. SPSS software v29.0 (https://www.ibm.com/cn-zh/products/spss-statistics) to perform normality and variance homogeneity tests on the samples. Statistical tests were selected based on data characteristics. When both normality (assessed via Shapiro–Wilk test) and homogeneity of variances (assessed via Levene’s test) were satisfied, an independent samples t-test was used. If normality was met but variances were unequal, Welch’s t-test was applied. If neither normality or homogeneity of variances was violated, the non‑parametric Mann‑Whitney U test was employed. For three samples, Šidák’s multiple comparisons test was used following a significant ordinary one-way ANOVA, as the data met assumptions of normality and homogeneity of variances. Dunn’s test with p-value adjustment was used for post hoc pairwise comparisons following a significant Kruskal–Wallis H test, as the data were non-normally distributed. Differences were considered statistically when p < 0.05 and p < 0.01.
Data availability
Raw data from scRNA-seq have been deposited at GSA-human. Raw data reported in this paper will be shared by the lead contact upon request. Raw data of scRNA-seq at [https://ngdc.cncb.ac.cn/gsa-human/s/FdDsn5B5] and [https://ngdc.cncb.ac.cn/gsa-human/s/z66g0yYU].
References
Dimou L, Simon C, Kirchhoff F, Takebayashi H, Götz M. Progeny of Olig2-expressing progenitors in the gray and white matter of the adult mouse cerebral cortex. J Neurosci. 2008;28:10434–42.
Foerster S, Hill MFE, Franklin RJM. Diversity in the oligodendrocyte lineage: Plasticity or heterogeneity? Glia. 2019;67:1797–805.
Sim FJ, McClain CR, Schanz SJ, Protack TL, Windrem MS, Goldman SA. CD140a identifies a population of highly myelinogenic, migration-competent and efficiently engrafting human oligodendrocyte progenitor cells. Nat Biotechnol. 2011;29:934–41.
Yamanaka S. Pluripotent Stem Cell-Based Cell Therapy-Promise and Challenges. Cell Stem Cell. 2020;27:523–31.
Douvaras P, Wang J, Zimmer M, Hanchuk S, O'Bara MA, Sadiq S, et al. Efficient generation of myelinating oligodendrocytes from primary progressive multiple sclerosis patients by induced pluripotent stem cells. Stem cell reports 2014;3:250–9.
Kang SM, Cho MS, Seo H, Yoon CJ, Oh SK, Choi YM, et al. Efficient induction of oligodendrocytes from human embryonic stem cells. Stem Cells. 2007;25:419–24.
Hu BY, Du ZW, Zhang SC. Differentiation of human oligodendrocytes from pluripotent stem cells. Nat Protoc. 2009;4:1614–22.
Dietz KC, Polanco JJ, Pol SU, Sim FJ. Targeting human oligodendrocyte progenitors for myelin repair. Exp Neurol. 2016;283:489–500.
Azari H. Isolation and enrichment of defined neural cell populations from heterogeneous neural stem cell progeny. Methods Mol Biol. 2022;2389:111–23.
Azari H. Isolation and enrichment of defined neural cell populations from heterogeneous neural stem cell progeny. Methods Mol Biol. 2013;1059:95–106.
Zhou H, He Y, Wang Z, Wang Q, Hu C, Wang X, et al. Identifying the functions of two biomarkers in human oligodendrocyte progenitor cell development. J Transl Med. 2021;19:188.
Jäkel S, Agirre E, Mendanha Falcão A, van Bruggen D, Lee KW, Knuesel I, et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature. 2019;566:543–7.
Chamling X, Kallman A, Fang W, Berlinicke CA, Mertz JL, Devkota P, et al. Single-cell transcriptomic reveals molecular diversity and developmental heterogeneity of human stem cell-derived oligodendrocyte lineage cells. Nat Commun. 2021;12:652.
Liu X, Dong C, Liu K, Chen H, Liu B, Dong X, et al. mTOR pathway repressing expression of FoxO3 is a potential mechanism involved in neonatal white matter dysplasia. Hum Mol Genet. 2022;31:2508–20.
Ge C, Liu D, Sun Y. The promotive effect of activation of the Akt/mTOR/p70S6K signaling pathway in oligodendrocytes on nerve myelin regeneration in rats with spinal cord injury. Br J Neurosurg. 2024;38:284–92.
Ge C, Li C. Targeting the AKT/mTOR/p70S6K pathway for oligodendrocyte differentiation and myelin regeneration in neurological disorders. Curr Neurovasc Res. 2023;20:453–63.
Saito Y, Chikenji T, Ozasa Y, Fujimiya M, Yamashita T, Gingery A, et al. PDGFR signaling mediates hyperproliferation and fibrotic responses of subsynovial connective tissue cells in idiopathic carpal tunnel syndrome. Sci Rep. 2017;7:16192.
McNamara NB, Munro DAD, Bestard-Cuche N, Uyeda A, Bogie JFJ, Hoffmann A, et al. Microglia regulate central nervous system myelin growth and integrity. Nature. 2023;613:120–9.
Snaidero N, Velte C, Myllykoski M, Raasakka A, Ignatev A, Werner HB, et al. Antagonistic functions of MBP and CNP establish cytosolic channels in CNS myelin. Cell Rep. 2017;18:314–23.
Nave KA, Werner HB. Myelination of the nervous system: mechanisms and functions. Annu Rev Cell Dev Biol. 2014;30:503–33.
Kim HG, Moon WJ, Han J, Choi JW. Quantification of myelin in children using multiparametric quantitative MRI: a pilot study. Neuroradiology 8. 2017;59:1043–51.
van Bruggen D, Agirre E, Castelo-Branco G. Single-cell transcriptomic analysis of oligodendrocyte lineage cells. Curr Opin Neurobiol. 2017;47:168–75.
Bechler ME, Byrne L, Ffrench-Constant C. CNS myelin sheath lengths are an intrinsic property of oligodendrocytes. Curr Biol. 2015;25:2411–6.
Marques S, Zeisel A, Codeluppi S, van Bruggen D, Mendanha Falcão A, Xiao L, et al. Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science. 2016;352:1326–9.
Marques S, van Bruggen D, Vanichkina DP, Floriddia EM, Munguba H, Väremo L, et al. Transcriptional convergence of oligodendrocyte lineage progenitors during development. Dev Cell. 2018;46:504–.e507.
Ye D, Wang Q, Yang Y, Chen B, Zhang F, Wang Z, et al. Identifying Genes that Affect Differentiation of Human Neural Stem Cells and Myelination of Mature Oligodendrocytes. Cell Mol Neurobiol 2022 Cell Mol Neurobiol. 2023;43:2337–58.
Lu Y, Yang Y, Wang Z, Wang C, Du Q, Wang Q, et al. Isolation and culture of human oligodendrocyte precursor cells from neurospheres. Brain Res Bull. 2015;118:17–24.
Svensson V, Natarajan KN, Ly LH, Miragaia RJ, Labalette C, Macaulay IC, et al. Power analysis of single-cell RNA-sequencing experiments. Nat Methods. 2017;14:381–7.
Ding J, Adiconis X, Simmons SK, Kowalczyk MS, Hession CC, Marjanovic ND, et al. Systematic comparison of single-cell and single-nucleus RNA-sequencing methods. Nat Biotechnol. 2020;38:737–46.
Wang F, Flanagan J, Su N, Wang LC, Bui S, Nielson A, et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J Mol Diagnostics. 2012;14:22–29.
Ruff CA, Ye H, Legasto JM, Stribbell NA, Wang J, Zhang L, et al. Effects of adult neural precursor-derived myelination on axonal function in the perinatal congenitally dysmyelinated brain: optimizing time of intervention, developing accurate prediction models, and enhancing performance. J Neurosci. 2013;33:11899–915.
Merino D, Weber TS, Serrano A, Vaillant F, Liu K, Pal B, et al. Barcoding reveals complex clonal behavior in patient-derived xenografts of metastatic triple negative breast cancer. Nat Commun. 2019;10:766.
Rodriguez-Fraticelli AE, Wolock SL, Weinreb CS, Panero R, Patel SH, Jankovic M, et al. Clonal analysis of lineage fate in native haematopoiesis. Nature. 2018;553:212–6.
Haradhvala NJ, Leick MB, Maurer K, Gohil SH, Larson RC, Yao N, et al. Distinct cellular dynamics associated with response to CAR-T therapy for refractory B cell lymphoma. Nat Med. 2022;28:1848–59.
Zheng C, Wang J, Zhang J, Hou S, Zheng Y, Wang Q. Myelin and lymphocyte protein 2 regulates cell proliferation and metastasis through the Notch pathway in prostate adenocarcinoma. Transl Androl Urol. 2021;10:2067–77.
Tiane A, Schepers M, Riemens R, Rombaut B, Vandormael P, Somers V, et al. DNA methylation regulates the expression of the negative transcriptional regulators ID2 and ID4 during OPC differentiation. Cell Mol Life Sci. 2021;78:6631–44.
Kahm YJ, Kim RK, Jung U, Kim IG. Epithelial membrane protein 3 regulates lung cancer stem cells via the TGFbeta signaling pathway. Int J Oncol. 2021;59:80.
Liu X, Dong C, Liu K, Chen H, Liu B, Dong X, et al. mTOR pathway repressing expression of FoxO3 is a potential mechanism involved in neonatal white matter dysplasia. Hum Mol Genet. 2022;31:2508–20.
Shao Q, Zhao M, Pei W, Pu Y, Liu M, Liu W, et al. Pinocembrin promotes OPC differentiation and remyelination via the mTOR signaling pathway. Neurosci Bull. 2021;37:1314–24.
Cervenka J, Tyleckova J, Kupcova SH, Vodickova KK, Poliakh I, Valekova I, et al. Proteomic characterization of human neural stem cells and their secretome during in vitro differentiation. Front Cell Neurosci. 2020;14:612560.
Fasciani I, Pluta P, Gonzalez-Nieto D, Martinez-Montero P, Molano J, Paino CL, et al. Directional coupling of oligodendrocyte connexin-47 and astrocyte connexin-43 gap junctions. Glia. 2018;66:2340–52.
Fang C, Sun J, Wei L, Gao F, Qian J. Oscillating field stimulation promotes recovery from spinal cord injury in rats by regulating the differentiation of endogenous neural stem cells. Exp Ther Med. 2021;22:979.
Zheng J, Sun X, Ma C, Li BM, Luo F. Voluntary wheel running promotes myelination in the motor cortex through Wnt signaling in mice. Mol Brain. 2019;12:85.
Itoh F, Asao H, Sugamura K, Heldin CH, ten Dijke P, Itoh S. Promoting bone morphogenetic protein signaling through negative regulation of inhibitory Smads. EMBO J. 2001;20:4132–42.
Zi Z. Molecular engineering of the TGF-β Signaling Pathway. J Mol Biol. 2019;431:2644–54.
Wang C, Luan Z, Yang Y, Wang Z, Wang Q, Lu Y, et al. High purity of human oligodendrocyte progenitor cells obtained from neural stem cells: suitable for clinical application. J Neurosci Methods. 2015;240:61–66.
Hao Y, Hao S, Andersen-Nissen E, Mauck WM 3rd, Zheng S, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184:3573–.e3529.
Liu S, Wang Z, Zhu R, Wang F, Cheng Y, Liu Y. Three differential expression analysis methods for RNA sequencing: limma, EdgeR, DESeq2. J Vis Exp. 2021.
Yu G. Gene ontology semantic similarity analysis using GOSemSim. Methods Mol Biol. 2020;2117:207–15.
Zahid KR, Su M, Khan ARR, Han S, Deming G, Raza U. Systems biology based meth-miRNA-mRNA regulatory network identifies metabolic imbalance and hyperactive cell cycle signaling involved in hepatocellular carcinoma onset and progression. Cancer cell Int. 2019;19:89.
Corti S, Locatelli F, Papadimitriou D, Donadoni C, Salani S, Del Bo R, et al. Identification of a primitive brain-derived neural stem cell population based on aldehyde dehydrogenase activity. Stem Cells. 2006;24:975–85.
Kirkcaldie MT, Collins JM. The axon as a physical structure in health and acute trauma. J Chem Neuroanat. 2016;76:9–18.
Acknowledgements
We would like to thank Editage (www.editage.cn) for English language editing.
Funding
This work was supported by the National Key Research and Development Program of China (2017YFA0104200). The recipient was Zuo Luan. All authors approved the Funding.
Author information
Authors and Affiliations
Contributions
YD, Z-HP, Y-YX, LZ, and WQ conceived of the presented idea and designed the experiments. YD, Z-HP, Q-SQ, Y-YX, WQ, W-ZY, ZF, W-XH, ZY, and L-JL performed the experiments. YD analyzed the experimental data and write the manuscript. Y-YX, LZ, and WQ provided technical and material support. All authors read and approved the final paper. All authors approved the author’ contributions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing interests. The funding body only provided the financial means to allow the authors to carry out the study, and played no role in the design of the study and collection, analysis, and interpretation of data and in writing the manuscript.
Ethics approval and consent to participate
This study was approved by the Ethics Committees of Beijing Center for Physical & Chemical Analysis (SWDW-01-03). All animal operations were approved by the Experimental Animal Welfare and Ethics Committee of Beijing Physical and Chemical Analysis and Testing Center (Ethics number: 210520-SWDWF-003).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ye, D., Zhou, H., Qu, S. et al. Multi-omics reveals heterogeneity and functional populations of oligodendrocyte progenitor cells induced by human neural stem cells. Cell Death Discov. 12, 112 (2026). https://doi.org/10.1038/s41420-026-02971-w
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41420-026-02971-w