Fig. 1: A highly reliable PAM prediction algorithm.
From: Automated identification of sequence-tailored Cas9 proteins using massive metagenomic data
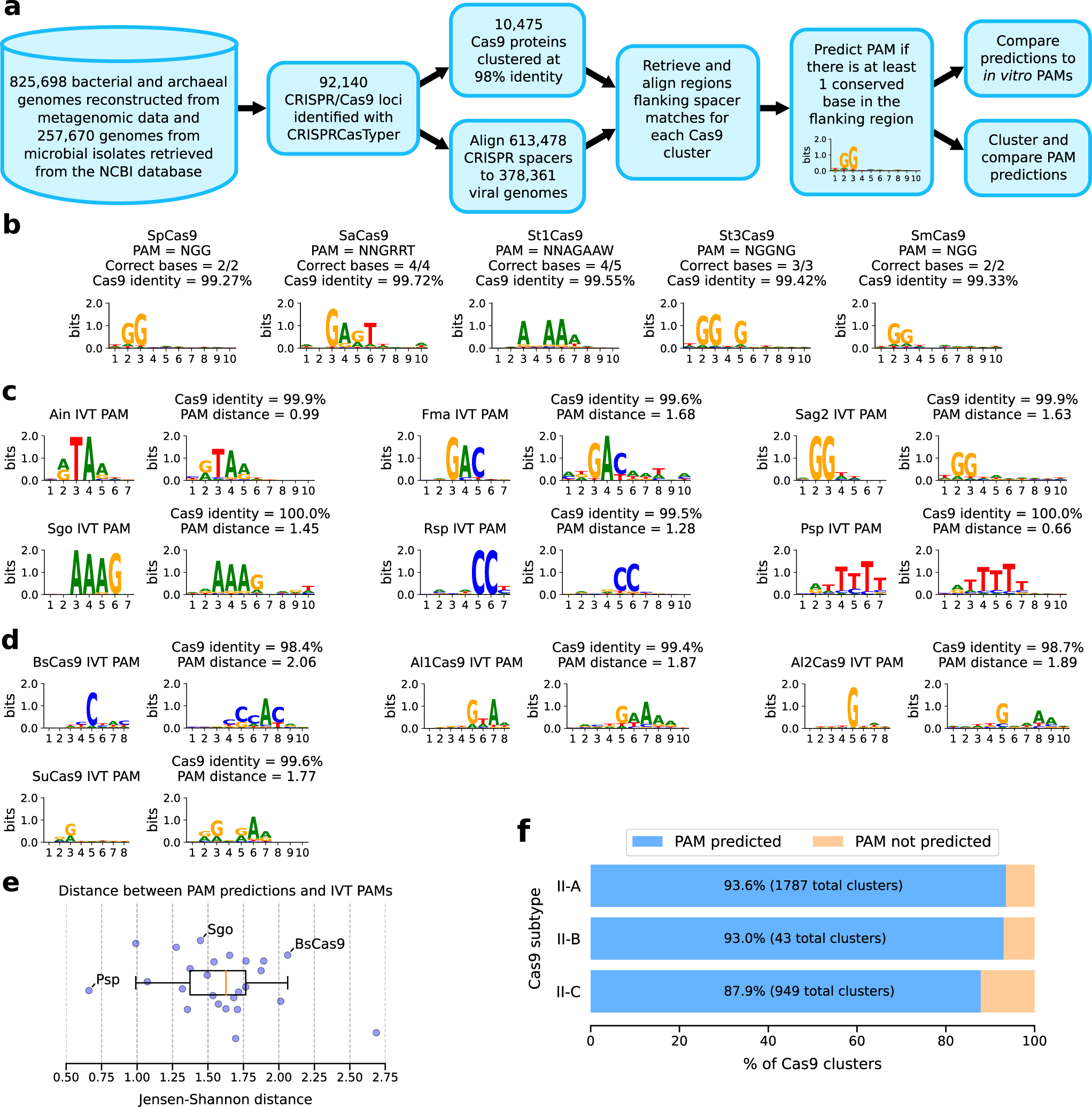
a Schematic description of the pipeline used to generate PAM predictions. b Predicted PAM for Cas9 proteins with experimentally verified PAM. Sequence identity (%) between the reported Cas9 and the Cas9 identified in our dataset is also shown. c In vitro PAMs (left logos) and predicted PAM (right logos) for selected Cas9 variants from Gasiunas et al.19 (additional orthologs are in Supplementary Fig. 2). A distance measure between predicted and verified PAMs is also shown. d In vitro and predicted PAMs (left and right logo respectively) for selected Cas9 variants from our dataset. e Boxplot of distance between in vitro and predicted PAMs for all tested orthologs. Most orthologs have a low (<2 bits) PAM distance. Central line, median; box limits, upper and lower quartiles; whiskers, ×1.5 interquartile range; n = 25 independent PAMs. f Fraction of Cas9 clusters with more than 10 mapped spacers and a predicted PAM, for each Cas9 subtype.
