Abstract
Detecting intermediate-sized structural variants (SVs) remains challenging in diagnostics, as tools for single-nucleotide and copy-number variants, particularly read-depth-based methods, are often insufficient. GRIDSS addresses this gap by integrating paired-end mapping, split-read analysis, and assembly-based approaches. However, its use in targeted sequencing and diagnostic workflows remains complex. NGS panel data from 9726 patients with suspected hereditary cancer were analyzed using GRIDSS. A filtering strategy was developed to prioritize clinically relevant germline SVs. Multiple parameter settings were tested to optimize performance. The initial dataset of 1,307,592 variants was reduced to 89 candidates after applying the selected filtering strategy. Of these, 24 had been previously detected by routine callers and were not further analyzed. Among the remaining 65, 13 were considered likely true positives after visual inspection using IGV. Experimental validation was performed by Sanger/Nanopore long-read sequencing for these variants, all of which were confirmed. Eight were classified as (likely) pathogenic, including two frameshift duplications in MSH6, one splicing variant in BARD1, and five mobile element insertions in APC, BRCA2, and PALB2. Altogether, GRIDSS implementation increased diagnostic yield while maintaining feasibility for diagnostic workflows.

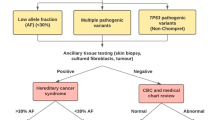
Comprehensive workflow scheme for germline structural variant detection and results in our diagnostic setting.
Similar content being viewed by others
Introduction
Short-read NGS panels are widely used in routine diagnostics due to their optimal balance between cost and efficiency [1]. Different bioinformatics tools are available to detect single nucleotide variants (SNVs), small insertions or deletions (indels, <20 bp) and structural variants (SVs). For SV detection on panel data, most programs rely exclusively on read depth, by comparing coverage changes across genomic windows to determine whether the observed coverage aligns with the expected levels.
While these approaches effectively detect many variants, they have notable limitations. SNV and indel callers rely on reads aligning to both sides of a variant, a requirement that becomes increasingly difficult for large SVs when using short-read sequencing [2]. As SV size increases, fewer reads fully span the breakpoints, reducing the caller’s confidence and decreasing the detected variant allele fraction (VAF). In some cases, this can result in missed detections. Besides, read depth-based methods fail to identify variants significantly shorter than the genomic window used for coverage analysis or those that do not alter copy number, such as balanced SVs, and insertions of elements that do not align the target region or genome. Although there is no consensus definition for the term ‘intermediate size variants’, our study defines them as variants ranging from 20 bp to 1 kb.
Structural variants represent a significant source of genomic variation that may play a role in numerous diseases with a genetic etiology, including cancer predisposition. Therefore, improving detection methods is crucial to uncover these variants and fully understand their role in human diseases. In addition to the read depth approach, there are three other main strategies to call SVs from short-read data: paired-end mapping, split reads and de novo assembly [3]. Paired-end mapping analyzes the distribution of paired-end reads to identify discrepancies in their expected distance, orientation or order. The split-read strategy detects SVs by identifying individual reads in which adjacent segments align to distinct, non-contiguous or differently oriented genomic locations, reflecting a breakpoint. These reads often contain misaligned bases that are soft-clipped by the aligner. When soft-clipped bases are highlighted in genome browsers, split reads become easier to identify and interpret through visual inspection. Lastly, de novo assembly methods reconstruct genomic sequences without a reference genome. Reads are assembled into longer contiguous sequences known as contigs, which are then compared to the reference genome or another assembly. Each method has its own strengths and limitations; thus, relying on a single approach can lead to incomplete detection of some variant types. To address this issue, recent tools have combined multiple strategies to improve detection capabilities. One such tool is GRIDSS (Genome Rearrangement IDentification Software Suite) [4], which integrates paired-end mapping, split reads and de novo assembly, offering a more comprehensive approach to increase variant identification [5].
However, implementing GRIDSS in routine diagnostics poses certain challenges. Originally designed for Illumina sequencing data in a whole-genome context, GRIDSS has not yet been validated for exon-targeted data. Additionally, it reports variants in a breakend notation, which, although comprehensive in describing structural variation, can be difficult to interpret. Furthermore, GRIDSS returns many variants, and while the authors have developed a code to filter somatic variants, no specific guidance or tools are available for filtering germline variants, which further complicates its application in germline diagnostic settings.
This study aims to determine whether structural variants were previously overlooked due to the exclusive use of a read depth-based method for SV identification, by adapting GRIDSS for use with gene panel data. In addition, the study seeks to provide practical filters to identify germline SVs with high clinical impact, making the process suitable for routine diagnostic practice.
Subjects and methods
Study cohort
A total of 9726 patients with suspected hereditary cancer were included in this study. All of them were referred to the Molecular Diagnostics Service at the Catalan Institute of Oncology (ICO) and provided informed written consent for both diagnostic and research purposes. The study protocol was approved by the Ethics Committee of the Catalan Institute of Oncology–Bellvitge University Hospital (PR278/19).
Routine diagnostics genetic testing
DNA was extracted from peripheral blood leukocytes, and genetic testing was performed using the custom NGS ICO-IMPPC Hereditary Cancer Panel (I2HCP), which includes all exons and flanking intron-exon boundaries (approximately ±150 bp) of 122–165 hereditary cancer-associated genes, depending on the panel version [6]. For the diagnostic analysis of SNVs, indels, and CNVs, the target region was defined as all coding exons plus their immediate ±20 bp flanking sequences, which were covered at a minimum depth of 30x. To improve the detection of structural variants in this study, however, we extended the target region to include up to ±150 bp of flanking intronic sequence.
Samples were paired-end sequenced on three different platforms: 5552 samples were sequenced on a NextSeq platform (read length = 150 bp, average coverage = 595x), 2699 on a HiSeq platform (read length = 250 bp, average coverage = 868x), and 1475 on a MiSeq platform (read length = 300 bp, average coverage = 494x) (Illumina, San Diego, CA, US). Two variant callers were used: VarScan for SNVs and short indels and DECoN for CNVs [6, 7].
Bioinformatics pipeline
GRIDSS
GRIDSS (v2.13.2) [4] was installed, and the core script was executed on each BAM file using the default parameters to generate a GRIDSS SV VCF file. All BAM files were aligned to the GRCh37 human reference genome. Additionally, Repeat Masker (v.4.1.5) was configured to enable the use of the gridss_annotate_vcf_repeatmasker script, which annotates the VCF file by identifying and classifying inserted sequences based on RepeatMasker’s database.
Variant filtering pipeline
The filter_gridss pipeline was developed in the R statistical computing environment (R v4.2.1), using functions from packages available in R/Bioconductor or CRAN. The source code is available at the GitHub repository: https://github.com/emunte/gridss_filter.
For each VCF file, a dataset dedicated to two-breakend variants was created by pairing events by their mate ID, ensuring that each variant was represented by a single row. Variants lacking a matching mate ID were stored separately in a one-breakend dataset. This separation allowed for the application of distinct filtering criteria to each dataset.
The following filters were applied uniformly to both datasets: (1) exclusion of identical variants (defined as those with identical breakpoints and alternative allele) detected in ten or more samples; (2) exclusion of variants located in deep intronic regions (beyond ±150 bp from exon boundaries) of phenotype-related genes, as well as in any intergenic region, or region (coding or non-coding) of genes not associated with the patient’s phenotype, in compliance with Catalan Health Service guidelines [8]. POLE and POLD1 genes were also excluded, given that loss of proofreading activity in these genes is typically caused by missense pathogenic variants [9]; (3) exclusion of variants with a VAF below 10%, calculated as described in the GRIDSS documentation; (4) exclusion of variants located in regions annotated as simple repeat or low-complexity by RepeatMasker.
Additional dataset-specific filters were then applied. For the two-breakend dataset: (5) exclusion of highly similar variants (defined as those with identical breakpoints) present in fifteen or more samples, as these likely represent a recurrent event/artifact in complex genomic regions; (6) exclusion of indels shorter than 21 bp, as such events were presumed to have been previously captured by short indel callers. For the one-breakend dataset: (5) exclusion of highly similar variants (as defined above), with the additional criterion that insertions were also considered highly similar if the transposable element overlapped another element of the same class within a ± 4 bp window; (6) restriction of the analysis to variants annotated as transposable elements by RepeatMasker.
Filter parameter optimization
Prior to the pipeline implementation, three filters were tested using five different values to establish the optimal thresholds for our dataset. The minimum VAF (filter 3) was evaluated at 5%, 8%, 10%, 12%, and 15%, while the frequency thresholds for exclusion of identical (filter 1) and highly similar variants (filter 5) were tested at 5, 8, 10, 12, and 15 samples. All valid combinations of these values were assessed (Supplementary Table S1). Since filter 1 is more restrictive than filter 5, the threshold for filter 5 was required to be equal to or greater than that of filter 1.
Selection of candidates by visual inspection
Following the pipeline, SVs that passed the filters were visually inspected in Integrative Genomics Viewer (IGV) to identify likely true positive calls. Coverage patterns, read pair orientation and soft-clipped bases were examined, and the exact breakpoints were extracted for subsequent experimental validation.
Experimental validation
Variants selected at the visual inspection step were validated in genomic DNA by Sanger sequencing. PCR reactions were performed using primers flanking the expected breakpoints, with DreamTaq DNA Polymerase or Phusion High-Fidelity DNA Polymerase (Thermo Fisher Scientific, Waltham, MA, US) according to the manufacturers’ protocols. PCR products were purified with ExoSAP-IT and sequenced on an AB3500 Genetic Analyzer using BigDye™ Terminator v3.1 kit (Thermo Fisher Scientific). Primer sequences and PCR conditions are available upon request.
A Long Interspersed Nuclear Element (LINE) insertion in the APC gene was validated through long-read sequencing. The analysis followed the Comprehensive Germline Cancer Panel Workflow provided by Oxford Nanopore Technologies. Briefly, genomic DNA libraries were prepared using the Native Barcoding Kit 24 V14 (SQK-NBD114.24) and sequenced on a PromethION platform with an R10.4.1 flow cell (Oxford Nanopore Technologies, Oxford, UK). Enrichment of a panel of 258 hereditary cancer genes was performed via adaptive sampling, and the analysis was run in EPI2ME platform with the wf-hereditary-cancer workflow, which uses Sniffles2 tool for SV calling. Alignments were visualized and interpreted manually with IGV version 2.18.4.
mRNA RT-PCR and sequencing was performed in those patients harboring SVs predicted to disrupt splicing, as previously described [10]. Briefly, total RNA was isolated using TRIzol reagent from cultured peripheral blood lymphocytes treated with and without puromycin, and reverse transcribed with iScript cDNA Synthesis kit (Bio-Rad Laboratories, Hercules, CA, US). cDNA amplification was performed using exonic primers that encompassed the region of interest with DreamTaq DNA Polymerase (Thermo Fisher Scientific), and PCR products were purified and sequenced on an AB3500 Genetic Analyzer (ThermoFisher Scientific).
Variant classification
Variants were classified following the American College of Medical Genetics and Genomics and the Association for Molecular Pathology (ACMG/AMP) guidelines [11]. Gene-specific guidelines and other guidance developed by ClinGen Variant Curation Expert Panels and the ClinGen Sequence Variant Interpretation Working Group (SVI WG) were used when applicable (accessible at: https://cspec.genome.network/cspec/ui/svi/ and https://clinicalgenome.org/working-groups/sequence-variant-interpretation/). As current guidelines do not address the application of PM2 criterion for SVs, PM2_supporting was applied to mobile element insertions (MEIs) whenever the variant was absent from the gnomAD SV v4.0 dataset.
Results
Optimization of filtering parameters
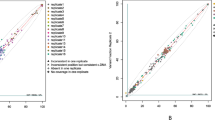
To reduce variant burden while preserving clinically relevant findings, our filtering strategy included a VAF threshold, recurrence across samples, genomic context and variant class (see Materials and Methods, variant filtering pipeline section). We also explored the potential use of read depth as a filtering parameter by examining the distribution of all detected variants across sequencing platforms (Supplementary Fig. 1). However, no consistent association was observed between read depth and variant reliability, and both true and false positives were distributed across a wide range of depths. Given the lack of a clear discriminatory pattern and the risk of discarding true positive variants, read depth was not implemented as a filter in our pipeline.
To ensure optimal performance of the filtering pipeline for our dataset, different threshold values for the minimum VAF and the threshold number of identical or highly similar variants were tested (Supplementary Table S1). Parameter combinations were evaluated independently for the two-breakend and one-breakend datasets. To enable evaluation across all combinations, the most permissive thresholds were initially applied, and all variants retained under this configuration were visually inspected in IGV; filtering was then refined to define the final parameter set.
Combinations that filtered out at least one of the confirmed candidate variants (defined as those passing all filtering steps and subsequently validated by visual inspection and/or experimental confirmation; see Validated variants in clinical context) were excluded. In the two-breakend dataset, 27 out of 75 combinations (36%) were discarded, including all scenarios with a minimum VAF threshold of 15% and those where the threshold for highly similar variants was set at 5 or 8 samples (Supplementary Fig. 2A). In the one-breakend dataset, all combinations with a minimum VAF threshold of 12% or 15% were discarded (20/75 = 26.7%) (Supplementary Fig. 2B). Notably, thresholds for the number of identical or highly similar variants did not impact the detection of true positive variants for the combinations tested.
Among the parameter combinations that retained all confirmed candidate variants, we selected thresholds that could be consistently applied to both datasets. The final configuration (minimum VAF = 10%, recurrence in <10 samples for identical variants and <15 samples for highly similar variants) ensured retention of all true positives while keeping the number of false positive calls within acceptable limits (Supplementary Fig. 2).
Variant filtering strategy
The filter_gridss script was developed to prioritize germline variants from the VCF files generated by the gridss_annotate_vcf_repeatmasker script. BAM files of 9726 samples were analyzed using GRIDSS and subsequently filtered using the filter_gridss script: 5552 were sequenced on a NextSeq platform (average coverage = 595x), 2699 on a HiSeq (average coverage = 868x) and 1475 on a MiSeq (average coverage = 494x). The initial two-breakend dataset contained 798,536 variants (298,630 from NextSeq, 398,912 from HiSeq and 100,994 from MiSeq), while the one-breakend dataset included 509,056 variants (443,148 from NextSeq, 50,519 from HiSeq and 15,389 from MiSeq). After applying the filters described in the Methods section, 79 two-breakend variants and 10 one-breakend variants passed the filtering criteria. For a detailed breakdown of the number of variants filtered at each step, refer to Fig. 1. Of the remaining 89 variants, 24 had been previously detected by routine diagnostic callers (7 short indels detected by VarScan and 17 CNVs detected by DECoN; Supplementary Table S2) and were therefore disregarded. A total of 65 variants were retained for visual inspection.
A Two-breakend dataset; B One-breakend dataset. Each box represents a filtering step, with arrows indicating the flow of variants through the pipeline. The number of variants retained or removed is shown for each step. The final filter in both panels is shaded in light yellow to indicate that it is a dataset-specific filter. bp base pairs, ROI region of interest, VAF variant allele frequency.
Visual inspection of candidate variants
Visual inspection involved analyzing coverage patterns to detect regions with abrupt read-depth shifts and identifying the exact positions of breakpoints by examining soft-clipped bases. Additionally, evaluating pair orientation using the read pair IGV option, along with aligning soft-clipped bases through IGV-assisted BLAT, provided a clearer understanding of the nature of many variants. Of the 65 variants visually inspected, 52 showed no evidence of structural variation and were discarded. Thirteen variants remained for experimental validation (Table 1).
Validated variants in clinical context
Thirteen variants were experimentally tested and all were successfully validated (eight two-breakend and five one-breakend; Table 1). Of these, nine were considered to have potential clinical relevance (patient IDs 2, 5, 6 and 8–13; Table 2), while four were of limited clinical significance (patient IDs 1, 3, 4 and 7).
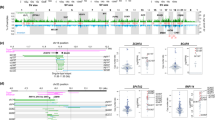
Among two-breakend variants, two pathogenic frameshift duplications in the MSH6 gene were identified: c.3834_3862dup (p.(Lys1288Thrfs*49)) and c.3922_3979dup (p.(Asn1327Thrfs*11)), leading to a diagnosis of Lynch syndrome. These findings were consistent with loss of MSH6 expression in the tumors of both patients. Additionally, the BARD1 c.1865_1903+274del variant was identified in a breast cancer patient. Since this deletion encompassed the canonical donor site of exon 9, mRNA RT-PCR and sequencing were performed to assess its impact on splicing. Two alternative transcripts were detected: (1) the predominant transcript caused the skipping of exons 8 and 9 (r.1678_1903del), resulting in a frameshift predicted to trigger nonsense-mediated decay (p.Met560Glyfs*2); (2) the minor transcript ( ~ 10%) resulted in the skipping of exon 9 (r.783_806del), an in-frame alteration that removed a central region within the BRCT1 domain (p.Val604_Trp635delinsGly) (Supplementary Fig. 3). Consequently, this variant was classified as likely pathogenic. Lastly, an in-frame duplication in the PALB2 gene was identified in a breast cancer patient (c.739_891dup; p.(Thr247_Thr297dup)). However, with the current information, it was classified as a variant of uncertain significance (VUS) and did not impact clinical management. The remaining four variants were considered to have limited clinical relevance. Three were classified as VUS: two EPCAM variants located in regions not associated with MSH2 inactivation [12], and one deep intronic MSH6 deletion with no predicted impact on splicing according to the SpliceAI tool (https://spliceailookup.broadinstitute.org/). Finally, one deep intronic ATM variant was classified as likely benign based on its population frequency and the absence of any predicted effect on splicing.
Among the five one-breakend variants, four were Alu insertions: one in the PALB2 gene identified in a breast cancer patient diagnosed at age 38, one in the ATM gene found in a prostate cancer patient diagnosed at age 53, and two in the BRCA2 gene, both identified in breast cancer patients with two tumor diagnoses each (ages 33 and 57 in patient 11, and ages 46 and 51 in patient 12). The BRCA2 variant detected in patient 11 corresponds to the well-known Portuguese founder Alu insertion c.156_157insAlu [13]. The fifth case involved a six-kb long LINE1 element insertion in the APC gene, found in a patient diagnosed with classical familial adenomatous polyposis at age 14. His family history included his mother’s diagnosis of colorectal polyposis at age 38 and his brother’s diagnosis at age 18 (Table 2; Supplementary Fig. 4).
Across the 9726 samples analyzed in this study, 1267 individuals (13.0%) were previously found to carry (likely) pathogenic variants, accounting for a total of 1311 pathogenic variants, since 40 individuals harbored more than one pathogenic variant. The implementation of this strategy led to the identification of eight additional (likely) pathogenic variants, representing a 0.61% (8/1311) relative increase in the total number of pathogenic variants detected.
Discussion
Increasing the diagnostic yield
Our study aimed to adapt GRIDSS for detecting germline SVs of intermediate size from targeted NGS data, addressing a gap in routine diagnostics. By implementing a customized filtering pipeline, we identified eight (likely) pathogenic variants, increasing the diagnostic yield from 13.48% to 13.56% (a 0.61% relative improvement). In terms of colorectal cancer susceptibility, we diagnosed two patients with Lynch syndrome and one patient with familial adenomatous polyposis. Additionally, we identified four (likely) pathogenic variants in breast cancer susceptibility genes and one pathogenic variant in a prostate cancer patient, highlighting the role of SVs in the missing heritability of cancer. These diagnoses are of high clinical value, involving high- to moderate-risk genes with well-established management, surveillance, and treatment protocols. Furthermore, other family members may benefit from predictive testing, allowing for personalized risk management and prevention strategies.
Five (likely) pathogenic variants from our dataset were MEIs, including Alu and LINE elements. While their detection is challenging due to their repetitive nature and ubiquitous presence in the genome, MEIs could account for up to 0.3% of all disease-causing variants [14]. Therefore, our findings further reinforce the importance of incorporating mobile element detection strategies into routine diagnostic pipelines.
Filtering strategy
When developing a new tool for clinical practice, achieving an optimal balance between sensitivity and specificity is crucial. Our filtering strategy was designed to reduce variant burden while prioritizing clinically relevant findings. To this end, we evaluated three parameters at multiple thresholds: the minimum VAF and the recurrence of identical or highly similar variants across samples. Read depth was also explored as a potential filter, but the distribution varied widely across platforms, with both true variants and false positives spanning a broad coverage range and showing no consistent trend. No threshold could be defined without risking the loss of true positive events. We therefore discarded this filter, as it would have reduced sensitivity without substantially lowering the variant validation burden (with the selected filters only 65 variants needed visual inspection, and 13 of them, experimental confirmation). Nevertheless, in other datasets with different characteristics or larger sets of validated positives, depth-based thresholds could still be useful.
We acknowledge that our stringent VAF threshold may hinder the detection of clinically significant events, such as mosaic variants or insertions located in challenging genomic regions, where supporting reads may be sparse or poorly aligned. The filter on identical variants is intended to reduce false positive calls, sequencing artifacts and common polymorphisms; however, it also entails the risk of excluding recurrent or founder pathogenic variants that other tools might miss. Likewise, we implemented a filter to discard highly similar variants, assuming these would likely represent the same underlying event miscalled multiple times. In the context of a retrospective study, while these recurrence-based filters may exclude the most frequent SVs, their application significantly reduces the burden of manual review. Without recurrence filtering, the two-breakend dataset would contain 926 variants after filtering and the one-breakend dataset 264, compared to 79 and 10 variants, respectively. This corresponds to a 91.5% reduction in the number of variants requiring visual inspection for the two-breakend dataset and a 96.2% reduction for the one-breakend dataset.
Among the combinations that retained all confirmed candidate variants, the most efficient option in terms of false positive burden used thresholds of VAF ≥ 10%, recurrence in <5 samples for identical variants, and <10 samples for highly similar variants, resulting in 43 false positive variant calls (Supplementary Figure 1A; Supplementary Table S1). However, we opted for a slightly more permissive configuration (identical: <10; highly similar: <15) to try to maximize sensitivity while maintaining a manageable workload, which resulted in a modest 8.5% increase in false positives (47 in total). In contrast, relaxing these filters further would have resulted in 60 to 100 false positive variants being retained, representing an increase of 27.7% to 112.8%, which was deemed excessive for routine implementation.
We note that the set of experimentally validated variants used to optimize our filtering strategy is not exhaustive, and additional true variants may exist beyond those detected in this study.
GRIDSS limitations
While GRIDSS is a powerful tool, its performance can be influenced by sequencing quality. In our dataset, an AluYb8 insertion in BRCA2 was identified in patient 23. Although typically one proband per family is studied, two additional family members were also included in the study: her sister, diagnosed with breast cancer at age 36, and a distant cousin, diagnosed with breast cancer at age 37. Initially, the Alu insertion was not detected in either relative. However, upon further inspection of the VCF files, the variant was found in the sister’s data but had failed to meet the GRIDSS quality threshold of 1500 (quality score = 918) and had been discarded. There were few reads supporting this variant (VAF = 6.4%), but visual inspection suggested that the variant was present. In contrast, the variant was absent in the cousin’s data, confirming her non-carrier status. It is plausible that other SVs with low quality parameters may have remained undetected due to this same issue.
GRIDSS performance was also influenced by the sequencing platform, particularly regarding the balance between one-breakend and two-breakend calls. A key distinguishing feature among platforms is read length, which may impact the ability to resolve both breakpoints of a SV. MiSeq and HiSeq, with longer reads (300 bp and 250 bp, respectively), enabled more accurate mapping of both breakpoints, resulting in a higher proportion of two-breakend calls (87% and 89%, respectively). In contrast, the shorter read length of NextSeq (150 bp) may hinder breakpoint resolution, contributing to a higher proportion of one-breakend calls (60%). Notably, MiSeq also showed the lowest overall detection rate, with an average of 79 variants called per sample, compared to 134 for NextSeq and 167 for HiSeq. This reduced sensitivity is likely attributable to the lower coverage obtained in MiSeq runs.
It is important to note that GRIDSS cannot detect SVs when both breakpoints lie outside captured regions. While this is usually not an issue for whole-genome sequencing, it can limit detection in targeted panels covering only exons and adjacent intronic regions. In such cases, several scenarios may affect variant calls: (1) if one breakpoint is inside and the other outside the captured region, the variant may still be detected with an altered VAF; (2) if one breakpoint is near the edge of a low-coverage region and the other outside, the variant may be called as a one-breakend event and filtered out under our strategy, since it would not meet criteria for a MEI. In contrast, read depth-based methods may still detect some of these variants if at least one exon is captured.
A clear example illustrating the complementarity of GRIDSS and depth-based callers is the BARD1 c.1865_1903+274del variant. In our cohort, this variant was identified in two patients: in one, it was detected by both DECoN and GRIDSS, and is reported as previously identified by routine callers (Supplementary Table S2); in the other, only GRIDSS detected the variant, representing a novel finding. This case highlights how GRIDSS can uncover SVs missed by conventional callers, particularly in the intermediate-size range.
Given the strengths and limitations of each approach, combining tools with complementary strategies is the most effective way to maximize detection of all SV types and sizes.
GRIDSS in a clinical context
The methodology applied in this study entails the limitations previously discussed, and a more lenient filtering approach could have increased the detection rate. However, the modular nature of our strategy allows for threshold adjustments tailored to both the cohort characteristics and available resources for variant visualization and validation. This flexibility could enable laboratories to adapt the tool to their needs, balancing sensitivity with time/cost investment, and reducing variants to a manageable number. For smaller datasets or cases with high clinical suspicion, alternative strategies may maximize variant detection. For instance, our focus on variants in phenotype-related genes inherently limits the identification of incidental findings, which might be desirable in certain diagnostic contexts.
The application of GRIDSS in routine diagnostics holds great potential. In our study, the filtering strategy was developed within a retrospective framework, where recurrence-based filters were essential to keep the manual review workload feasible. In a prospective diagnostic setting, however, far fewer variants are analyzed per run, allowing manual inspection without recurrence filters. As additional samples are processed over time, an in-house database could be built to correct for panel- or laboratory-specific artifacts and common polymorphisms, further reducing the number of variants requiring review and allowing recurrence filters to be applied more selectively.
In summary, our approach has enhanced the detection of clinically relevant SVs. The restrictive filtering strategy presented here has demonstrated practical applicability in a diagnostic setting, providing a reliable and adaptable method for identifying high-impact variants in clinical practice.
Data availability
The variants reported in this study have been submitted to ClinVar and will be accessible with the following IDs: SCV007113783 (APC variant), SCV007113784 and SCV007113785 (ATM variants), SCV007108624 and SCV007108625 (BRCA2 variants), SCV007108621 to SCV007108623 (MSH6 variants), SCV007113786 and SCV007113787 (PALB2 variants) and SCV007108617 to SCV007108619 (EPCAM and BARD1 variants). Mobile element insertion sequences are available in GenBank under accession numbers PQ998981 and PX241358–PX241361.
Code availability
The source code is available at the GitHub repository: https://github.com/emunte/gridss_filter.
References
Rehm HL. Disease-targeted sequencing: A cornerstone in the clinic. Nat Rev Genet. 2013;14:239–40.
Mahmoud M, Gobet N, Cruz-Dávalos DI, Mounier N, Dessimoz C, Sedlazeck FJ. Structural variant calling: the long and the short of it. Genome Biol. 2019;20:268.
Escaramís G, Docampo E, Rabionet R. A decade of structural variants: description, history and methods to detect structural variation. Brief Funct Genomics. 2015;14:305–14.
Cameron DL, Schröder J, Penington JS, Do H, Molania R, Dobrovic A, et al. GRIDSS: sensitive and specific genomic rearrangement detection using positional de Bruijn graph assembly. Genome Res. 2017;27:2050–60.
Barbitoff YA, Ushakov MO, Lazareva TE, Nasykhova YA, Glotov AS, Predeus AV. Bioinformatics of germline variant discovery for rare disease diagnostics: current approaches and remaining challenges. Brief Bioinform. 2024;25:bbad508.
Castellanos E, Gel B, Rosas I, Tornero E, Santín S, Pluvinet R, et al. A comprehensive custom panel design for routine hereditary cancer testing: preserving control, improving diagnostics and revealing a complex variation landscape. Sci Rep. 2017;7:39348.
Moreno-Cabrera JM, del Valle J, Castellanos E, Feliubadaló L, Pineda M, Brunet J, et al. Evaluation of CNV detection tools for NGS panel data in genetic diagnostics. Eur J Hum Genet. 2020;28:1646–55.
Feliubadaló L, López-Fernández A, Pineda M, Díez O, del Valle J, Gutiérrez-Enríquez S, et al. Opportunistic testing of BRCA1, BRCA2 and mismatch repair genes improves the yield of phenotype driven hereditary cancer gene panels. Int J Cancer. 2019;145:2682–91.
Mur P, García-Mulero S, del Valle J, Magraner-Pardo L, Vidal A, Pineda M, et al. Role of POLE and POLD1 in familial cancer. Genet Med. 2020;22:2089–2100.
Rofes P, Menéndez M, González S, Tornero E, Gómez C, Vargas-Parra G, et al. Improving Genetic Testing in Hereditary Cancer by RNA Analysis: Tools to Prioritize Splicing Studies and Challenges in Applying American College of Medical Genetics and Genomics Guidelines. J Mol Diagn. 2020;22:1453–68.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Ligtenberg MJL, Kuiper RP, Geurts Van Kessel A, Hoogerbrugge N. EPCAM deletion carriers constitute a unique subgroup of Lynch syndrome patients. Fam Cancer. 2013;12:185–93.
Peixoto A, Santos C, Pinheiro M, Pinto P, Soares MJ, Rocha P, et al. International distribution and age estimation of the Portuguese BRCA2 c.156-157insAlu founder mutation. Breast Cancer Res Treat. 2011;127:671–9.
Qian Y, Mancini-DiNardo D, Judkins T, Cox HC, Brown K, Elias M, et al. Identification of pathogenic retrotransposon insertions in cancer predisposition genes. Cancer Genet. 2017;216–217:159–69.
Acknowledgements
We thank CERCA Program / Generalitat de Catalunya for their institutional support. We also wish to thank all patients, families and members of the ICO Hereditary Cancer Program.
Funding
This study has been funded by Instituto de Salud Carlos III through the projects PI23/00017, PI19/00553, PI16/00563, IMP/00009, PID2019-111254RB-I00, PID2020-112595RB-I00, CIBERONC [CB16/12/00234] and PMP22/00064, implemented under the NextGenerationEU funds, which finance the actions of the Mecanismo de Recuperación y Resiliencia. Also supported by the Government of Catalonia (Secretariat for Universities and Research of the Department of Business and Knowledge; grant 2021SGR01112).
Author information
Authors and Affiliations
Contributions
E.M., P.R., L.F., J.V. and C.L. conceived, designed, and planned the study; E.M., P.R., M.M-C., X.M., O.C., A.A., M.A-B., B. M-B., E.N., G.V-P., R.C., J.M.M-C., D.C., M.P., L.F., J.V. and C.L. contributed to the acquisition, analysis, and/or interpretation of the molecular data; A.S. and M.S. provided samples and clinical data; E.M. and P.R. drafted the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The research was conducted in accordance with the principles of the Declaration of Helsinki, and ethical approval was obtained from the ethics committee of Bellvitge Biomedical Research Institute (IDIBELL; PR278/19). Informed written consent for both diagnostic and research purposes was obtained from all participants.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Munté, E., Rofes, P., Millán-Castillo, M. et al. Optimizing GRIDSS for clinical use: A targeted NGS filtering strategy for germline structural variant detection. Eur J Hum Genet (2026). https://doi.org/10.1038/s41431-026-02016-x
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41431-026-02016-x