Abstract
Mitonuclear discordance between species is readily documented in marine fishes. Such discordance may either be the result of past natural phenomena or the result of recent introgression from previously seperated species after shifts in their spatial distributions. Using ancient DNA spanning five millennia, we here investigate the long-term presence of Pacific bluefin tuna (Thunnus orientalis) and albacore (Thunnus alalunga) -like mitochondrial (MT) genomes in Atlantic bluefin tuna (Thunnus thynnus), a species with extensive exploitation history and observed shifts in abundance and age structure. Comparing ancient (n = 130) and modern (n = 78) Atlantic bluefin MT genomes from most of its range, we detect no significant spatial or temporal population structure, which implies ongoing gene flow between populations and large effective population sizes over millennia. Moreover, we identify discordant MT haplotypes in ancient specimens up to 5000 years old and find that the frequency of these haplotypes has remained similar through time. We therefore conclude that MT discordance in the Atlantic bluefin tuna is not driven by recent introgression. Our observations provide oldest example of directly observed MT discordance in the marine environment, highlighting the utility of ancient DNA to obtain insights in the long-term persistence of such phenomena.
Similar content being viewed by others
Introduction
Discordance between mitochondrial (MT) and nuclear gene phylogenies is commonly observed in eukaryotes and can result from incomplete lineage sorting (ILS) or introgression from another species (Kimball et al. 2021; Tamashiro et al. 2019; Platt et al. 2018). Although frequency of the phenomenon across biological systems remains debated, the increased use of next-generation sequencing across non-model taxa has revealed mitonuclear discordance to be a more common phenomenon in nature than previously thought (Dagilis et al. 2022). The majority of documented mitonuclear discordance in animals has been explained as the result of introgression from a closely related species (Sloan et al. 2017; Pons et al. 2014; Toews and Brelsford 2012). Typically inherited maternally in vertebrates, the non-recombining introgressed MT genome remains largely intact over time (Seixas et al. 2018; Brown 2008). The presence of introgressed MT haplotypes can cause significant bias when using mitogenomic data to describe a species demographic properties or evolutionary history. Even rare hybridization events can result in the presence of whole MT haplotypes that do not accurately reflect the typical history or demography of the taxon. For example, the presence of introgressed MT haplotypes may dominate genealogies with recent dispersal history and thereby overshadow genetic signals from past dispersal events (Sloan et al. 2017; Ballard and Whitlock 2004). Presence of heterospecific haplotypes will also affect population genomic analyses by inflating measures of genetic diversity and divergence (Oosting et al. 2023; Rodriguez and Krug 2022; Wang et al. 2022; Hawks 2017). Avoiding such inflation is important because these statistics can influence management choices (Willi et al. 2022; Hohenlohe et al. 2021; Kardos et al. 2021) and increased measures of genetic diversity or effective population size may exaggerate the genetic robustness of a truly vulnerable population.
Marine fish hybridize according to their ecologies and life history strategies, thus the rate of hybridization and proportion of introgression will vary according to migration behaviour, spawning site overlap, fecundity, spawning ontology, and offspring survival (Montanari et al. 2016; Gardner 1997; Hubbs 1955). In the economically important redfish (Sebastes spp.), high proportions of introgressive hybridization (15% of all samples) have been found between two species (S. fasciatus and S. mentella) that live sympatrically in hybrid zones and yet maintain their morphology, resembling one of the parent species (Benestan et al. 2021; Roques et al. 2001). Likewise, introgression has been observed in European seabass (Dicentrarchus labrax) (Duranton et al. 2020; Vandeputte et al. 2019), capelin (Mallotus villosus) (Cayuela et al. 2020; Colbeck et al. 2011), European anchovy (Engraulis encrasicolus) (Le Moan et al. 2016), Australasian snapper (Chrysophrys auratus) (Oosting et al. 2023) and Atlantic and Pacific herring (Clupea harengus and C. pallasii) (Semenova 2020).
Formation of hybrid zones after recent range shifts induced by contemporary climate change have already been observed in a number of species (Kersten et al. 2023; Ottenburghs 2021; Taylor and Larson 2019; Ryan et al. 2018; Garroway et al. 2010) including marine fish (Muhlfeld et al. 2014; Potts et al. 2014). The formation of such hybrid zones can have both deleterious and advantageous effects. For instance, in trout, warmer freshwater temperatures and lower precipitation is expected to increase introgressive hybridization between native European brown trout (Salmo trutta) and released non-native brown trout in Mediterranean rivers, potentially leading to loss of local genetic variants (Vera et al. 2023). Yet in rainbowfish (Melanotaenia spp.), it has been suggested that introgressive hybridization contributes to climate change resiliency by incorporating potentially adaptive genetic variation (Brauer et al. 2023; Turbek and Taylor 2023). Regardless of the evolutionary consequences, knowledge about the timing of the introgression is necessary to understand if it is anthropogenic impacts that increase rates of hybridization, thereby positively or negatively altering the adaptive potential of species (Xuereb et al. 2021; Hoffmann and Sgrò 2011).
Atlantic bluefin tuna (Thunnus thynnus, Linneaus 1758) is a highly migratory marine predatory fish distributed across the Atlantic Ocean (SCRS 2023; Nøttestad et al. 2020; Block 2019). Atlantic bluefin exhibits strong natal homing behaviour (Brophy et al. 2016; Boustany et al. 2008; Block et al. 2005) and is therefore managed as two separate stocks: the larger Eastern stock spawning predominantly in the Mediterranean, and a smaller Western stock spawning predominantly in the Gulf of Mexico (ICCAT 2023). Recent studies, however, have demonstrated weak genetic divergence in Atlantic bluefin and the existence of a previously unknown spawning ground in the Slope Sea where the stocks seem to interbreed (Diaz-Arce et al. 2024; Aalto et al. 2023; Andrews et al. 2021; Rodríguez‐Ezpeleta et al. 2019), thereby challenging the assumption of two reproductively isolated populations. After severe international overfishing during the last century, the Eastern Atlantic bluefin stock has at present recovered due to strict management measures and favourable oceanographic conditions in the recent decade (ICCAT 2022a, 2022b) followed by improved recruitment with a series of very strong year classes (i.e. individuals born during the same spawning season) (ICCAT 2023; Reglero et al. 2018; Garcia et al. 2013). Nonetheless, the heavy exploitation (Andrews et al. 2022; Block 2019; MacKenzie et al. 2009), lead to shifts in its age structure and foraging behaviour (Andrews et al. 2023a; Di Natale 2015; MacKenzie et al. 2014; Worm and Tittensor 2011). These distributional changes, as well as the establishment of potentially new spawning grounds may impact the potential for introgression between different species.
The phylogeny within the Thunnus genus has been debated and was only recently resolved (Díaz-Arce et al. 2016; Santini et al. 2013; Viñas and Tudela 2009; Chow et al. 2006; Alvarado Bremer et al. 1997; Chow and Kishino 1995). The Atlantic bluefin was previously thought to be a subspecies of Northern bluefin tuna together with Pacific bluefin (Thunnus orientalis, Temminck and Schlegel 1844). The bluefins are now regarded as distinct species forming a monophyletic group (Ciezarek et al., (2019); Díaz-Arce et al. 2016; Chow et al. 2006) (see Fig. S1), with non-overlapping ranges (Tseng et al. 2011), with the albacore tuna (Thunnus alalunga, Bonnaterre 1788) consistently appearing as sister-species. Yet in mitochondrial phylogenies, the Pacific bluefin is more closely related to albacore tuna (Gong et al. 2017; Viñas and Tudela 2009; Chow et al. 2006) than the Atlantic bluefin. Albacore tuna is found in both the Pacific, Indian and Atlantic Oceans, including the Mediterranean Sea, typically preferring warmer waters than the Pacific and Atlantic bluefins, but with largely overlapping ranges and spawning areas (Saber et al. 2015; Chow and Ushiama 1995).
Pacific bluefin- and albacore-like MT genomes have been observed in the Atlantic bluefin and Atlantic bluefin- and albacore-like MT genomes have been observed the Pacific bluefin, but no bluefin-like MT genomes have been found in albacore (e.g. Diaz-Arce et al. 2024; Chow and Kishino 1995). The presence of the discordant MT genomes has been explained by introgression (Viñas et al. 20032011; Viñas and Tudela 2009; Rooker et al. 2007; Chow et al. 2006; Alvarado Bremer et al. 2005; Carlsson et al. 2004; Chow and Kishino 1995; Chow and Inoue 1993). In the Atlantic bluefin, the rates of Pacific bluefin- and albacore-like MT genomes are similar at around 2–5% (Viñas and Tudela 2009; Rooker et al. 2007). Nonetheless, it is unclear if these rates are stable over longer periods of time. In addition to recent distributional shifts likely caused by high fishing pressures, it is possible that climate warming has contributed to novel opportunities for introgression in recent decades. The distribution of Atlantic bluefin over the last century has fluctuated with temperature (Faillettaz et al. 2019; Ravier and Fromentin 2004), and ocean warming has been implicated in altering migration patterns, spawning ontology, and habitats of the Atlantic bluefin (Diaz-Arce et al. 2024; Fiksen and Reglero 2022; Faillettaz et al. 2019; Muhling et al. 2011). Determining the frequency of discordant MT genomes in the past can therefore shed light on the drivers of such phenomena in modern populations.
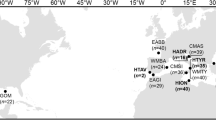
Here, we use DNA extracted from ancient Atlantic bluefin specimens to directly investigate the past occurance of MT discordance and to elucidate potential changes in population structure and genetic diversity over time (Kersten et al. 2023). Fish bones have physiological qualities that may increase the likelihood of finding well preserved DNA (Ferrari et al. 2021; Kontopoulos et al. 2019; Szpak 2011) allowing for whole genome sequencing (Star et al. 2017), even from very limited amounts of bone (e.g. <10 mg) (Atmore et al. 2023). Here we use such ancient DNA (aDNA) methods to analyze MT genomes from 130 ancient and 78 modern Atlantic bluefin spanning a period of approximately 5000 years (Fig. 1). By sampling before and after the period of heavy exploitation (1970–2007) and predating anthropogenic climate change, we investigate spatiotemporal patterns of genetic diversity.
The equal-distance line (45°W) separates the Eastern and Western stocks for management purposes. Sample locations of modern (squares, white boxes) and ancient tuna (circles, brown boxes) used in this study are indicated on the map. Arrows indicate the main migration routes of adult Atlantic bluefin (adapted from Fromentin et al. 2014). GOM Gulf of Mexico, NOR Norway, WMED Western Mediterranean, CMED Central Mediterranean, EMED Eastern Mediterranean. YoY young-of-the-year.
Methods
Collection, extraction, and sequencing of ancient samples from Norway
38 Neolithic (ca. 3000 BCE) tuna bones from the south of Norway were obtained from three archaeological excavations at Jortveit from 2018 to 2020. Bones were found at varying depths (42–130 cm) in six of nine total trenches and were estimated to be from 3700–2500 BCE based on radiocarbon dating of wood and charcoal from the sediment profiles, as well as directly dated bone harpoons. Three of the bones were also directly radiocarbon dated to the period approximately 3400–2800 BCE (Nielsen 2020a, 2020b, 2020c; Nielsen and Persson 2020).
All laboratory work prior to PCR was performed in a dedicated aDNA laboratory at the University of Oslo, following strict anti-contamination protocols (Llamas et al. 2017; Gilbert et al. 2005). All samples were extracted using a standard extraction protocol adapted from Dabney et al. (2013) after a pre-digestion step (DD from Damgaard et al. 2015) or mild bleach treatment and pre-digestion (BleDD from Boessenkool et al. 2017) as described in Ferrari et al. (2021) (Table S1). Dual-indexed sequencing libraries were built as double stranded, blunt-ended libraries following Meyer and Kircher (2010) and Kircher et al. (2011) with modifications or as single stranded libraries following the Santa Cruz Reaction (SCR) protocol (Kapp et al. 2021) (Table S1). Libraries were sequenced on the Illumina HiSeq 4000 or NovaSeq 6000 (SP Flow Cell) platforms at the Norwegian Sequencing Centre with paired-end 150 bp reads and demultiplexed allowing zero mismatches in the index tag. For additional details, see supplementary section 1.2.
Ancient specimens from the Mediterranean
92 individuals from archaeological excavations and zoological collections throughout the Mediterranean region dating from 100 to 1941 CE were obtained from Andrews et al. (2024) as BAM files (Table S2). For additional details about the samples and archaeological sites, see supplementary section 1.8. These samples were prepared and extracted in the Ancient DNA Laboratory of the Department of Cultural Heritage (University of Bologna, Ravenna Campus, Italy), following strict criteria for aDNA analysis as per the Norwegian samples, and sequenced as single-stranded libraries (Kapp et al. 2021) at Macrogen facilities (Seoul, South Korea/Amsterdam, Netherlands) on a HiSeq X (100 bp paired-end) Illumina sequencing platform. Reads were processed using the Paleomix pipeline v.1.2.14 (Schubert et al. 2014) with settings described below (see “Bioinformatic processing of ancient and modern sequence data”), yielding an average of 28% endogenous DNA and 11-fold MT coverage (Table S8).
Collection, extraction, and sequencing of modern samples
Modern tuna tissue samples of migratory, foraging adults from Norway (NOR) (n = 38) were collected by the Norwegian Institute of Marine Research (IMR), from commercial catch off the coast of Møre og Romsdal, Western Norway (Table S3) (Supplementary section 1.3). The modern samples from Norway were all extracted in the modern DNA isolation laboratories at the University of Oslo, using the DNeasy Blood and Tissue kit (Qiagen) and following the manufacturer’s protocol.
Modern larvae or young-of-the-year (YoY) specimens (GOM: Gulf of Mexico, WMED: Western Mediterranean Balearic Islands, CMED: Central Mediterranean Sicily, EMED: Eastern Mediterranean Levantine Sea, n = 40, Table S4) were collected from each of the major Atlantic bluefin spawning sites (Fig. 1) between 2013 and 2018. Juvenile albacore samples from the Bay of Biscay were caught by commercial vessels trolling in the Bay of Biscay between June and September of 2010 (Table S5). Larvae and tissue samples from each specimen were preserved in 96% ethanol and stored at −20 °C until further processing. Modern spawning site and albacore samples were extracted at the University of Bologna by a modified salt-based extraction protocol, as per Cruz et al. (2017), using SSTNE extraction buffer (Blanquer 1990), and treated with RNase to remove residual RNA.
For the Norwegian samples, libraries were built using the TruSeq DNA Nano200 preparation kit (Illumina). Modern spawning site extracts, along with albacore extracts, underwent single stranded library preparation following the SCR library protocol (Kapp et al. 2021). Sequencing and demultiplexing, allowing for zero mismatches, was performed at the Norwegian Sequencing Centre on a combination of the HiSeq 4000 and NovaSeq 6000 (SP Flow Cell) Illumina sequencing platforms with paired-end 150 bp reads for all samples.
Raw sequence data of Pacific bluefin whole genome (Suda et al. 2019) were downloaded from DDBJ (accession no DRA008331) (Table S6) and used for interspecific population structure analyses.
Bioinformatic processing of ancient and modern sequence data
Both modern and ancient reads were processed using the Paleomix pipeline v.1.2.14 (Schubert et al. 2014). All reads were aligned to a draft nuclear (NCBI BioProject: PRJNA408269) and MT reference genome (GenBank accession nr NC_014052.1) with BWA-mem v.0.7.17 for mapping. Only the MT BAMfiles were further processed in GATK v.4.1.4.0 following GATK best practices (McKenna et al. 2010). Filtered VCFs were indexed using Tabix v.0.2.6 (Li 2011) and consensus sequences created as individual fasta files in BCFtools v.1.9 (bcftools consensus -H 1). Outgroup sequences were downloaded from GenBank (Clark et al. 2016) and curated using SeqKit v. 0.11.0 (restart -i) (Shen et al. 2016) so that all sequences started at position 1 in the D-loop, to correspond with the sample sequences. After renaming the fasta headers to their appropriate sample-IDs using BBMap v.38.50b (Bushnell 2014) and combining the files to a multiple sequence alignment (MSA), the joint fasta files were aligned using MAFFT v.7.453 (Katoh et al. 2002) (--auto). For additional details, see supplementary section 1.4.
Population genomic analyses
After an investigation and creation of datasets (supplementary section 1.5), genetic population structure was investigated using Principal component analyses (PCA). A map of missing loci and base variants diverging from the reference genome, was created to assess missing genotypes in both ancient and modern samples and better visualize introgressed specimens. All plots were created with R 4.3 in RStudio (Rstudio Team 2021), using various packages for data loading, analyses, and visualization (supplementary section 1.1).
Genetic diversity was investigated using a range of standard population genetic measurements (number of haplotypes (Nh), haplotype diversity (hD) number of segregating sites (S), nucleotide diversity (π) (Nei 1987), Tajima’s D (TD) (Tajima 1989), and Fu and Li’s F statistic (F) (Fu and Li 1993)) using Fitchi (Matschiner 2016), DnaSP v.6 (Rozas et al. 2017) and the R-package pegas (Paradis 2010). To account for differences in sample sizes across sites when calculating π and TD, an additional analysis using 1000 bootstrap replicates and subsampling five individuals per round without replacement, was performed in pegas on datasets where the total sample size was over five.
Phylogenetic relationships were investigated using both ML and Bayesian approaches. ML trees with 100 nonparametric bootstrap replicates were created in IQTREE v. 1.6.12 (Nguyen et al. 2015). ModelFinder Plus (MFP) (Kalyaanamoorthy et al. 2017) was used to search all available models, and best-fit models were selected according to the Bayesian Information Criterion (BIC) (Schwarz 1978). Bayesian trees were created in BEAST 2 v.2.6.4 (R. Bouckaert et al. 2014), using the Yule model prior under a strict clock with mutation rate 3.6 × 10−8 substitutions per site per year as per Donaldson and Wilson (1999), running MCMC over 800,000,000 generations and sampling once every 1000 generations (supplementary section 1.6). The final trees in all phylogenetic analyses were visualized and curated in FigTree v.1.4.4 (Rambaut 2018).
Evolutionary relationships were visualized using haplotype networks created in Fitchi (--haploid -p) using the ML trees generated in IQTREE (described above) (supplementary section 1.7)
Genetic distance between sample locations was assessed using measures of absolute (dxy) and relative (ΦST) divergence, calculated using DnaSP v.6 (“DNA divergence between populations”, all sites) and Arlequin v.3.5 (Excoffier and Lischer 2010) respectively. In Arlequin, pairwise ΦST was calculated via a distance matrix computed by Arlequin based on Tamura and Nei (1993) and assuming no rate heterogeneity, as suggested by bModelTest (R. R. Bouckaert and Drummond 2017) (implemented in BEAST 2 v.2.6.4 (R. Bouckaert et al. 2014)). To test the significance of ΦST, p-values were generated in Arlequin using 1000 permutations.
Results
DNA yield and library success
A total of 1.7 billion sequencing reads were obtained for the 38 ancient samples from Norway. These specimens had remarkable DNA preservation with 100% library success and yielding, on average, 24% endogenous DNA and 20-fold MT coverage (Table S7). The reads showed postmortem degradation patterns expected for authentic aDNA (Fig. S2). A total of 3.1 billion sequencing reads were obtained for the 84 modern specimens, resulting in 711-fold MT coverage on average for the 78 Atlantic bluefin specimens (Tables S9 and S10) and 221-fold MT coverage on average for the six albacore samples (Table S11). The Pacific bluefin raw sequence data from Suda et al. (2019) yielded 3322-fold MT coverage (Table S12). After stringent filtering, 186 out of 208 specimens (~90%) were kept for further analyses (Tables S7, S8).
Detecting discordant MT genomes
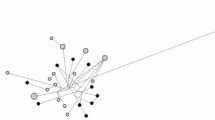
Out of 186 samples analyzed, seven ancient and four modern individuals had MT haplotypes that clustered closely with albacore or Pacific bluefin in the PCA, haplotype network and phylogenies. The PCA reveals three distinct clusters (Fig. 2) with PC1 separating an Atlantic bluefin cluster from a Pacific bluefin and albacore cluster and PC2 separating the latter two speciese. Within the Pacific bluefin cluster, we observe two modern (both NOR) and four ancient (two Norway 3000BCE, one Istanbul 800–1200CE and one Sardinia 1500–1700CE) Atlantic bluefin specimens. Within the albacore cluster, we observe two modern (one NOR and one WMED) and three ancient (one Istanbul - 800–1200CE and two Sicily – 900–1200CE) Atlantic bluefin specimens. The PCA-clusters are reiterated in the haplotype network (Fig. 2). ML and Bayesian phylogenetic analyses provided full statistical support (bootstrap = 100, posterior probability = 1) for the three species as monophyletic groups with the same six Pacific-like and five albacore-like haplotypes again clustering with their respective species (Fig. 2, see also Fig. S6).
A Three species specific clusters detected in ancient and modern Atlantic tuna specimens. The PCA shows three species specific clusters and PCA eigenvalues are shown in the bottom left corner. Modern modern Pacific tuna (blue) and albacore (grey) specimens are included as controls. B Relative abundance of haplotypes per location within each PCA-cluster is visualized as pie-charts, with the number of samples from each location indicated on the slices. C Haplotype network showing three species specific haplotypes. Haplotypes separated by seven or fewer substitutions were collapsed into single nodes. D Interspecific phylogeny of specimens with posterior probability support for the species clades (see also Fig. S6). Colours are representative of the spatiotemporal cohorts listed in the legend of panel (A).
Spatiotemporal population structure
We find no significant mitogenomic differentiation between any of the temporal cohorts. We also observe no spatial differences in the level of genetic variation between any of the sampling locations for Atlantic bluefin. Atlantic bluefin individuals from both management stocks and across the Eastern stock range and spawning areas are scattered across the intraspecific PCA (Fig. S8) and haplotype network (Fig. S9). The sampling locations are also distributed along the entire phylogeny within the Atlantic bluefin group (Fig. 2, see also Figs. S6, S10). The intraspecific Atlantic bluefin haplotype network reveals a star-like pattern with more recent MT haplotypes deriving from an ancestral, central haplotype (Fig. S9).
Genetic divergence and diversity influenced by introgression
Measures of pairwise genetic distance between Atlantic bluefin sampling locations show low levels of absolute (dxy) and relative (Φst) divergence, either excluding (Fig. 3a) or including discordant MT haplotypes (Fig. 3b). Genetic differentiation increases when divergent MT haplotypes are included, which are not present at each location or temporal cohort. In all cases, levels of Φst remained low and non-significant (Fig. S7) across all populations. Including individuals with discordant MT haplotypes increased values of nucleotide diversity π and S (Table S15). The number of haplotypes (hD) is not impacted; most sample locations only contained unique specimens (N = Nh) therefore leading to a hD of 1, meaning 100% probability of obtaining unique samples during random sampling. Tajima’s D (TD) was also not affected by the inclusion of introgressed individuals and was significantly negative for most locations and temporal cohorts and when analyzing all specimens jointly (Table S15).
Pairwise population divergence is presented as a heatmap showing absolute (dxy) and relative (ΦST) divergence between populations when (A) excluding and (B) including the discordant MT genomes. Divergence is increased when including discordant MT genomes. Locations containing discordant MT genomes are highlighted with darker shading in panel (B). The nucleotide diversity within each population is shown on the diagonal. P-values for ΦST can be found in supplementary (Fig. S7).
Frequency of discordant MT haplotypes over time
We observe discordant MT genomes in Atlantic bluefin throughout a 5000-year chronology (Fig. 4). The earliest observation is the presence of two Pacific bluefin-like MT genomes in the Neolithic (ca. 3000 BCE) in Norway. Pacific bluefin-like MT genomes are further found in early medieval Istanbul (800–1200 CE), late-medieval Sardinia (1500–1700 CE) and modern Norway. Albacore-like MT haplotypes are found in early medieval Istanbul (800–1200 CE) and Sicily (900–1200 CE), modern Western Mediterranean and modern Norway.
Individual tuna specimens (n = 208) (circle or star) are grouped according to their age, determined by archaeological context. Specimens were either modern (n = 78), or ancient (n = 130) and dated by archaeological context. Uncertainty in the age range of ancient specimens is depicted beneath their respective sample sets (light shading).
Discussion
We here present a 5000-year chronology of MT discordance in the Atlantic bluefin tuna. The observation of divergent MT haplotypes in the Neolithic (ca. 3000 BCE), turn of the millennium (800–1200 CE), and in present day populations indicates that it is long-term natural phenomena rather than recent spatial shifts that explains their presence. Moreover, our results show that the frequency of MT discordance in the Atlantic bluefin has remained stable over millennia despite shifts in abundance and distribution of Atlantic bluefin populations.
Evidence of mitonuclear discordance through time
We obtain similar proportions of for Pacific bluefin- and albacore-like MT haplotypes at 2.6% (2/78 individuals) each as reported by previous studies in our modern samples. Including our ancient samples in the calculation, the proportion of MT discordance remains astonishingly stable with 3.2% Pacific bluefin-like haplotypes (6/186) and 2.7% albacore-like haplotypes (5/186) across all samples. In total, six Pacific-like and five albacore-like haplotypes consistently cluster with their respective species through all interspecific analyses. We do not observe albacore-like haplotypes from the Neolithic period, but because of the low frequency of introgression compared to the sample size we speculate this is likely due to sampling stochasticity rather than their lack of presence at the time. Still, it cannot be excluded that Neolithic climate conditions drove albacore populations away from areas inhabited by Atlantic bluefin.
The observation of discordant MT haplotypes relies on the assumption that the sampled archaeological bones used in this study all stem from Atlantic bluefin individuals. The nuclear DNA of the ancient Mediterranean samples used in this study, has been analyzed as part of Andrews et al. (2024) where all samples, including the individuals with diverging MT haplotypes cluster together with the modern Mediterranean and Norwegian Atlantic bluefin individuals. Some of the ancient Norwegian samples from Jortveit were also included in these analyses, however the two individuals with Pacific-like MT genomes were excluded from the analyses due to bad preservation of the nuclear genome. We therefore cannot be certain that these two individuals are in fact Atlantic bluefin vertebrae, as the Pacific and Atlantic bluefin vertebrae are morphologically diffucult to distinguish. However, no migration of Pacific bluefin into the Atlantic Ocean has ever been observed and there is no known range overlap. The optimal temperature range of Pacific bluefin is around 15–20 °C (Kitagawa et al. 2006), while the ocean surface temperature in Neolithic Norway peaked at around 8 °C in mid-august (Risebrobakken et al., (2011)). We therefore find it highly unlikely that these two individuals stem from migratory Pacific bluefin.
Should the mitonuclear discordance be driven by past introgressive hybridization from both albacore and Pacific bluefin, the location and timing of such events remain to be investigated. While the albacore has overlapping ranges and spawning areas with both bluefin species, the Pacific and Atlantic bluefins are geographically separated with no documented migration. The potential migration of Pacific bluefin into the Atlantic Ocean has been hypothesized to occur via the Indian Ocean and following the Agulhas current around the tip of Africa (Alvarado Bremer et al. 2005). Whether this represents a contemporary migration route or a historical process where past, stronger currents might have facilitated admixture is unclear (Alvarado Bremer et al. 2005). The similar proportion of mitonuclear discordance from both albacore and Pacific bluefin and the lack of divergent MT haplotypes in the albacore makes the range-overlapping albacore an unlikely carrier of MT haplotypes between the bluefins in the case of introgression. Given that Pacific bluefin also contains Atlantic bluefin-like MT haplotypes, incomplete lineage sorting (ILS) can also explain their presence. The Thunnus genus is thought to have diverged rapidly within the last 6–10 million years, with a more recent speciation of the Pacific and Atlantic bluefins only around 400,000 years ago (Ciezarek et al., (2019); Díaz-Arce et al. 2016; Santini et al. 2013). While introgressive hybridization is the likely origin of albacore-like haplotypes in both bluefin species (Ciezarek et al., (2019)), observed gene-tree versus species-tree discordance did not deviate from expectations under ILS in the same study (Ciezarek et al., (2019)). These results indicate that the observed patterns of Pacific-like MT haplotypes in the Atlantic bluefin population, and vice versa, may be a result of ILS rather than introgressive hybridization. Larger genomic databases are required to furhter delineate between these two hypotheses.
While our historical investigation focuses on the eastern Atlantic with samples from the Mediterranean and Norway, discordant MT genomes from albacore have also been found in the Gulf of Mexico (1% frequency) and the Slope Sea (6% frequency). Because such MT genomes in Atlantic bluefin were first observed in the eastern Atlantic, their presence in the western Atlantic has been hypothesized to be introduced via gene flow from the Mediterranean (Diaz-Arce et al. 2024). An increase in gene flow from the Mediterranean into the Gulf of Mexico and Slope Sea will likely erode genetic differences between the two management stocks (Diaz-Arce et al. 2024). Direct observations of hybridization events within the Western Atlantic bluefin stock have not been made, although albacore is known to spawn across tropical waters including the South-West Sargasso Sea as well as the Mediterranean (NOAA 2023; ICCAT 2016, 2020). Future studies could monitor the frequencies of MT discordance in the Gulf of Mexico to disentangle the recently suggested changes in demographic patterns (Diaz-Arce et al. 2024) and possibly locate the origins of contemporary introgression events.
Discordant MT haplotypes impact estimates of mitogenomic differentiation
The presence of discordant haplotypes increases measures of genetic diversity (S and π), which is driven by the high number of diverging bases in the introgressed MT genomes (Fig. S5). The inclusion of these diverging haplotypes also influences the measures of genetic divergence (dxy and ΦST) between locations and temporal cohorts, consistently increasing the absolute genetic diversity (dxy) and altering the pattern of relative genetic divergence (ΦST). Tajima’s D (TD) was not consistently affected by the inclusion of introgressed individuals, although in some cases TD changed value and lost or attained significance when divergent haplotypes were included (Table S15). Considering these results, one needs to be aware of highly diverging haplotypes when extrapolating population genetic statistics from subsamples of natural populations containing highly diverging haplotypes. The low frequency of divergent haplotypes in Atlantic bluefin causes stochasticity at low sample sizes, and we find that their inclusion inflates population genomic statistics that are commonly used for management and population viability assessments (e.g. Dapporto et al. 2022; Hohenlohe et al. 2021; Zhang et al. 2020).
Spatiotemporal population structure
We find no significant divergence and no pattern of mitogenomic differentiation between any of the spatial or temporal cohorts. Ancient and modern samples largely intermixed in all analyses, suggesting mitogenomic stability and temporal continuity through time. Similar observations in other species, such as Atlantic cod (Gadus morhua) (Martínez-García et al. 2021) and New Zealand snapper (Chrysophrys auratus) (Oosting et al. 2023) emphasize the low power of the MT genome to observe spatiotemporal differentiation in wide ranging fish species. The regular presence of identical haplotypes across sampling locations and temporal cohorts emphasizes the lack of mitogenomic variation and informative markers for population structure in this species. Although we cautiously removed identical samples from the same archaeological excavations, the presence of identical samples across cohorts that were processed in different laboratories shows that identical MT haploptypes can be observed regularly.
Population genetic statistics confirmed low mitogenomic variation with no significant divergence between any of the spatiotemporal cohorts and no temporal loss of genetic diversity despite heavy exploitation (Fig. 3A, Table S15). Across datasets, TD was negative and often significant, suggesting an excess of rare variants in the datasets. This is indicative of either positive selection or recent population expansion (Fijarczyk and Babik 2015; Delph and Kelly 2014). Population expansion is further corroborated in the intraspecific haplotype network, where newer haplotypes are derived from a shared central haplotype forming a star-like pattern (Fig. S9). These results highlight robust preservation of the MT genome despite centuries of human exploitation.
Conclusion
Atlantic bluefin tuna has experienced significant changes in distribution linked to sea surface temperature oscillations during the past centuries (Faillettaz et al. 2019; Muhling et al. 2011; Ravier and Fromentin 2004), alongside intense exploitation (Andrews et al. 2022; Block 2019), biomass depletion, range contraction, trophic niche loss (Andrews et al. 2021; Di Natale 2015; Tangen 2009), followed by recovery, increased biomass, and range expansion of the Eastern Atlantic bluefin stock during the last decade (ICCAT 2023; Nøttestad et al. 2020). Despite such extensive spatial shifts in distribution over time, we show that the presence of MT discordance is a long-term natural phenomenon in Atlantic bluefin. The stable frequency over time suggests that this phenomenon is robust against recent spatial shifts due to anthropogenic impacts. By providing a baseline observation, our study highlight the utility of aDNA to obtain temporal insights in the long-term persistence of such phenomenon.
Data availability
All mitochondrial BAM files used in this study have been uploaded in the European Nucleotide Archive (ENA) and can be accessed at https://www.ebi.ac.uk/ena/browser/view/PRJEB74135. See supplementary csv containing filenames, project accession number and ENA identifier for all samples.
References
Aalto EA, Dedman S, Stokesbury MJW, Schallert RJ, Castleton M, Block BA (2023) Evidence of bluefin tuna (Thunnus thynnus) spawning in the Slope Sea region of the Northwest Atlantic from electronic tags. ICES J Mar Sci 80:861–877. https://doi.org/10.1093/icesjms/fsad015
Alvarado Bremer JR, Naseri I, Ely B (1997) Orthodox and unorthodox phylogenetic relationships among tunas revealed by the nucleotide sequence analysis of the mitochondrial DNA control region. J Fish Biol 50:540–554. https://doi.org/10.1111/j.1095-8649.1997.tb01948.x
Alvarado Bremer JR, Viñas J, Mejuto J, Ely B, Pla C (2005) Comparative phylogeography of Atlantic bluefin tuna and swordfish: The combined effects of vicariance, secondary contact, introgression, and population expansion on the regional phylogenies of two highly migratory pelagic fishes. Mol Phylogenetics Evolut 36:169–187. https://doi.org/10.1016/j.ympev.2004.12.011
Andrews AJ, Di Natale A, Bernal-Casasola D, Aniceti V, Onar V, Oueslati T, Theodropoulou T, Morales-Muñiz A, Cilli E, Tinti F (2022) Exploitation history of Atlantic bluefin tuna in the Eastern Atlantic and Mediterranean—Insights from ancient bones. ICES J Mar Sci 79:247–262. https://doi.org/10.1093/icesjms/fsab261
Andrews AJ, Puncher GN, Bernal-Casasola D, Di Natale A, Massari F, Onar V, Toker NY, Hanke A, Pavey SA, Savojardo C, Martelli PL, Casadio R, Cilli E, Morales-Muñiz A, Mantovani B, Tinti F, Cariani A (2021) Ancient DNA SNP-panel data suggests stability in bluefin tuna genetic diversity despite centuries of fluctuating catches in the eastern Atlantic and Mediterranean. Sci Rep. 11:20744. https://doi.org/10.1038/s41598-021-99708-9
Andrews AJ, Eriksen EF, Star B, Præbel K, Natale AD, Malca E, Zapfe G, Onar V, Aniceti V, Carenti G, Piquès G, Nielsen SV, Persson P, Piattoni F, Fontani F, Atmore LM, Kersten O, Tinti F, Cilli E, Cariani A (2024) Ancient DNA reveals historical demographic decline and genetic erosion in the Atlantic bluefin tuna (p. 2024.09.14.613028). bioRxiv. https://doi.org/10.1101/2024.09.14.613028
Andrews AJ, Pampoulie C, Di Natale A, Addis P, Bernal-Casasola D, Aniceti V, Carenti G, Gómez-Fernández V, Chosson V, Ughi A, Von Tersch M, Fontanals-Coll M, Cilli E, Onar V, Tinti F, Alexander M (2023) Exploitation shifted trophic ecology and habitat preferences of Mediterranean and Black Sea bluefin tuna over centuries. Fish Fisheries. https://doi.org/10.1111/faf.12785
Atmore LM, Ferrari G, Martínez-García L, van der Jagt I, Blevis R, Granado J, Häberle S, Dierickx K, Quinlan LM, Lõugas L, Makowiecki D, Hufthammer AK, Barrett JH, Star B (2023) Ancient DNA sequence quality is independent of fish bone weight. J Archaeol Sci 149:105703. https://doi.org/10.1016/j.jas.2022.105703
Ballard JWO, Whitlock MC (2004) The incomplete natural history of mitochondria. Mol Ecol 13:729–744. https://doi.org/10.1046/j.1365-294X.2003.02063.x
Benestan LM, Rougemont Q, Senay C, Normandeau E, Parent E, Rideout R, Bernatchez L, Lambert Y, Audet C, Parent GJ (2021) Population genomics and history of speciation reveal fishery management gaps in two related redfish species (Sebastes mentella and Sebastes fasciatus). Evolut Appl 14:588–606. https://doi.org/10.1111/eva.13143
Blanquer A (1990) Phylogéographie intraspécifique d’un poisson marin, le flet Platichthys flesus L. (Heterosomata): Polymorphisme des marqueurs nucléaires et mitochondriaux. (PhD Thesis, University of Montipelier, France)
Block BA, Teo SLH, Walli A, Boustany A, Stokesbury MJW, Farwell CJ, Weng KC, Dewar H, Williams TD (2005) Electronic tagging and population structure of Atlantic bluefin tuna. Nature 434:1121–1127. https://doi.org/10.1038/nature03463
Block BA (2019) The Future of Bluefin Tunas: Ecology, Fisheries Management, and Conservation. JHU Press.
Boessenkool S, Hanghøj K, Nistelberger HM, Der Sarkissian C, Gondek AT, Orlando L, Barrett JH, Star B (2017) Combining bleach and mild predigestion improves ancient DNA recovery from bones. Mol Ecol Resour 17:742–751. https://doi.org/10.1111/1755-0998.12623
Bouckaert R, Heled J, Kühnert D, Vaughan T, Wu C-H, Xie D, Suchard MA, Rambaut A, Drummond AJ (2014) BEAST 2: A Software Platform for Bayesian Evolutionary Analysis. PLOS Comput Biol 10:e1003537. https://doi.org/10.1371/journal.pcbi.1003537
Bouckaert RR, Drummond AJ (2017) bModelTest: Bayesian phylogenetic site model averaging and model comparison. BMC Evolut Biol 17:42. https://doi.org/10.1186/s12862-017-0890-6
Boustany AM, Reeb CA, Block BA (2008) Mitochondrial DNA and electronic tracking reveal population structure of Atlantic bluefin tuna (Thunnus thynnus). Mar Biol 156:13–24. https://doi.org/10.1007/s00227-008-1058-0
Brauer CJ, Sandoval-Castillo J, Gates K, Hammer MP, Unmack PJ, Bernatchez L, Beheregaray LB (2023) Natural hybridization reduces vulnerability to climate change. Nat Clim Change 13:Article 3. https://doi.org/10.1038/s41558-022-01585-1
Brophy D, Haynes P, Arrizabalaga H, Fraile I, Fromentin JM, Garibaldi F, Katavic I, Tinti F, Karakulak FS, Macías D, Busawon D, Hanke A, Kimoto A, Sakai O, Deguara S, Abid N, Santos MN (2016) Otolith shape variation provides a marker of stock origin for north Atlantic bluefin tuna (Thunnus thynnus). Mar Freshw Res 67:1023–1036. https://doi.org/10.1071/MF15086
Brown KH (2008) Fish mitochondrial genomics: Sequence, inheritance and functional variation. J Fish Biol 72:355–374. https://doi.org/10.1111/j.1095-8649.2007.01690.x
Bushnell B (2014) BBMap: A Fast, Accurate, Splice-Aware Aligner (LBNL-7065E). Lawrence Berkeley National Lab. (LBNL), Berkeley, CA (United States). https://www.osti.gov/biblio/1241166
Carlsson J, McDOWELL JR, Díaz-Jaimes P, Carlsson JEL, Boles SB, Gold JR, Graves JE (2004) Microsatellite and mitochondrial DNA analyses of Atlantic bluefin tuna (Thunnus thynnus thynnus) population structure in the Mediterranean Sea. Mol Ecol 13:3345–3356. https://doi.org/10.1111/j.1365-294X.2004.02336.x
Cayuela H, Rougemont Q, Laporte M, Mérot C, Normandeau E, Dorant Y, Tørresen OK, Hoff SNK, Jentoft S, Sirois P, Castonguay M, Jansen T, Praebel K, Clément M, Bernatchez L (2020) Shared ancestral polymorphisms and chromosomal rearrangements as potential drivers of local adaptation in a marine fish. Mol Ecol 29:2379–2398. https://doi.org/10.1111/mec.15499
Chow S, Ushiama H (1995) Global population structure of albacore (Thunnus alalunga) inferred by RFLP analysis of the mitochondrial ATPase gene. Mar Biol 123:39–45. https://doi.org/10.1007/BF00350321
Chow S, Nakagawa T, Suzuki N, Takeyama H, Matsunaga T (2006) Phylogenetic relationships among Thunnus species inferred from rDNA ITS1 sequence. J Fish Biol 68:24–35. https://doi.org/10.1111/j.0022-1112.2006.00945.x
Chow S, Inoue S (1993) Intra and interspesific restriction fragment length polymorphism in mitochondrial genes of Thunnus tuna species.
Chow S, Kishino H (1995) Phylogenetic relationships between tuna species of the genus Thunnus (Scombridae: Teleostei): Inconsistent implications from morphology, nuclear and mitochondrial genomes. J Mol Evolut 41(6). https://doi.org/10.1007/BF00173154
Ciezarek AG, Osborne OG, Shipley ON, Brooks EJ, Tracey SR, McAllister JD, Gardner LD, Sternberg MJE, Block B, Savolainen V (2019) Phylotranscriptomic insights into the diversification of endothermic Thunnus Tunas. Mol Biol Evolut 36:84–96. https://doi.org/10.1093/molbev/msy198
Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW (2016) GenBank. Nucleic Acids Res 44:D67–D72. https://doi.org/10.1093/nar/gkv1276
Colbeck GJ, Turgeon J, Sirois P, Dodson JJ (2011) Historical introgression and the role of selective vs. Neutral processes in structuring nuclear genetic variation (AFLP) in a circumpolar marine fish, the capelin (Mallotus villosus). Mol Ecol 20:1976–1987. https://doi.org/10.1111/j.1365-294X.2011.05069.x
Cruz VP, Vera M, Pardo BG, Taggart J, Martinez P, Oliveira C, Foresti F (2017) Identification and validation of single nucleotide polymorphisms as tools to detect hybridization and population structure in freshwater stingrays. Mol Ecol Resour 17:550–556. https://doi.org/10.1111/1755-0998.12564
Dabney J, Knapp M, Glocke I, Gansauge M-T, Weihmann A, Nickel B, Valdiosera C, Garcia N, Paabo S, Arsuaga J-L, Meyer M (2013) Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proc Natl Acad Sci 110:15758–15763. https://doi.org/10.1073/pnas.1314445110
Dagilis AJ, Peede D, Coughlan JM, Jofre GI, D’Agostino ERR, Mavengere H, Tate AD, Matute DR (2022) A need for standardized reporting of introgression: Insights from studies across eukaryotes. Evolut Lett 6:344–357. https://doi.org/10.1002/evl3.294
Damgaard PB, Margaryan A, Schroeder H, Orlando L, Willerslev E, Allentoft ME (2015) Improving access to endogenous DNA in ancient bones and teeth. Sci Rep. 5:11184. https://doi.org/10.1038/srep11184
Dapporto L, Menchetti M, Vodă R, Corbella C, Cuvelier S, Djemadi I, Gascoigne-Pees M, Hinojosa JC, Lam NT, Serracanta M, Talavera G, Dincă V, Vila R (2022) The atlas of mitochondrial genetic diversity for Western Palaearctic butterflies. Glob Ecol Biogeogr 31:2184–2190. https://doi.org/10.1111/geb.13579
Delph LF, Kelly JK (2014) On the importance of balancing selection in plants. N. Phytologist 201: https://doi.org/10.1111/nph.12441
Di Natale A (2015) Review of the historical and biological evidences about a population of Bluefin tuna (Thunnus Thynnus L.) in the Eastern Mediterranean and the Black Sea. ICCAT. https://www.iccat.int/Documents/CVSP/CV071_2015/n_3/CV071031098.pdf
Díaz-Arce N, Arrizabalaga H, Murua H, Irigoien X, Rodríguez-Ezpeleta N (2016) RAD-seq derived genome-wide nuclear markers resolve the phylogeny of tunas. Mol Phylogenetics Evolut 102:202–207. https://doi.org/10.1016/j.ympev.2016.06.002
Díaz-Arce N, Gagnaire P-A, Richardson DE, Walter III JF, Arnaud-Haond S, Fromentin J-M, Brophy D, Lutcavage M, Addis P, Alemany F, Allman R, Deguara S, Fraile I, Goñi N, Hanke AR, Karakulak FS, Pacicco A, Quattro JM, Rooker JR, Rodríguez-Ezpeleta N (2024) Unidirectional trans-Atlantic gene flow and a mixed spawning area shape the genetic connectivity of Atlantic bluefin tuna. Mol Ecol 33:e17188. https://doi.org/10.1111/mec.17188
Donaldson KA, Wilson RR (1999) Amphi-panamic geminates of snook (percoidei: centropomidae) provide a calibration of the divergence rate in the mitochondrial DNA control region of fishes. Mol Phylogenetics Evolut 13:208–213. https://doi.org/10.1006/mpev.1999.0625
Duranton M, Allal F, Valière S, Bouchez O, Bonhomme F, Gagnaire P-A (2020) The contribution of ancient admixture to reproductive isolation between European sea bass lineages. Evolut Lett 4:226–242. https://doi.org/10.1002/evl3.169
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Faillettaz R, Beaugrand G, Goberville E, Kirby RR (2019) Atlantic Multidecadal Oscillations drive the basin-scale distribution of Atlantic bluefin tuna. Sci Adv 5:eaar6993. https://doi.org/10.1126/sciadv.aar6993
Ferrari G, Cuevas A, Gondek-Wyrozemska AT, Ballantyne R, Kersten O, Pálsdóttir AH, van der Jagt I, Hufthammer AK, Ystgaard I, Wickler S, Bigelow GF, Harland J, Nicholson R, Orton D, Clavel B, Boessenkool S, Barrett JH, Star B (2021) The preservation of ancient DNA in archaeological fish bone. J Archaeol Sci 126:105317. https://doi.org/10.1016/j.jas.2020.105317
Fijarczyk A, Babik W (2015) Detecting balancing selection in genomes: Limits and prospects. Mol Ecol 24:3529–3545. https://doi.org/10.1111/mec.13226
Fiksen Ø, Reglero P (2022) Atlantic bluefin tuna spawn early to avoid metabolic meltdown in larvae. Ecology 103:e03568. https://doi.org/10.1002/ecy.3568
Fromentin J-M, Reygondeau G, Bonhommeau S, Beaugrand G (2014) Oceanographic changes and exploitation drive the spatio-temporal dynamics of Atlantic bluefin tuna (Thunnus thynnus). Fish Oceanogr 23:147–156. https://doi.org/10.1111/fog.12050
Fu YX, Li WH (1993) Statistical tests of neutrality of mutations. Genetics 133:693–709. https://doi.org/10.1093/genetics/133.3.693
García A, Cortés D, Quintanilla J, Rámirez T, Quintanilla L, Rodríguez JM, Alemany F (2013) Climate-induced environmental conditions influencing interannual variability of Mediterranean bluefin (Thunnus thynnus) larval growth. Fish. Oceanogr. 22:273–287. https://doi.org/10.1111/fog.12021
Gardner JPA (1997) Hybridization in the Sea. In J. H. S. Blaxter & A. J. Southward (Eds.), Advances in Marine Biology (Vol. 31, pp. 1–78). Academic Press. https://doi.org/10.1016/S0065-2881(08)60221-7
Garroway CJ, Bowman J, Cascaden TJ, Holloway GL, Mahan CG, Malcolm JR, Steele MA, Turner G, Wilson PJ (2010) Climate change induced hybridization in flying squirrels. Glob Change Biol 16:113–121. https://doi.org/10.1111/j.1365-2486.2009.01948.x
Gilbert MTP, Bandelt H-J, Hofreiter M, Barnes I (2005) Assessing ancient DNA studies. Trends Ecol Evolut 20:541–544. https://doi.org/10.1016/j.tree.2005.07.005
Gong L, Liu L-Q, Guo B-Y, Ye Y-Y, Lü Z-M (2017) The complete mitochondrial genome characterization of Thunnus obesus (Scombriformes: Scombridae) and phylogenetic analyses of Thunnus. Conserv Genet Resour 9:379–383. https://doi.org/10.1007/s12686-017-0688-2
Hawks J (2017) Introgression Makes Waves in Inferred Histories of Effective Population Size. Hum Biol 89:67–80
Hoffmann AA, Sgrò CM (2011) Climate change and evolutionary adaptation. Nature 470:Article 7335. https://doi.org/10.1038/nature09670
Hohenlohe PA, Funk WC, Rajora OP (2021) Population genomics for wildlife conservation and management. Mol Ecol 30:62–82. https://doi.org/10.1111/mec.15720
Hubbs CL (1955) Hybridization between Fish Species in Nature. Syst Biol 4:1–20. https://doi.org/10.2307/sysbio/4.1.1
ICCAT (2016) Report of the 2016 ICCAT North and South Atlantic albacore stock assessment meeting. ICCAT. https://www.iccat.int/Documents/Meetings/Docs/2016_ALB_REPORT_ENG.pdf
ICCAT (2020) 2020 Advice to the commission - 5.1 ALB – Atlantic Albacore. ICCAT. https://www.iccat.int/documents/scrs/execsum/alb_eng.pdf
ICCAT (2022a) Recommendation By ICCAT Amending The Recommendation 21-08 Establishing A Multi-Annual Management Plan For Bluefin Tuna In The Eastern Atlantic And The Mediterranean (Rec-22-08).
ICCAT (2022b) Recommendation By ICCAT Establishing A Management Procedure For Atlantic Bluefin Tuna To Be Used For Both The Western Atlantic And Eastern Atlantic And Mediterranean Management Areas (Rec 22-09).
ICCAT. (2023) REPORT for biennial period, 2022-23 PART I (2022)—Vol. 3 Annual Reports. ICCAT. https://www.iccat.int/Documents/BienRep/REP_TRILINGUAL_22-23_I_3.pdf
IMR (2021) Makrellstørje. Havforskningsinstituttet (Institute of Marine research), Bergen, Norway. https://www.hi.no/hi/temasider/arter/makrellstorje
Kalyaanamoorthy S, Minh BQ, Wong TK, von Haeseler A, Jermiin LS (2017) ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589. https://doi.org/10.1038/nmeth.4285
Kapp JD, Green RE, Shapiro B (2021) A fast and efficient single-stranded genomic library preparation method optimized for ancient DNA. J Heredity 112:241–249. https://doi.org/10.1093/jhered/esab012
Kardos M, Armstrong EE, Fitzpatrick SW, Hauser S, Hedrick PW, Miller JM, Tallmon DA, Funk WC (2021) The crucial role of genome-wide genetic variation in conservation. Proc Natl Acad Sci 118:e2104642118. https://doi.org/10.1073/pnas.2104642118
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066. https://doi.org/10.1093/nar/gkf436
Kersten O, Star B, Krabberød AK, Atmore LM, Tørresen OK, Anker-Nilssen T, Descamps S, Strøm H, Ulf S. Johansson, Sweet PR, Jakobsen KS, Boessenkool S (2023) Hybridization of Atlantic puffins in the Arctic coincides with 20th-century climate change. Sci Adv 9(40), https://doi.org/10.1126/sciadv.adh1407
Kimball RT, Guido M, Hosner PA, Braun EL (2021) When good mitochondria go bad: Cyto-nuclear discordance in landfowl (Aves: Galliformes). Gene 801:145841. https://doi.org/10.1016/j.gene.2021.145841
Kircher M, Sawyer S, Meyer M (2011) Double indexing overcomes inaccuracies in multiplex sequencing on the Illumina platform. Nucleic Acids Res 40:e3. https://doi.org/10.1093/nar/gkr771
Kitagawa T, Kimura S, Nakata H, Yamada H (2006) Thermal adaptation of pacific bluefin tuna Thunnus orientalis to temperate waters. Fish Sci 72:149–156. https://doi.org/10.1111/j.1444-2906.2006.01129.x
Kontopoulos I, Penkman K, McAllister GD, Lynnerup N, Damgaard PB, Hansen HB, Allentoft ME, Collins MJ (2019) Petrous bone diagenesis: A multi-analytical approach. Palaeogeogr, Palaeoclimatol, Palaeoecol 518:143–154. https://doi.org/10.1016/j.palaeo.2019.01.005
Li H (2011) Tabix: Fast retrieval of sequence features from generic TAB-delimited files. Bioinformatics 27:718–719. https://doi.org/10.1093/bioinformatics/btq671
Llamas B, Valverde G, Fehren-Schmitz L, Weyrich LS, Cooper A, Haak W (2017) From the field to the laboratory: Controlling DNA contamination in human ancient DNA research in the high-throughput sequencing era. STAR: Sci Technol Archaeol Res 3:1–14. https://doi.org/10.1080/20548923.2016.1258824
MacKenzie BR, Mosegaard H, Rosenberg AA (2009) Impending collapse of bluefin tuna in the Northeast Atlantic and Mediterranean. Conserv Lett 2:26–35. https://doi.org/10.1111/j.1755-263X.2008.00039.x
MacKenzie BR, Payne MR, Boje J, Høyer JL, Siegstad H (2014) A cascade of warming impacts brings bluefin tuna to Greenland waters. Glob Change Biol 20:2484–2491. https://doi.org/10.1111/gcb.12597
Martínez-García L, Ferrari G, Oosting T, Ballantyne R, van der Jagt I, Ystgaard I, Harland J, Nicholson R, Hamilton-Dyer S, Baalsrud HT, Brieuc MSO, Atmore LM, Burns F, Schmölcke U, Jakobsen KS, Jentoft S, Orton D, Hufthammer AK, Barrett JH, Star B (2021) Historical Demographic Processes Dominate Genetic Variation in Ancient Atlantic Cod Mitogenomes. https://doi.org/10.17863/CAM.71522
Matschiner M (2016) Fitchi: Haplotype genealogy graphs based on the Fitch algorithm. Bioinformatics 32:1250–1252. https://doi.org/10.1093/bioinformatics/btv717
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. https://doi.org/10.1101/gr.107524.110
Meyer M, Kircher M (2010) Illumina Sequencing Library Preparation for Highly Multiplexed Target Capture and Sequencing. Cold Spring Harbor Protocols 2010(6):pdb.prot5448. 10.1101/pdb.prot5448
Le Moan A, Gagnaire P-A, Bonhomme F (2016) Parallel genetic divergence among coastal–marine ecotype pairs of European anchovy explained by differential introgression after secondary contact. Mol Ecol 25:3187–3202. https://doi.org/10.1111/mec.13627
Montanari SR, Hobbs J-PA, Pratchett MS, van Herwerden L (2016) The importance of ecological and behavioural data in studies of hybridisation among marine fishes. Rev Fish Biol Fish 26:181–198. https://doi.org/10.1007/s11160-016-9420-7
Muhlfeld CC, Kovach RP, Jones LA, Al-Chokhachy R, Boyer MC, Leary RF, Lowe WH, Luikart G, Allendorf FW (2014) Invasive hybridization in a threatened species is accelerated by climate change. Nat Clim Change 4:Article 7. https://doi.org/10.1038/nclimate2252
Muhling BA, Lee S-K, Lamkin JT, Liu Y (2011) Predicting the effects of climate change on bluefin tuna (Thunnus thynnus) spawning habitat in the Gulf of Mexico. ICES J Mar Sci 68:1051–1062. https://doi.org/10.1093/icesjms/fsr008
Nei M (1987) Molecular Evolutionary Genetics. Columbia Univ. Press, New York
Nguyen L-T, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evolution 32:268–274. https://doi.org/10.1093/molbev/msu300
Nielsen SV, Persson P (2020) The Jortveit farm wetland: A Neolithic fishing site on the Skagerrak coast, Norway. J Wetl Archaeol 0:1–24. https://doi.org/10.1080/14732971.2020.1776495
Nielsen SV (2020a) Myrfunn fra Jortveit. Rapport 1 av 3. Jortveit, 172/2, Grimstad, Agder. https://www.duo.uio.no/handle/10852/85693
Nielsen SV (2020b) Myrfunn fra Jortveit. Rapport 2 av 3. Jortveit, 172/2, Grimstad, Agder. https://www.duo.uio.no/handle/10852/85694
Nielsen SV (2020c) Myrfunn fra Jortveit. Rapport 3 av 3. Jortveit, 172/2, Grimstad, Agder. https://www.duo.uio.no/handle/10852/85695
NOAA (2023) North Atlantic Albacore Tuna | NOAA Fisheries (New England/Mid-Atlantic,Southeast). NOAA. https://www.fisheries.noaa.gov/species/north-atlantic-albacore-tuna
Nøttestad L, Boge E, Ferter K (2020) The comeback of Atlantic bluefin tuna (Thunnus thynnus) to Norwegian waters. Fish Res 231:105689. https://doi.org/10.1016/j.fishres.2020.105689
Oosting T, Martínez-García L, Ferrari G, Verry AJF, Scarsbrook L, Rawlence NJ, Wellenreuther M, Star B, Ritchie PA (2023) Mitochondrial genomes reveal mid-Pleistocene population divergence, and post-glacial expansion, in Australasian snapper (Chrysophrys auratus). Heredity 130:Article 1. https://doi.org/10.1038/s41437-022-00579-1
Ottenburghs J (2021) The genic view of hybridization in the Anthropocene. Evolut Appl 14:2342–2360. https://doi.org/10.1111/eva.13223
Paradis E (2010) pegas: An R package for population genetics with an integrated–modular approach. Bioinform 26:419–420. https://doi.org/10.1093/bioinformatics/btp696
Platt II RN, Faircloth BC, Sullivan KAM, Kieran TJ, Glenn TC, Vandewege MW, Lee Jr. TE, Baker RJ, Stevens RD, Ray DA (2018) Conflicting Evolutionary Histories of the Mitochondrial and Nuclear Genomes in New World Myotis Bats. Syst Biol 67:236–249. https://doi.org/10.1093/sysbio/syx070
Pons J-M, Sonsthagen S, Dove C, Crochet P-A (2014) Extensive mitochondrial introgression in North American Great Black-backed Gulls (Larus marinus) from the American Herring Gull (Larus smithsonianus) with little nuclear DNA impact. Heredity 112:Article 3. https://doi.org/10.1038/hdy.2013.98
Potts WM, Henriques R, Santos CV, Munnik K, Ansorge I, Dufois F, Booth AJ, Kirchner C, Sauer WHH, Shaw PW (2014) Ocean warming, a rapid distributional shift, and the hybridization of a coastal fish species. Glob Change Biol 20:2765–2777. https://doi.org/10.1111/gcb.12612
Rambaut A (2018) FigTree v1.4.4. GitHub. https://github.com/rambaut/figtree/releases
Ravier C, Fromentin J-M (2004) Are the long-term fluctuations in Atlantic bluefin tuna (Thunnus thynnus) population related to environmental changes? Fish Oceanogr 13:145–160. https://doi.org/10.1111/j.1365-2419.2004.00284.x
Reglero P, Balbin R, Abascal FJ, Medina A, Alvarez-Berastegui D, Rasmuson L, Mourre B, Saber S, Ortega A, Blanco E, de la Gandara F, Alemany FJ, Ingram GW, Hidalgo M (2018) Pelagic habitat and offspring survival in the eastern stock of Atlantic bluefin tuna. ICES J Marine Sci, https://doi.org/10.1093/icesjms/fsy135.
Risebrobakken B, Dokken T, Smedsrud LH, Andersson C, Jansen E, Moros M, Ivanova EV (2011) Early Holocene temperature variability in the Nordic Seas: The role of oceanic heat advection versus changes in orbital forcing. Paleoceanography, 26(4). https://doi.org/10.1029/2011PA002117
Rodriguez AK, Krug PJ (2022) Ecological speciation by sympatric host shifts in a clade of herbivorous sea slugs, with introgression and localized mitochondrial capture between species. Mol Phylogenetics Evolut 174:107523. https://doi.org/10.1016/j.ympev.2022.107523
Rodríguez‐Ezpeleta N, Díaz‐Arce N, Walter JF, Richardson DE, Rooker JR, Nøttestad L, Hanke AR, Franks JS, Deguara S, Lauretta MV, Addis P, Varela JL, Fraile I, Goñi N, Abid N, Alemany F, Oray IK, Quattro JM, Sow FN, Arrizabalaga H (2019) Determining natal origin for improved management of Atlantic bluefin tuna. Front Ecol Environ 17:439–444. https://doi.org/10.1002/fee.2090
Rooker JR, Alvarado Bremer JR, Block BA, Dewar H, de Metrio G, Corriero A, Kraus RT, Prince ED, Rodríguez-Marín E, Secor DH (2007) Life History and Stock Structure of Atlantic Bluefin Tuna (Thunnus thynnus). Rev Fish Sci 15:265–310. https://doi.org/10.1080/10641260701484135
Roques Sé, SÉvigny J-M, Bernatchez L (2001) Evidence for broadscale introgressive hybridization between two redfish (genus Sebastes) in the North-West Atlantic: A rare marine example. Mol Ecol 10:149–165. https://doi.org/10.1046/j.1365-294X.2001.01195.x
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A (2017) DnaSP 6: DNA Sequence Polymorphism Analysis of Large Data Sets. Mol Biol Evolut 34:3299–3302. https://doi.org/10.1093/molbev/msx248
RStudio Team (2021) RStudio: Integrated Development for R. RStudio, PBC, Boston, MA. http://www.rstudio.com/
Ryan SF, Deines JM, Scriber JM, Pfrender ME, Jones SE, Emrich SJ, Hellmann JJ (2018) Climate-mediated hybrid zone movement revealed with genomics, museum collection, and simulation modeling. Proc Natl Acad Sci 115:E2284–E2291. https://doi.org/10.1073/pnas.1714950115
Saber S, de Urbina JO, Gómez-Vives MJ, Macías D (2015) Some aspects of the reproductive biology of albacore Thunnus alalunga from the Western Mediterranean Sea. J Mar Biol Assoc U Kingd 95:1705–1715. https://doi.org/10.1017/S002531541500020X
Santini F, Carnevale G, Sorenson L (2013) First molecular scombrid timetree (Percomorpha: Scombridae) shows recent radiation of tunas following invasion of pelagic habitat. Ital J Zool 80:210–221. https://doi.org/10.1080/11250003.2013.775366
Schubert M, Ermini L, Sarkissian CD, Jónsson H, Ginolhac A, Schaefer R, Martin MD, Fernández R, Kircher M, McCue M, Willerslev E, Orlando L (2014) Characterization of ancient and modern genomes by SNP detection and phylogenomic and metagenomic analysis using PALEOMIX. Nat Protoc 9:Article 5. https://doi.org/10.1038/nprot.2014.063
Schwarz G (1978) Estimating the Dimension of a Model. Ann Stat 6:461–464
SCRS (2023) REPORT for biennial period, 2022-23 PART I (2022)—Vol. 2. ICCAT. https://www.iccat.int/Documents/BienRep/REP_EN_22-23-I-2.pdf
Seixas FA, Boursot P, Melo-Ferreira J (2018) The genomic impact of historical hybridization with massive mitochondrial DNA introgression. Genome Biol 19:91. https://doi.org/10.1186/s13059-018-1471-8
Semenova AV (2020) Introgressive Hybridization in the Secondary Contact Area of the Atlantic Herring Clupea harengus and the Pacific Herring C. pallasii (Clupeidae): Ecological Basis, Geographical Structure, and Temporal Variability of the Hybridization Zone. J Ichthyol 60:626–642. https://doi.org/10.1134/S0032945220030169
Shen W, Le S, Li Y, Hu F (2016) SeqKit: A Cross-Platform and Ultrafast Toolkit for FASTA/Q File Manipulation. PLOS ONE 11:e0163962. https://doi.org/10.1371/journal.pone.0163962
Sloan DB, Havird JC, Sharbrough J (2017) The on-again, off-again relationship between mitochondrial genomes and species boundaries. Mol Ecol 26:2212–2236. https://doi.org/10.1111/mec.13959
Star B, Boessenkool S, Gondek AT, Nikulina EA, Hufthammer AK, Pampoulie C, Knutsen H, André‚ C, Nistelberger HM, Dierking J, Petereit C, Heinrich D, Jakobsen KS, Stenseth NC, Jentoft S, Barrett JH (2017) Ancient DNA reveals the Arctic origin of Viking Age cod from Haithabu, Germany. Proc Natl Acad Sci 114:9152–9157. https://doi.org/10.1073/pnas.1710186114
Suda A, Nishiki I, Iwasaki Y, Matsuura A, Akita T, Suzuki N, Fujiwara A (2019) Improvement of the Pacific bluefin tuna (Thunnus orientalis) reference genome and development of male-specific DNA markers. Sci Rep. 9:Article 1. https://doi.org/10.1038/s41598-019-50978-4
Szpak P (2011) Fish bone chemistry and ultrastructure: Implications for taphonomy and stable isotope analysis. J Archaeol Sci 38:3358–3372. https://doi.org/10.1016/j.jas.2011.07.022
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595. https://doi.org/10.1093/genetics/123.3.585
Tamashiro RA, White ND, Braun MJ, Faircloth BC, Braun EL, Kimball RT (2019) What are the roles of taxon sampling and model fit in tests of cyto-nuclear discordance using avian mitogenomic data? Mol Phylogenetics Evolut 130:132–142. https://doi.org/10.1016/j.ympev.2018.10.008
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526. https://doi.org/10.1093/oxfordjournals.molbev.a040023
Tangen M (2009) The Norwegian Fishery for Atlantic Bluefin tuna. ICCAT Collect Vol Sci Pap ICCAT 63:79–93. https://www.iccat.int/Documents/CVSP/CV063_2009/n_1/CVOL63010079.pdf 2009
Taylor SA, Larson EL (2019) Insights from genomes into the evolutionary importance and prevalence of hybridization in nature. Nat Ecol Evolut 3:Article 2. https://doi.org/10.1038/s41559-018-0777-y
Toews DPL, Brelsford A (2012) The biogeography of mitochondrial and nuclear discordance in animals. Mol Ecol 21:3907–3930. https://doi.org/10.1111/j.1365-294X.2012.05664.x
Tseng M-C, Shiao J-C, Hung Y-H (2011) Genetic identification of Thunnus orientalis, T. thynnus, and T. maccoyii by a cytochrome b gene analysis. Environ Biol Fishes 91:103–115. https://doi.org/10.1007/s10641-010-9764-0
Turbek SP, Taylor SA (2023) Hybridization provides climate resilience. Nat Clim Change 13:Article 3. https://doi.org/10.1038/s41558-022-01586-0
Vandeputte M, Gagnaire P-A, Allal F (2019) The European sea bass: A key marine fish model in the wild and in aquaculture. Anim Genet 50:195–206. https://doi.org/10.1111/age.12779
Vera M, Aparicio E, Heras S, Abras A, Casanova A, Roldán M-I, García-Marin J-L (2023) Regional environmental and climatic concerns on preserving native gene pools of a least concern species: Brown trout lineages in Mediterranean streams. Sci Total Environ 862:160739. https://doi.org/10.1016/j.scitotenv.2022.160739
Viñas J, Tudela S (2009) A validated methodology for genetic identification of Tuna species (Genus Thunnus). PLoS ONE 4:e7606. https://doi.org/10.1371/journal.pone.0007606
Viñas J, Gordoa A, Fernández-Cebrián R, Pla C, Vahdet Ü, Araguas RM (2011) Facts and uncertainties about the genetic population structure of Atlantic bluefin tuna (Thunnus thynnus) in the Mediterranean. Implications for fishery management. Rev Fish Biol Fish 21:527–541. https://doi.org/10.1007/s11160-010-9174-6
Viñas J, Pla C, Tawil MY, Hattour A, Farrugia AF (2003) Mitochondrial Genetic Characterization Of Bluefin Tuna (Thunnus Thynnus) From Three Mediterranean (Libya, Malta, Tunisia); And One Atlantic Locations (Gulf Of Cadiz). 7.
Wang P, Dong N, Wang M, Sun G, Jia Y, Geng X, Liu M, Wang W, Pan Z, Yang Q, Li H, Wei C, Wang L, Zheng H, He S, Zhang X, Wang Q, Du X (2022) Introgression from Gossypium hirsutum is a driver for population divergence and genetic diversity in Gossypium barbadense. Plant J 110:764–780. https://doi.org/10.1111/tpj.15702
Willi Y, Kristensen TN, Sgrò CM, Weeks AR, Ørsted M, Hoffmann AA (2022) Conservation genetics as a management tool: The five best-supported paradigms to assist the management of threatened species. Proc Natl Acad Sci 119:e2105076119. https://doi.org/10.1073/pnas.2105076119
Worm B, Tittensor DP (2011) Range contraction in large pelagic predators. Proc Natl Acad Sci 108:11942–11947. https://doi.org/10.1073/pnas.1102353108
Xuereb A, D’Aloia CC, Andrello M, Bernatchez L, Fortin M-J (2021) Incorporating putatively neutral and adaptive genomic data into marine conservation planning. Conserv Biol 35:909–920. https://doi.org/10.1111/cobi.13609
Zhang Q, Sun C, Zhu Y, Xu N, Liu H (2020) Genetic diversity and structure of the round-tailed paradise fish (Macropodus ocellatus): Implications for population management. Glob Ecol Conserv 21:e00876. https://doi.org/10.1016/j.gecco.2019.e00876
Acknowledgements
This work was supported by the European Union’s Horizon 2020 Research and Innovation Programme under the Marie Skłodowska-Curie grant agreement No. 813383 (SeaChanges) and the 4-OCEANS Synergy grant agreement no. 951649. The European Research Agency is not responsible for any use that may be made of the information this work contains. Special thanks goes to the Norwegian Sequencing Centre (University of Oslo; https://www.sequencing.uio.no) for the sequencing of genomic libraries. Computation was performed using the resources and assistance from SIGMA2. The collection of recent samples in Norway was financed by the Institute of Marine Research. The collection of larva samples in the Gulf of Mexico was funded by the NOAA RESTORE Science Programme, BLOOFINZ-GoM #NOAA-NOS-NCCOS-2017-2004875; Cooperative Institute for Marine and Atmospheric Studies #NA20OAR4320472. This work is a contribution to the https://tunaarchaeology.org/ project. We are grateful to Ørjan Sørensen at the Institute of Marine Research (IMR) for his contributions to the collection and shipping of the modern Atlantic bluefin samples from Norway and providing sample metadata. We thank Verónica Gómez-Fernández (Instituto Nacional de Investigaciones Científicas y Ecológicas, Spain), Gabriele Carenti (CEPAM, CNRS, Université Côte d’Azur, France) and Darío Bernal-Casasola (Department of History, Geography and Philosophy, Faculty of Philosophy and Letters, University of Cádiz, Spain) for sample collection. We thank Leif Nøttestad (IMR) and Mark Ravinet (CEES, University of Oslo) for comments and stimulating scientific discussions.
Author information
Authors and Affiliations
Contributions
Author contributions following CRediT (https://credit.niso.org/). EFE: Conceptualization, Data curation, Formal Analysis, Investigation, Methodology, Visualization, Writing—original draft, Writing—review & editing. AJA: Data curation, Investigation, Methodology, Writing—review & editing. SVN: Resources, Writing—review & editing. PP: Writing—review & editing. EM: Resources, Writing—review & editing. VO: Resources. VA: Resources. GP: Resources. FP: Data curation, methodology, Writing—review & editing. FF: Investigation. MW: Resources, Writing-review & editing. KF: Resources, Writing-review & editing. OK: Investigation, Writing-review & editing. GF: Investigation, Writing-review & editing. AC: Resources, Funding acquisition, Writing-review & editing. FT: Resources, Funding acquisition, Writing-review & editing. EC: Resources, Funding acquisition. LMA: Software, Supervision, Writing—review & editing. BS: Conceptualization, Funding acquisition, Project administration, Software, Supervision, Writing—original draft, Writing—review & editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Research Ethics Statement
This research did not involve human participants, human material, or human data, nor did it involve experiments with animals. All animal material (fish bones/skin/muscle) was obtained from animals that were already deceased. Archaeological material was obtained in collaboration with local partners, with all necessary permissions and approvals from relevant authorities, if required.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Giorgio Bertorelle.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Eriksen, E.F., Andrews, A.J., Nielsen, S.V. et al. Five millennia of mitonuclear discordance in Atlantic bluefin tuna identified using ancient DNA. Heredity 134, 175–185 (2025). https://doi.org/10.1038/s41437-025-00745-1
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41437-025-00745-1
This article is cited by
-
Delineating and identifying species from mitochondrial DNA only: a cautionary tale from bats
Mammalian Biology (2025)