Abstract
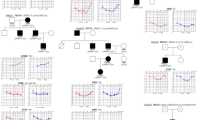
Hearing loss (HL) is one of the most common sensory defects, of which X-linked nonsyndromic hearing loss (NSHL) accounts for only 1–2%. While a COL4A6 variant has been reported in a single Hungarian family with NSHL associated with inner ear malformation, causative role of COL4A6 variants and their phenotypic consequences in NSHL remain elusive. Here we report two families in which we identified a male member with X-linked HL. Each has inherited a rare hemizygous COL4A6 variant from their respective mothers, NM_001287758.1: c.3272 G > C (p.Gly1091Ala) and c.951 + 1 G > C. An in vitro minigene splicing assay revealed that c.951 + 1 G > T leads to skipping of exon 15, strongly suggesting a pathogenic role for this variant in the HL phenotype. The p.Gly1091Ala variant is classified as a variant of unknown significance based on the variant interpretation guidelines. This report provides evidence for variants in the COL4A6 gene resulting in X-linked NSHL. It highlights the importance of in-depth genetic studies in all family members in addition to the proband, especially in multiplex families, to determine the precise etiology of HL.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Corvino V, Apisa P, Malesci R, Laria C, Auletta G, Franze A. X-linked sensorineural hearing loss: a literature review. Curr Genomics. 2018;19:327–38.
Knebelmann B, Breillat C, Forestier L, Arrondel C, Jacassier D, Giatras I, et al. Spectrum of mutations in the COL4A5 collagen gene in X-linked Alport syndrome. Am J Hum Genet. 1996;59:1221.
Plant KE, Green PM, Vetrie D, Flinter FA. Detection of mutations in COL4A5 in patients with Alport syndrome. Hum Mutat. 1999;13:124–32.
Wang F, Ding J, Yu Lx, Yang Jy. Detection of COL4A5 gene mutations in Alport syndrome by analyzing cDNA of skin fibroblasts. J Beijing Med Univ. 2002;3:219–224.
Mothes H, Heidet L, Arrondel C, Richter KK, Thiele M, Patzer L, et al. Alport syndrome associated with diffuse leiomyomatosis: COL4A5‐COL4A6 deletion associated with a mild form of Alport nephropathy. Nephrol Dial Transplant. 2002;17:70–4.
Uliana V, Marcocci E, Mucciolo M, Meloni I, Izzi C, Manno C, et al. Alport syndrome and leiomyomatosis: the first deletion extending beyond COL4A6 intron 2. Pediatr Nephrol. 2011;26:717–24.
Sá MJN, Fieremans N, de Brouwer AP, Sousa R, e Costa FT, Brito MJ, et al. Deletion of the 5′ exons of COL4A6 is not needed for the development of diffuse leiomyomatosis in patients with Alport syndrome. J Med Genet. 2013;50:745–53.
Kashtan CE. Alport Syndrome. In: Adam MP, Ardinger HH, Pagon RA, et al., editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993-2021.
Zhou J, Ding M, Zhao Z, Reeders ST. Complete primary structure of the sixth chain of human basement membrane collagen, alpha 6(IV). Isolation of the cDNAs for alpha 6(IV) and comparison with five other type IV collagen chains. J Biol Chem. 1994;269:13193–9.
Pandya A, Arnos KS, Xia XJ, Welch KO, Blanton SH, Friedman TB, et al. Frequency and distribution of GJB2 (connexin 26) and GJB6 (connexin 30) mutations in a large North American repository of deaf probands. Genet Med. 2003;5:295–303.
Arnos KS, Welch KO, Tekin M, Norris VW, Blanton SH, Pandya A, et al. A comparative analysis of the genetic epidemiology of deafness in the United States in two sets of pedigrees collected more than a century apart. Am J Hum Genet. 2008;83:200–7.
del Castillo FJ, del Castillo I, Moreno-Pelayo MA, Brownstein Z, Marlin S, Adina Q, et al. Prevalence and evolutionary origins of the del(GJB6-D13S1830) mutation in the DFNB1 locus in hearing-impaired subjects: a multicenter study. Am J Hum Genet. 2003;73:1452–8.
Pandya A, Xia X, Erdenetungalag R, Amendola M, Landa B, Radnaabazar J, et al. Heterogenous point mutations in the mitochondrial tRNA Ser (UCN) precursor coexisting with the A1555G mutation in deaf students from Mongolia. Am J Hum Genet. 1999;65:1803–6.
Yan D, Tekin D, Bademci G, Foster J, Cengiz FB, Kannan-Sundhari A, et al. Spectrum of DNA variants for non-syndromic deafness in a large cohort from multiple continents. Hum Genet. 2016;135:953–61.
Booth KT, Azaiez H, Kahrizi K, Wang D, Zhang Y, Frees K, et al. Exonic mutations and exon skipping: lessons learned from DFNA5. Hum Mutat. 2018;39:433–40.
Oza AM, DiStefano MT, Hemphill SE, Cushman BJ, Grant AR, Siegert RK, et al. Expert specification of the ACMG/AMP variant interpretation guidelines for genetic hearing loss. Hum Mutat. 2018;39:1593–613.
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM, Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet. 2014;46:310.
Schwarz JM, Cooper DN, Schuelke M, Seelow D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods. 2014;11:361.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248.
Ng PC, Henikoff S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res. 2003;31:3812–4.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–23.
Richard G, White TW, Smith LE, Bailey RA, Compton JG, Paul DL, et al. Functional defects of Cx26 resulting from a heterozygous missense mutation in a family with dominant deaf-mutism and palmoplantar keratoderma. Hum Genet. 1998;103:393–9.
Putcha GV, Bejjani BA, Bleoo S, Booker JK, Carey JC, Carson N, et al. A multicenter study of the frequency and distribution of GJB2 and GJB6 mutations in a large North American cohort. Genet Med. 2007;9:413.
Mani RS, Ganapathy A, Jalvi R, Srisailapathy CS, Malhotra V, Chadha S, et al. Functional consequences of novel connexin 26 mutations associated with hereditary hearing loss. Eur J Hum Genet. 2009;17:502.
Yuan Y, Huang D, Yu F, Zhu X, Kang D, Yuan H, et al. A de novo GJB2 (connexin 26) mutation, R75W, in a Chinese pedigree with hearing loss and palmoplantar keratoderma. Am J Med Genet Part A. 2009;149:689–92.
Lee JY, In S, Kim HJ, Jeong S, Choung YH, Kim YC. Hereditary palmoplantar keratoderma and deafness resulting from genetic mutation of Connexin 26. J Korean Med Sci. 2010;25:1539–42.
Weegerink NJ, Pennings RJ, Huygen PL, Hoefsloot LH, Cremers W, Kunst HP. Phenotypes of two Dutch DFNA3 families with mutations in GJB2. Ann Otol Rhinol Laryngol. 2011;120:191–7.
Pang X, Chai Y, Sun L, Chen D, Chen Y, Zhang Z, et al. Characterization of spectrum, de novo rate and genotype-phenotype correlation of dominant GJB2 mutations in Chinese hans. PLoS ONE. 2014;9:e100483.
Deng Y, Chen Y, Reuss L, Altenberg GA. Mutations of connexin 26 at position 75 and dominant deafness: essential role of arginine for the generation of functional gap-junctional channels. Hear Res. 2006;220:87–94.
Marziano NK, Casalotti SO, Portelli AE, Becker DL, Forge A. Mutations in the gene for connexin 26 (GJB2) that cause hearing loss have a dominant negative effect on connexin 30. Hum Mol Genet. 2003;12:805–12.
Inoshita A, Iizuka T, Okamura H, Minekawa A, Kojima K, Furukawa M, et al. Postnatal development of the organ of Corti in dominant-negative Gjb2 transgenic mice. Neuroscience 2008;156:1039–47.
Inoshita A, Karasawa K, Funakubo M, Miwa A, Ikeda K, Kamiya K. Dominant negative connexin26 mutation R75W causing severe hearing loss influences normal programmed cell death in postnatal organ of Corti. BMC Genet. 2014;15:1.
Rost S, Bach E, Neuner C, Nanda I, Dysek S, Bittner RE, et al. Novel form of X-linked nonsyndromic hearing loss with cochlear malformation caused by a mutation in the type IV collagen gene COL4A6. Eur J Hum Genet. 2014;22:208.
Acknowledgements
This study was supported by NIH Grant R01DC009645 from the National Institutes of Health/National Institute on Deafness and Other Communication Disorders.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
O’Brien, A., Aw, W.Y., Tee, H.Y. et al. Confirmation of COL4A6 variants in X-linked nonsyndromic hearing loss and its clinical implications. Eur J Hum Genet 30, 7–12 (2022). https://doi.org/10.1038/s41431-021-00881-2
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-021-00881-2
This article is cited by
-
Collagen IV in Gould syndrome and Alport syndrome
Nature Reviews Nephrology (2025)
-
Identification of known and novel genetic variants in sensorineural hearing loss: insights from whole exome sequencing in Indian families
Molecular Biology Reports (2025)
-
Identification of a novel intronic variant in COL4A2 gene associated with fetal severe cerebral encephalomalacia and subdural hemorrhage
BMC Medical Genomics (2024)
-
Genomic and phenotypic landscapes of X-linked hereditary hearing loss in the Chinese population
Orphanet Journal of Rare Diseases (2024)
-
New year, new issue
European Journal of Human Genetics (2022)