Abstract
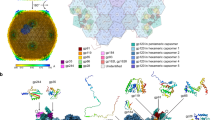
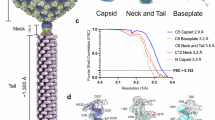
Jumbo phage phiKZ, a key model for studying phage nucleus formation and bacterial defense mechanisms, possesses a highly complex tail machine that is essential for infection. Here, we present the structural atlas of the intact jumbo phage phiKZ by cryo-EM, thereby identifying 40 constituent proteins and unveiling its modular architecture. The virion, with a length of approximately 360 nm, is comprised of an icosahedral capsid of 2520 polypeptide chains from 11 proteins, and a massive tail machine of over 900 polypeptide chains from 29 proteins. The tail features a unique, multi-layered neck and a highly elaborate baseplate. The neck is reinforced by whisker-like proteins and anchors the contractile tail, which terminates in the baseplate. The baseplate is constituted by a central hub, an inner periphery of interlocking wedge heterotrimers and hexagonal rings, and an outer periphery with a striking hexagonal star configuration. This intricate peripheral region of the baseplate serves as an extended platform for twelve peripheral fibers, which mediate host cell adsorption. Our findings provide a structural framework for understanding jumbo phage assembly and infection, thus contributing to the foundation for future functional studies and rational engineering of these phages for potential therapeutic applications.
Similar content being viewed by others
Data availability
The cryo-EM maps have been deposited at the Electron Microscopy Data Bank under accession codes EMD-67125 (overall tail), EMD-67123 (unit1), EMD-67124 (unit2), EMD-67095 (portal-adaptor), EMD-67102 (neck), EMD-67096 (sheath-tube), EMD-67121 (baseplate C6) and EMD-67098 (baseplate C3), EMD-67097 (docker), and EMD-67111 (outer peripheral). The atomic models have been deposited at the Protein Data Bank under accession codes 9XPE (portal-adaptor), 9XPS (neck), 9XQS (baseplate C6), 9XPF (sheath-tube), 9XPH (baseplate C3), 9XPG (docker), and 9XQD (outer peripheral). All other data are available in the main text or the supplementary materials.
References
Hendrix, R. W. Jumbo bacteriophages. Curr. Top. Microbiol. Immunol. 328, 229–240 (2009).
Harding, K. R., Kyte, N. & Fineran, P. C. Jumbo phages. Curr. Biol. 33, R750–r751 (2023).
Yuan, Y. & Gao, M. Jumbo Bacteriophages: an overview. Front. Microbiol. 8, 403 (2017).
Chaikeeratisak, V. et al. Assembly of a nucleus-like structure during viral replication in bacteria. Science 355, 194–197 (2017).
Chaikeeratisak, V. et al. The phage nucleus and tubulin spindle are conserved among large pseudomonas phages. Cell Rep. 20, 1563–1571 (2017).
Mendoza, S. D. et al. A bacteriophage nucleus-like compartment shields DNA from CRISPR nucleases. Nature 577, 244–248 (2020).
Kokontis, C., Klein, T. A., Silas, S. & Bondy-Denomy, J. Multi-interface licensing of protein import into a phage nucleus. Nature 639, 456–462 (2025).
Malone, L. M. et al. A jumbo phage that forms a nucleus-like structure evades CRISPR-Cas DNA targeting but is vulnerable to type III RNA-based immunity. Nat. Microbiol. 5, 48–55 (2020).
Guan, J. & Bondy-Denomy, J. Intracellular organization by jumbo bacteriophages. J. Bacteriol. 203, e00362–20 (2020).
Guan, J. et al. Bacteriophage genome engineering with CRISPR-Cas13a. Nat. Microbiol. 7, 1956–1966 (2022).
González, B. et al. Phage G structure at 6.1Å resolution, condensed DNA, and host identity revision to a Lysinibacillus. J. Mol. Biol. 432, 4139–4153 (2020).
Effantin, G. et al. Cryo-electron microscopy three-dimensional structure of the jumbo phage ΦRSL1 infecting the phytopathogen Ralstonia solanacearum. Structure 21, 298–305 (2013).
Neumann, E. et al. 3D structure of three jumbo phage heads. J. Gen. Virol. 101, 1219–1226 (2020).
Hua, J. et al. Capsids and genomes of jumbo-sized bacteriophages reveal the evolutionary reach of the HK97 fold. mBio 8, e01579-17 (2017).
Ouyang, R. et al. High-resolution reconstruction of a Jumbo-bacteriophage infecting capsulated bacteria using hyperbranched tail fibers. Nat. Commun. 13, 7241 (2022).
Yang, Y. et al. Capsid structure of bacteriophage ΦKZ provides insights into assembly and stabilization of jumbo phages. Nat. Commun. 15, 6551 (2024).
Hu, B., Margolin, W., Molineux, I. J. & Liu, J. Structural remodeling of bacteriophage T4 and host membranes during infection initiation. Proc. Natl. Acad. Sci. USA 112, E4919–E4928 (2015).
Krylov, V. et al. Phage phiKZ-the first of the giants. Viruses 13, 149 (2021).
Mesyanzhinov, V. V. et al. The genome of bacteriophage phiKZ of Pseudomonas aeruginosa. J. Mol. Biol. 317, 1–19 (2002).
Yuping, L. et al. Jumbo phage killer immune system targets early infection of nucleus-forming phages. Cell 188, 2127–2140.e2121 (2025).
Fokine, A. et al. A three-dimensional cryo-electron microscopy structure of the bacteriophage phiKZ head. J. Mol. Biol. 352, 117–124 (2005).
Lecoutere, E. et al. Identification and comparative analysis of the structural proteomes of phiKZ and EL, two giant Pseudomonas aeruginosa bacteriophages. Proteomics 9, 3215–3219 (2009).
Thomas, J. A. et al. Extensive proteolysis of head and inner body proteins by a morphogenetic protease in the giant Pseudomonas aeruginosa phage φKZ. Mol. Microbiol. 84, 324–339 (2012).
Fokine, A. et al. Cryo-EM study of the Pseudomonas bacteriophage phiKZ. Structure 15, 1099–1104 (2007).
Li, X. et al. Near-atomic resolution structure determination of a cypovirus capsid and polymerase complex using cryo-EM at 200kV. J. Mol. Biol. 429, 79–87 (2017).
Chen, W. et al. Structural changes of a bacteriophage upon DNA packaging and maturation. Protein Cell 11, 374–379 (2020).
Chen, W., Zheng, J., Zhou, J., Cheng, L. & Liu, H. A data-processing strategy of asymmetric reconstruction for tailed phages by cryo-electron microscopy. J. Struct. Biol. 217, 108262 (2025).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Jamali, K. et al. Automated model building and protein identification in cryo-EM maps. Nature 628, 450–457 (2024).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
He, J., Lin, P., Chen, J., Cao, H. & Huang, S. Y. Model building of protein complexes from intermediate-resolution cryo-EM maps with deep learning-guided automatic assembly. Nat. Commun. 13, 4066 (2022).
Fokine, A., Miroshnikov, K. A., Shneider, M. M., Mesyanzhinov, V. V. & Rossmann, M. G. Structure of the bacteriophage phi KZ lytic transglycosylase gp144. J. Biol. Chem. 283, 7242–7250 (2008).
Subramanian, S., Kerns, H. R., Braverman, S. G. & Doore, S. M. The structure of Shigella virus Sf14 reveals the presence of two decoration proteins and two long tail fibers. Commun. Biol. 8, 222 (2025).
Sun, L. et al. Cryo-EM structure of the bacteriophage T4 portal protein assembly at near-atomic resolution. Nat. Commun. 6, 7548 (2015).
Taylor, N. M. et al. Structure of the T4 baseplate and its function in triggering sheath contraction. Nature 533, 346–352 (2016).
Jiang, F. et al. Cryo-EM structure and assembly of an extracellular contractile injection system. Cell 177, 370–383.e315 (2019).
Sycheva, L. V. et al. Crystal structure and location of gp131 in the bacteriophage phiKZ virion. Virology 434, 257–264 (2012).
Leiman, P. G., Chipman, P. R., Kostyuchenko, V. A., Mesyanzhinov, V. V. & Rossmann, M. G. Three-dimensional rearrangement of proteins in the tail of bacteriophage T4 on infection of its host. Cell 118, 419–429 (2004).
Antonova, D. et al. Genomic transfer via membrane vesicle: a strategy of giant phage phiKZ for early infection. J. Virol. 98, e0020524 (2024).
Mozumdar, D. et al. Characterization of a lipid-based jumbo phage compartment as a hub for early phage infection. Cell Host Microbe 32, 1050–1058.e1057 (2024).
Mozumdar, D., A Agard, D. & Bondy-Denomy, J. The complex developmental mechanisms of nucleus-forming jumbo phages. Curr. Opin. Microbiol. 88, 102676 (2025).
Li, F. et al. High-resolution cryo-EM structure of the Pseudomonas bacteriophage E217. Nat. Commun. 14, 4052 (2023).
Kanamaru, S. et al. Structure of the cell-puncturing device of bacteriophage T4. Nature 415, 553–557 (2002).
Wu, W., Thomas, J. A., Cheng, N., Black, L. W. & Steven, A. C. Bubblegrams reveal the inner body of bacteriophage φKZ. Science 335, 182 (2012).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Kivioja, T., Ravantti, J., Verkhovsky, A., Ukkonen, E. & Bamford, D. Local average intensity-based method for identifying spherical particles in electron micrographs. J. Struct. Biol. 131, 126–134 (2000).
Thuman-Commike, P. A. & Chiu, W. Improved common line-based icosahedral particle image orientation estimation algorithms. Ultramicroscopy 68, 231–255 (1997).
Fuller, S. D., Butcher, S. J., Cheng, R. H. & Baker, T. S. Three-dimensional reconstruction of icosahedral particles-the uncommon line. J. Struct. Biol. 116, 48–55 (1996).
Pettersen, E. F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Goddard, T. D. et al. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci. A Publ. Protein Soc. 27, 14–25 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D. Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D. Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
This research was supported by the National Natural Science Foundation of China (32430020 and 12034006 to H.L., 32401014 to H.X., 32200994 to W.C., 32371263 to L.C.), the National Science and Technology Major Project of China (2023ZD0500501 to H.L.), Major Fundamental Research Program of Hunan Province (2025ZYJ004 to H.L.), Natural Science Foundation of Hunan Province, China (2024JJ6304 to H.X.), and the Science and Technology Innovation Program of Hunan Province (2024RC3150 to W.C.). We thank the Cryo-EM center of the Shanxi Academy of Advanced Research and Innovation (SAARI) for providing facilities and technical support.
Author information
Authors and Affiliations
Contributions
H.L. and H.X. conceived, initiated, and coordinated the project; H.X. produced the phiKZ sample and prepared cryo-EM grids; H.X. and Y.P. collected the cryo-EM data; H.X., Z.P., J.Z., Y.C., Y.T., W.C., and H.L. processed the cryo-EM data; H.X., Z.P., J.Z., T.L., and S.H. built and refined the structure model; H.X., Z.P., and H.L. wrote the manuscript. All authors discussed the experiments and results and read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Sarah Doore, who co-reviewed with Hailey Kerns, and the other, anonymous, reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiao, H., Peng, Z., Zhou, J. et al. Structural atlas of the intact jumbo phage phiKZ. Nat Commun (2026). https://doi.org/10.1038/s41467-026-71561-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-71561-2