Abstract
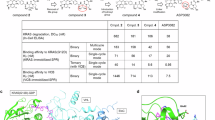
KRAS, a frequently mutated oncogene, has been challenging to target therapeutically. Although covalent inhibitors like sotorasib against KRASG12C have been developed, their efficacy is often limited by acquired resistance. Targeted protein degradation offers a potential solution but has largely relied on large PROTAC molecules. Here, we report DJX-A-KM, a small-molecule degrader of KRASG12C, designed by incorporating an acrylamide warhead into the MRTX849 scaffold. It induces potent and sustained degradation of KRASG12C in cells and in vivo. Mechanistic investigation reveal that degradation is mediated by the ubiquitin-proteasome system, facilitated by covalent engagement with a E3 ligase, FBXO28, at cysteine 98. Antiproliferation assays demonstrate its potent inhibitory effects across multiple KRASG12C-mutant cancer models. This strategy also enables the development of pan-KRAS degraders against a broader spectrum of KRAS mutations. Our work presents a small-molecule degrader recruiting FBXO28 and provides a blueprint for exploring E3 ligases in protein degradation.
Similar content being viewed by others
Data availability
The Proteomics raw data have been deposited in a ProteomeXchange partner repository under PXD067337 and PXD072213. Source Data including uncropped and unprocessed scans of all blots and gels, reported means/averages in box plots, bar charts, and tables, including all quantified replicates are provided with this paper as a Source Data file.
References
Reiner, D. J. & Lundquist, E. A. Small GTPases. WormBook 2018, 1–65 (2018).
Land, H., Parada, L. F. & Weinberg, R. A. Tumorigenic conversion of primary embryo fibroblasts requires at least two cooperating oncogenes. Nature 304, 596–602 (1983).
Hobbs, G. A., Der, C. J. & Rossman, K. L. RAS isoforms and mutations in cancer at a glance. J. Cell Sci. 129, 1287–1292 (2016).
Hunter, J. C. et al. Biochemical and structural analysis of common cancer-associated KRAS mutations. Mol. Cancer Res. 13, 1325–1335 (2015).
Kim, H. J. et al. Oncogenic KRAS: signaling and drug resistance. Cancers 13, 5599 (2021).
Li, S., Balmain, A. & Counter, C. M. A model for RAS mutation patterns in cancers: finding the sweet spot. Nat. Rev. Cancer 18, 767–777 (2018).
Milburn, M. V. et al. Molecular switch for signal transduction: structural differences between active and inactive forms of protooncogenic ras proteins. Science 247, 939–945 (1990).
Ostrem, J. M. et al. K-Ras(G12C) inhibitors allosterically control GTP affinity and effector interactions. Nature 503, 548–551 (2013).
Simanshu, D. K., Nissley, D. V. & McCormick, F. RAS proteins and their regulators in human disease. Cell 170, 17–33 (2017).
Timar, J. & Kashofer, K. Molecular epidemiology and diagnostics of KRAS mutations in human cancer. Cancer Metastasis Rev. 39, 1029–1038 (2020).
Canon, J. et al. The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity. Nature 575, 217–223 (2019).
Skoulidis, F. et al. Sotorasib for lung cancers with KRAS p.G12C mutation. N. Engl. J. Med. 384, 2371–2381 (2021).
Hallin, J. et al. The KRASG12C inhibitor MRTX849 provides insight toward therapeutic susceptibility of kras-mutant cancers in mouse models and patients. Cancer Discov. 10, 54–71 (2020).
Jänne, P. A. et al. Adagrasib in non-small-cell lung cancer harboring a KRASG12C mutation. N. Engl. J. Med. 387, 120–131 (2022).
Shi, Y. et al. Glecirasib in KRASG12C-mutated nonsmall-cell lung cancer: a phase 2b trial. Nat. Med. 31, 894–900 (2025).
Liu, J., Kang, R. & Tang, D. The KRAS-G12C inhibitor: activity and resistance. Cancer Gene Ther. 29, 875–878 (2022).
Tsai, Y. S. et al. Rapid idiosyncratic mechanisms of clinical resistance to KRASG12C inhibition. J. Clin. Invest. 132, e155523 (2022).
Prenen, H. et al. SO-39 evaluation of acquired resistance to sotorasib in KRAS p.G12C-mutated colorectal cancer: exploratory plasma biomarker analysis of CodeBreaK 100. Ann. Oncol. 33, S373 (2022).
Koga, T. et al. KRAS secondary mutations that confer acquired resistance to KRASG12C inhibitors, sotorasib and adagrasib, and overcoming strategies: insights from in vitro experiments. J. Thorac. Oncol. 16, 1321–1332 (2021).
Akhave, N. S., Biter, A. B. & Hong, D. S. Mechanisms of resistance to KRASG12C-targeted therapy. Cancer Discov. 11, 1345–1352 (2021).
Foley, K. P. et al. Chemically induced chaperone-mediated protein degradation (CHAMP) of KRAS(G12C). Cancer Res. 83, 1649 (2023).
Gong, R. H. et al. Combination of artesunate and WNT974 induces KRAS protein degradation by upregulating E3 ligase ANACP2 and β-TrCP in the ubiquitin-proteasome pathway. Cell Commun. Signal. 20, 34 (2022).
Popow, J. et al. Targeting cancer with small-molecule pan-KRAS degraders. Science 385, 1338–1347 (2024).
Sakamoto, K. M. et al. Protacs: chimeric molecules that target proteins to the Skp1-Cullin-F box complex for ubiquitination and degradation. Proc. Natl. Acad. Sci. USA 98, 8554–8559 (2001).
Li, K. & Crews, C. M. PROTACs: past, present and future. Chem. Soc. Rev. 51, 5214–5236 (2022).
Henning, N. J. et al. Discovery of a covalent FEM1B recruiter for targeted protein degradation applications. J. Am. Chem. Soc. 144, 701–708 (2022).
Samarasinghe, K. T. G. & Crews, C. M. Targeted protein degradation: a promise for undruggable proteins. Cell Chem. Biol. 28, 934–951 (2021).
Oleinikovas, V. et al. From thalidomide to rational molecular glue design for targeted protein degradation. Annu. Rev. Pharmacol. Toxicol. 64, 291–312 (2024).
Rodriguez-Gonzalez, A. et al. Targeting steroid hormone receptors for ubiquitination and degradation in breast and prostate cancer. Oncogene 27, 7201–7211 (2008).
Zhang, D. et al. Targeted degradation of proteins by small molecules: a novel tool for functional proteomics. Comb. Chem. High. Throughput Screen. 7, 689–697 (2004).
Shi, Y. et al. Targeting KRAS: from metabolic regulation to cancer treatment. Mol. Cancer 24, 9 (2025).
Zeng, M. et al. Exploring targeted degradation strategy for oncogenic KRASG12C. Cell Chem. Biol. 27, 19–31 (2020).
Yang, F. et al. Efficient targeted oncogenic KRASG12C degradation via first reversible-covalent PROTAC. Eur. J. Med. Chem. 230, 114088 (2022).
Yang, N. et al. Discovery of highly potent and selective KRASG12C degraders by VHL-recruiting PROTACs for the treatment of tumors with KRASG12C-mutation. Eur. J. Med. Chem. 261, 115857 (2023).
Yokoo, H., Naito, M. & Demizu, Y. Investigating the cell permeability of proteolysis-targeting chimeras (PROTACs). Expert Opin. Drug Discov. 18, 357–361 (2023).
Kagiou, C. et al. Alkylamine-tethered molecules recruit FBXO22 for targeted protein degradation. Nat. Commun. 15, 5409 (2024).
Basu, A. A. et al. A CRISPR activation screen identifies FBXO22 supporting targeted protein degradation. Nat. Chem. Biol. 20, 1608–1616 (2024).
Lim, M. et al. DCAF16-based covalent handle for the rational design of monovalent degraders. ACS Cent. Sci. 10, 1318–1331 (2024).
Pei, J. et al. Piperlongumine conjugates induce targeted protein degradation. Cell Chem. Biol. 30, 203–213 (2023).
Shergalis, A. G. et al. CRISPR screen reveals BRD2/4 molecular glue-like degrader via recruitment of DCAF16. ACS Chem. Biol. 18, 331–339 (2023).
Zhang, X. et al. DCAF11 supports targeted protein degradation by electrophilic proteolysis-targeting chimeras. J. Am. Chem. Soc. 143, 5141–5149 (2021).
Tao, Y. et al. Targeted protein degradation by electrophilic PROTACs that stereoselectively and site-specifically engage DCAF1. J. Am. Chem. Soc. 144, 18688–18699 (2022).
Luo, M. et al. Chemoproteomics-enabled discovery of covalent RNF114-based degraders that mimic natural product function. Cell Chem. Biol. 28, 559–566 (2021).
Li, Y. D. et al. Template-assisted covalent modification underlies activity of covalent molecular glues. Nat. Chem. Biol. 20, 1640–1649 (2024).
Spradlin, J. N. et al. Harnessing the anti-cancer natural product nimbolide for targeted protein degradation. Nat. Chem. Biol. 15, 747–755 (2019).
Zhang, X., Crowley, V. M., Wucherpfennig, T. G., Dix, M. M. & Cravatt, B. F. Electrophilic PROTACs that degrade nuclear proteins by engaging DCAF16. Nat. Chem. Biol. 15, 737–746 (2019).
Nie, D. Y. et al. Recruitment of FBXO22 for targeted degradation of NSD2. Nat. Chem. Biol. 20, 1597–1607 (2024).
He, X. et al. Covalent recruitment of NEDD4 for targeted protein degradation: rational design of small molecular degraders. J. Am. Chem. Soc. 147, 21512–21525 (2025).
Ouyang, X. et al. Proteome-wide ligand and target discovery by using β-nitrostyrene electrophiles: supporting targeted protein degradation. Angew. Chem. Int. Ed. 64, e202504813 (2025).
Shah, R. R. et al. Structure-guided design and optimization of covalent VHL-targeted sulfonyl fluoride PROTACs. J. Med. Chem. 67, 4641–4654 (2024).
Qin, Z. et al. Discovering covalent inhibitors of protein-protein interactions from trillions of sulfur(VI) fluoride exchange-modified oligonucleotides. Nat. Chem. 15, 1705–1714 (2023).
Zhao, Q. et al. Broad-spectrum kinase profiling in live cells with lysine-targeted sulfonyl fluoride probes. J. Am. Chem. Soc. 139, 680–685 (2017).
Yu, Z. et al. Simultaneous covalent modification of K-Ras(G12D) and K-Ras(G12C) with tunable oxirane electrophiles. J. Am. Chem. Soc. 145, 20403–20411 (2023).
Liang, L., Zhang, Z., You, Q. & Guo, X. Recent advances in the design of small molecular drugs with acrylamides covalent warheads. Bioorg. Med. Chem. 112, 117902 (2024).
Scattolin, T., Gharbaoui, T. & Chen, C. Y. A nucleophilic deprotection of carbamate mediated by 2-mercaptoethanol. Org. Lett. 24, 3736–3740 (2022).
Snead, D. R. et al. Development of Adagrasib’s commercial manufacturing route. Org. Process Res. Dev. 27, 530–538 (2023).
RayzeBio Inc. et al. Conjugates Comprising Covalent Binders for Targeting Intracellular KRASG12C Proteins, WIPO, WO2023014979 (2023).
Shindo, N. & Ojida, A. Recent progress in covalent warheads for in vivo targeting of endogenous proteins. Bioorg. Med. Chem. 47, 116386 (2021).
Koo, T. Y. et al. N-Acryloylindole-alkyne (NAIA) enables imaging and profiling new ligandable cysteines and oxidized thiols by chemoproteomics. Nat. Commun. 14, 3564 (2023).
Fell, J. B. et al. Identification of the clinical development candidate MRTX849, a covalent KRASG12C inhibitor for the treatment of cancer. J. Med. Chem. 63, 6679–6693 (2020).
Troup, R. I. et al. Current strategies for the design of PROTAC linkers: a critical review. Explor. Target Antitumor Ther. 1, 273–312 (2020).
Bond, M. J. et al. Targeted degradation of oncogenic KRASG12C by VHL-recruiting PROTACs. ACS Cent. Sci. 6, 1367–1375 (2020).
Cepeda, D. et al. CDK-mediated activation of the SCF(FBXO28) ubiquitin ligase promotes MYC-driven transcription and tumourigenesis and predicts poor survival in breast cancer. EMBO Mol. Med. 5, 1067–1086 (2013).
Phillips, E. et al. PFKFB4 interacts with FBXO28 to promote HIF-1α signaling in glioblastoma. Oncogenesis 11, 57 (2022).
Qiao, X. et al. FBXO28 suppresses liver cancer invasion and metastasis by promoting PKA-dependent SNAI2 degradation. Oncogene 42, 2878–2891 (2023).
Wang, X., Lawson, J. D. & Marx, M. A. Azaquinazoline pan-KRAS Inhibitors, WIPO, WO2022132200 (2022).
Yang, J. et al. A pan-KRAS degrader for the treatment of KRAS-mutant cancers. Cell Discov. 10, 70 (2024).
Acknowledgements
This work was supported by funds from Natural Science Foundation of China (22377033, 22377037, 22577038, 82473109), the Science and Technology Program of Guangdong Province (2023B1515120023, 2025A0505080018, 2024A1515013002, 2023A0505050149), Guangdong Basic and Applied Basic Research Foundation (2023B1515040016, 2025A1515012357, 2024A1515013266, 2026B1515020070). Science and Technology Project in Guangzhou (202102070001). Huizhou-Hong Kong-Macao Cooperation Development Fund (2025EQ010043). The authors sincerely thank to everyone who made scientific contributions to this work.
Author information
Authors and Affiliations
Contributions
Conception: J.D., S.S., L.H., F.X., W.H., C.H., Z.Z., T.L., Y.T., and Z.L. Experimental design and execution: J.D., S.S., L.H., F.X., W.H., C.H., Z.Z., T.L., Y.T., and Z.L. Data analysis, interpretation, and generation of figures: J.D., S.S., L.H., F.X., C.H., Z.Z., T.L., Y.T., and Z.L. Authored, edited and substantively revised the manuscript: J.D., S.S., L.H., F.X., T.L., Y.T., and Z.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Deng, J., Shen, S., Huang, L. et al. Small-molecule degraders for oncogenic KRASG12C and pan-KRAS mutations. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71093-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71093-9