Abstract
Arid ecosystems, characterized by severe water scarcity, play a crucial role in preserving Earth’s biodiversity and resources. The Tarim Basin in Northwestern China, a typical arid region isolated by the Tianshan Mountains and expansive deserts, provides a special study area for investigating how plant response and adaptation to such environments. Tamarix hispida, a species well adapted to saline-alkaline and drought conditions, dominates in the saline-alkali lands of the Tarim Basin. This study aims to examine the genetic diversity and environmental adaptation of T. hispida in the Tarim Basin. Genomic SNPs for a total of 160 individuals from 17 populations were generated using dd-RAD sequencing approach. Population genetic structure and genetic diversity were analyzed by methods including ADMIXTURE, PCA, and phylogenetic tree. Environmental association analysis (EAA) was performed using LFMM and RDA analyses. The results revealed two major genetic lineages with geographical substitution patterns from west to east, indicating significant gene flow and hybridization. Environmental factors such as Precipitation Seasonality (bio15) and Topsoil Sand Fraction (T_SAND) significantly shaped allele frequencies, supporting the species’ genetic adaptability. Several genes associated with environmental adaptation were identified and annotated, highlighting physiological and metabolic processes crucial for survival in arid conditions. The study highlights the role of geographical isolation and environmental factors in shaping genetic structure and adaptive evolution. The identified adaptive genes related to stress tolerance emphasize the species’ resilience and highlight the importance of specific physiological and metabolic pathways.
Similar content being viewed by others
Introduction
Arid regions are characterized by severe water shortage and are mainly located in the 25°–35° latitudinal zones on both sides of the equator. Drought causes significant changes in the composition of animal and plant communities and soil microbial structures (Vásquez-Dean et al. 2020). The Tarim Basin in northwestern China is a typical arid region with strong temporal and spatial environmental dynamics driven by regional tectonic movements. Since the Quaternary period, the Tarim Basin has been continuously undergoing aridification, with alternating periods of wetter and drier conditions. The surrounding aeolian sediments are mainly caused by climate change due to the uplift of the Tibetan Plateau and adjacent highlands (Zhu et al. 2014). As the largest inland basin in China, the Tarim Basin has experienced intense mountain uplift and differential subsidence. Phylogeographic studies of drought-tolerant species such as Nitraria sphaerocarpa and Gymnocarpos przewalskii show that the Tarim Basin, isolated by the Tianshan Mountains and desert barriers, is an independent biogeographic unit from other areas in northwestern China (Su and Zhang 2013; Zhang et al. 2020). The extreme aridity (annual precipitation <50 mm), early drought (about 6 million years ago) and geographical isolation of the basin make it an ideal site for genetic structure analysis to answer important questions in phylogeography (Yang et al. 2022).
In arid regions, plants are subjected to extreme environmental conditions, such as drought, high temperatures, nutrient deficiency, and salinity. To cope with these challenges, desert plants have evolved various adaptations that minimize water loss and enhance soil moisture retention. Some species have developed deep root systems to access groundwater reserves, while others have evolved thick cuticles or waxy layers to reduce evaporation. Additionally, specialized water storage tissues aid in survival during periods of drought (Mohanta et al. 2024). These adaptations enable desert plants to flourish and maintain the balance of their ecosystems under harsh conditions. Furthermore, plants may adjust their growth patterns and physiological mechanisms, such as altering leaf size, shape, and photosynthetic rates, to adapt to environmental changes (Bastola et al. 2021; Kirschner et al. 2021).
Tamarix is a genus with high salinity and drought tolerance, wide distribution, and rich genetic diversity, which has become an important research object in arid regions (Wang et al. 2021; Zhang et al. 2022; Liu et al. 2022). Genetic diversity studies have revealed the extensive environmental adaptability of Tamarix and its complex evolutionary history. Studies on Tamarix species along the Yellow River showed that T. austromongolica and T. chinensis had frequent gene flow and little genetic differentiation among populations (Liang et al. 2019). Using SLAF-seq, researchers found both shared ancestry and genetic differentiation among Tamarix populations from different areas (Li et al. 2024). SSR markers analysis in T. taklamakanensis in the Tarim Basin showed moderate genetic diversity but fragmented distribution, low allelic diversity, and bottleneck effects were considered to be the main reasons for the observed genetic structure (Su et al. 2018).
Tamarix hispida Willd., a member of the Tamaricaceae family, predominantly inhabits desert regions along river floodplains, sedimentation plains, and lake basin edges, as well as in salt-alkaline meadows and sand dunes. This species is known for its resilience to drought and salinity. It is densely covered with short, straight, single-celled hairs, and its flowers exhibit a range of colors from purple to bright red; accompanied by unique morphological features (Gaskin 2003). In China, T. hispida is primarily found in Xinjiang, Qinghai, Gansu, Ningxia, and Inner Mongolia, with additional populations in Central Asia, Iran, Afghanistan, and Mongolia. As a key natural assemblage species in the arid northwestern region of China, T. hispida plays a crucial role in restoring soil salinity and heavy metal pollution, thereby supporting ecological stability (Pang et al. 2022; Xie et al. 2023). Most species within the Tamarix genus share similar flowering periods and mating behaviors, leading to morphological and genetic homogeneity that poses challenges for population genetic studies. However, T. hispida’s unique morphological features, such as flower color and dense hair coverage, serve as distinct identification characteristics (Terrones and Juan 2023). Compared to other Tamarix species, T. hispida exhibits more stable morphological traits and a narrower range of flowering overlap, making it an ideal subject for population genetic studies.
Current research on T. hispida predominantly centers on plant physiology, phylogenetics, and the development and application of saline-alkali land (Wang et al. 2018; Zhao et al. 2021). However, investigations into its evolutionary responses to environmental shifts and adaptive evolution remain scant. Given its fragmented distribution pattern within its geographical range, particularly in the Tarim Basin, analyzing its genetic diversity, genetic structure, and the regulatory mechanisms of population adaptive evolution is essential. This analysis can elucidate the local adaptive strategies of various populations under diverse environmental conditions. Such a study is significant as it provides a theoretical foundation for understanding the molecular mechanisms driving the local adaptive evolution of T. hispida in response to environmental factors. Furthermore, it serves as a benchmark for adaptive evolution research in other species within arid saline-alkali environments. In this context, the application of dd-RAD sequencing represents a novel approach, allowing for a detailed and comprehensive analysis of the genetic mechanisms underlying the adaptation of T. hispida populations to local environmental changes. Therefore, we aim to test the following scientific hypotheses: (1) The genetic diversity and genetic structure of the T. hispida population may be linked to the geographical environment, given its distribution characteristics; (2) Genomic methods can be utilized to analyze the genetic mechanisms underlying the adaptation of T. hispida populations to local environmental changes.
Materials and methods
Plant sampling and DNA extraction
A total of 160 wild T. hispida individuals were collected from 17 locations in Xinjiang, China, adhering to a protocol that required one individual per 10 meters within each population (Fig. 1, Table S2). The leaf samples were air-dried on-site using silica gel and subsequently sent to Shanghai Personalbio Technology Co., Ltd. for total DNA extraction via a modified CTAB method (Porebski et al. 1997). The quality of the DNA was evaluated using 0.8% agarose gel electrophoresis, while DNA quantification was conducted using a UV spectrophotometer.
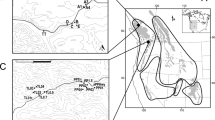
A The geographic locations and lineage distributions of 17 populations of Tamarix hispida are depicted. The pie chart illustrates the average proportions of three distinct genetic lineages identified by the ADMIXTURE tool in each population, with blue representing lineage A, red indicating lineage B, and green denoting lineage C. B Results of ADMIXTURE phylogenetic typing. The blue bar represents the likelihood Q value derived from lineage A, the red bar represents the likelihood Q value derived from lineage B, and the green bar represents the likelihood Q value derived from lineage C Based on Q value ≥ 0.5, all populations are divided into W, E, and C populations. C. Results of Principal Component Analysis (PCA). D Results of Maximum Likelihood (ML) phylogenetic tree. The numbers near the nodes are the support rates (%) for the nodes.
Library construction, sequencing, and SNP filtering
Whole-genome DNA was digested with a restriction enzyme (DpnII_MspI), and the fragments of defined length were recovered for library construction by Double Digest RADseq (dd-RAD) (Table S1). Next-generation sequencing (NGS) on an Illumina NovaSeq platform was performed for paired-end (PE) sequencing at an average depth of 1×. The raw data were filtered using fastp (https://github.com/OpenGene/fastp) with a sliding window method to obtain high quality data. The filtered data were aligned to the T. chinensis reference genome (http://gigadb.org/dataset/view/id/102417) using the BWA software (0.7.12-r1039) mem algorithm (Li and Durbin 2009). Preliminary SNP data were generated by Picard v1.107 (http://www.psc.edu/index.php/user-resources/software/picard) and GATK (Zhu et al. 2014). The filtering criteria were as follows: minimum allele frequency = 0.05, linkage disequilibrium (LD) (R² > 0), deviation from Hardy-Weinberg equilibrium at a significance level of 0.1, and removal of SNPs with individual missingness using VCFtools software (max-missing 1 –mac 3) and Plink software (–maf 0.05 –geno 0 –hwe 0.1 –indep-pairwise 50 5 0.05). A total of 62,940 SNP sites from 160 individuals in 17 populations were used for subsequent analyses.
Population genetic structure analysis
To examine the population genetic structure of T. hispida, a clustering analysis was conducted on the 160 individuals using ADMIXTURE software (Patterson et al. 2012). The genetic clustering values (K) were initially evaluated from 1 to 10, and the optimal K value was determined based on cross-validation error (CV). IQ-Tree software, utilizing SNP sites, constructed a phylogenetic tree of the 160 individuals to investigate kinship relationships across the 17 populations at an individual level (Nguyen et al. 2015). The ModelFinder algorithm within IQ-Tree identified the most suitable nucleotide substitution model, TVM + F + G4, through 1000 iterations. Furthermore, principal component analysis (PCA) based on SNP variation was performed using GCTA software (Yang et al. 2011).
Population genetic statistics
Genetic diversity indices, such as observed heterozygosity (Ho), expected heterozygosity (He), inbreeding coefficient (Fis), and nucleotide diversity (Pi), were determined using Plink software (Purcell et al. 2007). Fst values and Tajima’s D were calculated for individual populations of T. hispida with VCFtools to evaluate genetic differentiation among diverse populations (Danecek et al. 2011).
Population historical dynamics
To ensure consistency in sample sizes across different groups, we carefully selected four individuals from each population in groups W and C. Additionally, one individual was removed from the HJ population in group E to minimize any potential deviation in the calculation results. Filtered SNPs were further screened at 10 kb intervals using VCFtools, resulting in a total of 12,762 SNPs. Logistic regression was performed with the DIYABC software and Approximate Bayesian Computation (ABC) (Cornuet et al. 2010) with a 95% confidence interval. One percent of the observed data that had the closest match to the simulated data was used to estimate posterior probabilities (PP) for each evolutionary event, and the event with the highest PP was selected as optimal.
Environmental and soil factor association analysis
To examine the effect of environmental and soil factors on population genetic variation, Redundancy Analysis (RDA) (Capblancq and Forester 2021) and Gradient Forest Analysis (GF) (Bay et al. 2018) were used. Nineteen bioclimatic variables were obtained from the PaleoClim (www.PaleoClim.org), and climate data for each population were extracted using the Raster package in R. Correlation analysis was performed on the environmental variables to reduce multicollinearity problems. Pearson correlation analysis was conducted in R to retain only highly correlated variables with a Pearson correlation coefficient greater than 0.8, while taking into account the importance of various environmental factors for the species (Cleophas and Zwinderman 2018). Nine soil factors were extracted from HWSD (Harmonized World Soil Database) and subjected to Pearson analysis after extracting soil data based on sample points’ latitude and longitude using ArcGIS v10.8. Finally, five climate factors and nine soil factors were selected for Gradient Forest analysis.
In the RDA, the Gradient Forest analysis pinpointed six paramount environmental and soil factors that significantly influence genetic variation among populations. Genetic distances were determined by transforming Fst values into Fst/(1-Fst) values. Additionally, geographic distances between populations were calculated utilizing the R package geosphere. A Mantel test, incorporating 1,000 permutations, was performed using the R package Vegan to investigate the correlation between genetic and geographic distances among T. hispida populations (Legendre and Fortin 2010).
The Latent Factor Mixture Model (LFMM) is a technique utilized to investigate environmental adaptation traits within the genome and analyze the pattern of environmental adaptation by pinpointing SNP sites highly correlated with environmental variables (Frichot et al. 2013). Six key environmental factors influencing the genetic variation of T. hispida were derived in the Gradient Forest (GF) analysis. The optimal number of latent factors was selected to align with the optimal number of groups (K = 3) identified in the ADMIXTURE analysis. Initially, the VCF file was converted to a ped file using VCFtools, followed by the utilization of the LEA package and tidyverse package in R to read genotype and environmental data (Frichot and François 2015). The ‘lfmm’ function was then applied to conduct LFMM analysis. To manage false positive discovery rate (FDR), the Q-value (q < 0.01) method was employed, leading to the identification of significant candidate genes.
GO enrichment analysis
Adaptive loci identified from RDA and LFMM were extracted from the reference genome for Gene Ontology (GO) enrichment analysis. The web-based tool eggNOG-Mapper (http://eggnog-mapper.embl.de/) was employed for GO enrichment analysis (Cantalapiedra et al. 2021).
Result
Population structure
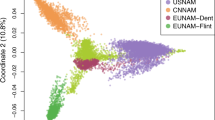
The admixture analysis identified three genetic groups (k = 3) based on the minimized cross-validation (CV) error value. Figure 1 illustrated a geographic replacement pattern of genetic structure along the longitudinal axis in the admixture results. Using Q = 0.5 as the threshold, the population was divided into three lineages: W (WPE, SL, ATS, YPH, ZP, MGT, BC), E (BH, HJ, GG), and the remaining populations as C. Principal component analysis (PCA) revealed that PC1 explained 2.23% and PC2 explained 3.29% of the total genetic variation. Notably, individuals from the three lineages showed distinct genetic differentiation with some mixing characteristics at the lineage boundaries. The findings were supported by the ML tree, emphasizing the higher genetic purity of the western population (W) compared to the genetic admixture observed in the central (C) and eastern (E) populations.
Genetic diversity
Genetic diversity analysis indicated a range of Pi values from 0.328 to 0.394 (Table 1), reflecting notable DNA sequence variations among individuals and a high level of genetic diversity. Tajima’s D values fell within the range of 0.05 to 0.28, suggesting a relatively high genetic diversity within the population without extreme polymorphism. The Fis values ranged from −0.128 to −0.0241 across all populations, indicating reduced genetic similarity at this locus with increased heterozygosity, potentially influenced by gene flow, selective pressures, or other factors. Geographically proximal populations exhibited clustered maximum and minimum values of Ho and He, showcasing varying levels of genetic diversity across different geographical locations. The majority of Fst values were between 0.05 and 0.1, signifying some genetic disparities among populations with restricted gene flow.
Demographical history
The DIYABC analysis examining the population history suggests that Model 1 is the most likely scenario, with a posterior probability of 0.803 and an error rate of 0.073 (Fig. 2, Tables S3 and S4). According to the model diagram, around 7 ka BP, the T. hispida population experienced a split, giving rise to two branches: the W and E populations. Roughly 10 ka years after this split, genetic mixing between the W and E populations resulted in the differentiation and emergence of the C population, impacting the current distribution patterns of these populations.
Demographic history of Tamarix hispida.
Genetic environment association
Among the 14 factors analyzed using GF analysis, bio15 (Precipitation Seasonality (Coefficient of Variation)), bio18 (Precipitation of Warmest Quarter), and bio8 (Mean Temperature of Wettest Quarter) were identified as the most influential environmental determinants affecting allele frequency shifts in the T. hispida population (Fig. 3A, Table S5). In contrast, soil factors such as T_SAND, T_SILT, and T_PH_H2O had a minor impact. While both environmental and soil factors play a role, environmental factors were found to be more significant. The remaining eight ecological factors had a smaller effect on genetic variability due to their lower weights.
Panel A displays the R2-weighted importance of environmental variables that elucidate genetic gradients, while Panel B shows the cumulative importance of allelic change across the initial six environmental gradients.
The ecological implications of the identified environmental factors are noteworthy. Precipitation seasonality (bio15) is a critical factor as it impacts water availability throughout different seasons, which is essential for the survival and reproduction of T. hispida. Variations in water availability can introduce selective pressures that favor alleles associated with drought resistance or water use efficiency. Additionally, bio18, which represents precipitation during the warmest quarter, plays a crucial role in determining water supply during the hottest part of the year, directly influencing plant growth and stress tolerance. Moreover, bio8, the mean temperature of the wettest quarter, influences the thermal environment during the peak of water availability, thereby affecting growth rates and metabolic processes.
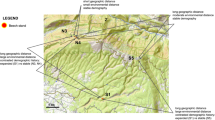
Figure 3B shows the cumulative trend of allele frequency changes along environmental gradients. Notably, RDA1 (33.4%) and RDA2 (17.4%) together explain half of the genetic variability, demonstrating a direct relationship between each ecological factor’s contribution to genetic variance as shown by RDA and the results from GF analysis. In the integrated analysis of isolation by distance (IBD) and isolation by environment (IBE), a significant correlation was found between genetic differentiation across the 17 populations and the observed geographical distances (R² = 0.066, p = 0.0025), indicating that geographical distance explains a portion of the genetic diversity observed (Fig. 4).
Mantel tests were conducted to assess the correlation between genetic distance and both geographic and environmental distances.
GO enrichment
The LFMM analysis identified 34 loci, while the RDA identified 141 adaptive loci, with 17 loci identified by both methods. Genomic annotation and GO enrichment analysis for T. hispida revealed 8 adaptive loci with annotations (Table 2). Among these loci, At3g25060, a mitochondrial Pentatricopeptide Repeat (PPR) protein, plays a role in mitochondrial gene splicing, RNA editing, and RNA stability, which are crucial for mitochondrial function and energy production, enhancing the plant’s ability to adapt to varying environmental conditions. Another identified locus, the Protein EARLY FLOWERING, is involved in regulating plant flowering time in response to environmental cues, which can optimize reproductive success in different climatic conditions. Furthermore, a 10 kDa polypeptide in Photosystem II (PSII) is essential for photosynthesis and growth directly impacting the plant’s ability to convert light energy into chemical energy, thereby affecting growth rates and biomass accumulation. The Protein Kinase Superfamily, particularly Ser/Thr/Tyr kinases, is crucial for plant growth and responses to stress, playing a pivotal role in signal transduction pathways that mediate responses to biotic and abiotic stresses, thus enhancing survival and adaptability. Lastly, the Vacuolar protein sorting-associated protein 41 homolog (VPS41) is responsible for vacuolar transport and developmental regulation in plants, which is essential for maintaining cellular homeostasis and facilitating growth and development under stress conditions.
The identified genes and their respective roles highlight specific adaptive traits that enable T. hispida to thrive in diverse and changing environments. Genes involved in mitochondrial function, photosynthesis, and stress response contribute to enhanced metabolic efficiency and resilience. Additionally, genes regulating flowering time and developmental processes allow the plant to synchronize its life cycle with environmental conditions, optimizing reproductive success and survival.
Discussion
The genetic structure of T. hispida populations and their geographical distribution patterns
This study aims to investigate the genetic structure of T. hispida populations within the Tarim basin and their geographical distribution patterns. By analyzing genomic SNPs data from various populations, we identified three major lineages (A, B, C) of T. hispida populations, showing a west to east geographical substitution phenomenon. This discovery supports theories of interspecific hybridization and gene introgression, highlighting the significant role of geographical substitution in shaping genetic patterns (Rieseberg et al. 2000; Wang et al. 2019).
Similar patterns of geographical barriers influencing genetic differentiation have been observed in various plant and animal populations. For example, research on Triportheus albus in the Amazon basin has shown how river systems and mountain ranges can impact genetic structure along geographical gradients (Queiroz et al. 2017). Alpine plant community studies have also highlighted the effects of altitude gradients on genetic diversity and population structure (Stöcklin et al. 2009). In the Tarim Basin, research on Triplophysa yarkandensis has revealed high genetic variation and significant interpopulation differentiation, emphasizing the role of geographical and environmental factors in species differentiation (Zhou et al. 2021). Furthermore, investigations on oasis edge species downstream of the Tarim River have further underscored the crucial influence of environmental factors on species differentiation (Zhang et al. 2022; Wang et al. 2023). Our study supports the view that the genetic structure and population differentiation of T hispida are significantly influenced by the unique environmental conditions of the Tarim Basin. This further emphasizes the overall significance of geographical and environmental barriers in shaping genetic structure and species differentiation.
Phylogenetic analysis, along with phylogenetic statistical parameters, has further supported the recent divergence of central Tamarix populations, with east and west populations showing incomplete segregation. This suggests that geographical isolation may have driven the alternate evolution of different lineages within the same species, consistent with prior studies (Liu et al. 2019; Tóth et al. 2019). The discovery of Lineage A, potentially influenced by gene introgression from other species, is remarkable. Gene introgression involves the incorporation of exogenous genes into the gene pool of the target species through hybridization, blending with native genes (Xiao et al. 2023). While the exact form and mechanism of hybridization remain unclear, previous research has highlighted its significant role in Tamarix speciation (Sun et al. 2016). Genetic structure analysis of T. chinese indicates that, despite relatively low population differentiation, genetic clusters linked to geographical origin can be distinguished, suggesting interpopulation hybridization events that impact genetic structure (Jiang et al. 2023). In the case of T. hispida, hybridization plays a crucial role in shaping its genetic structure and adaptive evolution. Research indicates that hybridization can introduce beneficial alleles from different lineages, increasing genetic diversity and enabling populations to adapt to varying environmental conditions (Porretta and Canestrelli 2023). This process is particularly significant for T. hispida, which thrives in diverse and often challenging environments. The incorporation of exogenous genes through hybridization may enhance traits such as drought tolerance, salt tolerance, and disease resistance, thereby strengthening the species’ adaptability and evolutionary potential (Singhal et al. 2021). Furthermore, hybridization could promote the development of new phenotypic traits that offer advantages in specific ecological niches, further supporting the survival and proliferation of T. hispida across diverse habitats (Rosser et al. 2024). The introduction of new genetic material through hybridization may have bolstered the adaptability and evolutionary potential of the Tamarix genus, essential for species survival and propagation in diverse environments.
The evolution of T. hispida populations adapted to the environment of the Tarim Basin
This study has examined the adaptive evolution of T. hispida populations in the Tarim basin, emphasizing the importance of functional genes in extreme environmental conditions. Our research has identified key genes related to environmental adaptation, specifically linked to plant physiological processes and metabolic pathways. Genes like the 10 kDa polypeptide of PSII and the Ser/Thr/Tyr kinases of the Protein Kinase Superfamily are crucial for the plant’s photochemical processes, response to abiotic stress, and adaptation to the unique conditions of the Tarim basin (Ljungberg et al. 1984; Hanks and Hunter 1995). Comparative studies in other species reveal similar genetic adaptations to harsh environments. For instance, the green alga Chlorella ohadii displays distinctive structural adaptations in its PSII complex, enabling it to endure high light intensities without suffering photodamage. These adaptations involve specific subunit configurations that shield the oxygen-evolving complex and the PSII core from harm (Fadeeva et al. 2023). These discoveries offer a more comprehensive understanding of the molecular mechanisms underpinning T. hispida’s ability to thrive in extreme environments, shedding light on the consistent nature of these adaptive strategies across various species and environmental conditions (Fig. 5).
White circles represent different individuals, while small black dots represent different Single Nucleotide Polymorphism (SNP) sites. The environmental factors considered in the analysis include Bio 8 (Mean Temperature of the Wettest Quarter), Bio 15 (Precipitation Seasonality, Coefficient of Variation), and Bio 18 (Precipitation of the Warmest Quarter).
Utilizing the GF analysis, our investigation has identified precipitation as the key environmental factor influencing the growth of T. hispida. This sheds light on how precipitation impacts the niche of T. hispida. A narrow niche breadth may indicate T. hispida’s strong reliance on specific precipitation conditions, while a broader niche suggests the species’ ability to adapt to a wider range of precipitation patterns. Additionally, precipitation interacts with other environmental factors like temperature and soil moisture, collectively shaping the growth and distribution of T. hispida. In nature, many plants and animals have developed adaptive strategies for extreme conditions. For example, the desert plant Haloxylon ammodendron accumulates salts and improves water-use efficiency to survive in arid and saline soils (Song et al. 2005; Yue et al. 2014). Similarly, research on desert mammals has uncovered shared genetic and physiological pathways linked to adaptations to water and food scarcity and extreme temperatures. These comparative studies offer a fresh perspective on understanding the adaptability of T. hispida populations, suggesting potential adaptations to animal feeding habits that contribute to their evolutionary process.
The demographical history suggests that the initial divergence occurred approximately 7 thousand years ago (ka BP), characterized by a Tamarix generation time of 1-2 years (Jiang et al. 2023). During this period, the climate presented a bit wet conditions, such as the brief cold-wet climate recorded at Lake Lop Nur (Wang and Jia 2017). In response to address the impacts of climate change, certain populations began to disperse to various environments, including the eastern region of Southern Xinjiang. Over the following 1 ka, the climate transitioned to warmer and drier conditions, leading to a decrease in sand areas and the stabilization of sand dunes along the banks of the Tarim River and its tributaries, as well as in the lowlands and lakes. This resulted in the expansion of river-alluvial regions and the gradual stabilization of the eastern populations. This process likely facilitated an increase in species diversity and the expansion of ecological niches. Similar adaptive patterns can be observed in desert foxes, which utilize interspecific hybridization to adapt to the Sahara desert, and in camels, whose genomes exhibit adaptations and evolution to desert environments (Wu et al. 2014; L Rocha et al. 2023). Around 3 ka BP, as warmer climates set in and land and water resources were overexploited, the Tarim basin experienced increased aridity, with alternating warm and cold climates. Human activities may have significantly impacted the ecosystem, leading to alterations in species distribution and genetic diversity. As climate variability persisted, populations continued to disperse to different environments and stabilize in various regions of the Tarim basin, potentially fostering an increase in species diversity and the expansion of ecological niches (Carvalho C da et al. 2017).
With the increasing impact of global climate change, the distribution and genetic makeup of T. hispida populations are likely to experience significant alterations. Projections for future climate conditions indicate rising temperatures and more erratic precipitation patterns, potentially affecting the availability and quality of habitats for T. hispida. These changes could result in shifts in the species’ geographical range, as populations may move towards more suitable climates. As a consequence of habitat fragmentation, gene flow between populations may decrease, leading to higher genetic drift and a decline in genetic diversity (Pauls et al. 2013). Moreover, the stress induced by climate change could promote hybridization events as populations interact at shifting range boundaries, potentially introducing new genetic variations that could facilitate adaptation (Aitken et al. 2008). However, if the pace of environmental change surpasses the species’ ability to adapt, there is a risk of local extinctions and subsequent loss of genetic diversity. It is crucial to continuously monitor and conduct research to comprehend and address the impacts of future climate change on T. hispida’s ecological roles and survival.
Despite the novel insights into the genetic structure of T. hispida populations provided by this study, it is important to acknowledge the limitations of the sample size, which may impact the generalizability of the results. To address this limitation, future research should consider expanding the sample size to validate and strengthen our findings. Our study has highlighted the significant impact of environmental factors on the genetic variability of T. hispida, a discovery that is crucial for understanding the adaptive evolution of species. With the rapid pace of global climate change, investigating how T. hispida populations respond to these shifts is essential for predicting and conserving biodiversity. By combining genomic and ecological approaches, we can uncover the molecular mechanisms through which T. hispida populations adapt to extreme environments, offering a solid scientific basis for ecosystem conservation and management.
Conclusion
In summation, this study has shed light on the genetic structure and geographical distribution patterns of T. hispida populations in the Tarim basin, highlighting the significant influence of environmental factors on species differentiation. These results provide new insights into the adaptation of T. hispida populations and suggest avenues for future research. Additionally, investigating the effects of human activities on the Tarim basin ecosystem and developing conservation and restoration strategies are vital areas for further study.
Data availability
All the original sequencing data in omics have been uploaded to the GSA public database (https://ngdc.cncb.ac.cn/gsub/) with the accession number CRA017951. https://ngdc.cncb.ac.cn/gsub/submit/gsa/subCRA028863/finishedOverview.
References
Aitken SN, Yeaman S, Holliday JA, Wang T, Curtis-McLane S (2008) Adaptation, migration or extirpation: climate change outcomes for tree populations. Evol Appl 1:95–111
Bastola AK, Soffiatti P, Behl M, Lendlein A, Rowe NP (2021) Structural performance of a climbing cactus: making the most of softness. J R Soc Interface 18:rsif.2021.0040, 20210040
Bay RA, Harrigan RJ, Underwood VL, Gibbs HL, Smith TB, Ruegg K (2018) Genomic signals of selection predict climate-driven population declines in a migratory bird. Science 359:83–86
Cantalapiedra CP, Hernández-Plaza A, Letunic I, Bork P, Huerta-Cepas J (2021) eggNOG-mapper v2: Functional Annotation, Orthology Assignments, and Domain Prediction at the Metagenomic Scale. Mol Biol Evol 38:5825–5829
Capblancq T, Forester BR (2021) Redundancy analysis: a Swiss Army Knife for landscape genomics. Methods. Ecol Evol 12:2298–2309
Carvalho C da S, Ballesteros-Mejia L, Ribeiro MC, Côrtes MC, Santos AS, Collevatti RG (2017) Climatic stability and contemporary human impacts affect the genetic diversity and conservation status of a tropical palm in the Atlantic Forest of Brazil. Conserv Genet 18:467–478
Cleophas TJ, Zwinderman AH (2018) Bayesian Pearson Correlation Analysis. In: Cleophas TJ, Zwinderman AH (eds) Modern Bayesian Statistics in Clinical Research. Springer International Publishing, Cham, p 111–118
Cornuet J-M, Ravigné V, Estoup A (2010) Inference on population history and model checking using DNA sequence and microsatellite data with the software DIYABC (v1.0). Bmc Bioinforma 11:401
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA et al. (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158
Fadeeva M, Klaiman D, Caspy I, Nelson N (2023) Structure of Chlorella ohadii Photosystem II Reveals Protective Mechanisms against Environmental Stress. Cells 12:1971
Frichot E, François O (2015) LEA: An R package for landscape and ecological association studies. Methods Ecol Evol 6:925–929
Frichot E, Schoville SD, Bouchard G, François O (2013) Testing for Associations between Loci and Environmental Gradients Using Latent Factor Mixed Models. Mol Biol Evol 30:1687–1699
Gaskin JF (2003) Tamaricaceae. In: Kubitzki K, Bayer C (eds) Flowering Plants · Dicotyledons: Malvales, Capparales and Non-betalain Caryophyllales, The Families and Genera of Vascular Plants. Springer, Berlin, Heidelberg, p 363–368
Hanks SK, Hunter T (1995) The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification1. FASEB J 9:576–596
Jiang Z, Yang A, Zhang H, Wang W, Zhang R (2023) Population structure and genetic diversity of Tamarix chinensis as revealed with microsatellite markers in two estuarine flats. Peerj 11:e15882
Kirschner GK, Xiao TT, Blilou I (2021) Rooting in the Desert: A Developmental Overview on Desert Plants. Genes-Basel 12:709
L Rocha J, Silva P, Santos N, Nakamura M, Afonso S, Qninba A et al. (2023) North African fox genomes show signatures of repeated introgression and adaptation to life in deserts. Nat Ecol Evol 7:1267–1286
Legendre P, Fortin M-J (2010) Comparison of the Mantel test and alternative approaches for detecting complex multivariate relationships in the spatial analysis of genetic data. Mol Ecol Resour 10:831–844
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760
Li Z, Wang D, Sun M, Cheng B, Zhang G, Xu X (2024) Development and genetic analysis of Tamarix chinensis single nucleotide polymorphism sites based on specific locus amplified fragment sequencing simplified genome technology. Genet Resour Crop Evol 71:549–558
Liang H, Liu C, Li Y, Wang Y, Kong Y, Quan J et al. (2019) Low population genetic differentiation in two Tamarix species (Tamarix austromongolica and Tamarix chinensis) along the Yellow River. Genetica 147:13–22
Liu Y, Dietrich CH, Wei C (2019) Genetic divergence, population differentiation and phylogeography of the cicada Subpsaltria yangi based on molecular and acoustic data: an example of the early stage of speciation? Bmc Evol Biol 19:5
Liu JN, Fang H, Liang Q, Dong Y, Wang C, Yan L et al. (2022) Genomic analyses provide insights into the evolution and salinity adaptation of halophyte Tamarix chinensis. Gigascience 12:giad053
Ljungberg U, Åkerlund H-E, Andersson B (1984) The release of a 10-kDa polypeptide from everted photosystem II thylakoid membranes by alkaline tris. Febs Lett 175:255–258
Mohanta TK, Mohanta YK, Kaushik P, Kumar J (2024) Physiology, genomics, and evolutionary aspects of desert plants. J Adv Res 58:63–78
Nguyen LT, Schmidt HA, Von Haeseler A, Minh BQ (2015) IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol Biol Evol 32:268–274
Pang X, Suo J, Liu S, Xu J, Yang T, Xiang N et al. (2022) Combined transcriptomic and metabolomic analysis reveals the potential mechanism of seed germination and young seedling growth in Tamarix hispida. BMC Genomics 23:109
Patterson N, Moorjani P, Luo Y, Mallick S, Rohland N, Zhan Y et al. (2012) Ancient Admixture in Human History. Genetics 192:1065–1093
Pauls SU, Nowak C, Bálint M, Pfenninger M (2013) The impact of global climate change on genetic diversity within populations and species. Mol Ecol 22:925–946
Porebski S, Bailey LG, Baum BR (1997) Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol Biol Rep. 15:8–15
Porretta D, Canestrelli D (2023) The ecological importance of hybridization. Trends Ecol Evol 38:1097–1108
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D et al. (2007) PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am J Hum Genet 81:559–575
Queiroz LJ, de, Torrente-Vilara G, Quilodran C, Doria CR, da C, Montoya-Burgos JI (2017) Multifactorial genetic divergence processes drive the onset of speciation in an Amazonian fish. Plos One 12:e0189349
Rieseberg LH, Baird SJE, Gardner KA (2000) Hybridization, introgression, and linkage evolution. In: Doyle JJ, Gaut BS (eds) Plant Molecular Evolution. Springer Netherlands, Dordrecht, p 205–224
Rosser N, Seixas F, Queste LM, Cama B, Mori-Pezo R, Kryvokhyzha D et al. (2024) Hybrid speciation driven by multilocus introgression of ecological traits. Nature 628:811–817
Singhal S, Roddy AB, DiVittorio C, Sanchez-Amaya A, Henriquez CL, Brodersen CR et al. (2021) Diversification, disparification and hybridization in the desert shrubs Encelia. N. Phytol 230:1228–1241
Song J, Feng G, Tian C, Zhang F (2005) Strategies for Adaptation of Suaeda physophora, Haloxylon ammodendron and Haloxylon persicum to a Saline Environment During Seed-Germination Stage. Ann Bot-Lond 96:399–405
Stöcklin J, Kuss P, Pluess AR (2009) Genetic diversity, phenotypic variation and local adaptation in the alpine landscape: case studies with alpine plant species. Bot Helv 119:125–133
Su Z, Zhang M (2013) Evolutionary response to Quaternary climate aridification and oscillations in north-western China revealed by chloroplast phylogeography of the desert shrub Nitraria sphaerocarpa (Nitrariaceae). Biol J Linn Soc 109:757–770
Su Z, Zhuo L, Li W, Liu H, Kang X, Jiang X (2018) Genetic Diversity of Tamarix taklamakanensis in the Tarim Basin. J Desert Res 38:1041
Sun L, Yang R, Zhang B, Zhang G, Wu X, Zhang W et al. (2016) Phylogenetic relationships among species of Tamarix (Tamaricaceae) in China. Biochem Syst Ecol 69:213–221
Terrones A, Juan A (2023) First dated phylogenetic reconstruction of the genus Tamarix (Tamaricaceae): biogeographical implications and hybridization processes in the Mediterranean Basin. Bot J Linn Soc 203:209–226
Tóth EG, Tremblay F, Housset JM, Bergeron Y, Carcaillet C (2019) Geographic isolation and climatic variability contribute to genetic differentiation in fragmented populations of the long-lived subalpine conifer Pinus cembra L. in the western Alps. Bmc Evol Biol 19:190
Vásquez-Dean J, Maza F, Morel I, Pulgar R, González M (2020) Microbial communities from arid environments on a global scale. A systematic review. Biol Res 53:29
Wang X, Chen L, Ma J (2019) Genomic introgression through interspecific hybridization counteracts genetic bottleneck during soybean domestication. Genome Biol 20:22
Wang Z, He Z, Xu X, Shi X, Ji X, Wang Y (2021) Revealing the salt tolerance mechanism of Tamarix hispida by large-scale identification of genes conferring salt tolerance. Tree Physiol 41:2153–2170
Wang J, Jia H (2017) Sediment record of environmental change at Lake Lop Nur (Xinjiang, NW China) from 13.0 to 5.6 cal ka BP. Chin J Oceano Limn 35:1070–1078
Wang Y, Li J, Qian K, Ye M (2023) Response of Plant Species Diversity to Flood Irrigation in the Tarim River Basin, Northwest China. Sustainability 15:1243
Wang L, Zhang C, Wang Y, Wang Y, Yang C, Lu M et al. (2018) Tamarix hispida aquaporin ThPIP2;5 confers salt and osmotic stress tolerance to transgenic Tamarix and Arabidopsis. Environ Exp Bot 152:158–166
Wu H, Guang X, Al-Fageeh MB, Cao J, Pan S, Zhou H et al. (2014) Camelid genomes reveal evolution and adaptation to desert environments. Nat Commun 5:5188
Xiao H, Liu Z, Wang N, Long Q, Cao S, Huang G et al. (2023) Adaptive and maladaptive introgression in grapevine domestication. P Natl Acad Sci Usa 120:e2222041120
Xie Q, Liu B, Dong W, Li J, Wang D, Liu Z et al. (2023) Comparative transcriptomic and metabolomic analyses provide insights into the responses to NaCl and Cd stress in Tamarix hispida. Sci Total Environ 884:163889
Yang M, Chang H, Qin X, Burr GS, Liu W (2022) Aridification in the Asian Interior Recorded by Mineral Assemblages in Tarim Basin since the Late Miocene and Its Link to Global Cooling. Minerals 12:1543
Yang J, Lee SH, Goddard ME, Visscher PM (2011) GCTA: A Tool for Genome-wide Complex Trait Analysis. Am J Hum Genet 88:76–82
Yue D, Zheng X-J, Li Y, Yan (2014) Dynamics of water usage in Haloxylon ammodendron in the southern edge of the Gurbantünggüt Desert. Chin J Plant Ecol 38:1214–1225
Zhang Y, Ma H, Zhou T, Zhu Z, Zhang Y, Zhao X et al. (2022) ThASR3 confers salt and osmotic stress tolerances in transgenic Tamarix and Arabidopsis. Bmc Plant Biol 22:586
Zhang Q, Sun C, Chen Y, Chen W, Xiang Y, Li J et al. (2022) Recent Oasis Dynamics and Ecological Security in the Tarim River Basin, Central Asia. Sustainability 14:3372
Zhang H-X, Wang Q, Jia S-W (2020) Genomic Phylogeography of Gymnocarpos przewalskii (Caryophyllaceae): Insights into Habitat Fragmentation in Arid Northwestern China. Diversity 12:335
Zhao X, Wang R, Zhang Y, Li Y, Yue Y, Zhou T et al. (2021) Comprehensive analysis of the stress associated protein (SAP) gene family in Tamarix hispida and the function of ThSAP6 in salt tolerance. Plant Physiol Bioch 165:1–9
Zhou X, Yi S, Zhao W, Zhou Q, Shen J, Li D et al. (2021) Genetic Diversity and Population Differentiation of Kashgarian Loach (Triplophysa yarkandensis) in Xinjiang Tarim River Basin. Biology 10:734
Zhu P, He L, Li Y, Huang W, Xi F, Lin L et al. (2014) OTG-snpcaller: An Optimized Pipeline Based on TMAP and GATK for SNP Calling from Ion Torrent Data (V Brusic, Ed.). Plos One 9:e97507
Zhu B, Yu J, Qin X, Rioual P, Zhang Y, Xiong H (2014) Formation and evolution of sand deserts in Xinjiang, Northwest China: II. The palaeo-environmental reconstruction. J Geogr Sci 24:539–559
Acknowledgements
This research was financially supported by the Third Xinjiang Scientific Expedition Program (grant number: 2022xjkk0202), the National Natural Science Foundation of China (grant number: 32170391 and 31960229), and “Western Light” program of the Chinese Academy of Sciences (grant number: 2022-XBQNXZ-007).
Author information
Authors and Affiliations
Contributions
HXZ and XJS designed the study. HWT, JZ, and WHM collected the data and interpreted the results. HWT analyzed the data and drafted the manuscript. HXZ revised the manuscript. HXZ and XJS obtained the funding for the project. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Research ethics statement
The materials used in this study were not related to protected species and were within the limits of the relevant national laws.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Pår Ingvarsson.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tian, H., Zhang, H., Shi, X. et al. Population genetic diversity and environmental adaptation of Tamarix hispida in the Tarim Basin, arid Northwestern China. Heredity 133, 298–307 (2024). https://doi.org/10.1038/s41437-024-00714-0
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41437-024-00714-0
This article is cited by
-
Telomere to telomere level genome assembly of the Yarkand hare (Lepus yarkandensis)
Scientific Data (2026)
-
Unveiling the genetic landscape of Taverniera cuneifolia (Roth) Arn. in India: Population structure and conservation implications for a medicinal Fabaceae species with restricted global distribution
Genetic Resources and Crop Evolution (2025)