Abstract
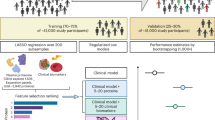
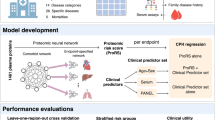
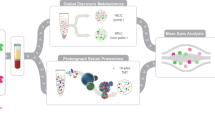
Multi-omics technologies, such as metabolomics and proteomics, offer deep molecular perspectives that could enhance risk prediction, but large-scale studies integrating both are scarce. Here we show the predictive values of these two omics across 17 incident diseases in 23,776 UK Biobank participants with complete baseline for 159 NMR-based metabolites and 2,923 Olink affinity-based proteins. We found that adding omics data significantly improved risk prediction for all 17 diseases compared to clinical predictors alone. Proteomics-only models generally outperformed metabolomics-only models for 16 of the 17 diseases, and integrating both omics added little prediction power over proteomics-only models. Furthermore, we identified key omics features, including both well-established (e.g., KLK3/PSA for prostate cancer) and potential novel ones (e.g., PRG3 for skin cancer). We further connected diseases with medication and socioeconomic factors through key proteins, highlighting the clinical utility of omics data for enhancing individual risk prediction, providing molecular insights into disease mechanisms, and potentially guiding future therapeutic development.
Similar content being viewed by others
Acknowledgements
This research has been conducted using the UKB Resource under Application Number 25953. We thank the UKB participants and research team for enabling this study. Y.L. discloses support for the research of this work from Funder R01AR083790, Funder U01HG011720, and Funder R01HL146500. All the other authors declare no relevant funding.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Du, J., Zhou, M., Wang, H. et al. Multi-omics integration predicts the incidence of 17 diseases in the UK Biobank. Nat Commun (2026). https://doi.org/10.1038/s41467-026-73017-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-73017-z