Abstract
Liquid‒liquid phase separation (LLPS) is a phenomenon relevant in the multicomponent settings of many biological processes, including compartmentation, pathological conditions such as Alzheimer’s disease, and protein assembly. LLPS also plays a key role in spider silk fiber formation. Many spider silk fibers display properties such as elasticity in combination with high mechanical strength, which result in an outstanding toughness exceeding that of steel or Kevlar. A thorough understanding of the natural silk spinning process is thus vital for translation to artificial spinning techniques to achieve biomimetic fibers with properties superior to those of other fibrous materials. This focus review summarizes the milestones of research on spider silk assembly, starting from two initial theories, i.e., the liquid crystal theory and the micelle theory, followed by evidence for the importance of LLPS in this process. Ex vivo studies and experiments utilizing recombinant spider silk proteins have highlighted the importance of LLPS during spider silk assembly. Here, we provide a consolidated view of the previously separate theories as a concerted, transitional concept, and describe practical implications showcasing the importance of this unifying concept for technical silk spinning.
Similar content being viewed by others
Introduction
Liquid‒liquid phase separation (LLPS) is a physical process that often occurs in multicomponent mixtures. In LLPS, solutions spontaneously separate into two immiscible liquid phases, one with a low concentration and the other with a high concentration of one or more individual components. The driving forces for LLPS include temperature as well as pressure changes, pH, charge neutralization, the specific polarity of the constituting components, and other surface characteristics, e.g., hydropathicity [1]. The process is widely known to occur in polymer-diluent systems, with applications, for example, in waste and pollution control or in the food processing industry [1]. In biological and biomedical research, phase separation is known to be an important compartmentation principle in physiological processes, including RNA metabolism, ribosome biogenesis, DNA repair mechanisms, and signal transduction [2]. LLPS has also been reported to occur under pathological conditions, especially in neurodegenerative diseases such as Alzheimer’s disease [1, 3]. The LLPS of proteins has recently received much attention since LLPS is essential for the formation of biological adhesives, e.g., in sandcastle worm cement or during the formation of mussel attachment adhesives underwater, as well as in hard biomolecular composites such as the squid beak [4]. Moreover, it has been reported that LLPS of spider silk proteins is important for fiber assembly [5]. A common feature of proteins undergoing LLPS seems to be the presence of intrinsically disordered protein regions (IDPs), which favor phase separation to increase their concentration without structural changes [6]. Weak but highly abundant noncovalent intra- and intermolecular interactions are known to be the driving forces for LLPS in biological systems, leading the proteins to condense in a concentrated liquid phase as a transition state for further processing or reactions [6]. In this manner, specific preassembly processes guided by intra- and intermolecular interactions are permitted that would not occur without LLPS [7].
The natural spider silk spinning process is highly sophisticated and involves liquid‒liquid and liquid‒solid transitions of the underlying spider silk proteins ( = spidroins) in response to multiple biological, biochemical, and mechanical cues [8]. Thus, a meticulously coordinated process is highly important to prevent premature solidification of the spidroin solution in the spider’s body, potentially obstructing fiber formation and subsequent usage. However, research on spiders or silk collected from spiders through milking is hampered by the cannibalistic and territorial behavior of most spider species, and only low quantities of silk can be acquired at a time by forced silking. Thus, elucidation of the mechanism underlying silk spinning from an aqueous solution has been performed mainly with rehydrated or recombinantly produced spider silk proteins [9].
On the basis of the work conducted in the past 20 years on elucidating the assembly mechanism for major ampullate spidroins (MaSps), we herein provide a topical overview, highlighting the most recent advances.
Liquid crystal theory and micelle theory
Low-concentration solutions of certain amphipathic block copolymers form nanometer-sized, approximately spherical aggregates in a colloidal suspension [10]. However, at higher polymer concentrations, self-assembly processes lead to the formation of cubical or cylindrical nanocrystalline structures, so-called liquid crystals (LCs), that feature liquid-like behavior [7, 11]. Polymers showing such behavior have various technical applications, e.g., in LC displays or holographic systems [12]. In 1999, Vollrath and Knight reported that silk proteins in the spider’s major ampullate gland and spinning duct also present LC characteristics, with the long axes of the spidroins aligning parallel to one another [13, 14]. This finding is in accordance with findings in block copolymers. Strikingly, the concentration of spidroins in spider glands can reach 50 wt%, a spidroin concentration that cannot be easily established in vitro without gelation or solidification of the protein solution. Vollrath and Knight hypothesized that the reduction in viscosity through a nematic phase preorientation of spidroins permits their specific alignment during elongational flow throughout the spinning duct, which is essential for the development of the outstanding mechanical properties of spider silk fibers. According to their observations, the following mechanism was proposed for in vivo spider silk spinning: the LC spidroin solution is drawn through an S-shaped duct to form an elastic thread, whereby the inner core and the thin coat of the thread are formed in two different zones. The inner core is made up of short, narrow spidroin filaments, whereas the shell of the thread is composed of hexagonal columnar LCs. The epithelial layer of the duct allows increased exchange of water and sodium ions for kosmotropic potassium ions, as well as acidification, together initiating fiber assembly. Simultaneously, the decreasing width of the tapered spinning duct promotes specifically aligned states of the spidroin molecules. The spidroins are first oriented in parallel but perpendicular to the epithelium before they turn to become parallel to the epithelial walls. Next, the molecules form bilayered disks, an orientation called the cellular optical texture, which can further elongate and align to form mainly β-sheet-rich structures [15]. Finally, the silk thread is actively drawn from the exit spigot with the help of a clamp-like valve (see Fig. 1) [14].
Schematic depiction of the putative spider silk formation process in vivo and two theories explaining the experimental observations, namely, the liquid crystal theory [14] and the micelle theory [16]. Spidroins consisting of a long repetitive core region flanked by nonrepetitive N- and C-terminal domains are excreted into the spider’s gland tail, wherein the N-terminal domains are in a monomeric state, while the C-terminal domains adopt a dimeric structure. In the ampulla, according to the liquid crystal theory, spidroin molecules form a nematic liquid crystalline phase under storage conditions. During fiber formation along the spinning duct, the spidroin molecules experience ion exchange, a gradual pH drop, water removal, and elongational flow, resulting in a specific assembly process of the spidroins from a cellular optical texture to β-sheet-rich nanofibrils. In contrast, the micelle theory claims that the dimeric spidroin molecules assemble into micellar constructs due to hydrophilic/hydrophobic interactions under storage conditions in the ampulla. These micellar structures coalesce into larger condensates that are elongated and compressed as they pass through the spinning duct. In combination with the pH decrease, ion exchange, and water removal, molecular interactions lead to the formation of β-sheet-rich nanofibrils. In both theories, the last step involves the transfer of the liquid-like nanofibrils to a solid-state by drawing the silk thread from the exit spigot (i.e., applying shear forces)
Despite the groundbreaking insights into spider silk spinning processes, the proposed LC behavior of spidroins in spider glands cannot fully explain how the proteins remain soluble at high concentrations and cannot be replicated in vitro. Moreover, many processing steps during silk formation were not understood at the time of publication, further resulting in problems when trying to artificially replicate the process.
Interestingly, during their work with rehydrated silk fibroin from the silkworm Bombyx mori, Jin and Kaplan reported micelle-like structures formed due to the hydrophilic-hydrophobic-hydrophilic character of silk fibroins, increasing their water solubility and preventing premature solidification [16]. The authors proposed that the micellar constructs coalesce and assemble into larger globules upon increased protein concentration. Physical shear subsequently lead to elongation and alignment of these globules during the drawing process in the spinning duct, an essential condition for the formation of insoluble β-sheet-rich silk threads (see Fig. 1).
Elucidating the mechanism of spider silk formation
Ex vivo studies
The best-studied type of spider silk is the major ampullate (MA) silk of orb-weaving spiders, which is used as dragline silk and for the construction of web radii [17]. To date, in the European Garden Spider Araneus diadematus, two structurally similar MaSps have been identified, with slightly different amino acid compositions [18]. MaSps are composed of a repetitive core and can contain 3500 amino acids flanked by nonrepetitive (NR) amino (N)- and carboxyl (C)-terminal domains of ~130 and ~110 amino acids, respectively [19]. Both NR domains are highly conserved among species and have a five-helix structure. Under storage conditions, the C-terminal domains are present as parallel dimers, whereas the N-terminal domains remain in a monomeric state. These conformational characteristics and the potential ability of N-terminal domains to dimerize later in response to acidification are vital for spinning dope storage and correct silk fiber assembly [20]. In the core region, poly-A, poly-GA, GPGQQ, GPGGX, and GGX sequences can be found, wherein the poly-(G)A sequences contribute to mechanical stability via β-sheet formation, whereas the structural features of the other sequences allow flexibility and elasticity of the spider silk fiber through assembly into β-spirals, β-turns, and 31 helices (see Fig. 2A) [21].
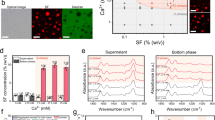
Structure, dynamics and assembly of native ex vivo L. hesperus major ampullate spidroins (MaSps). A Schematic organization of MaSps, the prevailing amino acid motifs of the repetitive core domain, and their corresponding secondary structure. n = 20–100. B 1H/15N HSQC NMR spectrum of isotopically enriched MaSps illustrates the disordered, unstructured nature of the repetitive core regions, as indicated by the poor chemical shift dispersion ( < 0.5 ppm) in the 1H dimension. Reprinted from [22] with permission from Elsevier. C NMR relaxation measurements show that the spidroins exhibit rapid subnanosecond backbone dynamics in the major ampullate gland. The NOE values are negative at 500 MHz (black) and moderately positive at 800 MHz (red), which is consistent with this interpretation. Reprinted from [22] with permission from Elsevier. D NMR diffusion measurements of spidroins within intact MA glands show restricted diffusion. The diffusion coefficient as a function of the interpulse delay (∆) shows exponential decay, indicating a diffusion restriction length scale of ~250–300 nm. This finding supports the silk protein micelle theory, which states that silk proteins are clustered in micelle-like preassemblies in the gland environment. Adapted from [23]. E 1H/15N HSQC NMR spectrum of isotopically enriched MaSp incubated in 4 M urea to disrupt micelles showing chemical shift perturbations for residues that are associated with micelle stabilization. The largest perturbations are observed for the amino acid motifs GAA and AAG (circled), while poly-A shows small perturbations, and the changes in GGQ are negligible. Adapted with permission from [24]. Copyright 2021 American Chemical Society. F CryoEM tomography imaging reveals that silk protein micelles are best described as complex micellar nanoparticles composed of assembled subdomains with an open core occupied by solvent. Adapted from [23]. G Significant chemical shift perturbations in 13C occur for alanine Cα and Cβ sites in the GAA and AAG motifs, which terminate the poly-A runs that bridge the GGX regions. This is consistent with the disruption of hydrophobic interactions as micelles disassemble in the denaturant. Adapted with permission from [24]. Copyright 2021 American Chemical Society
Ex vivo biophysical studies of MA glands excised directly from black widow spiders (Latrodectus hesperus) have provided significant insight into the structural and dynamic state of spidroins in a native-like environment prior to fiber spinning. Importantly, because these experiments were conducted on whole glands immediately following dissection, the state of the protein was as close to the true storage state as possible to an in vivo experiment. Solution nuclear magnetic resonance spectroscopy (NMR) of intact isotope-enriched black widow MA glands has shown that spidroins behave like classic IDPs [22]. Conformation-dependent isotropic chemical shifts were consistent with a random coil structure (Fig. 2B), and NMR relaxation measurements (T1, T2, NOE) revealed rapid spidroin backbone dynamics on the subnanosecond timescale. The high degree of rapid backbone motion is readily assessed from the heteronuclear 1H-15N nuclear Overhauser effect (NOE) measured in two fields (800 MHz and 500 MHz), which illustrates the highly flexible nature of the spidroins stored in the gland prior to fiber spinning (Fig. 2C). Diffusion NMR and cryogenic electron microscopy (cryoEM) have been used to further explore the internal global dynamics and nanoscale organization of spidroins in the gland environment of black widow MaSp [23]. Measuring the diffusion behavior of the spidroins in the silk gland via NMR revealed that the silk proteins are predominantly confined within entangled volumes that are several hundred nanometers in diameter, suggesting the existence of spherical micelles (Fig. 2D), as postulated in the micelle theory [16]. 3D imaging of these nanostructures via cryo-EM tomography revealed that the micellar assemblies are morphologically complex, existing as hierarchical submicron particles composed of networks of ~50 nm subdomains with an open core occupied by solvent (Fig. 2F). When the native dope is physically sheared with a micropipette, mimicking the extrusion process occurring within the spinning duct, these subdomains within the hierarchical micelles transform, becoming elongated fibrils that remain assembled as interwoven networks in the parent micelle. These findings shed new light on the hierarchical organization of spidroins in the gland, supporting the micelle theory. However, significant questions remain concerning how other physiochemical conditions relevant to silk spinning, such as ions or pH, affect the spidroin superstructure during the process of fiber formation.
In a recent NMR and dynamic light scattering (DLS) study, native MaSp micelles extracted from L. hesperus silk glands were investigated following urea denaturation to monitor the molecular-level interactions responsible for spidroin micelle stabilization [24]. Highlights of the investigation included determination of the critical spidroin micelle concentration ( ~ 5 wt%) by diffusion NMR, monitoring of micelle size during urea disruption with DLS, and the use of 3D solution NMR to investigate isotope-enriched samples to identify important motifs that are perturbed during micelle disruption. The results from the 3D solution NMR were particularly intriguing, indicating that the GAA and AAG motifs flanking the poly-A runs were the most perturbed by urea, suggesting that these regions may be important for spidroin micelle stabilization (Fig. 2E). Although these results pointed to a disruption of hydrophobic interactions for alanine Cα and Cβ sites in these poly-A flanking motifs (Fig. 2G), the role of the scarcer amino acids (Y, S, R) in the GGX regions remains to be determined and should be the focus of future studies.
Engineered silk proteins
To address the challenges associated with working with spiders, the establishment of recombinant spidroin mimetics was crucial for further investigations of the fiber formation process and its translation to artificial spinning setups to obtain fibers for various applications. In 2004, we established an approach to recombinantly produce two bioengineered MaSps, eADF3 and eADF4, named after their natural blueprints ADF3 and ADF4 (see Fig. 3) [18, 21]. This approach not only allows the large-scale fabrication of spidroin mimics but also enables the customization of engineered proteins tailored for their intended application [25].
Schematic of the genetic engineering of spider silk proteins, exemplified by two major ampullate spidroin derivatives, namely, ADF3 and ADF4 (Araneus diadematus fibroin), from the European Garden Spider Araneus diadematus. Based on the general structure and amino acid consensus sequences of the repetitive core modules of ADF3 and ADF4, engineered spider silk protein mimics eADF3 and eADF4 are derived by a reverse translation approach that combines PCR-amplified authentic gene fragments and engineered synthetic DNA modules into gene cassettes. After the cloning of multimerized constructs and transformation into a bacterial host, such as E. coli, the engineered silk proteins are recombinantly produced, extracted, purified, and lyophilized for storage. Adapted from [46] with permission from Springer Nature
LLPS of eADF3, which is more hydrophilic than eADF4 is, results in a dense, highly concentrated and a less dense, less concentrated phase [26]. This LLPS process was shown to be highly dependent on the potassium phosphate (K-Pi) concentration and needed a slightly acidic or neutral pH. In contrast, high NaCl concentrations and pH > 8.5 inhibited LLPS. By adding more K-Pi, β-sheet-rich fibers could be drawn from the high-density phase but not the low-density phase. Given that the NaCl concentration is also high in the natural spider gland, this work provides evidence for one of the key factors that prevents premature spidroin solidification. Additionally, the importance of LLPS for colloidal assembly, combined with control over ionic strength and pH to form stable silk threads, was demonstrated, laying the foundation for artificial silk spinning. The role of solvent conditions was further analyzed in another study using a microfluidic device [27]. It was found that fiber formation occurred only when an elongational flow, a pH drop from 8 to 6, and an increase in K-Pi concentration to 500 mM were applied simultaneously. We hypothesized that a narrowing of solvent streamlines below the diameter of eADF3 spherical colloidal assemblies forces them into contact. The so-called dangling ends—hydrophilic side chains on the outside of the assemblies—can then interact to form larger globules with helical conformations, resulting in shear thickening of the spinning dope. At this point, the process is governed mainly by configurational changes in the C-terminal dimerization domain, leading to the exposure of hydrophobic sites. The addition of shear forces finally leads to the formation of water-insoluble β-sheet-rich fibers due to increased H-bond interactions in poly-A stretches. These findings are comparable to those of Jin and Kaplan for B. mori silk [16]. Interestingly, the second engineered protein, eADF4, did not form silk threads in this microfluidic device. Importantly, the eADF4 variant used did not contain any C-terminal domain, in contrast to the eADF3 variant used. However, the importance of the presence of such a C-terminal domain for assembly was not known at that time.
To gain a deeper understanding of spider silk fiber assembly at the molecular level and to mimic the natural spinning process in an artificial setup, various analysis techniques have been utilized [28], building on the pioneering work of Exler et al. [26]. The NR domains of MaSp are highly conserved between different spider species and are essential for silk fiber formation [29]. Circular dichroism (CD) spectroscopy demonstrated that the NR C-terminal domain exhibited a mostly α-helical conformation crucial for the formation of micellar assemblies, followed by LLPS, which yielded larger liquid-like condensates [18]. Rheological studies revealed that C-terminal domain-containing molecules responded with a significant change in viscosity to shear stress, whereas molecules with no NR domain showed no changes. The lack of a C-terminal domain suppressed correct fiber formation, and only aggregates with randomly aligned β-sheets were formed [30]. Thus, the C-terminal domain might play a vital role in protein storage as well as fiber formation, permitting the formation of a specific supramolecular assembly without premature solidification.
The role of the NR N-terminal domain in the fiber assembly process was first assessed by A. Rising and coworkers [19, 31]. In contrast to the C-terminal domain, which tends to undergo self-assembly and LLPS, the N-terminal domain is responsible for the pH dependency of fiber formation. Hereby, amide hydrogen/deuterium exchange in combination with mass spectrometry (MS), nondenaturing electrospray ionization MS, and analytical ultracentrifugation revealed that the N-terminal domain, which is monomeric under storage conditions, undergoes a structural change upon acidification below pH 6.4, forming antiparallel dimers through charge-mediated interactions of monomeric subunits [19]. Far-UV CD spectroscopy revealed that the five-helix secondary structure of the N-terminal domain is not influenced by changes in the salt concentration or pH. However, the significant changes in the tertiary structure of the N-terminal domain to antiparallel dimers became apparent via near-UV CD spectroscopy upon changes in salinity at pH 6 and were accompanied by greater structural stability than that at pH 7.2 [32]. High salt concentrations at pH 7.2 also strongly increased structural stability, with 1H-15N heteronuclear single quantum coherence spectroscopy (HSQC) revealing a monomeric structure [32]. Therefore, the N-terminal domain seems to have little importance during protein storage but is significantly involved in fiber assembly, as it promotes antiparallel dimerization of the formed supramolecular structures.
These studies, which used recombinant spidroins and various analysis techniques, demonstrated that highly concentrated spidroins adopt a micelle-like architecture that is induced mainly by interactions of their C-terminal domain. Folded, hydrophilic sequences of the otherwise mostly hydrophobic C-terminus are located on the outside of these micelles to increase their solubility. The longer hydrophobic sequences can be seen as spacers that provide mobility of the chains and are hypothesized to play an important role during LLPS of the micellar constructs into larger liquid-like condensates [7, 20]. In spiders, preoriented spidroins are directed through a tapered duct where the NaCl concentration, as well as the water content, continuously decreases in exchange for kosmotropic K-Pi during fiber assembly (see Fig. 4A) [7]. It is hypothesized that in the presence of mechanical shear forces, a configurational change in the C-terminal domain is promoted, leading to exposure of hydrophobic patches and, consequently, to the formation of elongated and aligned protein chains that form H-bond-mediated β-sheet-rich nanofibrillar assemblies (see Fig. 4B). A simultaneously occurring pH decrease in the duct additionally induces antiparallel dimerization of the N-terminal domains, connecting the spidroins to a fibrillar network necessary for correct fiber assembly (see Fig. 4C). The process of protein chain alignment and elongation to form nanofibrils can thus be seen as a transition from an LLPS-dominated state to a liquid‒liquid‒crystalline phase separation (LLCPS)-dominated state, whereby relevant intra- and intermolecular interactions can occur owing to the densely packed spidroins [7, 32, 33]. The final step of liquid‒solid phase separation (LSP) involves pulling of the fiber from the exit spigot.
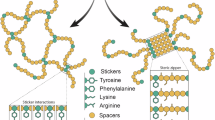
Overview of the role of the N- and C-terminal domains of major ampullate spidroins during storage and fiber formation. A General scheme of the spidroin assembly process from partially dimeric to micellar assemblies and larger droplet-like condensates, which are further compressed and elongated on their way through the spinning duct. Finally, a solid silk fiber is drawn from the exit spigot. B Schematic representation of the molecular interactions during the spider silk assembly process. After secretion by epithelial cells in the tail of the upper part of the gland, the C-terminal domains (green) of the spidroins are present as parallel dimers, often further stabilized by disulfide bridges at their interfaces. Owing to hydrophobic–hydrophilic interactions, the spidroins self-assemble into micelle-like structures, with the terminal domains spanning the sphere surface, whereas the more hydrophobic repetitive core domains are oriented toward the inside of the micelle-like constructs. At this stage, the N-terminal domains (blue) remain monomeric. In the distal part of the ampulla, the micellar preassemblies undergo LLPS to form a protein-rich phase with micrometer-sized liquid-like condensates, most likely triggered by a combination of pH changes, high protein concentrations, altered viscosity, and “sticker”-like interactions of amino acid residues such as tyrosine and arginine. Strikingly, the repetitive core domain remains intrinsically unstructured under these conditions. In the spinning duct, increasing acidification to pH < 6.5 leads to antiparallel dimerization of the N-terminal domains and, thus, the formation of a crosslinked spidroin network. Moreover, the repetitive core region now becomes more structured by forming helical secondary structures. Water removal and the exchange of chaotropic ions for kosmotropic phosphate in the spinning duct induce a configurational change in the C-terminal domains, allowing for a more packed alignment of the core regions and enhanced intra- and intermolecular interactions of poly-A motifs to form β-sheet-rich crystalline regions. Owing to the elongational flow in the tapered spinning duct, shear stress stretches the preassembled silk proteins to yield the final, optimally packed and oriented nanofibrils that are pulled as solid fibers from the spigot at the end of the spinning duct. C Rendered protein structures of monomeric N- and dimeric C- and N-terminal domains (adapted structures from the RCSB PDB data bank [47,48,49]). Both the N- and C-terminal domains have a five-helix bundle structure
Several research groups have attempted to understand the aqueous process of spinning to obtain fibers from soluble proteins via the use of recombinant proteins. However, no studies have provided a complete view of the assembly process across hierarchical length scales from soluble proteins in different dope preparations through the final spun fibrous state. A study by Malay et al. used recombinant MaSp2 to study spidroin assembly [34], supporting and highlighting the importance of LLPS. Another study demonstrated with simple engineered three-block molecules that coacervate formation is highly dependent on protein architecture [35]. Surprisingly, different methods of coacervate formation are possible even with the same protein, whereby the realized process depends on the environmental conditions. Thus, either liquid-like coacervates (LLCs) or solid-like coacervates can be formed depending on the phosphate ion concentration, but only LLCs permit the drawing of fibers [35]. Similarly, Lemetti et al. demonstrated a correlation between protein size and the phase separation mechanism, whereby all the studied proteins only assembled into coacervates above a certain concentration threshold. Below this threshold, smaller aggregates were formed [36]. Most importantly, the concentration threshold for coacervate formation was found to decrease with increasing protein sequence length, providing valuable insight for the bottom-up engineering of proteinaceous materials for specific applications. Recently, Landreh et al. published a comprehensive review focusing on the relevance of different amino acids for silk fiber formation and suggested that the fiber assembly process can be seen as a sequential mixture of LLPS and LSP, termed LLCPS [7]. In another recent study, a combination of biophysical methods, including solution and solid-state (SS) NMR, DLS, and light microscopy, was used to study the differences in eADF3 ((AQ)12NR3) spider silk dopes reconstituted under both biomimetic (preassembled) and classic conditions, as well as fibers produced after aqueous wet-spinning from both dopes [37]. This study provides a more complete picture of how silk spinning dope preparation differences dictate the downstream recombinant spider silk fiber structure and mechanics.
Light microscopy and DLS revealed distinct differences between classic spinning dopes (CSDs) and biomimetic spinning dopes (BSDs) at the micro- and nanoscales for the eADF3 system. In the BSD samples, liquid droplets between 5 and 25 µm in diameter were clearly detected via light microscopy, whereas micron-scale droplets were completely absent in the CSD samples (Fig. 5A). The main difference between the two dope preparations is that the phosphate ions in the BSD initiate LLPS, whereas the CSD contains Tris/HCl buffer and does not exhibit microscale LLPS. On the nanoscale, DLS revealed that the two dopes behaved differently during the concentration step, where the monomer/dimer equilibrium coalesces to form preassemblies in one step in CSD, whereas in BSD, this occurs gradually over time, with protein assemblies reaching the microscale (Fig. 5B).
Biophysical characterization of recombinant (AQ)12NR3 spider silk proteins in the dope phase and as spun fibers. A Light microscopy images of biomimetic spinning dopes (BSDs) (top, I-III) and classic spinning dopes (CSDs) (bottom, IV-VI), both at a concentration of 100 mg/ml. Regions from II, III, V and VI are enlarged to visualize the differences between the systems, where the BSD contains large droplets indicative of LLPS. B DLS of BSD (left) and CSD (right). Samples were taken after 0, 2, 4, and 6 h to follow the assembly process of the spinning solutions. After this, no further changes were observed, and the dope production was completed. During assembly, the concentrations gradually increased. For BSD, the concentrations at 0, 2, 4, and 6 hours were 27, 63, 96, and 177 mg/ml, respectively. For CSD, the concentrations at 0, 2, 4, and 6 hours were 25, 41, 106, and 197 mg/ml, respectively. These results are consistent with light microscopy observations (A), which revealed that dopes prepared with phosphate buffer create droplets significantly larger in size than those created with CSD. C 1H-15N HSQC NMR spectrum of fully labeled 13C/15N-(AQ)12NR3 BSD collected at 850 MHz. The bolded amino acids indicate resonance identities with di- and tripeptide repeats on the basis of 3D experiments [37]. Conformation-dependent chemical shifts illustrate that the repetitive core of the silk protein maintains intrinsic disorder in the condensed phase. D 1H-13C CP-MAS SS-NMR spectrum of sparsely labeled 13C/15N-(AQ)12NR3 lyophilized protein (black) and fibers spun from BSD (blue) and CSD (red) dopes. The full spectrum (top) and magnified image of the low ppm region (bottom) are shown. Fibers spun from both dopes have identical β-sheet formation rates of 77% for poly-A domains according to spectral deconvolutions [37]. E 2D 1H-13C HETCOR SSNMR spectrum collected with frequency-switched Lee‒Goldburg (FSLG) homonuclear proton decoupling for CSD (blue) and BSD (red) fibers. Slices were extracted at the tyrosine side chain resonance to illustrate the differences between the 1H chemical shifts of one of the tyrosine aromatic rings for the two fibers. A downfield 1H chemical shift in the BSD fibers is observed, illustrating differences in aromatic ring packing for fibers spun from the two dopes. The drift correction offset glitch due to FSLG homonuclear decoupling is indicated with an asterisk (*). Adapted with permission from [37]. Copyright 2023 American Chemical Society
To investigate the eADF3 conformational structure in CSD and BSD dopes and spun fibers, solution and SSNMR were used to characterize the 13C/15N-isotope-enriched samples. Solution NMR revealed that the repetitive core of the protein maintained intrinsic disorder in both dopes, with some subtle but distinct differences observed (Fig. 5C). BSD exhibited sharper resonances for numerous environments in the 13C solution NMR spectra, illustrating a protein preordering effect in the BSD together with notable differences in the tyrosine aromatic ring due to hydrogen bonding differences at the C-OH moiety and presumably perturbations in aromatic ring packing interactions between the two dopes. Interestingly, 13C CP-MAS SSNMR revealed that poly-A β-sheet formation was identical in fibers spun from both dope preparations, with 77% β-sheet content observed from spectral deconvolutions (Fig. 5D). However, a distinct difference in at least one of the tyrosine aromatic rings was revealed in the final fibers by SSNMR, where a large downfield chemical shift was observed for the 1H aromatics in the 1H-13C HETCOR SSNMR spectra when comparing the BSD and CSD fibers (Fig. 5E). These results pointed to the participation of tyrosine residues in spider silk assembly, where clear differences were observed between both the dopes and the spun fibers. This highlights the importance of tracking the non-β-sheet domains in spider silk formation, as both systems display similar poly-A β-sheet conversions but distinct differences in tyrosine interactions.
Applications of artificial spider silk fibers
Natural spider silk fibers are known for their outstanding properties, including their exceptional mechanical properties, unparalleled toughness, high biocompatibility, slow biodegradation, low immunogenicity, and, in some cases, microbe-repellent features [15, 38, 39]. The ability to produce artificial spider silk with similar properties has high application potential in various fields, such as textile technology and biomedicine. By exploring different methods, engineered spidroins have been processed into fibers via solvent extrusion, wet spinning, electrospinning, and microfluidic approaches (see Fig. 6A) [15, 40, 41]. In these methods, spider silk proteins are simply dissolved in a suitable solvent followed by a (classic) concentration step, e.g., polyethylene glycol is used to remove water without considering the biochemical prerequisites existing in vivo. Since such dopes have long been used in artificial spider silk formation, methods involving these dopes are termed classic approaches. In contrast, following the relevance of LLPS in spider silk formation, biomimetic dopes that did not require a concentration step were developed to produce bioengineered man-made fibers with mechanical properties resembling those of natural dragline fibers (see Fig. 6B) [40]. This successful approach uses the BSD and its LLPS, as depicted above. Subsequently, LPS was induced by ion exchange of chaotropic NaCl with comparatively kosmotropic K-Pi and acidification to at least pH 5.7. Two different processing routes have been developed: one is based on a spinning channel with a semipermeable membrane to mimic the epithelial cells in the spinning duct of a spider, while the second utilizes a wet spinning approach to generate continuous fibers [15, 40]. Concomitant shearing and mechanical stretching forces align the spidroins in the fiber, leading to intermolecular cross-linkages between adjacent fibrils, while at the same time, water molecules are displaced and removed. This nature-mimicking process resulted in the first engineered spider silk fiber reported with a nature-identical toughness, and an industrial scale-up of the wet-spinning process was commercialized under the brand name Biosteel® by the company AMSilk GmbH.
Comparison of classic and biomimetic fiber spinning methods. A Classic spinning methods include dry spinning, electrospinning, and wet spinning of silk proteins from a concentrated solution to form randomly aligned fibers. Often, organic and highly volatile solvents are used. B Bioinspired spinning methods require aqueous solvents and recapitulate the natural spinning mechanism by utilizing changes in pH and ion concentration to induce LLPS. Biomimetic spinning (left) induces LLPS prior to the spinning process via the dialysis of a spidroin solution against a phosphate-containing buffer, resulting in a low-concentration and a high-concentration micellar phase. Bioinspired microfluidic spinning (right) utilizes a microfluidic system. Citrate-phosphate buffer (CPB) and negative pressure induce LLPS and fiber formation under continuous flow spinning conditions, mimicking the native spider silk formation process (adapted from [42]). Furthermore, both methods can be combined by using a preassembled micellar spidroin phase for microfluidic spinning (middle). C Stress‒strain curves of microfluidically spun recombinant spider silk fibers from classic (CSD, colored dashed lines) and biomimetic (BSD, colored solid lines) spinning dopes in comparison to those of natural spider silk fibers (solid black line). The C- and N-terminal NR domains (CTD and NTD) attached to eADF3 and eADF4 are vital for fiber assembly. Pink lines depict the results for fibers spun from eADF3/eADF4 homo- and heterodimer dopes derived in vivo in E. coli. The green lines indicate the results from the spinning of in vitro-produced mixed dopes. The orange and blue lines show the stress‒strain curves of eADF4 and eADF3 homomeric dopes. The mechanical properties of the BSD fibers clearly exceeded those of the CSD fibers. Microfluidic spinning of a BSD consisting of an eADF3/eADF4 homo- and dimeric mixture prepared in vivo (solid pink line) resulted in mechanical properties comparable to those of natural spider silk fibers. Adapted from [43]; parts of the figure were prepared in BioRender
Other groups subsequently reported the processing of man-made biotech fibers using different spidroins. Recently, a microfluidic device approach demonstrated ion-induced LLPS of recombinant spidroins, and parallel control over pH-driven fibrillation and shear-dependent induction of β-sheet formation allowed spinning of fibers with a ß-sheet content comparable to that of native dragline silk (see Fig. 6B) [42]. Since the whole fiber formation process was performed in a continuous flow system, the native spinning mechanism was closely mimicked. Interestingly, by combining the heterodimeric constructs of both engineered spider silk proteins eADF3 and eADF4 as in vitro mixtures or as in vivo assemblies produced in E. coli, it was possible to imitate the natural fiber properties even more closely by utilizing both the BSD approach and a microfluidic spinning setup. The resulting biomimetic fibers outperformed both fibers made of homodimers of only single spidroin variants and mixtures provided by the classic spinning dope approach and presented mechanical properties close to those of the natural MA silk of A. diadematus (see Fig. 6C) [43]. These findings support the importance of MaSp interaction at the (supra)molecular level in preassemblies, the contribution of intermolecular interactions of terminal domains to controlling solubility, and self-assembly into higher-order protein structures to yield spider silk fibers. Taken together, the reported approach resulted in the toughest man-made protein fibers to date, strongly indicating that the mechanical performance increases with the extent of LLPS to preassemble the spidroins in the dope [44, 45].
Conclusion and outlook
The liquid crystal, micelle, and LLPS theories were previously viewed as individual concepts that all describe at least parts of native spider silk assembly. As in vivo studies are rare due to the challenges of working with live spiders, the complete silk-spinning mechanism remains unresolved. However, evidence strongly indicates that all three concepts can be seen as constituting a concerted, transitional mechanism for fiber assembly. Micellar function, LLPS to form micrometer-sized liquid-like condensates, and nematic phase generation most likely occur alongside each other in the upper gland parts (tail and ampulla). Next, ion exchange, pH drop, and elongational shear stress upon drawing through the tapered spinning duct induce configurational changes in the terminal domains, elongation, and alignment of the preassembled spidroins. The final assembly into β-sheet-rich nanofibrils and drawing from the exit spigot yields a solid fiber. The highly conserved C- and N-terminal NR domains play a vital role in correct fiber formation. This was further supported by ex vivo studies on native MA glands, which revealed the highly dynamic nature of the spidroins in the ampulla that behave like IDPs but self-assemble into micelle-like superstructures. Recent studies that combine biophysical methods have allowed the tracking of LLPS in recombinant spider silk systems and the determination of how LLPS influences silk protein structure and organization across length scales, including by structure determination in the final wet-spun fiber. These studies have highlighted the importance of scarcer residues with a polar sidechain, such as tyrosine, in LLPS and silk assembly, with the role of several other residues remaining to be elucidated. Another important, open-ended question regarding LLPS and the assembly of spidroins into fibers is the role of the different proteins in the process, as spider silk is a complex multicomponent system. We anticipate that combining studies on recombinant MaSp1 and MaSp2 constructs together with the native system will allow elucidation of each protein’s role as well as protein‒protein interactions important in LLPS and spider silk formation more generally. Even though the current knowledge of this process has allowed the fabrication of artificially spun silk fibers with native-like mechanical properties, deeper insights into this system will help yield materials with high toughness and tensile strength, high biocompatibility, weak immune response induction, and no cytotoxicity for various applications.
References
Xu Z, Wang W, Cao Y, Xue B. Liquid-liquid phase separation: Fundamental physical principles, biological implications, and applications in supramolecular materials engineering. Supramol Mater. 2023;2:100049. https://doi.org/10.1016/j.supmat.2023.100049.
Banani SF, Lee HO, Hyman AA, Rosen MK. Biomolecular condensates: Organizers of cellular biochemistry. Nat Rev Mol Cell Biol. 2017;18:285–98. https://doi.org/10.1038/nrm.2017.7.
Alberti S, Gladfelter A, Mittag T. Considerations and challenges in studying liquid-liquid phase separation and biomolecular condensates. Cell. 2019;176:419–34. https://doi.org/10.1016/j.cell.2018.12.035.
Sun Y, Lim ZW, Guo Q, Yu J, Miserez A. Liquid–liquid phase separation of proteins and peptides derived from biological materials: Discovery, protein engineering, and emerging applications. MRS Bull. 2020;45:1039–47. https://doi.org/10.1557/mrs.2020.301.
Leppert A, Chen G, Lama D, Sahin C, Railaite V, Shilkova O, et al. Liquid-liquid phase separation primes spider silk proteins for fiber formation via a conditional sticker domain. Nano Lett. 2023;23:5836–41. https://doi.org/10.1021/acs.nanolett.3c00773.
Abbas M, Lipiński WP, Wang J, Spruijt E. Peptide-based coacervates as biomimetic protocells. Chem Soc Rev. 2021;50:3690–705. https://doi.org/10.1039/D0CS00307G.
Landreh M, Osterholz H, Chen G, Knight SD, Rising A, Leppert A. Liquid-liquid crystalline phase separation of spider silk proteins. Commun Chem. 2024;7:260 https://doi.org/10.1038/s42004-024-01357-2.
Eisoldt L, Smith A, Scheibel T. Decoding the secrets of spider silk. Mater Today. 2011;14:80–6. https://doi.org/10.1016/s1369-7021(11)70057-8.
Bittencourt DMdC, Oliveira P, Michalczechen-Lacerda VA, Rosinha GMS, Jones JA, Rech EL. Bioengineering of spider silks for the production of biomedical materials. Front Bioeng Biotechnol. 2022;10:958486. https://doi.org/10.3389/fbioe.2022.958486.
Jaramillo-Cano D, Camargo M, Likos CN, Gârlea IC. Dynamical properties of concentrated suspensions of block copolymer stars in shear flow. Macromol. 2020;53:10015–27. https://doi.org/10.1021/acs.macromol.0c01365.
Cheng VA, Walker LM. Transport of nanoparticulate material in self-assembled block copolymer micelle solutions and crystals. Faraday Discuss. 2016;186:435–54. https://doi.org/10.1039/c5fd00122f.
Rasing T, Muševič I Surfaces and interfaces of liquid crystals. ed. 1. Berlin, Germany; Heidelberg, Germany
Knight DP, Vollrath F. Liquid crystals and flow elongation in a spider’s silk production line. Proc R Soc B. 1999;266:519–23. https://doi.org/10.1098/rspb.1999.0667.
Vollrath F, Knight DP. Liquid crystalline spinning of spider silk. Nat. 2001;410:541–8. https://doi.org/10.1038/35069000.
Heim M, Keerl D, Scheibel T. Spider silk: From soluble protein to extraordinary fiber. Angew Chem Int Ed. 2009;48:3584–96. https://doi.org/10.1002/anie.200803341.
Jin H-J, Kaplan DL. Mechanism of silk processing in insects and spiders. Nat. 2003;424:1057–61. https://doi.org/10.1038/nature01809.
Thamm C, Scheibel T. Recombinant production, characterization, and fiber spinning of an engineered short major ampullate spidroin (masp1s). Biomacromolecules. 2017;18:1365–72. https://doi.org/10.1021/acs.biomac.7b00090.
Huemmerich D, Helsen CW, Quedzuweit S, Oschmann J, Rudolph R, Scheibel T. Primary structure elements of spider dragline silks and their contribution to protein solubility. Biochem. 2004;43:13604–12. https://doi.org/10.1021/bi048983q.
Landreh M, Askarieh G, Nordling K, Hedhammar M, Rising A, Casals C, et al. A ph-dependent dimer lock in spider silk protein. J Mol Biol. 2010;404:328–36. https://doi.org/10.1016/j.jmb.2010.09.054.
Shi Y-X, Zhu Y-J, Qian Z-G, Xia X-X. Artificial spider silk materials: from molecular design, mesoscopic assembly, to macroscopic performances. Adv Funct Mater. 2412793. https://doi.org/10.1002/adfm.202412793.
Bargel H, Scheibel T. A bio-engineering approach to generate bioinspired (spider) silk protein-based materials. Autom. 2024;72:657–65. https://doi.org/10.1515/auto-2024-0020.
Xu D, Yarger JL, Holland GP. Exploring the backbone dynamics of native spider silk proteins in black widow silk glands with solution-state nmr spectroscopy. Polymer. 2014;55:3879–85. https://doi.org/10.1016/j.polymer.2014.06.018.
Parent LR, Onofrei D, Xu D, Stengel D, Roehling JD, Addison JB, et al. Hierarchical spidroin micellar nanoparticles as the fundamental precursors of spider silks. PNAS. 2018;115:11507–12. https://doi.org/10.1073/pnas.1810203115.
Onofrei D, Stengel D, Jia D, Johnson HR, Trescott S, Soni A, et al. Investigating the atomic and mesoscale interactions that facilitate spider silk protein pre-assembly. Biomacromolecules. 2021;22:3377–85. https://doi.org/10.1021/acs.biomac.1c00473.
Saric M, Scheibel T. Engineering of silk proteins for materials applications. Curr Opin Biotechnol. 2019;60:213–20. https://doi.org/10.1016/j.copbio.2019.05.005.
Exler JH, Scheibel HuemmerichD. T. The amphiphilic properties of spider silks are important for spinning. Angew Chem Int Ed. 2007;46:3559–62. https://doi.org/10.1002/anie.200604718.
Rammensee S, Slotta U, Scheibel T, Bausch AR. Assembly mechanism of recombinant spider silk proteins. PNAS. 2008;105:6590–5. https://doi.org/10.1073/pnas.0709246105.
Slotta UK, Rammensee S, Gorb S, Scheibel T. An engineered spider silk protein forms microspheres. Angew Chem Int Ed. 2008;47:4592–4. https://doi.org/10.1002/anie.200800683.
Bauer J, Scheibel T. Dimerization of the conserved n-terminal domain of a spider silk protein controls the self-assembly of the repetitive core domain. Biomacromolecules. 2017;18:2521–8. https://doi.org/10.1021/acs.biomac.7b00672.
Hagn F, Eisoldt L, Hardy JG, Vendrely C, Coles M, Scheibel T, et al. A conserved spider silk domain acts as a molecular switch that controls fibre assembly. Nat. 2010;465:239–42. https://doi.org/10.1038/nature08936.
Askarieh G, Hedhammar M, Nordling K, Saenz A, Casals C, Rising A, et al. Self-assembly of spider silk proteins is controlled by a ph-sensitive relay. Nat. 2010;465:236–8. https://doi.org/10.1038/nature08962.
Hagn F, Thamm C, Scheibel T, Kessler H. Ph-dependent dimerization and salt-dependent stabilization of the n-terminal domain of spider dragline silk-implications for fiber formation. Angew Chem Int Ed. 2011;50:310–3. https://doi.org/10.1002/anie.201003795.
Eisoldt L, Thamm C, Scheibel T. Review the role of terminal domains during storage and assembly of spider silk proteins. Biopolymers. 2012;97:355–61. https://doi.org/10.1002/bip.22006.
Malay AD, Suzuki T, Katashima T, Kono N, Arakawa K, Numata K. Spider silk self-assembly via modular liquid-liquid phase separation and nanofibrillation. Sci Adv. 2020;6:eabb6030. https://doi.org/10.1126/sciadv.abb6030.
Mohammadi P, Aranko AS, Lemetti L, Cenev Z, Zhou Q, Virtanen S, et al. Phase transitions as intermediate steps in the formation of molecularly engineered protein fibers. Commun Biol. 2018;1:86 https://doi.org/10.1038/s42003-018-0090-y.
Lemetti L, Scacchi A, Yin Y, Shen M, Linder MB, Sammalkorpi M, et al. Liquid–liquid phase separation and assembly of silk-like proteins is dependent on the polymer length. Biomacromolecules. 2022;23:3142–53. https://doi.org/10.1021/acs.biomac.2c00179.
Stengel D, Saric M, Johnson HR, Schiller T, Diehl J, Chalek K, et al. Tyrosine’s unique role in the hierarchical assembly of recombinant spider silk proteins: From spinning dope to fibers. Biomacromolecules. 2023;24:1463–74. https://doi.org/10.1021/acs.biomac.2c01467.
Aigner TB, DeSimone E, Scheibel T. Biomedical applications of recombinant silk-based materials. Adv Mater. 2018;30:e1704636. https://doi.org/10.1002/adma.201704636.
Kumari S, Lang G, DeSimone E, Spengler C, Trossmann VT, Lücker S, et al. Engineered spider silk-based 2d and 3d materials prevent microbial infestation. Mater Today. 2020;41:21–33. https://doi.org/10.1016/j.mattod.2020.06.009.
Heidebrecht A, Eisoldt L, Diehl J, Schmidt A, Geffers M, Lang G, et al. Biomimetic fibers made of recombinant spidroins with the same toughness as natural spider silk. Adv Mater. 2015;27:2189–94. https://doi.org/10.1002/adma.201404234.
Jokisch S, Neuenfeldt M, Scheibel T Silk-based fine dust filters for air filtration. Adv Sustain Syst. 2017;1. https://doi.org/10.1002/adsu.201700079.
Chen J, Tsuchida A, Malay AD, Tsuchiya K, Masunaga H, Tsuji Y, et al. Replicating shear-mediated self-assembly of spider silk through microfluidics. Nat Commun. 2024;15:527. https://doi.org/10.1038/s41467-024-44733-1.
Saric M, Eisoldt L, Döring V, Scheibel T. Interplay of different major ampullate spidroins during assembly and implications for fiber mechanics. Adv Mater. 2021;33:2006499. https://doi.org/10.1002/adma.202006499.
Andersson M, Jia Q, Abella A, Lee X-Y, Landreh M, Purhonen P, et al. Biomimetic spinning of artificial spider silk from a chimeric minispidroin. Nat Chem Biol. 2017;13:262–4. https://doi.org/10.1038/nchembio.2269.
Välisalmi T, Bettahar H, Zhou Q, Linder MB. Pulling and analyzing silk fibers from aqueous solution using a robotic device. Int J Biol Macromol. 2023;250:126161. https://doi.org/10.1016/j.ijbiomac.2023.126161.
Doblhofer E, Heidebrecht A, Scheibel T. To spin or not to spin: Spider silk fibers and more. Appl Microbiol Biotechnol. 2015;99:9361–80. https://doi.org/10.1007/s00253-015-6948-8.
Jiang W, Askarieh G, Shkumatov A, Hedhammar M, Knight SD Crystal structure of an asymmetric dimer of the n-terminal domain of euprosthenops australis major ampullate spidroin 1. RCSB PDB. 2016;https://doi.org/10.2210/pdb6R9D/pdb, 10.1107/S2059798319007253, PDB ID 6R9D.
Schaal D, Bauer J, Schweimer K, Scheibel T, Roesch P, Schwarzinger S Amino-terminal domain of latrodectus hesperus masp1 with neutralized acidic cluster. RCSB PDB. 2016;https://doi.org/10.2210/pdb2n3e/pdb, PDB ID 2N3E.
Hagn F, Eisoldt L, Hardy JG, Vendrely C, Coles M, Scheibel T, et al. Structure of the c-terminal non-repetitive domain of the spider dragline silk protein adf-3. RCSB PDB. 2010;https://doi.org/10.2210/pdb2KHM/pdb, 10.1038/nature08936, PDB ID 2KHM.
Acknowledgements
TS, and GPH would like to acknowledge funding received from the Bavaria California Technology Center (BaCaTec) for the project “Understanding the natural spinning process of spider silk” (No. 2 [2019-1]). GPH acknowledges grant support from the Department of Defense Air Force Office of Scientific Research (DOD-AFOSR) under Award Nos. FA9550-14-1-0014, FA9550-17-1-0282, FA9550-20-1-0103 and FA9550-23-1-0616.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Article conception: LZ, HB; Literature collection: LZ, HB, TS, GPH; Manuscript writing: LZ, HB, GPH; Figure preparation: LZ, HB, GPH; Editing and critical revision of content: TS, GPH. All authors reviewed, edited and approved the manuscript before submission.
Corresponding author
Ethics declarations
Conflict of interest
TS is the founder and shareholder of AMSilk GmbH, Germany. The other authors declare that they have no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zeußel, L., Bargel, H., Holland, G.P. et al. Liquid‒liquid phase separation of spider silk proteins. Polym J 57, 831–843 (2025). https://doi.org/10.1038/s41428-025-01039-3
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41428-025-01039-3