Abstract
In the era of rapidly accumulating genomic data, largely driven by the broad use of whole-genome sequencing (WGS) in clinical settings, interpreting lesser-known genes with varied phenotypes remains challenging. PubMatcher is a new tool that simplifies bibliographic research for multiple genes at once and grants quick and easy access to relevant gene information. It helps users efficiently identify potential genotype-phenotype associations using PubMed complemented by additional data. By significantly reducing analysis time, PubMatcher supports the interpretation of novel or under-documented genes. Freely available for academic and non-commercial use, PubMatcher is a user-friendly and efficient solution for researchers, clinical scientists and clinical geneticists working on pan-genomics analyses.
Similar content being viewed by others
Introduction
Genomic sequencing advancements have led to an explosion of data, making the interpretation of variants in lesser-known genes a day-to-day challenge for geneticists. Key gene-phenotype associations often remain underrepresented in widely used databases involved in human disease, like Online Mendelian Inheritance in Man (OMIM) [1]. For example, OMIM may omit some gene-phenotype associations [2] or include them, but with an emphasis on symptoms different from those observed in some patients. To avoid this issue, PubMed or other databases can be useful to find the most relevant scientific publications regarding the link between a gene and a specific phenotype. This thorough approach to genomic data interpretation is time-consuming and potentially less accurate over time. This is especially true for whole-genome sequencing (WGS) analysis, where a significant number of variants located in non-OMIM morbid genes are retained by classical filters (such as “rare loss-of-function”, “rare homozygous missense for a recessive hypothesis”).
To address these challenges, we developed PubMatcher, a free online tool that simplifies the retrieval of gene-phenotype associations by querying multiple curated databases and PubMed simultaneously. PubMatcher uniquely supports batch format-free analysis, significantly reducing the time required to identify candidate genes relevant to a patient’s phenotype. We describe in this article the modus operandi of PubMatcher and its relevance in revealing non-obvious gene-phenotype associations.
Materials and methods
PubMatcher is a full-stack web application developed using Node.js version 18 [3, 4], with an Express.js backend and a Vue.js 3.5 frontend. Dependency management is handled via npm, and data persistence relies on a PostgreSQL 14 database. The application is fully containerized using Docker to ensure reproducibility and ease of deployment. Two types of inputs are required in Pubmatcher: one or more genes and one or more phenotypes (or relevant keywords) (Fig. 1). The ‘Extract from Text’ feature employs a client-side pattern-matching algorithm utilizing a cached dataset of HGNC symbols and aliases. The extraction logic prioritizes official symbols first, identifying matches within the input text using case-insensitive regular expressions with word boundary enforcement (\b). If an official symbol is not found, the algorithm scans for known aliases. Validated matches are automatically mapped to their current official HGNC symbol. To minimize false positives—a common challenge in text mining—aliases shorter than three characters are automatically excluded. Furthermore, the tool incorporates a user-controlled exclusion list (‘blacklist’) stored in the browser’s local storage, allowing users to persist the exclusion of specific problematic aliases or genes that may trigger false positives in their specific clinical context. The PubMatcher pipeline aggregates information using a hybrid approach: it performs real-time web scraping for PubMed (Keyword(s) + gene) and queries public APIs for UniProt [5], the International Mouse Phenotyping Consortium (IMPC) [6], and PanelApp [12]. To optimize performance, gnomAD constraint metrics [7], ClinVar, and Gene Curation Coalition (GenCC) data are accessed via locally stored datasets updated periodically, while API responses are managed with local caching strategies.
Search genes either manually or by using an “EXTRACT FROM TEXT” mode which consists of copy-pasting characters including gene names. Phenotypes can be incremented manually in the lower box, one or more, separated by commas. (version: January 2025).
The results page presents a summary of all the information collected (Fig. 2). Ensuring wide accessibility, PubMatcher is designed to be accessed via web browsers at https://pubmatcher.fr and the source code and documentation are available on GitHub (https://github.com/victormar1/PubMatcher). It does not require user registration, adhering to most journal’s guidelines for software tools.
(query: genes “DIDO1, MC4R, BRAT1, SLC12A5” and phenotype “obesity, diabetes”). Crosses indicate lack of information in gene. “GENE” column contains gene constraint metrics; “PUBMATCH” column contains the number of publications retrieved on PubMed and the title of the first publication; “FUNCTION” column contain the Uniprot function description of the protein and functional tags; “PHENOTYPE KO”column contains icons representing mouse symptoms after KO compiled within IMPC database; “CLINVAR LOOKUP” column contains the number and type (missense / loss-of-function) of either pathogenic / likely pathogenic variants or variants of unknown significance; “STATUS” column contains OMIM, Gene Curation Coalition and PanelApp England / Australia information.
Results are presented in an organized table format, where gene-phenotype pairs are listed with key metrics, such as constraint scores and publication count. The details of each query are described below.
Genes constraint metrics
PubMatcher obtains for each gene the following constraint metrics from the gnomAD v2.1 and v4 database [8] : pLi (probability of being loss-of-function intolerant), LOEUF (loss-of-function observed/expected upper bound fraction), MOEUF (missense observed/expected upper bound fraction) and missense Z-Score. LOEUF and MOEUF metrics indicate a gene’s tolerance to loss-of-function and missense variants, respectively, helping prioritize genes under selective constraint for clinical relevance. LOEUF and pLi values are highlighted based on constraint levels: dark red for low pLOEUF ( < 0,35 for v2 and <0,6 for v4) or high pLi ( > 0,9) and colored in green for low pLi ( < 0,1) [9]. Discrepancies between the two gnomAD versions are highlighted with an exclamation mark. To note, GnomAD v4 and v2 versions differ especially in terms of size, European ancestry proportions, technical and sequencing quality, cohort segmentation regarding the disease or non-disease status. Hence, metric values and interpretations of metrics may vary due to differences in cohort size (i.e., improve statistical power for same observed/expected ratio), the loci quality and also the different threshold recommended. Thus, concordance between metrics and improvement in statistical power for v4 could reinforce interpretation of gene constraint, but differences should be interpreted with caution.
PubMed
PubMed is a free online database providing access to a vast repository of biomedical research articles maintained by the National Center for Biotechnology Information (NCBI) and represents an “up-to-date” knowledge source for gene-phenotype associations [10]. PubMatcher includes the number of publications retrieved following a query, the title of the first publication in the list, and a link to access the query on PubMed and the related research articles. The PubMed research includes the association between a gene name and a phenotype. Moreover, the queries are cumulative for each gene-phenotype pair. An example of query is shown in Fig. 2, which includes five genes and two phenotypes. The PubMed query for each gene follows this pattern: (GENE AND PHENOTYPE_1) OR (GENE AND PHENOTYPE_2). Hovering over the title of the publication will display titles of other matching publications.
Uniprot
The UniProt database [5] provides information about protein functions, which may be relevant for genetic interpretation. PubMatcher requests the protein description and biological features keywords from UniProt using API access.
International mouse phenotyping consortium
The IMPC database [6] provides information about the consequences of gene knockouts in mice, which could suggest a gene’s involvement in human diseases. Different phenotypes are listed as presented on IMPC and specific symptoms can be displayed by mouseover.
Clinvar lookup
PubMatcher integrates data from ClinVar, a public database maintained by the NCBI that provides clinically relevant interpretations of genetic variants, including their pathogenicity, molecular consequences, and supporting evidence. For pathogenic and likely pathogenic small nucleotide variants, PubMatcher displays both the number of loss-of-function (LOF) variants—including frameshift, nonsense, and canonical splice site alterations—and the number of missense variants. Additionally, VUS are also reported to ensure no potentially relevant findings are overlooked.
Gene curation coalition, PanelApp & OMIM
PubMatcher integrates data from GenCC [11], PanelApp England and Australia [12], and OMIM [1] to provide comprehensive information on gene-disease associations, ensuring rapid and accurate curation of clinically relevant genes. GenCC aggregates gene-disease validity information from multiple expert-curated sources, facilitating the identification of genes with well-established evidence for their role in human diseases. PubMatcher displays the gene status from GeneCC. The number of genes listed in both PanelApp England and PanelApp Australia are mentioned in the PubMatcher output due to their significance in fast gene-disease curation. Links are provided for quick access to the relevant entries on the PanelApp websites. OMIM is a comprehensive, authoritative resource that catalogs human genes and genetic phenotypes, including their relationships to disease. PubMatcher integrates data from OMIM to indicate whether a gene is associated with a known morbid condition or phenotype.
Relevance of PubMatcher in human whole-genome sequencing analysis
We evaluated the relevance of the PubMatcher tool in WGS analyses of patients with rare diseases performed at the Auragen laboratory in Lyon, France. This laboratory is part of the French 2025 genomic project, which aims to expand genomic access in human healthcare [13, 14]. First, the proportion of variants filtered out by an example set of common WGS filters (detailed in Table S1) that were not located in OMIM morbid genes across 20 trio-based WGS analyses was assessed. Then, we present examples of variants revealed by PubMatcher in genes that proved potentially relevant for medical use after analyzing 100 WGS cases.
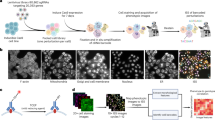
Whole-genome sequencing was performed following the recommendations of “France Médecine Génomique 2025” Plan. Genomic DNA extracted from whole blood was sequenced according to standard procedures for a Polymerase Chain Reaction-Free genome on a NovaSeq6000 instrument (Illumina, San Diego, California, USA). Sequencing data were aligned to the GRCh38p13 full assembly using bwa 0.7 + . Variants were called by several algorithms including GATK4 + , Bcftools1.10 + , Manta1.6 + , CNVnator0.4 + , and annotated using the variant effect predictor. Detected variants were prioritized using in-house procedures. Further details are available on request on http://www.auragen.fr.
Results
Variants in non-OMIM genes found by common WGS filters
PubMatcher is meant to quickly identify gene-phenotype associations using the most up-to-date sources. Although the OMIM database is regularly updated, the most recent phenotype-to-gene associations may be missing, potentially leading to the exclusion of relevant variants. Therefore, we evaluated the proportion of non-OMIM morbid genes in 20 WGS trios consisting of an affected patient and unaffected parents, using a classic filtering strategy (see Table S1 for filters’ details).
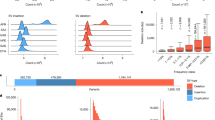
After applying these filters, the remaining variant counts ranged from 80 to 150 per sample, with a mean of 114. Among these, the mean proportions of variants mapping to OMIM morbid genes, OMIM non-morbid genes, and non-OMIM genes were 31%, 52%, and 18%, respectively (Fig. 3). These results confirm a high representation of non-morbid or non-OMIM genes (70%) post-filtering, underscoring the utility of PubMatcher for efficiently screening them.
Morbid: 30.7% (SD = 5.72), Non-Morbid: 51.3% (SD = 6.11), Non-OMIM: 18% (SD = 5.56).
Misannotated or non-annotated genes with relevant variants in 100 WGS analyses
We present examples of variants found in genes either not annotated in OMIM for the researched disease or with a non-syndromic form not specified in OMIM (Table 1). These relevant variants were identified in 15 out of 100 whole-genome sequences analyzed at the French laboratory Auragen (Lyon, France) for medical purposes. The genomes included in this study were selected solely based on their availability as trios and were analyzed in chronological order in those medical contexts: genodermatosis, chronic nephropathy, intellectual deficiency, or red blood cell diseases. None of those genomes were analyzed previously, hence they also include diagnosis on well know genes correctly annotated within OMIM database. Among the 15 genomes with relevant variants in incompletely annotated genes, only one also carried a pathogenic variant in a well-known gene (Table 1).
Integrating PubMatcher in genomic variant analysis workflows
PubMatcher is a tool that can be integrated early in the general workflow of genomic single nucleotide variant analysis. We propose a flowchart for data interpretation in a large-scale genomic approach (Fig. 4).
Proposed integration of PubMatcher in interpretation of pangenomic analysis.
Starting with a conventional filtering strategy (as described in Table S1), a rapid diagnosis can be made if a causative variant is identified—for example, a previously described ClinVar pathogenic variant that matches the patient’s clinical presentation. If such a variant is not found, a more thorough variant analysis is required to explore and report relevant genetic variants.
The tool can be used for gene screening across all identified variants, allowing for a quick exploration of the most recent scientific knowledge (via PubMed and PanelApp queries), gene constraint metrics, protein functions (Uniprot), and the consequences of mouse knockout models (IMPC). The Mode of Inheritance based on the family pedigree is also crucial. A recent publication from Chong et al. [9]. compiled five key criteria for retaining genes of interest, nearly all of which are integrated into the proposed flowchart that includes PubMatcher, except for gnomAD variant co-occurrence.
After analyzing the Pubmatcher output in the context of the patients’ phenotypes, some genes of interest may be retained. If the gene evidence level is sufficient, the variants located within can be interpreted via usual tools and reported if classified pathogenic or likely pathogenic (with additional exploration needed if it is a variant of unknown significance). Conversely, if the evidence level is low, a more research-focused approach, such as submitting to MatchMaker Exchange [15] (Genematcher, etc.) or conducting further fundamental post-genomic investigations, may be suggested.
Discussion
We believe PubMatcher is a significant advancement in clinical genomic research, addressing the need for more efficient interpretation of genomic data. By rapidly identifying relevant gene-phenotype associations—especially in lesser-known genes—PubMatcher increases both the speed and accuracy of genomic analyses. Indeed, the high proportion of candidates variants in filters located outside OMIM morbid genes (70%) is more quickly analyzed using PubMatcher than individual queries, because it offers a unique window to display all of this information for many genes. This approach also helps ensure that rare yet important variants are not missed, which is critical for their inclusion in broader research studies; given their rarity, these cases can provide invaluable insights into disease mechanisms and phenotypic diversity.
There are other tools which interrogate gene-function links such as Open Targets Platform. That tool, however, cannot query the most recent PubMed literature which is critical to obtain the most up-to-date information about gene-rare diseases relationship. For instance, the request of LEF1 gene (Table 1) does not rely with genodermatosis despite of articles published several years ago which demonstrated its involvement. Moreover, there is no easy-to-use batch mode available to analyze gene list. PubMatcher ’s main strength is its ability to display in one window the main information to screen the variants located on the non-OMIM morbid genes.
PubMatcher is a gene-level tool, which is complementary with variant-centered tools like Varsome or MobiDetails [16]. Indeed, WGS analysis in medical context needs to study gene and then variant relevance.
An important consideration is the inclusion of animal models, such as the mouse model, which provides invaluable insights into gene function and disease relevance due to its genetic similarity to humans. However, mouse models present limitations, including differences in gene expression and phenotypic responses. Expanding to other model organisms, such as zebrafish, could diversify the functional insights available to PubMatcher users, particularly for genes where murine models have limited general data or translational relevance.
The effectiveness of PubMatcher heavily depends on the quality and completeness of its external data sources. Attempts to incorporate alternative sources, such as Google Scholar, resulted in an overwhelming volume of unspecific and irrelevant data, highlighting PubMed as the most reliable and curated source for retrieving relevant literature. Advances in AI-driven text-mining tools, such as PubTator [17], offer promising avenues for improving data retrieval by extracting gene-disease relationships from biomedical literature. These tools could significantly enhance the exhaustivity of PubMatcher ’s results by identifying additional relevant publications that might otherwise remain undetected. However, current rate limitations (3 requests per second) within the PubTator API preclude its integration into PubMatcher at this stage.
PubMatcher has demonstrated effectiveness in identifying clinically relevant genes, thereby fulfilling its primary objective. Notably, several geneticists outside the development team have already integrated PubMatcher into their variant interpretation workflows, underscoring its reliability and practical utility and adaptability in clinical genomics. Further exploration of PubMatcher ’s applications in clinical settings could be beneficial. Another important consideration is the accessibility of the tool. While the current interface is user-friendly—particularly in term of input formatting, result clarity, and advanced features upon login (such as input history)—further simplifying the user experience and providing enhanced guidance and support would make the tool even more accessible to a wider audience.
Integrating artificial intelligence or machine learning could also boost PubMatcher ’s capabilities by adding features like gene scoring to rank the matches by their relevancy to the phenotype. Ongoing updates, as well as feedback from the user community, will be crucial for the tool’s continued development and for expanding its utility in the field of genomic research.
Conclusion
PubMatcher provides an effective solution for supporting genomic data analysis by seamlessly integrating bibliographic research into genomic interpretation workflows. This approach significantly enhances efficiency, particularly in identifying lesser-known yet clinically relevant gene-phenotype associations. As PubMatcher continues to evolve, improvements in data integration, interface design, and user-driven enhancements will further solidify its role as a valuable tool for both clinical diagnostics and genomic research.
Data availability
The human whole-genome sequencing data used in this study were obtained from the French national genomic medicine initiative, Plan France Médecine Génomique 2025 (PFMG2025). These sequencing data were generated and analyzed at the AURAGEN genomic sequencing center. Due to ethical and privacy restrictions, the raw sequencing data are not publicly available but as described in Abadie and al. (2025) [14], access request to molecular datasets can be found in online repositories on the website: https://pfmg2025.fr/ Additional data are available from the corresponding author on reasonable request.
Code availability
The source code for PubMatcher is freely available under the Massachusetts Institute of Technology (MIT) License. Project name: PubMatcher Project home page: https://github.com/victormar1/PubMatcher Operating system(s): Platform independent Programming language: JavaScript (Node.js & Vue.js). Other requirements: None License: Massachusetts Institute of Technology License. Any restrictions to use by non-academics: No specific restrictions, open-source license.
References
McKusick VA Mendelian Inheritance in Man. A catalog of human genes and genetic disorders. Johns Hopkins University Press. 1998;12.
Shakir A, Ripperger M, Jiang Z, Wierenga KJ. Inferred inheritance of MorbidMap genes without OMIM clinical synopsis. Genet Med. 2018;20:470–3.
Holowaychuk TJ tj/ejs [Internet]. 2024 [cité 22 avr 2024]. Disponible sur: https://github.com/tj/ejs.
nodejs/node [Internet]. Node.js; 2024 [cité 22 avr 2024]. Disponible sur: https://github.com/nodejs/node.
The UniProt Consortium. UniProt: the universal protein knowledgebase in 2023. Nucleic Acids Res. 2023;51:D523–31.
Groza T, Gomez FL, Mashhadi HH, Muñoz-Fuentes V, Gunes O, Wilson R, et al. The International Mouse Phenotyping Consortium: comprehensive knockout phenotyping underpinning the study of human disease. Nucleic Acids Res. 2023;51:D1038–45.
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alföldi J, Wang Q, et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581:434–43.
Chen S, Francioli LC, Goodrich JK, Collins RL, Kanai M, Wang Q, et al. A genomic mutational constraint map using variation in 76,156 human genomes. Nature. 2024;625:92–100.
Chong JX, Berger SI, Baxter S, Smith E, Xiao C, Calame DG, et al. Considerations for reporting variants in novel candidate genes identified during clinical genomic testing. Genet Med. 2024;26:101199.
Sayers EW, Bolton EE, Brister JR, Canese K, Chan J, Comeau DC, et al. Database resources of the national center for biotechnology information. Nucleic Acids Res. 2022;50:D20–6.
The Gene Curation Coalition: A global effort to harmonize gene-disease evidence resources - PubMed [Internet]. [cité 9 janv 2025]. Disponible sur: https://pubmed.ncbi.nlm.nih.gov/35507016/.
Martin AR, Williams E, Foulger RE, Leigh S, Daugherty LC, Niblock O, et al. PanelApp crowdsources expert knowledge to establish consensus diagnostic gene panels. Nat Genet. 2019;51:1560–5.
Sanlaville D, Vidaud M, Thauvin-Robinet C, Nowak F, Lethimonnier F. French Genomic Medicine Plan 2025 (PFMG2025): France enters the era of genomic medicine]. Rev Prat. 2021;71:1061–4.
Abadie C, Abderrahmane A, Abdous O, Abel C, Ackermann O, Acquaviva C, et al. PFMG2025–integrating genomic medicine into the national healthcare system in France. The Lancet Regional Health – Europe [Internet]. 2025 [cité 13 mars 2025];50. Disponible sur: https://www.thelancet.com/journals/lanepe/article/PIIS2666-7762(24)00352-1/fulltext.
Philippakis AA, Azzariti DR, Beltran S, Brookes AJ, Brownstein CA, Brudno M, et al. The matchmaker exchange: a platform for rare disease gene discovery. Hum Mutat. 2015;36:915–21.
Baux D, Van Goethem C, Ardouin O, Guignard T, Bergougnoux A, Koenig M, et al. MobiDetails: online DNA variants interpretation. Eur J Hum Genet. 2021;29:356–60.
Wei CH, Allot A, Lai PT, Leaman R, Tian S, Luo L, et al. PubTator 3.0: an AI-powered literature resource for unlocking biomedical knowledge. Nucleic Acids Res. 2024;52:W540–6.
Hassan A, Morice-Picard F, Marin V, Lasseaux Robine E, Lebreton L, Davaze-Schneider J. Hypohidrotic ectodermal dysplasia in a family: expanding spectrum of LEF1-related disorders. Clin Exp Dermatol. 2024;49:1725–6.
Dufour W, Alawbathani S, Jourdain AS, Asif M, Baujat G, Becker C, et al. Monoallelic and biallelic variants in LEF1 are associated with a new syndrome combining ectodermal dysplasia and limb malformations caused by altered WNT signaling. Genet Med. 2022;24:1708–21.
Lévy J, Capri Y, Rachid M, Dupont C, Vermeesch JR, Devriendt K, et al. LEF1 haploinsufficiency causes ectodermal dysplasia. Clin Genet. 2020;97:595–600.
De Franco E, Wakeling MN, Frew RD, Russ-Silsby J, Peters C, Marks SD, et al. A biallelic loss-of-function PDIA6 variant in a second patient with polycystic kidney disease, infancy-onset diabetes, and microcephaly. Clin Genet. 2022;102:457–8.
Al-Fadhli FM, Afqi M, Sairafi MH, Almuntashri M, Alharby E, Alharbi G, et al. Biallelic loss-of-function variant in the unfolded protein response gene PDIA6 is associated with asphyxiating thoracic dystrophy and neonatal-onset diabetes. Clin Genet. 2021;99:694–703.
Münch J, Engesser M, Schönauer R, Hamm JA, Hartig C, Hantmann E, et al. Biallelic pathogenic variants in roundabout guidance receptor 1 associate with syndromic congenital anomalies of the kidney and urinary tract. Kidney Int. 2022;101:1039–53.
Christians A, Kesdiren E, Hennies I, Hofmann A, Trowe MO, Brand F, et al. Heterozygous variants in the DVL2 interaction region of DACT1 cause CAKUT and features of Townes-Brocks syndrome 2. Hum Genet. 2023;142:73–88.
Yan H, Shi Z, Wu Y, Xiao J, Gu Q, Yang Y, et al. Targeted next generation sequencing in 112 Chinese patients with intellectual disability/developmental delay: novel mutations and candidate gene. BMC Med Genet. 2019;20:80.
Ha T, Morgan A, Bartos MN, Beatty K, Cogné B, Braun D, et al. De novo variants predicting haploinsufficiency for DIP2C are associated with expressive speech delay. Am J Med Genet A. 2024;194:e63559.
Smits DJ, Schot R, Popescu CA, Dias KR, Ades L, Briere LC, et al. De novo MCM6 variants in neurodevelopmental disorders: a recognizable phenotype related to zinc binding residues. Hum Genet. 2023;142:949–64.
Azad P, Caldwell AB, Ramachandran S, Spann NJ, Akbari A, Villafuerte FC, et al. ARID1B, a molecular suppressor of erythropoiesis, is essential for the prevention of Monge’s disease. Exp Mol Med. 2022;54:777–87.
Shen Y, Bassett MA, Gurumurthy A, Nar R, Knudson IJ, Guy CR, et al. Identification of a novel enhancer/chromatin opening element associated with high-level γ-globin gene expression. Mol Cell Biol. 2018;38:e00197-18.
Werren EA, Peirent ER, Jantti H, Guxholli A, Srivastava KR, Orenstein N, et al. Biallelic variants in CSMD1 are implicated in a neurodevelopmental disorder with intellectual disability and variable cortical malformations. Cell Death Dis. 2024;15:379.
Boonsawat P, Asadollahi R, Niedrist D, Steindl K, Begemann A, Joset P, et al. Deleterious ZNRF3 germline variants cause neurodevelopmental disorders with mirror brain phenotypes via domain-specific effects on Wnt/β-catenin signaling. Am J Hum Genet. 2024;111:1994–2011.
Gordon PM, Efthymiou S, Salpietro V, Fielding T, Borgione E, Scuderi C, et al. Human patient SFPQ homozygous mutation is found deleterious for brain and motor development in a zebrafish model [Internet]. bioRxiv; 2020 [cité 3 nov 2024]. p. 2020.03.18.993634. Disponible sur: https://www.biorxiv.org/content/10.1101/2020.03.18.993634v1.
Abramson J, Adler J, Dunger J, Evans R, Green T, Pritzel A, et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature. 2024;630:493–500.
Schmid CM, Gregor A, Costain G, Morel CF, Massingham L, Schwab J, et al. LHX2 haploinsufficiency causes a variable neurodevelopmental disorder. Genet Med. 2023;25:100839.
Acknowledgements
This research was made possible through access to the data generated by the 2025 French Genomic Medicine Initiative.
Funding
No financial assistance was received in support of the study.
Author information
Authors and Affiliations
Contributions
VM conceptualized the project, developed the software, performed data analyses, and wrote the main manuscript text and figures. HL and VD contributed to the development of the software used in the work. JT provided data access, provided scientific guidance, and have substantively revised the manuscript. DB assisted with software development, provided scientific guidance, and have substantively revised the manuscript. A-FR contributed scientific expertise and have substantively revised the manuscript. EL helped with conceptualization, promoted our work, provided scientific feedback, and have substantively revised the manuscript. PP contributed to study design, offered scientific input, and have substantively revised the manuscript. LL co-conceived the project, supervised the research, and co-wrote the manuscript. All authors reviewed and approved the final version of the manuscript and agree to be accountable for the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study involved genomic analyses conducted as part of routine clinical care for patients with rare diseases in France. As such, a clinical trial registration was not required, since all data reported were obtained during standard diagnostic procedures. In accordance with the French Bioethics Law (Law No. 2004-800, dated August 6, 2004), all patients provided written informed consent for diagnostic procedures and were specifically informed that any remaining biological material could be used for research purposes. The retrospective use of these data was approved by the Bordeaux University Hospital under registration number CHUBX2025RE0134.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Independent Researcher: Hugo Lannes, Victor Dumont.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Marin, V., Lannes, H., Dumont, V. et al. PubMatcher: a web app to support genomic data interpretation through simplified bibliographic research. Eur J Hum Genet (2026). https://doi.org/10.1038/s41431-026-02068-z
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41431-026-02068-z