Abstract
Vibrational spectroscopy is a powerful tool for spectral imaging of biological samples, thanks to its narrow bandwidth (10 cm⁻¹) compared to fluorescence. Single-molecule vibrational spectroscopy has been demonstrated with near-field amplification as in surface-enhanced Raman spectroscopy or fluorescence detection as in stimulated Raman excited fluorescence and bond-selective fluorescence-detected infrared-excited spectro-microscopy. However, these methods often require elaborate sample preparation or sometimes generate background signals when unintended processes lead to fluorescence emission. In response to these issues, we developed electronic resonance stimulated Raman scattering (ER-SRS) to achieve single-molecule sensitivity in far-field vibrational microscopy without relying on fluorescence detection. ER-SRS has encountered difficulties due to large electronic backgrounds. To overcome this, we employed Raman-amplified nonfluorescent molecular probe (RANMP) alongside our synchronously pumped, independently tunable double optical parametric oscillators for effective optimization of the signal-to-background ratio. The optimization of probe and light source allowed us to successfully detect ER-SRS signal from single particles in solution and from single molecules embedded in polymer matrix. ER-SRS combined with RANMP provides single-molecule sensitivity without fluorescence detection, enabling applications in biological and chemical imaging, particularly in multiplexed imaging.
Similar content being viewed by others
Introduction
Fluorescence microscopy has revolutionized biology with its unmatched molecular specificity and single-molecule sensitivity1,2,3,4. Single-molecule fluorescence microscopy has unveiled the individual trajectories of biological molecules, revealing the molecular mechanisms5,6. Fluorescence detection has driven advancements in cutting-edge technologies such as massively parallel DNA sequencing7 and super-resolution optical microscopy8,9,10,11. In recent years, fluorescence microscopy evolved to highly multiplexed imaging, opening new fields of spatial genomics, transcriptomics and proteomics6,12,13. However, fluorescence-based multiplexed imaging relies on iterative cycles of labeling, imaging and washing for each molecular target due to the broad bandwidth (1500 cm-1) of excitation and emission that limit its ability to spectrally distinguish different molecules14.
In contrast, vibrational transitions, such as Raman scattering and infrared (IR) absorption, have a much narrower bandwidth (10 cm−1) compared to fluorescence, making them highly suitable for simultaneously differentiating multiple substances. However, the intrinsic low intensity of Raman signals necessitates extensive signal amplification. Surface-enhanced Raman spectroscopy (SERS) enhances the Raman signal by over 107 using plasmonic nanoparticles, enabling single-molecule detection15,16,17. However, the preparation of SERS substrates can be complicated and requires precise control of nanoparticle fabrication and probe positioning for consistent results18,19.
Electronic pre-resonance stimulated Raman scattering (EPR-SRS, Fig. 1A) enhances Raman sensitivity by up to 1011 compared to spontaneous Raman without near-field effects20. However, it still requires fluorescence detection, such as stimulated Raman excited fluorescence21 (SREF, Supplementary Fig. 1A), for single-molecule sensitivity. Efficient SREF fluorophores must fulfill multiple requirements: (1) conjugation of vibrational modes to a chromophore, (2) large Raman cross-section, (3) high near-infrared (NIR) absorbance, and (4) high fluorescence quantum yields22. However, high quantum yields lead to high background, including anti-Stokes fluorescence23 from thermally excited populations, limiting the signal-to-background (S/B) ratios and practicality. Bond-selective fluorescence-detected infrared-excited spectro-microscopy24 (BonFIRE) also demonstrated single-molecule sensitivity (Supplementary Fig. 1A). However, BonFIRE suffered from anti-Stokes fluorescence and IR-induced photothermal background, requiring high-frequency modulation for background substraction24.
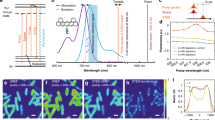
A Energy diagram of two-beam EPR-SRS (left) and ER-SRS (right) processes, where the frequency difference between the pump and Stokes beams is resonant to the vibrational transition. In ER-SRS (right), the pump beam is rigorously resonant to the electronic transition while in EPR-SRS (left), the pump beam is slightly off-resonant but still close to the electronic transition. B Two-beam ER-SRS instrument with synchronously pumped double OPO system with two independently tunable OPOs whose signal beams are each used as the pump and Stokes beams. C Absorption and emission spectra of Rho800 and the wavelength combinations of pump and Stokes used for collecting ER-SRS spectra. D ER-SRS spectra with various combinations of the pump and Stokes wavelengths indicated in (C). SR gain signal is plotted against the frequency differences between the pump and Stokes beams. Powers are set as 5 mW for pump and 5 mW for Stokes. Lines indicate spline interpolation. Error bars represent the mean ± standard deviation (n = 5). E Signal and background intensities in the ER-SRS spectra in (D) plotted against the pump wavelengths. Intensities are reported in arbitrary units (a.u.).
Our research aims to leverage electronic resonance stimulated Raman scattering (ER-SRS) to amplify Raman sensitivity without the aid of near-field effects or fluorescence detection (Fig. 1A). First demonstrated in 2018, ER-SRS has never demonstrated single-molecule sensitivity due to large electronic background and lack of suitable probes and light sources25. In this regard, our strategy combines nonfluorescent, highly conjugated Raman-active probe and frequency optimization for two-beam ER-SRS by using our unique, independently tunable double optical parametric oscillator (OPO) laser system (Fig. 1B and Supplementary Fig. 1B), enabling single-molecule detection.
Results
ER-SRS and EPR-SRS of Rhodamine 800 fluorophore
Rhodamine 800 (Rho800) serves as a key probe for EPR-SRS and SREF20,21,23. Utilizing our independently tunable double OPO system, we analyzed the ER-SRS and EPR-SRS spectra of Rho800 under various pump/Stokes wavelength combinations (Figs. 1C-E). The selected wavelengths meet these conditions: (1) \({\omega }_{{{\rm{p}}}}-{\omega }_{{{\rm{s}}}}=\,{\omega }_{{{\rm{v}}}}\); (2) \({|\omega }_{{{\rm{p}}}}-{\omega }_{{{\rm{abs}}},\max }|\) < 700 cm−1 for ER-SRS25; (3) 1400 cm−1 < \({|\omega }_{{{\rm{p}}}}-{\omega }_{{{\rm{abs}}},\max }|\) < 4200 cm−1 for EPR-SRS20, where \({\omega }_{{{\rm{p}}}}\) and \({\omega }_{{{\rm{s}}}}\) are pump and Stokes frequencies; \({\omega }_{{{\rm{abs}}},\max }\) and \({\omega }_{{{\rm{v}}}}\) are the frequencies at the absorption max and Raman shift of the probe, respectively. For Rho800, \({\omega }_{{{\rm{v}}}}\) = 2229.6 cm−1 (Supplementary Fig. 2A). Using our synchronized, yet independently tunable double OPO system with a tuning range of 660–960 nm, we optimized pump/Stokes combinations to maximize the S/B ratio (peak height/background). The pump wavelength was tuned around the absorption max (λmax = 700 nm) while adjusting the Stokes wavelength to achieve vibrational resonance \((\omega_{{{\rm{p}}}}-{\omega }_{{{\rm{s}}}}=\,{\omega }_{{{\rm{v}}}})\). When the pump matched λmax (black line, Fig. 1C), an inverted Raman peak was observed (black spectrum, Fig. 1D), consistent with prior findings for SiR845 fluorophore25. At λpump = 716 nm, within Rho800’s resonance region, the signal increased over 60-fold compared to the pre-resonant condition at (pump, Stokes) = (838.4 nm, 1031.2 nm) (Supplementary Fig. 2B), as previously demonstrated with conventional OPO20. However, significant background at λpump = 716 nm led to a low S/B ratio. Deviating λpump from λmax by >50 nm significantly improved the S/B ratio (Fig. 1E). For highly fluorescent probe like Rho800, ER-SRS failed to enhance the S/B ratio due to strong background from fluorescence and pump-probe processes20, illustrating the limitations of ER-SRS with conventional fluorophores as resonant Raman probes.
ER-SRS of Raman-amplified nonfluorescent molecular probe (RANMP)
We developed Raman-amplified nonfluorescent molecular probes (RANMPs) with (1) high far-red to NIR absorption for electronic resonance to the light source, (2) Raman-active groups conjugated to the chromophore for large Raman cross-section and electronic resonance, and (3) low fluorescence quantum yields to reduce background (Fig. 2A). The series–RANMP, RANMP-4F, RANMP-4Cl, and RANMP-T26–is based on an electron-rich core, bis(alkylsulfanyl)methylene-substituted 4,9-dihydro-s-indaceno[1,2-b:5,6-b’]dithiophene (IDT), and the electron-withdrawing terminal group 2-(3-oxo-2,3-dihydro-1H-inden-1-ylidene)malononitrile (INCN) containing Raman-active CN moieties. These molecules were synthesized following a previously reported procedure26 (Supplementary Fig. 3). The final compounds were obtained by coupling a formylated IDT core with INCN derivatives via a Knoevenagel condensation reaction using pyridine as a base, in approximately 80% yields for all compounds. The effective conjugation was adjusted by controlling intramolecular charge transfer (ICT) interactions in each molecule. This was achieved by incorporating halogen substituents into INCN group (RANMP-4F and RANMP-4Cl) or introducing a thiophene spacer between the core and INCN terminal moieties (RANMP-T), thereby finely modulating the absorption max. The ICT effect allows λmax to fall within the pump tuning range of 660–960 nm, facilitating resonance conditions (Supplementary Fig. 4A). In RANMP-T, the addition of a thiophene linker between the donor (D) and acceptor (A) units increases the effective conjugation length, resulting in a Raman peak shift of the CN groups (Supplementary Fig. 4B).
A Molecular structures of the RANMP series in which four nitrile groups conjugated to the highly conjugated structures with near-infrared absorption and fluorescence quantum yield below the detection limit of our instrumentation: RANMP, RANMP-4F, RANMP-4Cl, and RANMP-T. B Sample geometry of chromophore film for SRS detection in transillumination geometry. C Time trace of ER-SRS signal of RANMP-T. Blue curve is Gaussian fitting to the trace when the Stokes pulse precedes the pump. Red line is exponential fitting to the trace when pump pulse precedes Stokes. D Pump power dependence of ER-SRS signal. E ER-SRS spectra of the four RANMPs at the optimal combination of pump and Stokes wavelengths. Lines indicate spline interpolation. Error bars represent the mean ± standard deviation (n = 5), with individual replicates shown as purple dots. F Summary of ER-SRS spectra at the optimal condition in (E) in terms of the absorption max (λmax), wavelength difference of the pump wavelength from the absorption max (λmax − λpump), S/B ratio and signal enhancement in comparison to the EPR-SRS signal of Rho800 with fixed Stokes wavelength at 1031.2 nm as in previous studies. Intensities are reported in arbitrary units (a.u.).
From quantum chemical calculations, we found that the donor unit (IDT core) and the acceptor unit (INCN terminal groups) of the RANMP series are responsible mainly for the HOMO and LUMO, respectively (Supplementary Fig. 5). The natural transition orbitals (HONTO and LUNTO) represent the ICT character from the donor to the acceptor unit upon S0 → S1 transition (Supplementary Fig. 6A). These A-D-A type molecules have large Raman cross-sections due to the four CN groups in both terminals conjugated to the IDT core and INCN terminal group. The calculated Raman intensity of the CN vibrational mode in tetracyanoethylene (TCNE) is significantly lower—by a factor of 10 to 30—compared to those of the RANMP series, indicating that the influence of the conjugation on the Raman cross-section (Supplementary Fig. 7C and Supplementary Table 1). In addition, the Raman peak of the CN vibrational mode in the RANMP series depends on the molecular structures. The calculated Raman intensities of the CN vibrational modes increase in the order of RANMP, RANMP-4F, RANMP-4Cl, and RANMP-T (Supplementary Fig. 7C), which is consistent with the experimental results (Supplementary Fig. 4). The calculated Raman frequency is red-shifted in the order of RANMP-4F, RANMP-4Cl, RANMP, and RANMP-T, which correlates with the CN bond length of the RANMP series (Supplementary Fig. 7D). The RANMP series exhibits photoluminescence undetectable with our spectrophotometer, significantly improving the Raman S/B ratio. To understand the low photoluminescence of the RANMP series, the intersystem crossing rate constants (kISC) and radiative decay rate constants (kr) of the RANMP series and Rho800 were calculated (Supplementary Note 1)27. As shown in Table SN2 of Supplementary Note 1, the kISC / kr ratio of the RANMP series (ranging from 102 to 103) is significantly higher than that of Rho800 (7.3). Therefore, compared with Rho800, the RANMP series exhibit much faster ISC to the triplet state and more efficient quenching of the fluorescence of the S1 state, which explains the low photoluminescence. Note that fluorescence quenching by rapid ISC is a key strategy to substantially reduce fluorescence background in ER-SRS. As shown in Eq. (S1) in Supplementary Note 1, the ISC rate constant increases with a large spin-orbit coupling (SOC) constant and a small energy difference between singlet and triplet states (ΔEST). The SOC can be increased by introducing heavy atoms (e.g., S, Se, Br) into organic compounds, a concept widely employed in designing photosensitizers for photodynamic therapy28,29. For the RANMP series, the SOC constants are found to be large due to sulfur atoms (heavy atom effects) (Supplementary Table SN2 in Supplementary Note 1).
To optimize ER-SRS for the RANMP series, we measured the modulation transfer signal of RANMP-T film at (pump, Stokes) = (749 nm, 897.9 nm) by scanning the time delay between pump and Stokes pulses (Fig. 2B). When Stokes preceded pump (blue line, Fig. 2C), the intensity fits to a Gaussian peak centered at time zero, consistent with SRS convolved with a picosecond pulse. When pump precedes Stokes (red line, Fig. 2C), the signal decayed exponentially with a 4.26 ps lifetime, which is on the order of that of stimulated emission. This asymmetrical time delay dependence indicates pump-dominated background. Also, the signal exhibited nonlinear pump power dependence as in the previous literature25 (Fig. 2D). The Stokes power dependence of ER-SRS signal was largely linear below a saturation level (Supplementary Fig. 8).
Using a reduced pump power (500 µW) and a Stokes power below saturation (10 mW), we obtained ER-SRS spectra of RANMP, RANMP-4F, RANMP-4Cl and RANMP-T films (Fig. 2E, Supplementary Figs. 8-9). The double OPO system enabled flexible pump and Stokes tuning to optimize the S/B ratio. Under rigorous resonance, the overwhelming background signal resulted in inverted Raman peaks, whereas positive peaks appeared slightly off resonance (Supplementary Fig. 9). At the optimized conditions, the S/B values ranged from 0.1 to 1.2 (Fig. 2E), comparable to or exceeding those reported for SREF and BonFIRE. Also, ER-SRS signals showed 200-400x amplification compared to EPR-SRS of Rho800 [(pump, Stokes) = (838.4 nm, 1031.2 nm)] (Fig. 2F). ER-SRS spectra of RANMP and RANMP-T solutions in ortho-dichlorobenzene (o-DCB) under optimized conditions yielded S/B ratios of 0.1 and 0.8, respectively. These spectra showed approximately 577-fold and 1152-fold signal amplification compared to EPR-SRS of Rho800 for RANMP and RANMP-T solutions, respectively (Supplementary Figs. 9E–H). These results align with the theoretical predictions (10-100x amplification), given the four CN groups per RANMP molecule versus one CN in Rho80025.
Single-particle imaging of RANMP Pdots
Previously, we demonstrated that chromophores aggregated into polymer dot (Pdot) forms exhibit high resonance Raman scattering with suppressed fluorescence due to aggregation-induced quenching30. Likewise, we expected ER-SRS signals in RANMP-based Pdots to benefit from probe concentration and further fluorescence suppression. RANMP-4Cl and RANMP-T, showing the highest ER-SRS amplification (Fig. 2F), were fabricated into Pdots (Fig. 3A, Supplementary Fig. 10). Using independent wavelength tuning, we optimized the ER-SRS S/B ratios, achieving maxima at λmax + 37 nm for RANMP-T and λmax + 20 nm for RANMP-4Cl (Fig. 3B, C, Supplementary Fig. 11C). Under the optimized conditions, the S/B ratios were 1.2 and 0.9 for RANMP-4Cl and RANMP-T Pdots, respectively. The sharp signal enhancement observed at λmax + 77 nm (λpump = 784.4 nm) is attributed to the proximity of the pump wavelength to the second absorption maximum of the RANMP-T Pdot (778.2 nm), enhancing electronic resonance effect.
A Schematic of Pdot structure. B ER-SRS spectra of RANMP-T under various wavelength combinations of pump and Stokes for optimizing the S/B ratio. Powers are set as 1 and 8 mW for pump and Stokes beam, respectively. Pure SRG is calculated as peak height–background. Lines indicate spline interpolation. C Signal and background intensities in the ER-SRS spectra in (B) plotted against the pump wavelengths. Error bars in (B and C) represent the mean ± standard deviation (n = 3), with individual replicates shown as red open circles. D Two-color single-particle images of RANMP-T and RANMP-4Cl. Far left: brightfield image shows four particles with no way to identify the probe. Middle left: ER-SRS image under optimal condition for RANMP-T. Middle right: ER-SRS image under the optimal condition for RANMP-4Cl. Far right: Off-resonance condition in which pump-Stokes frequency difference is in between the vibrational frequencies of the probe. Powers are set as 1.2 and 16.8 mW for pump and Stokes beams, respectively. E ER-SRS spectra of RANMP-T and RANMP-4Cl Pdots. Cross-sectional profiles of the ER-SRS images of RANMP-T (F) and RANMP-4Cl (G) along the dotted lines in (D) and their fitting to Gaussian (lines). Intensities are reported in arbitrary units (a.u.).
Under the optimum ER-SRS conditions, we achieved dual-color single-particle imaging using the narrow linewidth of Raman scattering (Fig. 3D-G). In a mixture of RANMP-T and RANMP-4Cl Pdots, indistinguishable particles in the brightfield image became spectrally distinct at 2218.4 cm−1 (RANMP-T) and 2235.4 cm−1 (RANMP-4Cl) (Fig. 3D). At the optimal condition for RANMP-T [(pump, Stokes) = (744.4 nm, 891.6 nm)], RANMP-4Cl particles were nearly invisible, and vice versa for RANMP-4Cl [(pump, Stokes) = (760.2 nm, 915.8 nm)]. Off-resonance condition [(pump, Stokes) = (760.7 nm, 915.8 nm)] rendered all particles invisible. The Full Width at Half Maximum (FWHM) of the cross-sectional profile of the particle image was 361 nm for RANMP-4Cl and 356 nm for RANMP-T, closely matching the diffraction-limited resolution of 378 nm and particle size (Fig. 3F, G, Supplementary Fig. 10B). Each 5 μm x 5 μm field of view (FOV) was acquired in ~62 s, enabling two-color imaging under 3 min without noticeable bleaching. The imaging speed, limited by the sample piezo stage, could be improved with a Galvano scanner for beam scanning. The 17 cm-1 Raman shift difference between particles (Fig. 3E) allowed clear spectral distinction for successful two-color single-particle imaging.
Single-molecule imaging of individual molecules embedded in film
To detect single molecule at room temperature, we optimized ER-SRS for poly(methyl methacrylate) (PMMA) samples sparsely embedded with RANMP molecules. Surpassing the EPR-SRS detection limit of 250 nM20, we detected signals at 1 nM for RANMP-T (Figs. 4A and 4B), equivalent to ~6 molecules in the laser focal volume. To minimize bleaching, we employed a polyvinyl alcohol (PVA)31 layer above and below the PMMA (Fig. 4C and Supplementary Fig. 12A) and applied a minimal pump power of 300 μW at optimized wavelengths (Supplementary Fig. 12B). Under these conditions, we performed single-molecule imaging of RANMP-T (1 nM) in a PMMA matrix and successfully obtained single-molecule images and single-step photobleaching traces (Fig. 4D, E and Supplementary Fig. 12E). Orange and blue arrows in Fig. 4D correspond to photobleaching of M1 and M2 molecules, where abrupt photobleaching events in the time trace confirm their single-molecule nature. We then compared traces from a sample with no PVA layer (Supplementary Fig. 12E-G) to those from a PVA-PMMA-PVA configuration (Fig. 4C). Both samples exhibited clear single-step bleaching events and show diverse range of bleaching times (Fig. 4E and Supplementary Fig. 12E). For quantitative analysis, we constructed histograms of photobleaching durations and amplitude (Fig. 4F, G and Supplementary Fig. 12F, G). The distributions of photobleaching time and signal amplitude were well-fitted to single exponential32 and log-normal functions33, respectively, consistent with single-molecule behavior. The extracted time constants (τ) were 42.2 ms for the sample without PVA (n = 63) and 410.3 ms for the PVA-PMMA-PVA configuration (n = 32), indicating a nearly 10-fold increase in photobleaching lifetime—consistent to previous single-molecule studies that used PVA to suppress photobleaching34.
A ER-SRS spectrum of 1 nM RANMP-T embedded in PMMA film. The red line represents a fit using a Voigt profile. B SR gain intensity in various concentrations of RANMP-T. Error bars represent the mean ± standard deviation (n = 3), with individual replicates shown in purple open circles. C Sample geometry of RANMP-T film for ER-SRS single-molecule detection. D Single-molecule images (Top) of two molecules (M1 and M2) embedded in PMMA sandwiched by PVA films for photobleaching prevention and their corresponding time traces (bottom). Single-step bleaching events (arrows) indicate single-molecule nature of the imaged objects. E Five representative time traces with single-step photobleaching events. F Histogram of bleaching times obtained from 32 traces, including the 5 examples shown in (E). The decay constant (τ) is extracted by fitting the histogram with a single-exponential function (red line). G Histogram of the average amplitude in the 32 traces. The red line represents a fit to log-normal distribution. H Single-molecule time delay dependence. Measurements were performed in the sequence of 0 ps, +1 ps, and 0 ps. The recovery of the molecular signal upon returning to 0 ps verifies that the vanishment of signal at +1 ps is not due to photobleaching. I Single-molecule frequency difference dependence. Measurements were performed in the sequence of 2218.2 cm−1, 2236.3 cm−1 and 2218.2 cm−1. The pump and Stokes powers were set to Ppump = 300 μW and PStokes = 4 mW for (D, H and I), and to Ppump = 500 μW and PStokes = 3.3 mW for Figure E. Scale bars: 500 nm. Exposure times: 1 ms. Intensities are reported in arbitrary units (a.u.).
Single-molecule behaviors were also indicated in multi-step photobleaching event (Supplementary Fig. 12C) obtained at a concentration of 10 nM, 10 times more concentrated than our single step photobleaching results in Fig. 4. The multistep trace is consistent with multiple molecules within a focal spot where discrete emitters bleach in steps—demonstrating the sensitivity of our method to resolve individual bleaching events. Also, supplementary Fig. 12D presents a half-moon photobleaching pattern in an image constructed by raster scanning, which serves as a signature of single-molecule detection.
ER-SRS specificity was verified by collecting single-molecule images at two temporal overlaps of pump and Stokes pulses (Fig. 4H). A molecular signal was observed when two pulses arrived concurrently (left 0 ps, Fig. 4H). When the Stokes pulse preceded the pump only by 1 ps ( + 1 ps, Fig. 4H), the signal sharply attenuated, consistent with the bulk time-dependent measurements in Fig. 2C. Reverting the delay to 0 ps restored the signal (right 0 ps, Fig. 4H), confirming that the signal reduction was due to the short vibrational lifetime of nitrile groups ( < 1 ps)35.
To confirm vibrational resonance, we analyzed signal variations with pump-Stokes frequency differences (Fig. 4I). Signals appeared at the CN stretch mode (2218.2 cm⁻¹) but vanished when the pump wavelength tuned by 1 nm to a frequency difference of 2236.3 cm⁻¹. Restoring resonance at 2,218.2 cm⁻¹ recovered the signal, confirming the loss was due to off-resonance, not photobleaching. The abrupt photobleaching as well as sensitivity to temporal overlap and frequency difference validate the near-diffraction-limited features in Fig. 4D, H and I as single-molecule ER-SRS images.
Discussion
In this study, we successfully demonstrated single-molecule ER-SRS by optimizing both light sources and probes. The previous ER-SRS demonstration25 faced challenges in parameter optimization due to conventional SRS light source providing only a single tunable pump beam and a fixed Stokes beam (usually at 1031.2 nm). In contrast, using our independently tunable double OPO laser system, we optimized the wavelength combination for two-beam ER-SRS to maximize the S/B ratio for Raman-active molecules with CN-moieties absorbing NIR light. To further enhance the signal and suppress the background, we designed a series of four Raman-active nonfluorescent molecules (RANMP, RANMP-4F, RANMP-4Cl and RANMP-T) and pre-analyzed their optical properties using DFT and time-dependent (TD) DFT calculations, thereby facilitating effective ER-SRS signal enhancement. As a result, we experimentally realized over 100-fold signal amplification, achieving detection of single molecules via Raman scattering without fluorescence detection or near-field enhancement.
The fluorescence-free nature of ER-SRS enables the extension of single-molecule vibrational microscopy to tissue and animal imaging. While fluorescence enables SREF and BonFIRE to achieve single-molecule sensitivity, it introduces significant background noise: SREF signals are accompanied by anti-Stokes fluorescence from thermally excited populations, and BonFIRE is affected by both anti-Stokes fluorescence and photothermal-induced fluorescence change. These background signals necessitate confocal detection for imaging single cell layers in SREF and BonFIRE, and additional strategies to suppress out-of-focus signal and background from thick samples for deep-tissue imaging. In contrast, the nonlinear nature of ER-SRS inherently provides optical sectioning, offering significant advantages for in vivo imaging, as extensively demonstrated in SRS imaging36,37. ER-SRS also facilitates the development of probes for single-molecule vibrational microspectroscopy. In SREF, achieving an optimal S/B requires stringent electronic pre-resonance conditions, limiting the selection of fluorophores. While BonFIRE provides greater flexibility in dye selection, it is constrained by the tuning range of OPO and relies on NIR fluorophores, which often exhibit low photoluminescence quantum yields (PLQYs)—for instance, Rho800 has a PLQY of 17%. Shifting absorption and emission into the NIR wavelengths typically employ strong donor-acceptor conjugation to induce intramolecular charge transfer (ICT). However, this approach often leads to strong ICT quenching, making it challenging to develop highly fluorescent NIR dyes. By contrast, designing nonfluorescent NIR absorbing molecules is more feasible. The A-D-A molecular design motif used in this study effectively extends conjugation to meet the resonance condition for ER-SRS, where strong ICT interaction suppresses PL emission, optimizing the S/B ratio38,39,40.
Photobleaching, a common challenge in fluorescence microscopy, also occurs in ER-SRS due to the use of electronic resonance. To mitigate this, single-molecule images in Fig. 4 were obtained using a protective PVA layer above and below the probe-containing PMMA layer. When no PVA layer was applied, ER-SRS signals from single molecules were still detectable but photobleaching occurred more rapidly (Supplementary Fig. 12E-G). While fast transfer to the triplet state of RANMP series can promote photobleaching, the photobleaching lifetime of ER-SRS signals from a RANMP-T film was significantly longer than that of EPR-SRS signal from a Rho800 film (Supplementary Fig. 13). Our attempts to measure the triplet-state lifetime of RANMP-T film were unsuccessful, as no phosphorescence signal was detectable, suggesting a short triplet-state lifetime. This implies that the triplet state of RANMP, likely in the NIR region, is highly susceptible to thermal quenching. Efficient triplet-state quenching may therefore contribute to the slower photobleaching observed in the ER-SRS signal of RANMP. To further suppress triplet-mediated radical formation leading to photobleaching, strategies used in fluorescence imaging—such as adding reducing agents, antioxidants and triplet state quenchers—could be adapted for ER-SRS41.
ER-SRS with RANMP achieved single-molecule detection from conventional film and solution samples using standard optical microscopes, highlighting a strong potential for advanced biological imaging using existing labeling methods and instruments. The RANMP in this study are organic small molecules, a widely used class of biological probes. Many strategies used in fluorescence sensors can be applied to RANMP. For instance, the incorporation of organelle-targeting moieties—such as cationic groups for mitochondria or basic groups for lysosomes—into the RANMP framework holds strong potential to convert these hydrophobic probes into lipophilic, organelle-specific markers42. Additionally, introducing hydrophilic functional groups, such as phosphates, could broaden the applicability of RANMP probes by improving their solubility. While the planar core of RANMP facilitates π–π stacking and potential aggregation, the bulky alkyl chains (R1 and R2 in Fig. 2A) can be strategically modified to further suppress undesired stacking. (Supplementary Fig. 6B). For ER-SRS multiplexed imaging, the primary requirement is to synthesize more probes with nonfluorescent, NIR-absorbing conjugated molecules with Raman-active moieties such as nitrile, alkynes and azido groups. Raman shifts of these probes can be tuned through various strategies, including incorporating isotopes into vibrating groups43, extending polyene structures44 and exploiting various core structures of conjugated molecules45. In this study, the thiophene spacer inserted between the core and INCN terminal moieties induced a blue shift in the CN vibrational frequency sufficient for two-color imaging (Fig. 3). This demonstrates an approach for synthesizing multiplex Raman probes through molecular design of conjugated structures. The development of multiplexed RANMP probes with single-molecule sensitivity expands the applicability of spectral identification of molecular targets in spatial omics46,47 and super-resolution microscopy48,49. The PSMA-encapsulated Pdot in this study also offers a convenient, general route for boosting the S/B of ER-SRS probes, for biocompatibility and live-cell applications and for efficient targeting in biological samples by using the carboxylic groups on the particle surface30. The S/B ratios of solution, film and single-molecule samples could be further enhanced by adopting stimulated Raman loss (SRL) detection, which effectively suppresses the pump-induced background that dominates in the current SRG scheme (red line, Fig. 2C). We anticipate that ER-SRS, combined with RANMP, will serve as a powerful tool for multiplexed imaging of individual molecules or nanoparticles within complex biological environments.
Methods
Independently tunable double OPO system
The ER-SRS microscopy setup is described in Fig. 1B and Supplementary Fig. 1B. Two OPOs are pumped by a common, powerful IR fiber laser with a frequency doubling unit (Angewandte Physik & Elektronik GmbH, Emerald Engine HP). The Emerald Engine generates an output pulse train at 1031.2 nm with a 2 ps pulse width and an 80 MHz repetition rate. The frequency-doubled beam at 516 nm is used to synchronously seed two OPOs, referred to as the high power (HP) OPO and the low power (LP) OPO. The tunable pump beam (660–960 nm, ~2 ps) from the HP OPO is modulated at 10 MHz by an electro-optic modulator (EOM) to achieve shot-noise limited detection sensitivity. The tunable Stokes beam (660-960 nm, ~2 ps) is pumped from the LP OPO.
ER-SRS microscopy
As in Fig. 1B and Supplementary Fig. 1B, the independently tuned pump and Stokes beams are combined at the dichroic mirror and then directed into a home-built sample-scanning microscope body (Cerna system, Thorlabs). To confirm the spatial overlap of the two beams, two position sensors were each installed on the near and far sides of the dichroic mirror. The temporal overlap of pump and Stokes pulses is precisely defined using an optical delay line, installed in the pump path, and verified with the sum-frequency generation signal of a BBO crystal. The sample is placed on a piezo-stage (PINano XY Piezo System, P-545.3C8S; PI) for X and Y axis scanning. All beams are focused on the sample using a 60X objective lens with NA 1.2 (UplanSApo; Olympus). A condenser lens with NA 1.4 (oil immersion, D-CUO; Nikon) is used to collect the transmitted beams. The output beams pass through two band-pass filters and a dichroic mirror, which transmits the Stokes beam but blocks the pump, measured as the stimulated Raman gain (SRG) signal. Different band-pass filters and dichroic mirrors are used for each wavelength combination (Supplementary Table 2). The transmitted Stokes beam is collected through a large area photodiode and amplified by a lock-in amplifier (APE GmbH). The signal is sent to a computer in which a home-built LabVIEW program records the ER-SRS intensity at each pixel. We measured SR gain due to the peak sensitivity of the Si photodiode around 960 nm. Many spectra in this study were obtained by averaging multiple measured spectra (five times in most cases). The spectra in Fig. 1D, Fig. 2E, Fig. 3B, Fig. 3C, Fig. 4B, Figure S9E and Figure S9F are presented with the mean ± standard deviation to indicate measurement variance. We did not use any spectral fitting to our spectra except for Fig. 4A, but used spline interpolation without any additional data processing. Figure 4A was the only spectrum that we fitted with a Voigt function50. For single-molecule imaging, we used a 1 ms exposure time per pixel to balance signal strength and imaging speed. This allows for rapid acquisition while minimizing sample drift and photodamage, making it suitable for sensitive or time-limited samples. For photobleaching analysis (Fig. 4E-G and Supplementary Fig. 12E-G), only traces lasting longer than 30 ms were included.
Polymer dots (Pdots) fabrication
The nanoprecipitation method was employed to form Pdots (Supplementary Fig. 10). 100 μL of RANMP-4Cl (or RANMP-T) solution in THF (1 mg/mL) was mixed with 400 μL of poly(styrene-co-maleic anhydride) (PSMA, Mn ≈ 1600 Da, ~1.3:1 styrene anhydride mol ratio) solution in THF (1 mg/mL). Subsequently, 4.5 mL of THF was added. The resulting THF solution was quickly transferred into 10 mL of deionized water under sonication. The water-THF mixture was then heated to 80 °C to facilitate the evaporation of THF and achieve nanoparticle formation until the volume was reduced to 2 mL under N2 bubbling. The concentrated Pdot solution was filtered through a 0.2 μM syringe filter and stored in a brown-colored vial.
To verify if the Pdot nanoparticles are produced at a concentration similar to the theoretical concentration of the Raman probe, a certain amount of RANMP-T Pdot nanoparticle solution was taken and quantified through an absorption coefficient calibration curve (Supplementary Fig. 14). Due to losses occurring during bubbling when evaporating THF, the experimental concentration has an error of approximately ~10% compared to the theoretical concentration.
Imaging sample preparation
Single-particle sample
A 6% w/w agarose gel was prepared by dissolving the appropriate amount of agarose powder (A4018, Sigma) in a 25 mM MES buffer. The mixture was heated to 95 °C to ensure complete dissolution. Subsequently, a glass slide was heated on a hot plate to 95 °C. Then, 10 μL of the Pdot solution and 10 μL of the agarose gel were sequentially added on top of the slide. A cover glass was carefully placed over the mixture and gently pressed to achieve full coverage.
Two-color Pdot sample
The same procedure was performed to prepare the single particle imaging sample. Before adding the agarose gel, 5 μL of the Pdot solution of RANMP-4Cl and 5 μL of the Pdot solution of RANMP-T were sequentially added.
Single-molecule sample
A glass substrate was washed in acetone, isopropyl alcohol and deionized water (DI) water for 15 minutes then UV-ozone plasma treatment was performed for 10 min. A 5% w/w poly(vinyl alcohol) (PVA)/ DI water solution was spin-coated onto a glass substrate at 1000 rpm for 120 s. Using a 1% w/w poly(methyl methacrylate) (PMMA)/CF solution, 1 nM of RANMP-T solution was prepared to form a single-molecule embedded polymer thin film layer. This solution was then spin-coated onto the PVA layer at 1500 rpm for 120 seconds. Afterward, the 5% w/w PVA layer was spin-coated onto the PMMA/RANMP-T layer at 1000 rpm for 120 s.
Material characterization
The 1H spectra were measured by a Bruker Advance III system operating at 500 MHz. UV-Vis-NIR spectra were measured by a JASCO V-770 spectrophotometer. The hydrodynamic diameters of Pdots were measured in aqueous solution using Zetasizer Nano S90 (Malvern Panalytical). Spontaneous Raman spectra were obtained by HORIBA LabRAM HR Evolution Raman spectrometer (Japan). The laser beam, 532 nm, was focused on the samples at the microscope stage by a 100x/0.90 NA objective lens (Plan N, Olympus, Japan). The laser conditions were kept consistent with 1% power, a 5 s acquisition time and 5 accumulations.
Synthesis
RANMP, RANMP-4F, RANM -4Cl, and RANMP-T were synthesized following a previously reported procedure26 (Supplementary Figs. 3, 15-18). RANMP, RANMP-4F, and RANMP-4Cl were synthesized via Vilsmeier–Haack formylation of a bis(alkylsulfanyl)methylene-substituted IDT core, followed by Knoevenagel condensation with (halogenated) INCN. In contrast, RANMP-T was prepared through stannylation of the bis(alkylsulfanyl)methylene-substituted IDT core, subsequent Stille coupling with an octyl-substituted thiophene spacer, and a final Knoevenagel condensation with INCN.
RANMP 1H NMR (500 MHz, CDCl3): δ 8.92 (s, 2H), 8.89 (s, 2H), 8.64 (s, 2H), 8.59–8.54 (m, 2H), 7.89–7.82 (m, 2H), 7.72–7.65 (m, 4H), 3.14 (m, 8H), 1.80–1.68 (m, 4H), 1.56–1.13 (m, 64H), 0.96–0.71 (m, 24H). RANMP-4F 1H NMR (500 MHz, CDCl3): δ 9.10–9.05 (m, 2H), 8.92 (s, 2H), 8.72 (s, 2H), 8.57–8.41 (m, 2H) 7.64 (t, J = 7.4 Hz, 2H), 3.21–3.09 (m, 8H), 1.79–1.70 (m, 4H), 1.53–1.11 (m, 64H), 0.93–0.72 (m, 24H). RANMP-4Cl 1H NMR (500 MHz, CDCl3): δ 9.11 (s, 2H), 8.92 (s, 2H), 8.72 (s, 2H), 8.53–8.43 (m, 2H), 7.64 (t, J = 7.4 Hz, 2H), 3.15 (m, J = 6.7 Hz, 8H), 1.81–1.66 (m, 4H), 1.52–1.15 (m, 64H), 0.91–0.72 (m, 24H). RANMP-T 1H NMR (500 MHz, CDCl3): δ 8.93 (s, 2H), 8.80 (s, 2H), 8.69 (d, J = 7.0 Hz, 2H), 8.47 (s, 2H), 7.94–7.90 (m, 2H), 7.78–7.68 (m, 6H), 3.15–3.09 (m, 8H), 2.93 (t, 3H), 1.85–1.68 (m, 4H), 1.52–1.14 (m, 88H), 0.91–0.73 (m, 30H).
Quantum chemical calculations
Density functional theory (DFT) and time-dependent (TD) DFT methods (B3LYP/6-31 G(d,p)) were used to obtain the optimized molecular structures in S0, frontier molecular orbitals (HOMO and LUMO), and natural transition orbitals (HONTO and LUNTO) of the molecules in chloroform (Supplementary Figs. 5-6). DFT calculations were performed on simplified structures in which the bulky alkyl chains (R1 and R2 in Fig. 2A) were shortened to methyl groups, retaining only the planar conjugated core. As a result, the calculations may overestimate molecular planarity and packing propensity. The full RANMP molecules contain bulky alkyl chains that can sterically hinder π–π stacking (Supplementary Fig. 6B). The optimized structures in S1 of molecules were obtained by using the TD-DFT method. Additionally, Raman spectra of molecules were calculated and analyzed to understand the dependence of Raman peak (CN vibrational mode) on the molecular structures (Supplementary Fig. 7). The intersystem crossing rate constants (kISC) and radiative decay rate constants (kr) of molecules were calculated as described in Supplementary Note 1. The vertical electronic transition energies (singlet and triplet states) and spin-orbit coupling (SOC) constants were obtained using the Tamm-Dancoff approximation (TDA)-DFT method (B3LYP/6-31 G(d,p)) implemented in ORCA.
Data availability
The data supporting the findings of this study consist primarily of spectroscopy datasets. Due to the absence of a standardized public repository that supports these data formats in a reusable manner, the data are not deposited in a public database. The data that support the findings of this study are available from the corresponding author upon request.
Code availability
The codes that support the plots and data analysis within this paper are available from the corresponding author upon request.
References
Ha, T. et al. Probing the interaction between two single molecules: fluorescence resonance energy transfer between a single donor and a single acceptor. Proc. Natl. Acad. Sci. USA 93, 6264–6268 (1996).
Yildiz, A. et al. Myosin V walks hand-over-hand: single fluorophore imaging with 1.5-nm localization. Science 300, 2061–2065 (2003).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–796 (2006).
Manley, S. et al. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat. Methods 5, 155–157 (2008).
Bintu, B. et al. Super-resolution chromatin tracing reveals domains and cooperative interactions in single cells. Science 362, eaau1783 (2018).
Metzker, M. L. Sequencing technologies—the next generation. Nat. Rev. Genet. 11, 31–46 (2010).
Sigal, Y. M. et al. Visualizing and discovering cellular structures with super-resolution microscopy. Science 361, 880–887 (2018).
Balzarotti, F. et al. Nanometer resolution imaging and tracking of fluorescent molecules with minimal photon fluxes. Science 355, 606–612 (2017).
Weisenburger, S. et al. Cryogenic optical localization provides 3D protein structure data with Angstrom resolution. Nat. Methods 14, 141–144 (2017).
Reinhardt, S. C. M. et al. Ångström-resolution fluorescence microscopy. Nature 617, 711–716 (2023).
Chen, K. H. et al. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Moffitt, J. R., Lundberg, E. & Heyn, H. The emerging landscape of spatial profiling technologies. Nat. Rev. Genet. 23, 741–759 (2022).
Lakowicz, J. R. Principles of Fluorescence Spectroscopy, 3rd edn (Springer, 2011).
Nie, S. & Emory, S. R. Probing single molecules and single nanoparticles by surface-enhanced Raman scattering. Science 275, 1102–1106 (1997).
Langer, J. et al. Present and future of surface-enhanced Raman scattering. ACS Nano 14, 28–117 (2020).
Laing, S., Gracie, K. & Faulds, K. Multiplex in vitro detection using SERS. Chem. Soc. Rev. 45, 1901–1918 (2016).
Jimenez de Aberasturi, D. et al. Surface enhanced Raman scattering encoded gold nanostars for multiplexed cell discrimination. Chem. Mater. 28, 6779–6790 (2016).
Kang, H. et al. Near-Infrared SERS nanoprobes with plasmonic Au/Ag hollow-shell assemblies for in vivo multiplex detection. Adv. Funct. Mater. 23, 3719–3727 (2013).
Wei, L. et al. Super-multiplex vibrational imaging. Nature 544, 465–470 (2017).
Xiong, H. et al. Stimulated Raman excited fluorescence spectroscopy and imaging. Nat. Photonics 13, 412–417 (2019).
Xiong, H. et al. Stimulated Raman excited fluorescence spectroscopy of visible dyes. J. Phys. Chem. Lett. 10, 3563–3570 (2019).
Xiong, H. & Min, W. Combining the best of two worlds: stimulated Raman excited fluorescence. J. Chem. Phys. 153, 210901 (2020).
Wang, H. et al. Bond-selective fluorescence imaging with single-molecule sensitivity. Nat. Photonics 17, 846–855 (2023).
Shi, L. et al. Electronic resonant stimulated Raman scattering micro-spectroscopy. J. Phys. Chem. B 122, 9218–9224 (2018).
Cho, S. J. et al. A-D-A type semiconducting small molecules with bis(alkylsulfanyl)methylene substituents and control of charge polarity for organic field-effect transistors. ACS Appl. Mater. Interfaces 12, 41842–41851 (2020).
Cho, E. et al. Impact of secondary donor units on the excited-state properties and thermally activated delayed fluorescence (TADF) efficiency of pentacarbazole-benzonitrile emitters. J. Chem. Phys. 153, 144708 (2020).
Nguyen, V.-N. et al. An emerging molecular design approach to heavy-atom-free photosensitizers for enhanced photodynamic therapy under hypoxia. J. Am. Chem. Soc. 141, 16243–16248 (2019).
Nguyen, V.-N. et al. Design and synthesis of efficient heavy-atom-free photosensitizers for photodynamic therapy of cancer. Chem. Commun. 56, 11489–11492 (2020).
Jeong, J.-E. et al. Resonant Raman-active polymer dot barcodes for multiplex cell mapping. ACS Nano 17, 4800–4812 (2023).
Kwon, J., Elgawish, M. S. & Shim, S.-H. Bleaching-resistant super-resolution fluorescence microscopy. Adv. Sci. 9, 2101817 (2022).
Wennmalm, S. & Rigler, R. On death numbers and survival times of single dye molecules. J. Phys. Chem. B 103, 2516–2519 (1999).
Mutch, S. A. et al. Deconvolving single-molecule intensity distributions for quantitative microscopy measurements. Biophys. J. 92, 2926–2943 (2007).
Gensch, T. et al. Single molecule blinking and photobleaching separated by wide-field fluorescence microscopy. J. Phys. Chem. A 109, 6652–6658 (2005).
Kocheril, P. A. et al. Nitrile vibrational lifetimes as probes of local electric fields. J. Phys. Chem. Lett. 15, 5306–5314 (2024).
Lu, F.-K. et al. Label-free DNA imaging in vivo with stimulated Raman scattering microscopy. Proc. Natl. Acad. Sci. USA 112, 11624–11629 (2015).
Fu, D. et al. In vivo metabolic fingerprinting of neutral lipids with hyperspectral stimulated Raman scattering microscopy. J. Am. Chem. Soc. 136, 8820–8828 (2014).
Santra, D. C. et al. Charge-transfer-induced fluorescence quenching of anthracene derivatives and selective detection of picric acid. Chem. Eur. J. 22, 2012–2019 (2016).
Bu, L. et al. An AIE and ICT based NIR florescent probe for cysteine and homocysteine. Dyes Pigm 136, 724–731 (2017).
Bednarik, S. et al. Tuning electron-accepting properties of phthalocyanines for charge transfer processes. Inorg. Chem. 63, 8799–8806 (2024).
Cordes, T. et al. Mechanisms and advancement of antifading agents for fluorescence microscopy and single-molecule spectroscopy. Phys. Chem. Chem. Phys. 13, 6699–6709 (2011).
Yang, Y., Bai, X. & Hu, F. Photoswitchable polyynes for multiplexed stimulated Raman scattering microscopy with reversible light control. Nat. Commun. 15, 2578 (2024).
Chen, Z. et al. Multicolor live-cell chemical imaging by isotopically edited alkyne vibrational palette. J. Am. Chem. Soc. 136, 8027–8033 (2014).
Hu, F. et al. Supermultiplexed optical imaging and barcoding with engineered polyynes. Nat. Methods 15, 194–200 (2018).
Miao, Y. et al. 9-Cyanopyronin probe palette for super-multiplexed vibrational imaging. Nat. Commun. 12, 4518 (2021).
Marx, V. Method of the Year: spatially resolved transcriptomics. Nat. Methods 18, 9–14 (2021).
Method of the Year 2024: spatial proteomics. Nat. Methods 21, 2195–2196 (2024).
Schueder, F. et al. Unraveling cellular complexity with transient adapters in highly multiplexed super-resolution imaging. Cell 187, 1769–1784 (2024).
Unterauer, E. M. et al. Spatial proteomics in neurons at single-protein resolution. Cell 187, 1785–1800 (2024).
Shi, L., Hu, F. & Min, W. Optical mapping of biological water in single live cells by stimulated Raman excited fluorescence microscopy. Nat. Commun. 10, 4764 (2019).
Acknowledgements
This work was supported by Samsung Science and Technology Foundation SSTF-BA2201-07 and a Korea University Grant for S.-H.S. This work was also supported by grants from the National Research Foundation of Korea (RS-2022-NR069080 for S.P., RS-2024-00334832 for H.Y.W. and RS-2024-00357947 for S.-H.S.)
Author information
Authors and Affiliations
Contributions
H.Y.W. and S.-H.S. conceived and designed the research. S.O. built the ER-SRS set-up and performed all the SRS spectroscopy and microscopy experiments. Y.E. and A.T. prepared polymer dots and film samples. S.O. and Y.E. collected and analyzed the data. H.Y.W. and S.-H.S. supervised the experiments. S.P. supervised theoretical and quantum chemical calculations and analyzed the results. H.Y.K. performed theoretical and quantum chemical calculations. The manuscript was written by S.O. and S.-H.S. with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Lu Wei and the other anonymous reviewers for their contribution to the peer review of this work. [A peer review file is available.]
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Oh, S., Eom, Y., Kim, H.Y. et al. Fluorescence-free single-molecule microscopy via electronic resonance stimulated Raman scattering. Nat Commun 17, 2720 (2026). https://doi.org/10.1038/s41467-026-69348-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69348-6