Abstract
Epitranscriptomic modifications are evolutionarily conserved changes to RNA and are present in all RNA types, including mRNA. Next to methylation, pseudouridine is also a prevalent modification found in mRNA and other RNA types. Pseudouridylation of RNA improves RNA stability, RNA stacking and translation fidelity. While m6A RNA methylation has been demonstrated to regulate the activation of various immune cell types, such as T cells, dendritic cells or NK cells, insight into the role of pseudouridine in the immune system is limited. Trub1 is one of 13 pseudouridine synthases (PUS) and has been suggested to co-install the majority of pseudouridine residues on mRNA alongside the activity of PUS7 and has been reported to install pseudouridine in tRNA. Against this background, we hypothesized that Trub1 may regulate immune cell development, homeostasis or functional differentiation. Using conditional and global Trub1-deficient mice, in vivo competition in mixed bone marrow chimaeras and high-parametric flow cytometry, we interrogated the role of Trub1 in T cells, B cells, NK cells and myeloid cells. Despite our rigorous analyses and the identification of Trub1-dependent pseudouridine residues in mitochondrial tRNAs, we did not identify a significant role for Trub1-mediated pseudouridylation in immune cells at steady state or during disease. We hence reason that the role of Trub1 in mRNA pseudouridylation has previously been overestimated in computational studies or that co-installations of pseudouridine at other positions in the same RNA molecules may compensate for the lack of Trub1-installed modifications.
Similar content being viewed by others
Introduction
Pseudouridylation, the process of converting uridine (U, 1-\(\beta\)-D-ribofuranosyluracil) into pseudouridine (Ψ, 5-\(\beta\)-D-ribofuranosyluracil), is one of the most prevalent post-transcriptional modifications among over 170 different types of RNA modifications [1, 2]. The presence of pseudouridine was first reported in ribosomal RNA (rRNA), transfer RNA (tRNA), small nuclear (snRNA) and subsequently in messenger RNA (mRNA) [3,4,5]. Owed to technological and methodological developments over the past decade, more than 9500 Ψ modification sites have been identified or computationally proposed in various RNA types across all organisms [6, 7]. Pseudouridylation is catalysed by 13 non-redundant pseudouridine synthases (PUS), a highly conserved family of enzymes critical in ensuring the proper function of RNA molecules [8, 9]. Pseudouridine synthases function either as ‘stand-alone’ writers [10] such as Trub1 or PUS7 or through an RNA-dependent mechanism involving H/ACA snoRNAs [11, 12]. The installation of pseudouridine facilitates RNA stability, splicing and translation fidelity by promoting structural rearrangements that enhance the function and integrity of various RNA species [13, 14].
Among the 13 PUS, Trub1 and PUS7 have been described to be together responsible for approximately 70% of all Ψ installations in mammalian mRNA [15, 16]. PUS7 is involved in a variety of biological processes and mutations have been demonstrated to be disease-causing in humans. Mutations in PUS7 impair tRNA and mRNA pseudouridylation and cause neurological disorders such as intellectual disability, developmental delay, microcephaly and autism spectrum disorders [17,18,19,20]. PUS7 has been found to regulate the activation of the PI3K/AKT/mTOR and Wnt/\(\beta\)-catenin signalling pathways in colorectal cancer cells [21, 22]. High expression of PUS7 also promotes cancer progression and poor prognosis in patients with other types of cancer, including non-small cell lung cancer, hepatocellular carcinoma, ovarian cancer and glioblastoma [23,24,25,26]. Moreover, normal human hematopoietic multilineage differentiation has been shown to be governed by the activity of PUS7 [27]. The function of Trub1, on the other hand, remains less understood. Recent research indicated the important role of Trub1 in tRNA and mRNA structures, which mediate RNA stability and function, contributing to the regulation of translation [28, 29]. Trub1-dependent pseudouridylation enhances the maturation of the let-7 microRNA family, a microRNA family well-known for its role in oncogenesis regulation, in which, for example, the KRAS proto-oncogene is regulated via let-7 [30, 31]. Kurimoto et al. revealed that knocking down Trub1 increases KRAS expression, while overexpressing Trub1 protein decreases KRAS expression and suppresses cell proliferation, effects that can be partially rescued by depletion of the let-7 family [31]. Further, low expression of Trub1 is also correlated with poor prognosis in renal clear carcinoma patients [32]. These insights emphasise the need for further research to fully elucidate the function of Trub1 and its potential implications in human health and disease.
The immune system is a complex network of various, functionally distinct cell types, with the ability to detect pathogens and to launch an effective immune response to eliminate pathogenic threats. The immune system fulfils all requirements for the necessity of fast-acting, dynamic regulatory mechanisms. Developing from hematopoietic stem cells via committed progenitors in different organs, immune cells are functionally very diverse and, importantly, plastic [33,34,35]. Further, immune challenges require fast functional differentiation and cell migration controlled at a molecular level by DNA modifications, RNA modifications (as shown for RNA m6A methylation [36,37,38,39]), transcriptional rewiring and extensive proteomic changes [40,41,42,43]. In line with these considerations, here we investigated a potential role of Trub1 in development, homeostasis and function of immune cell subsets, including T cells, B cells, NK cells and myeloid cells. We generated conditional and global Trub1-deficient mice and employed mixed bone marrow chimaeras to study the role of Trub1 in immune cells at steady state, in competition and during disease. Despite previous data suggesting relevance of Trub1 in mRNA stability in human cells [44] and despite describing Trub1-mediated pseudouridylation in several mitochondrial tRNAs, our data support the conclusion that the PUS Trub1 alone is insufficient to substantially regulate transcriptional and translational rewiring during development and differentiation of immune cells.
Material and methods
In silico analysis
Data on Trub1 mRNA expression across different immune cell subsets were obtained as expression values from Gene Skyline hosted by immgen.org.
Animals
Congenic CD45.1 mice (B6.SJL-Ptprca.Pepcb/BoyJ) were purchased from Jackson Laboratories (strain 002014) and crossed to Foxp3Thy1.1 reporter mice [45]. Trub1 genetrap heterozygous frozen embryos (C57BL/6N-Trub1tm1a(EUCOMM)Hmgu) were purchased from the MMRRC repository (UC Davis, University of California, CA, USA), then bred to mice with constitutive expression of flippase (B6.SJL-Tg(ACTFLPe)9205Dym/J, strain 005703, Jackson Laboratories) to create the conditional Trub1 allele (short Trub1flox). Trub1flox/+ mice were bred to CD4Cre transgenic mice for T cell-specific Trub1 deficiency (Tg(Cd4-cre)1Cwi [46], C57BL/6J background, Trub1Tcell-KO for short) or Foxp3CreYFP knockin mice for Treg-specific Trub1 deficiency (C57BL/6.129(Cg)-Foxp3tm4(Cre)Ayr/J [47], Trub1Foxp3-KO for short). Of note, the Foxp3YFPCre knock-in allele has been described to display promiscuous Cre recombinase activity for some conditional alleles [48]. In our mouse colony, we identified germline Cre recombinase activity resulting in mice with global Trub1 deficiency (Trub1All-KO for short).
Ethical approval
All methods were performed in accordance with the relevant guidelines and regulations. Specifically, sample sizes for mouse experiments were chosen based on power calculations and pilot data, in conjunction with the Animal Ethics Committee, to allow for robust sensitivity without excessive animal use. Mice were used at 8-12 weeks of age. Both male and female mice were used in the experiments. Mice were housed in specific pathogen-free conditions under a 12-h light/dark cycle in a temperature- and humidity-controlled room. Age- and sex-matched animals were used for each experiment. Animals were also co-housed when possible. No other randomization procedures were used. All animal procedures were approved by the KU Leuven Animal Ethics Committee (approvals: 150/2019, 008/2019, 228/2024, 74/2018, 060/2024).
Generation of mixed bone marrow chimera mice
Bone marrow was extracted from CD45.2 Trub1CD4/Foxp3/ALL-KO, Trub1WT or CD45.1 wild-type mice by crushing the femurs with a mortar and pestle. Cells were then resuspended in Tris-NH4Cl red blood cell (RBC) lysis buffer to lyse the RBCs. Mixed bone marrow chimeric mice were generated by co-transplanting an equal mixture of bone marrow from CD45.2 Trub1CD4/Foxp3/ALL-KO mice with CD45.1 wild-type mice (KO: WT). To control for strain differences, mice were also co-transplanted with an equal mixture of bone marrow from CD45.2 Trub1WT and CD45.1 wild-type mice (WT: WT). A total of 20 × 106 cells were injected intravenously into lethally irradiated mice (9.5Gy gamma-irradiation). Chimeric mice were analysed 8-10 weeks post-transplantation.
RNA isolation, reverse transcription and quantitative PCR
Total RNA was isolated from spleens and lymph nodes from Trub1-deficient mice and wild-type C57BL/6J mice using TRIzol reagent (Invitrogen). 1 μg of RNA was reverse-transcribed using Maxima H Minus First Strand cDNA Synthesis Kit (Thermo Scientific). Real-time qPCR was performed using Fast SYBR Green Master Mix (Applied Biosystems) with a StepOnePlus Real-Time PCR System (Applied Biosystems), according to the manufacturer’s instructions. The β-actin transcript was used as an internal control to normalise the transcript levels of Trub1. The following primers were used for qPCR reactions: Trub1-forward: 5’-TGAACCGGCTGAAGGAGAAG-3’, Trub1-reverse: 5’-CCAACCACCAGAACACCTTG-3’, β-actin-forward: 5’-CATCACTATTGGCAACGAGC-3’, β-actin-reverse: 5’-ACGCAGCTCAGTAACAGTCC-3’. Technical duplicates, as well as Reverse Transcriptase minus and no template control reactions, were performed alongside (data not shown). Relative quantification of Trub1 expression was evaluated using the ΔΔCt method [49]. The fold-change in the relative gene expression was determined by calculating the 2−ΔΔCt value.
Flow cytometry and cell sorting
Antibodies and fixable viability dye were purchased from eBioscience, Biolegend or BD Biosciences as indicated below.
Splenocytes were resuspended with Tris-NH4Cl red blood cell (RBC) lysis buffer to lyse the RBCs. Cells were then washed twice with PBS, stained with fixable viability dye (eBioscience) and, where applicable, stained for anti-CD69 (H1.2F3, Biolegend) at 4 °C for 20 min prior to fixation. Cells were then fixed using eBioscience™ Foxp3/Transcription Factor fixation/permeabilization kit (#00-5521-00, eBioscience) according to the manufacturer’s instructions. Cells were stained overnight at 4 °C with the panels covering: anti-CD45 (30-F11), anti-CD3 (145-2C11, 17A2,), anti-CD19 (B4), anti-CD11c (N418), anti-MHCII (M5/114.15.2), anti-CD172a (P84), anti-CD64 (X54-5/7.1), anti-PDCA-1 (927), anti-XCR-1 (ZET), anti-Ly6G (1A8), anti-CD44 (IM7), anti-ICOS (C398.4 A), anti-PD-1 (29 F.1A12), anti-KLRG1 (2F1/KLRG1), anti-IL-2 (JES6-5H4), anti-IL-4 (11B11), anti-IL-22 (Poly5164), anti-TGF-β (TW7-15B4), anti-TNF-α (MP6-XT22), anti-CD127 (S18006K), anti-Neuropilin-1 (3E12), anti-FoxP3 (FKJ-16s) (all from Biolegend); anti-CD3 (KT3), anti-CD19 (B4,6D5), anti-CD4 (RM4-5) (all from Bio-Rad); anti-CD4 (GK1.5), anti-CD103 (M290), anti-Ly6C (AL-21, anti-Siglec-F (E50-2440), anti-CD117 (c-kit) (2B8), anti-CD8α (53-6.7), anti-IgE (R35-72), anti-CD11b (M1/70), anti-CD95 (Jo2), anti-TCR-β (H57-597), anti-CD62L (MEL-14), anti-CTLA4 (UC10-4F10-11), anti-CD127 (SB/199), anti-GATA-3 (L50-823), anti-CD25 (PC61), anti-T-bet (4B10), anti-GITR (DTA-1), anti-IL-17 (TC11-18H10), anti-IL-6 (MP5-20F3), anti-IL11β (B122), anti-IFN-γ (XMG1.2), anti-RORγt (Q31-378) (all from BD Biosciences); anti-IgM (II/41), anti-NK1.1 (PK126), anti-Eomes (Dan11mag), anti-Neuropilin-1 (3DS304M), anti-ST2 (RMST2-2), anti-IL-10 (JES5-16A3), anti-IL-3 (MP2-8F8), anti-IL-13 (eBio13A), anti-IL-21 (FFA21), anti-RORγt (B2D), anti-Ki-67 (SolA15), anti-Helios (22F6), anti-GATA-3 (TWAJ), anti-IL11β (NJTEN3), anti-Ki-67 (16A8, SolA15) (all from eBioscience); anti-FoxP3 (REA788), anti-Helios (REA829) (both from Miltenyi Biotec); anti-Amphiregulin (polyclonal), anti-IL-5 (TRFK5) (both from R&D Systems).
To assess cytokine production, cells were stimulated in the presence of phorbol myristate acetate (50 ng/mL), ionomycin (1 µg/mL) and brefeldin A (2 µg/mL) (all Tocris Biotechne) for 4 h at 37 °C in RPMI supplemented with 10% of fetal bovine serum, 0.1 mM of non-essential amino acid, 50 UI/mL of penicillin G, 5 μg/mL streptomycin, 1 mM sodium pyruvate, 10 μg/mL β-mercaptoethanol (all from GIBCO, Thermo Fisher Scientific). Cells were then stained with fixable viability dye, followed by fixation with 2% Formaldehyde (VWR chemicals, Radnor, Pennsylvania, PA, USA) in PBS at room temperature for 30 min and permeabilized with 1x eBioscience permeabilization buffer (#00-8333-56, eBioscience) according to the manufacturer’s instructions prior to antibody staining.
Data were acquired on the BD FACSymphony A5 (BD Biosciences) and Aurora spectral analyser (Cytek). Data was analysed using FlowJo (v10.9.0, BD Biosciences) and FCS Express (v7.28.0019, De Novo Software). Median Fluorescence Intensity (MFI) is reported on marker-positive populations exclusively. Samples were excluded when the number of cells in the parent population was less than 100 or when the frequency of positive cells was less than 0.01%. In these cases, figure legends indicate where populations were removed from data presentation.
For cell sorting, splenic CD4 T cells were purified using the MojoSort™ Mouse CD4 T Cell Isolation Kit (Biolegend) according to the manufacturer’s instructions, followed by surface staining with anti-CD4 (GK1.5, BD Biosciences), anti-CD62L (MEL-14, BD Biosciences), anti-CD44 (IM7, Biolegend) and anti-CD90.1/Thy1.1 (HIS51, eBioscience) (when using congenic Foxp3Thy1.1 reporter mice) or CD25 (PC61, eBioscience) when using CD45.2 mice carrying conditional or knockout Trub1 alleles. DAPI (Thermo Scientific) was added for dead cell exclusion. Naïve CD4 T cells (gated as Thy1.1neg or CD25neg non-Treg) were defined as CD4+ CD44neg CD62L+ Thy1.1neg (or CD25neg). Treg were defined as CD4+ Thy1.1neg (or CD25neg) and post-sort stained for Foxp3 for additional purity confirmation. Cell sorting was performed on a BD FACSAria III instrument (BD Biosciences).
In vitro polarisation assay
96-well plates were pre-coated either overnight at 4 °C or 1 h at 37 °C with 5 µg/mL anti-CD3ε (145-2C11, Invitrogen) and 5 µg/mL anti-CD28 (37.51, Invitrogen). After washing the wells with PBS, 2 × 105 sort-purified naïve CD4 T cells were added and activated for 5 days in RPMI 1640 medium (Lonza) supplemented with 10% of fetal bovine serum, 0.1 mM of non-essential amino acid, 50 UI/mL of penicillin G, 5 μg/mL streptomycin, 1 mM sodium pyruvate, 10 μg/mL β-mercaptoethanol (all from GIBCO, Thermo Fisher Scientific). The following cytokine/neutralizing antibody cocktails were added to induce T helper cell polarization: TH1: IL-12p70 (10 ng/mL), IL-2 (1 ng/mL); TH2: anti-IFN-γ (10 μg/mL, XMG1.2, Invitrogen), IL-4 (10 ng/mL), IL-2 (1 ng/mL); TH17: anti-IFN-γ (10 μg/mL, XMG1.2, Invitrogen), IL-6 (40 ng/mL), TGF-β (1 ng/mL); Treg: IL-2 (20 ng/mL), TGF-β (5 ng/mL), anti-IFN-γ (10 µg/mL, XMG1.2, Invitrogen).
In vitro suppression of conventional T cells by Treg
Tregs from Trub1Foxp3-KO or from wild-type littermates were sort-purified as stated above. Conventional T cells (Tconv, CD4+ Thy1.1neg) were sort-purified from spleens of congenic CD45.1 Foxp3Thy1.1 reporter mice and were stained with CellTraceTM Violet (CTV, Thermo Fisher Scientific) as per the manufacturer’s protocol. 1 × 105 labelled Tconv were cultured in a U-bottom 96-well with CTV-unlabelled Tregs at different Treg: Tconv ratios (1:1, 1:2, 1:4, 1:8) in the presence of anti-CD3ε (1 μg/mL, 145-2C11, Invitrogen) and splenocytes from Rag1–/– mice (5 × 104 cells). After five days of co-culture, the cells were stained with fixable viability dye (eBioscience) and anti-CD45.1 (A20, eBioscience) and data were acquired on a BD FACSymphony A5 (BD Biosciences). Proliferation of CD45.1 Tconv was evaluated via dilution of CTV and labelled Tconv cultured for 5 days in the absence of Tregs were used as a control for proliferation.
MC38 tumour mouse model
The colon adenocarcinoma MC38 cell line was kindly provided by Prof. Gabriele Bergers, VIB-KU Leuven, Belgium and maintained in DMEM (Gibco) supplemented with 10% fetal bovine serum (Thermo Fisher), 1% glutamine, 100 U/mL penicillin and 100 U/mL streptomycin (All Gibco). 5 × 105 cells were injected subcutaneously in the left and right flanks of Trub1ALL-KO or wild-type littermates. Tumour width and length were followed at day 10, 12, 14 and 17 post-injection and tumour volume was calculated as: (width)2 x length x 0.52. For the measurement of tumour width and length, the operator was blinded to the experiment groups and the genotype of the animals. Mice were sacrificed 17 days after injection and for flow cytometry analysis, tumours, inguinal (tumour-draining), mesenteric, brachial and axillary lymph nodes (non-tumour draining) were isolated. Leukocytes from the lymph nodes were isolated via mechanical dissociation using frosted glass slides and were filtered through a 70 μm mesh. To obtain tumour-infiltrating leukocytes, tumours from both the left and right flanks were pooled and finely chopped. The pieces were transferred to a gentleMACSTM C Tube (Miltenyi Biotec) containing the enzyme mix with enzymes A, D and R (Tumour Dissociation Kit, Miltenyi Biotec). The enzyme mix was prepared as per the manufacturer’s instructions. The tissue was dissociated using the soft/medium program on a gentleMACSTM Octo Dissociator (Miltenyi Biotec). The single cell suspension was then filtered through a 70 μm mesh and RBC lysis was performed as described above. Tumour-infiltrating lymphocytes were further enriched via density gradient centrifugation using 40% v/v and 80% v/v Percoll (Cytiva). Antibody staining for flow cytometry-based analysis of the leukocytes and for assessing the cytokine production was performed as described above.
Influenza mouse model
Trub1KO or Trub1Tcell-KO mice or wild-type littermates received intranasal instillation of 10 plaque-forming units (PFU) of mouse-adapted influenza virus A/H3N2/Ishikawa/7/82 or PBS (10 μl per nostril) under inhalation anaesthesia with isoflurane (1.5–2% in 100% oxygen, Piramal Critical Care). Body weight and clinical score based on body weight, respiratory parameters and condition (adapted from [50]) were followed daily for up to 7 days post-inoculation. For the body weight measurement and evaluation of clinical score, the operator was blinded to the experiment groups and the genotype of animals. Mice were sacrificed 7 days after influenza infection. Spleen and bronchoalveolar lavage (BAL) fluid were collected. BAL fluid was collected after three lung lavages with 700 μL 0.9% NaCl (B. Braun). Immunophenotyping was performed on the cells obtained from BAL fluid and spleen via high-parameter flow cytometry as detailed above.
Micro-computed tomography (μCT) for influenza-infected mice
A whole-body small animal μCT scanner (SkyScan 1278, Bruker micro-CT. Kontich, Belgium) was used to acquire lung μCT data with the following parameters: 50kVP X-ray source, 1 mm aluminium X-ray filter, 350 μA current, 150 ms exposure time per projection, 0.9° increments over a total angle of 220° (adapted from [50]). µCT data was reconstructed, visualised and segmented using NRecon, DataViewer and CTan software, all provided by the manufacturer. Non-aerated lung volume (NALV), which is the imaging-derived biomarker of pulmonary lesions, was quantified based on the manual delineation of a region of interest (ROI), which resulted in a volume of interest (VOI) covering the entire lung and the predetermined threshold of 108-225 greyscale (adapted from [50]). For the imaging and visualisation, the operator was blinded to the experiment groups and the genotype of the animals.
Statistics
Statistical analyses were performed using GraphPad Prism, version 9.0 (GraphPad Software, San Diego, CA, USA) with a non-parametric Mann–Whitney U test due to non-normal data distribution. For the statistical analysis of clinical score progression, body weight loss and non-aerated lung volume in the influenza model and for tumour volume progression in the MC38 tumour model, two-way ANOVA with Tukey’s multiple comparison was used. p < 0.05 was considered significant. When linear mixed-effects models (LMMs) were applied and significance was assessed using ANOVA with F-tests, this is mentioned in the respective figure legend.
Results
Normal development and in vitro function of Treg in the absence of Trub1
The differential expression of Trub1 in peripheral- and thymic-induced Tregs (Fig. 1A) implored us to investigate the role of Trub1 in Treg development, function and activation. To achieve a Treg-specific knockout of Trub1, we intercrossed mice carrying a conditional knockout allele of Trub1 (Trub1flox) with Foxp3YFP-Cre mice, generating Trub1Foxp3-KO mice (Fig. 1B). To assess the role of Trub1 in Treg induction, we induced Treg in vitro (iTreg) from sort-purified naïve CD4 T cells from Trub1Foxp3-KO mice and wild-type littermates. The induction efficiency of iTreg was found to be similar in both groups, indicating that Treg induction is independent of Trub1 (Fig. 1C), as reduced Trub1 levels in peripherally-induced Treg may suggest (Fig. 1A). Furthermore, the suppressive capacity of Trub1-deficient Treg was assessed in vitro. To this end, congenic CD45.1+ conventional CD4 T cells were activated in the presence of Trub1-deficient Tregs (from Trub1Foxp3-KO mice) or Trub1-sufficient Tregs (from Trub1WT mice). The proliferation of conventional CD4 T cells remained comparable in cultures with Trub1-deficient Treg versus wild-type Treg at different Treg: Tconv ratios, indicating that the suppressive potential of Tregs is not hampered by Trub1 deficiency (Fig. 1D). Together, these findings indicate that Trub1-mediated pseudouridylation is dispensable for in vitro Treg induction and suppressive function.
A Expression of Trub1 mRNA compared via publicly available RNAseq data (immgen.org) between colonic Treg (enriched for peripherally-induced Treg, pTreg) and splenic Treg (enriched for thymus-derived Treg, tTreg) from 6-week-old C57BL/6 J mice. pTreg were defined as CD4+ FoxP3+ Nrp1lo whereas tTreg were defined as CD4+ FoxP3+ CD25hi. Expression values were normalised by DESeq2 on immgen.org. B Breeding scheme to obtain mice with conditional Trub1 deletion in Treg or all T cells. C In vitro Treg induction (iTreg) and D Treg-mediated suppression of CD4 Tconv, wild-type littermates used as control (n = 3 for Trub1WT and n = 3 for Trub1Foxp3-KO, with three technical replicates). Flow cytometric analysis of mBMC (n = 5 for WT: WT, n = 5 for KO: WT). E Frequency of CD4+ FoxP3+ Tregs of total CD4 T cells compared among CD45.2 Trub1WT or CD45.2 Trub1Foxp3-KO cells. F Frequency of CD45.2 Trub1WT or CD45.2 Trub1Foxp3-KO Tregs expressing surface markers and transcription factors CD25, CD69, CTLA-4, GATA-3, GITR, Helios, ICOS, Ki-67, Neuropilin-1 (Nrp-1), PD-1 and RORγt. G Frequency of Tregs of Trub1Foxp3-KO or Trub1WT origin expressing cytokines AREG (Amphiregulin), IL-2, IL-10, TGF-β and TNF-α. Data are plotted as mean ± SD and p values are calculated via an unpaired Mann–Whitney U test. CD45.2 wild-type bone marrow donors refer to Foxp3WT Trub1WT littermates. Refer to Fig. S2A for the gating strategy used for the analysis of surface markers, transcription factors and cytokines and refer to Fig. S3 for the median fluorescence intensity (MFI) of surface markers, transcription factors and cytokines presented in the figure.
Next, we sought to identify cell-intrinsic defects in Treg due to the absence of Trub1 in vivo. To this end, we generated 50:50 mixed bone marrow chimeras (mBMC) from congenically disparate mice, i.e., using bone marrow from CD45.2 Trub1Foxp3-KO mice and CD45.1 wild-type mice (KO: WT chimeras). To control for potential strain differences, we also co-transplanted bone marrow cells from CD45.2 Trub1Foxp3-WT (wild-type littermates of Trub1Foxp3-KO mice) and CD45.1 wild-type (WT: WT chimeras). We assessed the relative frequency of Tregs originating from wild-type or knockout CD45.2 donors and observed that Trub1 deficiency did not yield a competitive disadvantage (Fig. 1E). The expression of various markers on Tregs for activation and suppressive potential (surface and transcription factors) was then assessed employing flow cytometry and the frequency of marker-positive cells as well as per cell expression level was evaluated (Figs. 1F and S2A; S3A). Similarly, we compared cytokine production (Figs. 1G and S3B). We found that all assessed parameters in Trub1-deficient and -sufficient Tregs were comparable, altogether suggesting that deficiency of Trub1-mediated pseudouridylation upon expression of FoxP3 does not affect Treg function or homeostasis at steady state.
Normal development and differentiation of T cells in the absence of Trub1
We analysed publicly available RNAseq data (immgen.org) to assess Trub1 expression profiles across various T cell subsets and observed a dynamic expression of Trub1 across T cell differentiation, spanning from double-negative (DN) thymocytes to splenic naïve and activated CD4 and CD8 T cells (Fig. 2A). To elucidate the role of Trub1-mediated pseudouridylation in CD4 and CD8 T cell homeostasis and function, we generated T cell-specific Trub1-deficient mice (Trub1Tcell-KO for short) by crossing Trub1flox mice with CD4Cre transgenic mice (Fig. 1B). In these mice, Trub1 is conditionally ablated in all T cells from the thymic CD4+CD8+ double-positive developmental stage. First, we compared the capability of Trub1-deficient naïve CD4 T cells to differentiate into the CD4 T helper subsets TH1, TH2 and TH17 to that of naïve CD4 T cells from wild-type mice (Trub1WT, Trub1wtCD4cre) (Fig. 2B). As Trub1 mRNA levels seem decreased upon T cell activation (Fig. 2A), we also overexpressed Trub1 in CD4 T cells to investigate whether higher Trub1 levels impair or promote T helper cell polarization (Fig. S4A, B). TH subsets were defined based on the expression of a respective hallmark cytokine or the hallmark transcription factor. The efficiency of in vitro polarization into each TH subset was comparable between Trub1-deficient and -sufficient CD4 T cells, as well as comparing Trub1WT and Trub1OE CD4 T cells.
A Expression of Trub1 mRNA by mouse CD4neg CD8neg double-negative thymocyte subsets (DN1, DN2a, DN2b, DN3, DN4), double-positive thymocytes (CD4/CD8 DP), CD4 and CD8 single-positive thymocytes (CD4 SP and CD8 SP, respectively), as well as splenic naïve (nCD4) and activated (actCD4) CD4 T cells and naïve (nCD8), activated (actCD8) and central-memory (cmCD8) CD8 T cells. B In vitro T helper cell polarization from naïve T cells from Trub1Tcell-KO mice or wild-type littermates. T helper cell subsets were characterised via cytokine production (left) and hallmark transcription factor expression (right) (n = 7 for wild-type, n = 6 for Trub1Tcell-KO). C–H Flow cytometric analysis of mBMC. Frequency of naïve (nCD4/nCD8, CD62L+ CD44neg), activated (act, CD62Lneg CD44+) and central-memory (cm, CD62L+ CD44+) in (C) CD4 T cells and D CD8 T cells of WT: WT (dark grey, Trub1WT: CD45.1WT) and KO: WT (light grey, Trub1Tcell-KO: CD45.1WT) mBMC. Frequency of cell expressing the surface markers or transcription factors KLRG1, CD69, CD127, CTLA-4, Eomes, GATA-3, Helios, ICOS, Ki-67, Neuropilin-1 (Nrp-1) and PD-1 compared in E CD4 or F CD8 T cells from CD45.2 wild-type littermates in WT: WT mBMC (dark grey) and from Trub1Tcell-KO mice in KO: WT mBMC (light grey). Frequency of cytokine-producing CD45.2 wild-type T cells in WT: WT mBMC (dark grey) and CD45.2 Trub1Tcell-KO T cells in KO: WT (light grey). G CD4 T cells and H CD8 T cells producing the cytokines Amphiregulin (AREG), IL-2, IL-4, IL-10, IL-17, IFN-γ, TGF-β and TNF-α. For (C-H) n = 4 for WT: WT and n = 5 for KO: WT. Filled and unfilled symbols are representative of two independent experiments; wild-type littermates are CD4-Cre+Trub1WT. Data are plotted as mean ± SD. p-values for in vitro assays are calculated via unpaired Mann–Whitney U test and for mBMC experiments via F-test using mixed linear model. p-values reporting potential statistical significance (F-test p < 0.05) are not reported when similar trends were observed in the respective CD45.1 WT cells in these bone marrow chimeras. Wild-type littermates for in vitro assays are CD4-Cre+Trub1WT. Refer to Fig. S2B for the gating strategy used for the analysis of surface markers, transcription factors and cytokines. Refer to Fig. S4A, B for in vitro T helper cell polarization in Trub1OE CD4 T cells compared to Trub1WT CD4 T cells. Refer to Fig. S4C for surface marker and transcription factor expression within naïve, activated and central-memory CD4 T cells and CD8 T cells of WT:WT vs KO:WT mBMC defined in Fig. 2E, D. Refer to Fig. S4D, E for the median fluorescence intensity (MFI) of surface markers, transcription factors and cytokines presented in Fig. 2E–H.
Next, we investigated whether Trub1 deficiency causes cell-intrinsic defects in T cell development or homeostasis. To this end, we generated 50:50 mBMC from CD45.2+ Trub1Tcell-KO mice. As for Trub1Foxp3-KO 50:50 mBMC, also here, to control for strain differences between the Trub1Tcell-KO strain and the CD45.1 congenic wild-type strain (KO: WT), mBMC from wild-type littermates (Trub1Tcell-WT) and congenic CD45.1 mice were created (WT: WT). High-parameter flow cytometry was then used to analyse CD4 and CD8 T cell populations in these chimeras (Figs. 2C–H and S2B, S4C–E). Trub1-deficient T cells did not display significant shifts in frequencies of naïve, activated and central memory CD4 and CD8 T cells compared to the wild-type counterparts from the same strain (Fig. 2C, D). Furthermore, we neither observed changes in the expression of various markers of cell states and activation nor in the expression of several cytokines between CD4 and CD8 T cells originating from Trub1WT and Trub1Tcell-KO (Fig. 2E–H). Also, within naïve, effector or central-memory subsets of CD4 and CD8 T cells, we did not observe changes in the expression of markers indicative of cell states and activation (Fig. S4C). This was also the case for per cell levels of all analysed surface proteins, transcription factors and cytokines in CD4 and CD8 T cells (Fig. S4D, E). In all, this suggests that Trub1-mediated pseudouridylation is dispensable for CD4 or CD8 T cell function, homeostasis or competitive fitness.
Trub1 mediates site-specific pseudouridylation of mitochondrial tRNA
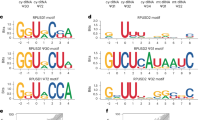
Previous studies, including our own, have reported that Cre recombinase expression from the FoxP3CreYFP knock-in allele can exhibit leakage, leading to sporadic recombination of floxed alleles in non-Treg cells. This is likely due to low-level expression from the Foxp3 locus during progenitor stages [48]. Here, promiscuous germline expression of Cre recombinase in Trub1Foxp3-KO mice generated a mouse strain with global deficiency of Trub1 (Trub1All-KO). Trub1 deficiency was confirmed by RT-qPCR, Western Blotting and HydraPsiSeq (Fig. S5). Profiles for NormUscore demonstrate that the absence of Trub1 markedly increases hydrazine-mediated cleavage at U55 in mt tRNAs, attesting to the absence of modification in the knock-out. While it has been reported previously that mammalian tRNAs (nuclear, cytoplasmic and mitochondrial) are pseudouridylated at different positions [51, 52], we identify here Trub1-mediated pseudouridylation in 7 mouse mitochondrial (mt) tRNAs, despite Trub1 being reported as primarily present in the nucleus [16, 28]. Interestingly, human TRUB1 has been reported to introduce the highly conserved pseudouridine in position 55 of four mt tRNAAsn, tRNAGlu, tRNAGln and tRNAPro required for efficient base-pairing and mitochondrial translation [29]. In mice, Trub1’s role in mt tRNA pseudouridylation was not previously studied and here we extend the list of mouse substrates for 7 mt tRNAs: tRNALeu, tRNAGln, tRNAAsn, tRNAIle, tRNAMet, tRNASer and tRNATyr.
Trub1 deficiency permits normal Bcell and myeloid cell development
In Trub1All-KO mice, we next explored the impact of Trub1 deficiency on B cells and myeloid cells. We analysed the frequencies of B cell subsets, conventional dendritic cells (cDC), NK and NKT cells, monocytes, basophils, eosinophils, neutrophils and macrophages, compared to wild-type littermates (Fig. 3A, B). Additionally, we also interrogated the cytokine production capacity of CD11b+ myeloid cells and CD19+ B cells, which appeared comparable among wild-type and Trub1-deficient mice at steady state (Fig. 3C, D).
Ex vivo flow cytometry analysis of Trub1ALL-KO and wild-type littermates. A Comparison of the frequency of CD19+ B cells and germinal centre B cells (CD95+ CD19+) in spleen and B frequency of NK and myeloid cell subsets. Frequency of cytokine-producing splenocytes among C CD11b+ myeloid cells and D CD19+ B cells (n = 5 for Trub1ALL-KO, n = 4 for Foxp3-CrenegTrub1WT wild-type littermates). Flow cytometry analysis of mixed bone marrow chimeras (mBMC) created with bone marrow cells of Trub1ALL-KO and CD45.1 wild-type mice (KO: WT) or of Trub1WT mice and CD45.1 wild-type mice (WT: WT). Shown are data of Trub1ALL-KO cells from KO: WT mBMC vs Trub1WT cells from WT: WT mBMC. E Frequencies of CD19+ B cells and germinal centre B cells (CD95+ CD19+). F Frequencies of myeloid cells and innate lymphocytes. Frequencies of cytokine-producing cells among G CD11b+ myeloid cells and H CD19+ B cells. For mBMC, n = 3 for WT: WT and n = 4 for KO: WT. Data are plotted as mean ± SD and p values are calculated via an unpaired Mann–Whitney U test. Wild-type littermates are Foxp3-CrenegTrub1WT. MΦ=macrophages (CD19neg CD11b+ CD64+), NK=natural killer cells (CD19neg CD64neg NK1.1+), NKT=natural killer T cells (CD3+ NK1.1+), cDC=conventional dendritic cells (NK1.1neg Ly-6neg CD11c+ MHC-II+), type 1 (cDC1) and type 2 (cDC2) are defined as XCR-1+ CD11c+ MHCII+ and CD172a+ CD11c+ MHCII+ respectively, Mono=monocytes (NK1.1neg Ly-6C+), Neu=neutrophils (NK1.1neg Ly-6G+), Eos=eosinophils (Siglec-F+), Bas=basophils (Ly-6neg CD11cneg MHC-IIneg c-kitneg IgE+). Refer to Fig. S2C for the gating strategy. Refer to Fig. S6 for expression levels of cytokines and flow cytometric analyses of B cells and myeloid cell subsets in lymph nodes and Peyer’s patches of the mBMC mice.
Subsequently, we sought to determine whether cell-intrinsic defects due to the absence of Trub1 in B cells and myeloid cells are masked in Trub1All-KO mice through compensatory mechanisms. Therefore, we generate 50:50 mBMC from CD45.2 Trub1All-KO mice, similar to those generated for Trub1Foxp3-KO and Trub1Tcell-KO. To control for strain differences between the Trub1All-KO strain and the congenic wild-type strain, bone marrow from wild-type littermates of CD45.2 Trub1All-KO mice was co-transplanted with congenic CD45.1 wild-type bone marrow. We evaluated the relative presence of B cell subsets and myeloid cell subsets in spleen, mesenteric and peripheral lymph nodes and Peyer’s patches (Figs. 3E, F and S6A–C) as well as cytokine expression profiles of CD19+ B cells and CD11b+ myeloid cells in spleen (Figs. 3G, H and S6D, E). Overall, Trub1 deficiency did not impact the frequency of myeloid and B cells or the production of various cytokines, neither intrinsically nor extrinsically, suggesting that it has no regulatory function in immune cell development or homeostasis in healthy mice.
Trub1 deficiency does not impair T cell-mediated antiviral response in influenza-infected mice
During an immune challenge, T cells undergo extensive proteomic and transcriptional rewiring to mount an effective immune response [51, 53]. We hypothesized that impaired RNA pseudouridylation may impact such rewiring, potentially impairing T cell activation, polarization or function in vivo during infection-induced immune responses. Here, we investigated the role of Trub1 in mounting an antiviral response by inoculating Trub1ALL-KO or Trub1Tcell-KO mice and their respective wild-type littermates with influenza A H3N2 virus and monitored weight loss along with clinical scoring daily for 7 days post-inoculation (Figs. 4A and S7, 8). We observed that the clinical scores evidenced severe influenza infection and were comparable between Trub1-deficient mice and their wild-type counterparts and both groups showed similar weight loss (Trub1WT = 11.84% ± 8.08% or Trub1ALL-KO = 14.16% ± 5.23% weight loss between d0 and d7) (Fig. 4B). Non-aerated lung volume (NALV) determined through μCT showed a significant increase in influenza-inoculated mice when compared to non-infected mice indicative of a similar lung pathology due to influenza infection (Fig. 4C). Using high-parameter flow cytometry, we performed immunophenotyping on the infiltrating T cells in the bronchoalveolar lavage (BAL) fluid and spleen. Total cell numbers of CD4 Tconv cells, CD8 T cells and Tregs appeared comparable in the BAL fluid of infected mice (Fig. S7A). Indicative of mounting an antiviral immune response upon infection, T cells in the BAL fluid of both infected Trub1ALL-KO and wild-type mice displayed a strong increase in CD8 T cells compared to mock-infected mice (Fig. 4D). This immune response was local as splenic T cells did not differ in the ratio of CD4 to CD8 T cells (Fig. 4D). Next, we measured the expression of several intracellular and surface proteins as well as cytokines with roles in activation, polarization or function in CD8 T cells, CD4 Tconv and Treg in BAL fluid (Fig. 4E–J). First, we assessed frequencies of naïve, activated and central-memory CD4 Tconv and CD8 T cells in BAL fluid of infected mice and found these comparable between Trub1ALL-KO and wild-type mice (Fig. S7B). Among CD4 and CD8 T cells, the expression of various interrogated markers and cytokines appeared normal in Trub1-deficient mice (Figs. 4E–J and S7C–E). This analysis was extended to the respective naïve, activated and central-memory subsets of CD4 Tconv and CD8 T cells, revealing no differences in the marker and cytokine expression in these subsets between Trub1-deficient and wild-type mice (Fig. S7F, G). Similar data were obtained upon influenza infection of Trub1Tcell-KO mice and respective wild-type littermates (Fig. S8). Hence, data from high-parameter flow cytometry combined with the clinical scoring suggest that Trub1 does not significantly impact the ability of mice to mount an antiviral response against influenza.
A Schematic overview of the timeline of the disease model. Trub1ALL-KO mice (n = 6) or wild-type littermates (n = 7) were infected with influenza A H3N2 or Trub1KO/WT littermates were mock-inoculated with PBS (n = 3). Clinical score and body weight were followed until the experimental endpoint of day 7. B Clinical score evolution and body weight loss compared to baseline. C μCT-derived biomarker of pneumonia non-aerated lung volume (NALV) compared in the three groups. D CD4-to-CD8 T cell ratio in the spleen and BAL fluid compared among the three groups. E–J Expression of surface markers and transcription factors CD69, CD103, CD127, CTLA-4, GATA-3, GITR, Helios, ICOS, Ki-67, KLRG1, Neuropilin-1 (Nrp1), PD-1, ST2 and T-bet along with cytokine production of Amphiregulin (AREG), IFN-γ, IL-1β, IL-2, IL-3, IL-4, IL-6, IL-10, IL-13, IL-17, IL-22, TGF-β and TNF-α compared between the BAL fluid of influenza-infected Trub1ALL-KO (n = 6) and Trub1WT (n = 7) mice in (E, F) CD8 T cells (for cytokine expression: n = 6 for Trub1WT), (G, H) conventional CD4 T cells (Tconv) and (I, J) Tregs (n = 6 for Trub1WT). Data are plotted as mean ± SD and p values are calculated via an unpaired Mann–Whitney U test. For the statistical analysis of clinical score progression and body weight loss, two-way ANOVA with Tukey’s multiple comparison was used and ns signifies p value > 0.05 in the data and **** p < 0.0002. Wild-type littermates are Foxp3CrenegTrub1WT, whereas the mock-inoculated controls are Foxp3Cre+Trub1WT. Samples are excluded from analysis if the number of events in the parent population is less than 100. Refer to Fig. S2B for the gating strategy used for the analysis of surface markers, transcription factors and cytokines. Refer to Fig. S7 for detailed analysis of cell numbers, surface markers, transcription factors and cytokines in naïve, activated and central memory T cell subsets and per cell expression levels and to Fig. S8 for the analysis of influenza infection in Trub1Tcell-KO vs Trub1WT mice.
Trub1 deficiency does not affect anti-tumour response in the MC38 tumour model
Finally, we employed Trub1All-KO mice and wild-type littermates to assess a possible role of Trub1-mediated pseudouridylation in another immune response context, i.e., the immunogenic MC38 tumour model. Upon inoculation with tumour cells, tumour volume was measured on days 10, 12, 14 and 17 and final tumour weight was assessed on day 17. Both parameters were comparable between Trub1-sufficient and -deficient mice (Fig. 5A, B). Next, to investigate whether Trub1 deficiency causes perturbation in tumour-infiltrating lymphocytes (TILs), we used flow cytometry to evaluate the frequency and composition of TILs as well as of tumoral myeloid cell subsets along with the subsets in tumour-draining and non-tumour-draining lymph nodes. The frequencies of Tregs, naïve (CD62L+) and antigen-experienced (CD62Lneg) CD4 and CD8 T cells were similar between wild-type and Trub1ALL-KO mice in tumours and lymph nodes (Fig. 5C). Moreover, expression of various markers associated with cell state and activation in Tregs, CD4 and CD8 T cells within the tumours, the tumour-draining and non-tumour draining lymph nodes were not affected by Trub1 deficiency (Figs. 5D–F and S9A). This was also consistent across naïve (CD62L+) and antigen-experienced (CD62Lneg) subsets of CD4 and CD8 T cells (Fig. S9B, C). Finally, the frequency of various B, NK and myeloid cell subsets in tumours was also found to be similar between Trub1All-KO and wild-type mice (Fig. 5G). Hence, the data obtained through phenotyping, combined with mean tumour volumes, indicates that Trub1 deficiency neither impairs nor enhances anti-tumour immune response against MC38 tumours.
A Tumour volume of MC38 tumours was assessed in Trub1ALL-KO and Trub1WT mice at days 10, 12, 14 and 17 after inoculation. B Tumour weight was compared between Trub1ALL-KO and Trub1WT mice at the experimental endpoint of 17 days. C Frequencies of CD62L+ and CD62Lneg CD4 T cells and CD8 T cells along with Tregs were measured in tumour-infiltrating lymphocytes (TILs), tumour-draining lymph nodes (DLNs) and non-tumour draining lymph nodes (NDLN) and compared between Trub1ALL-KO (light grey) and Trub1WT mice (dark grey). Frequency of cells expressing the surface markers and transcription factors CD103, CTLA-4, Eomes, PD-1, KLRG1, Helios, T-bet, GITR, CD69, Ki-67 in D conventional CD4 T cells, E CD8 T cells and F Tregs in TILs and DLNs of Trub1ALL-KO and Trub1WT mice. G Frequency of myeloid cell subsets, innate lymphocytes and B cells in TILs of MC38 tumours in Trub1ALL-KO and Trub1WT mice (NK=natural killer cells, CD19neg CD64neg NK1.1+; NKT=natural killer T cells, CD3+ NK1.1+; cDC=conventional dendritic cells, NK1.1neg Ly-6neg CD11c+ MHC-II+; Neu=neutrophils, NK1.1neg Ly-6G+; Eos=eosinophils, Siglec-F+; Bas=Basophils, Ly-6neg CD11cneg MHC-IIneg c-kitneg IgE+). For all data, n = 4 for both groups Trub1ALL-KO and Trub1WT. Data are plotted as mean ± SD and p values are calculated via unpaired Mann–Whitney U test. For the statistical analysis of tumour-volume progression, two-way ANOVA with Tukey’s multiple comparison was used and ns signifies p value > 0.05. Wild-type littermates are Trub1flox/WT. Refer to Fig. S2B for the gating strategy used for the analysis of surface markers, transcription factors and cytokines in T cells and refer to Fig. S2C for the gating strategy for the analysis of myeloid cells, innate lymphocytes and B cells. Refer to Fig. S9 for further flow cytometry analysis on CD4 and CD8 T cell subsets.
Discussion
Epitranscriptomic modifications and in particular METTL3-mediated RNA m6A methylation have been shown to play a significant role in immune cell activation, including T cells, dendritic cells, myeloid cells and NK cells and to play a role in the enhancement of Treg function [38, 52, 54,55,56,57,58,59]. We hypothesized that apart from m6A, other epitranscriptomic modifications such as pseudouridylation may regulate transcriptomic and proteomic rewiring during immune cell development and functional differentiation—processes requiring substantial molecular adaptation. In this study, we focused on the impact of deleting the pseudouridine synthase Trub1 on immune cell development, differentiation and function.
Trub1 has been suggested to be a key mRNA modifier, responsible for the installation of the majority of mRNA pseudouridines alongside PUS7 [15]. Unlike most other pseudouridine synthases, Trub1 has a well-defined consensus sequence with a hairpin structure [16]. Through computational analysis, we identified 1152 perfectly matched sites with a 4-basepair stem present in 719 mouse transcripts or 4188 sites perfectly matched to a 3-basepair stem present in 2351 mouse transcripts (data not shown). PUS activities are thought to be compartmentalised and Trub1 has been described to be mostly located in the nucleus [16]. Nevertheless, nuclear human TRUB1 has been described to have Ψ55 activity for cytoplasmic tRNAs (redundant with PUS10) and for mitochondrial (mt) tRNAs, thereby enhancing translation fidelity and mitochondrial oxidative phosphorylation [29, 52]. The importance of Ψ55 for tRNA structure and function is illustrated by its conservation from bacteria and yeast to human cells. Here, we identified that, also in mice, the installation of Ψ55 in several mt tRNAs is Trub1-dependent. While human TRUB1 can also install Ψ55 in cytoplasmic tRNA, it shares this activity with PUS10. Of note, in mice we did not observe changes in Ψ55 in cytoplasmic tRNAs upon Trub1 deficiency (data not shown) and it is hence attractive to speculate that also in mouse cells Trub1 and PUS10 compete for the same binding site in cytoplasmic tRNAs. The expression level of pseudouridine synthases is further likely to regulate the level of pseudouridine installation and notably, Trub1 is differentially expressed in immune cell subsets and during the cellular development of T cells. To test our hypothesis, we studied major immune subsets in secondary lymphoid organs—including T cells, B cells, dendritic cells, macrophages and NK cells ex vivo, in vitro and in competitive mixed bone marrow chimaeras.
Deficiency of Trub1 in all T cells or specifically in Treg neither altered the development nor homeostasis of conventional CD4 or CD8 T cells, nor of FoxP3+ Treg. This was further confirmed in competitive in vivo conditions in which Trub1-deficient and -sufficient T cells were found comparable with no apparent alterations in subset frequencies or expression of markers indicative of differentiation, activation or function. As healthy mice display low numbers of CD4 T helper subsets, we interrogated the ability of Trub1-deficient and -sufficient conventional CD4 T cells to differentiate into T helper cells (TH1, TH2 and TH17) and in vitro-induced Treg (iTreg). The polarization into either T helper population from Trub1-deficient CD4 T cells was comparable to that of the Trub1-sufficient counterpart. Of note, comparable polarization capacity in Trub1-overexpressing CD4 T cells further substantiates the conclusion that Trub1-mediated pseudouridylation is dispensable for in vitro T helper subset induction. While in vitro polarisations under strong polarization stimuli may mask subtle effects, we could also not find evidence for impaired T cell function in two in vivo disease models, namely a murine influenza model and an immunogenic subcutaneous cancer model. We therefore conclude that Trub1 activity in the presence of other pseudouridine synthases is not playing a non-redundant, critical role in T cell development, homeostasis and functional differentiation.
To extend our findings, we utilised Trub1ALL-KO mice and assessed the competitive fitness of Trub1-deficient and -sufficient B cells, NK cells and CD11b+ myeloid cells in mixed bone marrow chimaeras. We found that neither immune cell type was impacted in its development or homeostasis by the deficiency of Trub1 and all analyzed parameters were comparable between Trub1-deficient and -sufficient cells. These data, together with our data in T cells, question a relevant role for Trub1 in immune cells, particularly in the presence of other pseudouridine synthases. While individual pseudouridine synthase possesses unique RNA tropism, overlapping targeting of the same (m)RNA molecule by multiple synthases may compensate for the lack of modifications of individual uridines without major negative effects for the RNA life cycle, such as for stability or translation efficacy. Notably, the previously suggested role of Trub1 as a major mRNA modifier has been contested by a subsequent study [54]. Our findings align with the possibility of a potentially less prominent role of Trub1 in mRNA pseudouridylation. Nevertheless, pseudouridine is a highly prevalent RNA modification present in all RNA types, including mRNA, although to a lesser extent in the latter than m6A methylation. Further, pseudouridine synthases, including Trub1, are highly conserved enzymes. Hence, overlapping targeting may prevent loss of function consequences of individual co-targeting synthases. Double-deficiencies of Trub1 and other PUS distorting mRNA life cycle or Trub1 and PUS10 distorting translation efficacies may result in defects in development, homeostasis or function of immune cells and future research addressing co-deficiencies is necessary to confirm such a hypothesis and the (redundant) roles of individual pseudouridine synthases in immune cells.
Data availability
Sequencing data for HydraPsiSeq and the NormUscores have been deposited at Zenodo with the following (https://doi.org/10.5281/zenodo.18214254). The HydraPsiSeq bioinformatic pipeline can be accessed via GitHub (https://github.com/FlorianPichot/HydraPsiSeqPipeline) [60].
Code availability
Sequencing data for HydraPsiSeq and the NormUscores have been deposited at Zenodo with the following (https://doi.org/10.5281/zenodo.18214254). The HydraPsiSeq bioinformatic pipeline can be accessed via GitHub (https://github.com/FlorianPichot/HydraPsiSeqPipeline) [60].
References
Cohn WE, Volkin E. Nucleoside-5′-phosphates from ribonucleic acid. Nature. 1951;167:483–4.
Cappannini A, Ray A, Purta E, Mukherjee S, Boccaletto P, Moafinejad SN, et al. MODOMICS: a database of RNA modifications and related information. 2023 update. Nucleic Acids Res. 2024;52:D239–44.
Reddy R, Ro-Choi TS, Henning D, Shibata H, Choi YC, Busch H. Modified nucleosides of nuclear and nucleolar low molecular weight ribonucleic acid. J Biol Chem. 1972;247:7245–50.
Nurse K, Wrzesinski J, Bakin A, Lane BG, Ofengand J. Purification, cloning and properties of the tRNA psi 55 synthase from Escherichia coli. RNA. 1995;1:102–12.
Schwartz S, Bernstein DA, Mumbach MR, Jovanovic M, Herbst RH, León-Ricardo BX, et al. Transcriptome-wide mapping reveals widespread dynamic-regulated pseudouridylation of ncRNA and mRNA. Cell. 2014;159:148–62.
Zhang M, Jiang Z, Ma Y, Liu W, Zhuang Y, Lu B, et al. Quantitative profiling of pseudouridylation landscape in the human transcriptome. Nat Chem Biol. 2023;19:1185–95.
Sun WJ, Li JH, Liu S, Wu J, Zhou H, Qu LH, et al. RMBase: a resource for decoding the landscape of RNA modifications from high-throughput sequencing data. Nucleic Acids Res. 2016;44:D259–65.
Li X, Ma S, Yi C. Pseudouridine: the fifth RNA nucleotide with renewed interests. Curr Opin Chem Biol. 2016;33:108–16.
Roovers M, Hale C, Tricot C, Terns MP, Terns RM, Grosjean H, et al. Formation of the conserved pseudouridine at position 55 in archaeal tRNA. Nucleic Acids Res. 2006;34:4293–301.
Rintala-Dempsey AC, Kothe U. Eukaryotic stand-alone pseudouridine synthases—RNA modifying enzymes and emerging regulators of gene expression? RNA Biol. 2017;14:1185–96.
Ganot P, Bortolin ML, Kiss T. Site-specific pseudouridine formation in preribosomal RNA is guided by small nucleolar RNAs. Cell. 1997;89:799–809.
McMahon M, Contreras A, Ruggero D. Small RNAs with big implications: new insights into H/ACA snoRNA function and their role in human disease. Wiley Interdiscip Rev RNA. 2015;6:173–89.
Borchardt EK, Martinez NM, Gilbert WV. Regulation and function of RNA pseudouridylation in human Cells. Annu Rev Genet. 2020;54:309–36.
Rodell R, Robalin N, Martinez NM. Why U matters: detection and functions of pseudouridine modifications in mRNAs. Trends Biochem Sci. 2024;49:12–27.
Carlile TM, Martinez NM, Schaening C, Su A, Bell TA, Zinshteyn B, et al. mRNA structure determines modification by pseudouridine synthase 1. Nat Chem Biol. 2019;15:966–74.
Safra M, Nir R, Farouq D, Vainberg Slutskin I, Schwartz S. TRUB1 is the predominant pseudouridine synthase acting on mammalian mRNA via a predictable and conserved code. Genome Res. 2017;27:393–406.
Han ST, Kim AC, Garcia K, Schimmenti LA, Macnamara E, Network UD, et al. PUS7 deficiency in human patients causes a profound neurodevelopmental phenotype by dysregulating protein translation. Mol Genet Metab. 2022;135:221–9.
Shaheen R, Tasak M, Maddirevula S, Abdel-Salam GMH, Sayed ISM, Alazami AM, et al. PUS7 mutations impair pseudouridylation in humans and cause intellectual disability and microcephaly. Hum Genet. 2019;138:231–9.
Darvish H, Azcona LJ, Alehabib E, Jamali F, Tafakhori A, Ranji-Burachaloo S, et al. A novel PUS7 mutation causes intellectual disability with autistic and aggressive behaviors. Neurol Genet. 2019;5:e356.
Naseer MI, Abdulkareem AA, Jan MM, Chaudhary AG, Alharazy S, AlQahtani MH. Next-generation sequencing reveals novel homozygous frameshift in PUS7 and splice acceptor variants in the AASS gene leading to intellectual disability, developmental delay, dysmorphic features and microcephaly. Saudi J Biol Sci. 2020;27:3125–31.
Du J, Gong A, Zhao X, Wang G. Pseudouridylate synthase 7 promotes cell proliferation and invasion in colon cancer through activating PI3K/AKT/mTOR signaling pathway. Dig Dis Sci. 2022;67:1260–70.
Zhang Q, Fei S, Zhao Y, Liu S, Wu X, Lu L, et al. PUS7 promotes the proliferation of colorectal cancer cells by directly stabilizing SIRT1 to activate the Wnt/β-catenin pathway. Mol Carcinog. 2023;62:160–73.
Zhang G, Zhu Y, Tan Y, Chen B, Shan S, Zhang G, et al. Higher expression of pseudouridine synthase 7 promotes non-small cell lung cancer progression and suggests a poor prognosis. J Cardiothorac Surg. 2023;18:222.
Cui Q, Yin K, Zhang X, Ye P, Chen X, Chao J, et al. Targeting PUS7 suppresses tRNA pseudouridylation and glioblastoma tumorigenesis. Nat Cancer. 2021;2:932–49.
Li H, Chen L, Han Y, Zhang F, Wang Y, Han Y, et al. The identification of RNA modification gene PUS7 as a potential biomarker of ovarian cancer. Biology. 2021;10:1130.
Jin Z, Song M, Wang J, Zhu W, Sun D, Liu H, et al. Integrative multiomics evaluation reveals the importance of pseudouridine synthases in hepatocellular carcinoma. Front Genet. 2022;13:944681.
Guzzi N, Cieśla M, Ngoc PCT, Lang S, Arora S, Dimitriou M, et al. Pseudouridylation of tRNA-derived fragments steers translational control in stem cells. Cell. 2018;173:1204–16.e26.
Mukhopadhyay S, Deogharia M, Gupta R. Mammalian nuclear TRUB1, mitochondrial TRUB2 and cytoplasmic PUS10 produce conserved pseudouridine 55 in different sets of tRNA. RNA. 2021;27:66–79.
Jia Z, Meng F, Chen H, Zhu G, Li X, He Y, et al. Human TRUB1 is a highly conserved pseudouridine synthase responsible for the formation of Ψ55 in mitochondrial tRNAAsn, tRNAGln, tRNAGlu and tRNAPro. Nucleic Acids Res. 2022;50:9368–81.
Xuan Y, Wang L, Zhang L, Lv M, Li F, Gong Q. Structural basis of pri-let-7 recognition by human pseudouridine synthase TruB1. Biochem Biophys Res Commun. 2024;721:150122.
Kurimoto R, Chiba T, Ito Y, Matsushima T, Yano Y, Miyata K, et al. The tRNA pseudouridine synthase TruB1 regulates the maturation of let-7 miRNA. EMBO J. 2020;39:e104708.
Yu Qxin. The prognostic significance of TRUB1 in patients with renal clear cell carcinoma. Asian J Surg. 2024;47:1143–5.
Graf T. Differentiation plasticity of hematopoietic cells. Blood. 2002;99:3089–101.
Margraf A, Perretti M. Immune cell plasticity in inflammation: insights into description and regulation of immune cell phenotypes. Cell. 2022;11:1824.
Ogawa M, LaRue AC, Mehrotra M. Plasticity of hematopoietic stem cells. Best Pr Res Clin Haematol. 2015;28:73–80.
Batista PJ, Molinie B, Wang J, Qu K, Zhang J, Li L, et al. m6A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell. 2014;15:707–19.
Perlegos AE, Shields EJ, Shen H, Liu KF, Bonini NM. Mettl3-dependent m6A modification attenuates the brain stress response in Drosophila. Nat Commun. 2022;13:5387.
Qin Y, Li B, Arumugam S, Lu Q, Mankash SM, Li J, et al. m6A mRNA methylation-directed myeloid cell activation controls progression of NAFLD and obesity. Cell Rep. 2021;37:109968.
Di Timoteo G, Giuliani A, Setti A, Biagi MC, Lisi M, Santini T, et al. M6A reduction relieves FUS-associated ALS granules. Nat Commun. 2024;15:5033.
Cui L, Ma R, Cai J, Guo C, Chen Z, Yao L, et al. RNA modifications: importance in immune cell biology and related diseases. Sig Transduct Target Ther. 2022;7:334.
Zha L, Wang J, Cheng X. The effects of RNA methylation on immune cells development and function. FASEB J. 2022;36:e22552.
Belliveau NM, Footer MJ, Akdoǧan E, Van Loon AP, Collins SR, Theriot JA. Whole-genome screens reveal regulators of differentiation state and context-dependent migration in human neutrophils. Nat Commun. 2023;1 4:5770.
Elsabbagh RA, Rady M, Watzl C, Abou-Aisha K, Gad MZ. Impact of N6-methyladenosine (m6A) modification on immunity. Cell Commun Signal. 2022;20:140.
Dai Q, Zhang LS, Sun HL, Pajdzik K, Yang L, Ye C, et al. Quantitative sequencing using BID-seq uncovers abundant pseudouridines in mammalian mRNA at base resolution. Nat Biotechnol. 2023;41:344–54.
Liston A, Nutsch KM, Farr AG, Lund JM, Rasmussen JP, Koni PA, et al. Differentiation of regulatory Foxp3+T cells in the thymic cortex. Proc Natl Acad Sci USA. 2008;105:11903–8.
Lee PP, Fitzpatrick DR, Beard C, Jessup HK, Lehar S, Makar KW, et al. A critical role for Dnmt1 and DNA methylation in T cell development, function and survival. Immunity. 200;15:763–74.
Rubtsov YP, Rasmussen JP, Chi EY, Fontenot J, Castelli L, Ye X, et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity. 2008;28:546–58.
Franckaert D, Dooley J, Roos E, Floess S, Huehn J, Luche H, et al. Promiscuous Foxp3-cre activity reveals a differential requirement for CD28 in Foxp3+ and Foxp3− T cells. Immunol Cell Biol. 2015;93:417–23.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001;25:402–8.
Seldeslachts L, Vanderbeke L, Fremau A, Reséndiz-Sharpe A, Jacobs C, Laeveren B, et al. Early oseltamivir reduces risk for influenza-associated aspergillosis in a double-hit murine model. Virulence. 2021;12:2493–508.
Lin TY, Stone Y, Glatt S. Mechanistic insight into the pseudouridylation of RNA. RNA Biol. 2025;22:1–25.
Xu H, Kong L, Li M, Pisignano G, Cheng J, Feng F, et al. A comprehensive tRNA pseudouridine map uncovers targets dependent on human stand-alone pseudouridine synthases. Nat Cell Biol. 2025;27:2186–97.
Malviya V, Yshii L, Junius S, Garg AD, Humblet-Baron S, Schlenner SM. Regulatory T-cell stability and functional plasticity in health and disease. Immunol Cell Biol. 2023;101:112–29.
Jurgens AP, Popović B, Wolkers MC. T cells at work: how post-transcriptional mechanisms control T cell homeostasis and activation. Eur J Immunol. 2021;51:2178–87.
Wolf T, Jin W, Zoppi G, Vogel IA, Akhmedov M, Bleck CKE, et al. Dynamics in protein translation sustaining T cell preparedness. Nat Immunol. 2020;21:927–37.
Li HB, Tong J, Zhu S, Batista PJ, Duffy EE, Zhao J, et al. m6A mRNA methylation controls T cell homeostasis by targeting the IL-7/STAT5/SOCS pathways. Nature. 2017;548:338–42.
Wang H, Hu X, Huang M, Liu J, Gu Y, Ma L, et al. Mettl3-mediated mRNA m6A methylation promotes dendritic cell activation. Nat Commun. 2019;10:1898.
Song H, Song J, Cheng M, Zheng M, Wang T, Tian S, et al. METTL3-mediated m6A RNA methylation promotes the anti-tumour immunity of natural killer cells. Nat Commun. 2021;12:5522.
Liu Y, Yuan Y, Zhou Z, Cui Y, Teng Y, Huang H, et al. Mettl14-mediated m6A modification enhances the function of Foxp3+ regulatory T cells and promotes allograft acceptance. Front Immunol. 2022;13:1022015.
Marchand V, Pichot F, Neybecker P, Ayadi L, Bourguignon-Igel V, Wacheul L. et al. HydraPsiSeq: a method for systematic and quantitative mapping of pseudouridines in RNA. Nucleic Acids Res. 2020;48:e110–e110.
Acknowledgements
This work was supported by the KU Leuven tenure track starting grant (STG/16/034 to SMS), the Research Foundation Flanders (Fonds voor Wetenschappelijk Onderzoek, FWO) (grants G054722N to SMS, 11l7525N to VM and 1199825 N to LM) and KU Leuven research funding (C14/20/106 to SMS). We acknowledge the contributions of Adamantios Mavrogiannis (AM) posthumously. He sadly passed away while the manuscript was being prepared. We thank Prof. Dr. Gabrielle Bergers for providing the MC38 colon adenocarcinoma cell line. The authors further acknowledge the contributions of Oliver Burton (University of Cambridge) for supporting flow cytometry panel design, the KUL Flow and Mass Cytometry core facility for flow cytometry support and the KU Leuven animalium and Imaging facilities for mouse husbandry.
Author information
Authors and Affiliations
Contributions
PL, VM, CTP, AM, GV, LM, LS, IM and ViM executed experiments. VM, CTP, LM, LR, IM and ViM analysed experimental data. AM conducted in silico analyses from publicly available datasets. VM and AM prepared the figures. VM, CTP and SMS wrote the manuscript. SMS and PL designed the study, secured funding, supervised the research and together with VM, CTP and IM revised the manuscript. GVV and GV designed disease model experiments and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
SMS is the associate editor at Genes and Immunity. The other authors declare no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Malviya, V., Pham, C.T., Michiels, L. et al. Trub1-mediated pseudouridylation is dispensable for immune cell development and homeostasis. Genes Immun (2026). https://doi.org/10.1038/s41435-026-00393-3
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41435-026-00393-3