Abstract
Arabidopsis PHOSPHATE 1 (AtPHO1) and its closest homologue AtPHO1;H1 are phosphate transporters that load phosphate into the xylem vessel for root-to-shoot translocation. AtPHO1 and AtPHO1;H1 are prototypical members of the unique SPX–EXS family, whose structural and molecular mechanisms remain elusive. In this study, we determined the cryogenic electron microscopy structure of AtPHO1;H1 binding with inorganic phosphate (Pi) and inositol hexakisphosphate in a closed conformation. Further molecular dynamic simulation and AlphaFold prediction support an open conformation. AtPHO1;H1 forms a domain-swapped homodimer that involves both the transmembrane ERD1/XPR1/SYG1 (EXS) domain and the cytoplasmic SYG1/Pho81/XPR1 (SPX) domain. The EXS domain presented by the SPX–EXS family represents a novel protein fold, and an independent substrate transport pathway and substrate-binding site are present in each EXS domain. Two gating residues, Trp719 and Tyr610, are identified above the substrate-binding site to control opening and closing of the pathway. The SPX domain features positively charged patches and/or residues at the dimer interface to accommodate inositol hexakisphosphate molecules, whose binding mediates dimerization and enhances AtPHO1;H1 activity. In addition, a C-terminal tail is required for AtPHO1;H1 activity. On the basis of structural and functional analysis, a working model for Pi efflux mediated by AtPHO1;H1 and its homologues was postulated, suggesting a channel-like mechanism. This study not only reveals the molecular and regulatory mechanism underlying Pi transport mediated by the unique SPX–EXS family, but also provides potential for crop engineering to enhance phosphorus-use efficiency in sustainable agriculture.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The 3D cryo-EM density maps of AtPHO1;H1 and AtPHO1;H1EXS have been deposited in the Electron Microscopy Data Bank under the accession numbers EMD-60648 and EMD-61430. Coordinates for structure models have been deposited in the Protein Data Bank under the accession codes PDB 9IK4 and PDB 9JF8. Source data are provided with this paper.
References
Maathuis, F. J. M. Physiological functions of mineral macronutrients. Curr. Opin. Plant Biol. 12, 250–258 (2009).
Poirier, Y., Jaskolowski, A. & Clúa, J. Phosphate acquisition and metabolism in plants. Curr. Biol. 32, R623–R629 (2022).
López-Arredondo, D. L., Leyva-González, M. A., González-Morales, S. I., López-Bucio, J. & Herrera-Estrella, L. Phosphate nutrition: improving low-phosphate tolerance in crops. Annu. Rev. Plant Biol. 65, 95–123 (2014).
Yang, S. Y., Lin, W. Y., Hsiao, Y. M. & Chiou, T. J. Milestones in understanding transport, sensing, and signaling of the plant nutrient phosphorus. Plant Cell 36, 1504–1523 (2024).
Hao, D. L. et al. Roles of plastid-located phosphate transporters in carotenoid accumulation. Front. Plant Sci. 13, 1059536 (2022).
Wang, Y., Wang, F., Lu, H., Liu, Y. & Mao, C. Phosphate uptake and transport in plants: an elaborate regulatory system. Plant Cell Physiol. 62, 564–572 (2021).
Młodzinska, E. & Zboinska, M. Phosphate uptake and allocation — a closer look at Arabidopsis thaliana L. and Oryza sativa L. Front. Plant Sci. 7, 1198 (2016).
Nussaume, L. et al. Phosphate import in plants: focus on the PHT1 transporters. Front. Plant Sci. 2, 83 (2011).
Xu, L. et al. Identification of vacuolar phosphate efflux transporters in land plants. Nat. Plants 5, 84–94 (2019).
Liu, J. et al. A vacuolar phosphate transporter essential for phosphate homeostasis in Arabidopsis. Proc. Natl Acad. Sci. USA 112, E6571–E6578 (2015).
Pao, S. S., Paulsen, I. T. & Saier, M. H. Major facilitator superfamily. Microbiol. Mol. Biol. Rev. 62, 1–34 (1998).
Yamaji, N. et al. Reducing phosphorus accumulation in rice grains with an impaired transporter in the node. Nature 541, 92–95 (2017).
Zhu, W. et al. The mitochondrial phosphate transporters modulate plant responses to salt stress via affecting ATP and gibberellin metabolism in Arabidopsis thaliana. PLoS ONE 7, e43530 (2012).
Reimer, R. J. & Edwards, R. H. Organic anion transport is the primary function of the SLC17/type I phosphate transporter family. Pflugers Arch. 447, 629–635 (2004).
Daram, P. et al. PHT2;1 encodes a low-affinity phosphate transporter from Arabidopsis. Plant Cell 11, 2153–2166 (1999).
Wang, Y., Ribot, C., Rezzonico, E. & Poirier, Y. Structure and expression profile of the Arabidopsis PHO1 gene family indicates a broad role in inorganic phosphate homeostasis. Plant Physiol. 135, 400–411 (2004).
Poirier, Y., Thoma, S., Somerville, C. & Schiefelbein, J. Mutant of Arabidopsis deficient in xylem loading of phosphate. Plant Physiol. 97, 1087–1093 (1991).
Hamburger, D., Rezzonico, E., MacDonald-Comber Petetot, J., Somerville, C. & Poirier, Y. Identification and characterization of the Arabidopsis PHO1 gene involved in phosphate loading to the xylem. Plant Cell 14, 889–902 (2002).
Stefanovic, A. et al. Members of the PHO1 gene family show limited functional redundancy in phosphate transfer to the shoot, and are regulated by phosphate deficiency via distinct pathways. Plant J. 50, 982–994 (2007).
Stefanovic, A. et al. Over-expression of PHO1 in Arabidopsis leaves reveals its role in mediating phosphate efflux. Plant J. 66, 689–699 (2011).
Arpat, A. B. et al. Functional expression of PHO1 to the Golgi and trans-Golgi network and its role in export of inorganic phosphate. Plant J. 71, 479–491 (2012).
Vetal, P. V. & Poirier, Y. The Arabidopsis PHOSPHATE 1 exporter undergoes constitutive internalization via clathrin-mediated endocytosis. Plant J. 116, 1477–1491 (2023).
Vogiatzaki, E., Baroux, C., Jung, J. Y. & Poirier, Y. PHO1 exports phosphate from the chalazal seed coat to the embryo in developing Arabidopsis seeds. Curr. Biol. 27, 2893–2900 e2893 (2017).
Zimmerli, C. et al. PHO1 expression in guard cells mediates the stomatal response to abscisic acid in Arabidopsis. Plant J. 72, 199–211 (2012).
Huang, Y. et al. Abscisic acid modulates seed germination via ABA INSENSITIVE5-mediated PHOSPHATE1. Plant Physiol. 175, 1661–1668 (2017).
Secco, D., Baumann, A. & Poirier, Y. Characterization of the rice PHO1 gene family reveals a key role for OsPHO1;2 in phosphate homeostasis and the evolution of a distinct clade in dicotyledons. Plant Physiol. 152, 1693–1704 (2010).
Che, J. et al. Node-localized transporters of phosphorus essential for seed development in rice. Plant Cell Physiol. 61, 1387–1398 (2020).
Ma, B. et al. A plasma membrane transporter coordinates phosphate reallocation and grain filling in cereals. Nat. Genet. 53, 906–915 (2021).
Nguyen, N. N. T. et al. PHO1 family members transport phosphate from infected nodule cells to bacteroids in Medicago truncatula. Plant Physiol. 185, 196–209 (2021).
Chiou, T. J. & Lin, S. I. Signaling network in sensing phosphate availability in plants. Annu. Rev. Plant Biol. 62, 185–206 (2011).
Wild, R. et al. Control of eukaryotic phosphate homeostasis by inositol polyphosphate sensor domains. Science 352, 986–990 (2016).
Dong, J. et al. Inositol pyrophosphate InsP(8) acts as an intracellular phosphate signal in Arabidopsis. Mol. Plant 12, 1463–1473 (2019).
Zhu, J. et al. Two bifunctional inositol pyrophosphate kinases/phosphatases control plant phosphate homeostasis. eLife 8, e43582 (2019).
Zhou, J. et al. Mechanism of phosphate sensing and signaling revealed by rice SPX1–PHR2 complex structure. Nat. Commun. 12, 7040 (2021).
Guan, Z. et al. Mechanistic insights into the regulation of plant phosphate homeostasis by the rice SPX2–PHR2 complex. Nat. Commun. 13, 1581 (2022).
Wege, S. et al. The EXS domain of PHO1 participates in the response of shoots to phosphate deficiency via a root-to-Shoot Signal. Plant Physiol. 170, 385–400 (2016).
Li, X. et al. Control of XPR1-dependent cellular phosphate efflux by InsP(8) is an exemplar for functionally-exclusive inositol pyrophosphate signaling. Proc. Natl Acad. Sci. USA 117, 3568–3574 (2020).
Xu, C. et al. A phosphate-sensing organelle regulates phosphate and tissue homeostasis. Nature 617, 798–806 (2023).
Hsieh, Y. F. et al. cis-Golgi phosphate transporters harboring an EXS domain are essential for plant growth and development. Plant Physiol. 192, 1000–1015 (2023).
Liu, T. Y. et al. Identification of plant vacuolar transporters mediating phosphate storage. Nat. Commun. 7, 11095 (2016).
Luan, M. et al. A SPX domain vacuolar transporter links phosphate sensing to homeostasis in Arabidopsis. Mol. Plant 15, 1590–1601 (2022).
Pedersen, B. P. et al. Crystal structure of a eukaryotic phosphate transporter. Nature 496, 533–536 (2013).
Lee, Y. et al. Structure of the triose-phosphate/phosphate translocator reveals the basis of substrate specificity. Nat. Plants 3, 825–832 (2017).
Tsai, J. Y. et al. Structure of the sodium-dependent phosphate transporter reveals insights into human solute carrier SLC20. Sci. Adv. 6, eabb4024 (2020).
Schneider, S., Kuhlbrandt, W. & Yildiz, O. Complementary structures of the yeast phosphate transporter Pho90 provide insights into its transport mechanism.Structure 32, 979–988 (2024).
Holm, L. DALI and the persistence of protein shape. Protein Sci. 29, 128–140 (2020).
Ernst, O. P. et al. Microbial and animal rhodopsins: structures, functions, and molecular mechanisms. Chem. Rev. 114, 126–163 (2014).
Kovalev, K. et al. Structure and mechanisms of sodium-pumping KR2 rhodopsin. Sci. Adv. 5, eaav2671 (2019).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Giovannini, D., Touhami, J., Charnet, P., Sitbon, M. & Battini, J. L. Inorganic phosphate export by the retrovirus receptor XPR1 in metazoans. Cell Rep. 3, 1866–1873 (2013).
Legati, A. et al. Mutations in XPR1 cause primary familial brain calcification associated with altered phosphate export. Nat. Genet. 47, 579–581 (2015).
Lopez-Sanchez, U. et al. Characterization of XPR1/SLC53A1 variants located outside of the SPX domain in patients with primary familial brain calcification. Sci. Rep. 9, 6776 (2019).
Yan, R. et al. Human XPR1 structures reveal phosphate export mechanism. Nature 633, 960–967 (2024).
Lu, Y. et al. Structural basis for inositol pyrophosphate gating of the phosphate channel XPR1. Science 386, eadp3252 (2024).
Heuer, S. et al. Improving phosphorus use efficiency: a complex trait with emerging opportunities. Plant J. 90, 868–885 (2017).
Ma, B. et al. Genetic improvement of phosphate-limited photosynthesis for high yield in rice. Proc. Natl Acad. Sci. USA 121, e2404199121 (2024).
Carlson, M. L. et al. The Peptidisc, a simple method for stabilizing membrane proteins in detergent-free solution. eLife 7, e34085 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2, 19–25 (2015).
Hoover, W. G. Canonical dynamics – equilibrium phase–space distributions. Phys. Rev. A 31, 1695–1697 (1985).
Nose, S. A unified formulation of the constant temperature molecular-dynamics methods. J. Chem. Phys. 81, 511–519 (1984).
Parrinello, M. & Rahman, A. Polymorphic transitions in single-crystals – a new molecular-dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald – an N·log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Chatzikyriakidou, Y., Ahn, D. H., Nji, E. & Drew, D. The GFP thermal shift assay for screening ligand and lipid interactions to solute carrier transporters. Nat. Protoc. 16, 5357–5376 (2021).
Pravda, L. et al. MOLEonline: a web-based tool for analyzing channels, tunnels and pores (2018 update). Nucleic Acids Res. 46, W368–W373 (2018).
Acknowledgements
We thank J. Ma from Okayama University and K. Yi from the Chinese Agriculture Academy of Science for critical reading of the manuscript. We thank M. Zhang at the cryo-EM center of the Chinese Academy of Sciences (CAS) Interdisciplinary Research Center on Biology and Chemistry. This work was supported by grants from the National Natural Science Foundation of China (grant no. 32025020) and the CAS (grant nos. XDB0630100 and 317GJHZ2022023GC), grants from Shanghai Science and Technology Commission (grant no. 23310710100).
Author information
Authors and Affiliations
Contributions
S.F., Y.Y. and X.Z. designed and performed the bulk of the experiments. S.F. carried out protein expression and purification, and grid sample preparation. X.Z. and S.F. carried out cryo-EM data collection and structure determination supervised by P.Z. Y.Y. carried out electrophysiological experiments guided by Y.-F.W. M.Z. carried out molecular dynamic analysis. Z.Y., Y.Z., C.Z. and F.Y. contributed to protein purification and grid sample preparation. P.Z., S.F. and Y.-F.W. wrote the manuscript with input from other authors. P.Z. conceived the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Zhenfeng Liu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cryo-EM analysis of AtPHO1;H1.
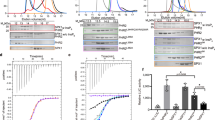
a, Gel filtration profile of a size-exclusion column (Superose 6 Increase 10/300 GL, Cytiva) and Coomassie-blue-stained SDS-PAGE analysis of AtPHO1;H1 in peptidiscs. Independent experiments have been repeated at least three times with similar results. b, Flowchart for data processing of AtPHO1;H1 and AtPHO1;H1EXS. c, Resolution map for AtPHO1;H1. d, Resolution map for AtPHO1;H1EXS. The color represents the local resolution in Å. e, The gold-standard Fourier shell correlation (FSC) curves of AtPHO1;H1 and AtPHO1;H1EXS. f, Structural superimposition of AtPHO1;H1 and AtPHO1;H1EXS. The N-subdomain of AtPHO1;H1 and AtPHO1;H1EXS are colored in blue and purple, respectively. The C-subdomain of AtPHO1;H1 and AtPHO1;H1EXS are colored in green and orange, respectively. The C-tail of AtPHO1;H1 and AtPHO1;H1EXS are colored in red and dark blue, respectively.
Extended Data Fig. 2 Representative EM densities of AtPHO1;H1.
a, EM density of EXS domain (Map contour level = 5.5 σ) and M-loop (Map contour level = 3 σ) in AtPHO1;H1. b, EM density of InsP6 in AtPHO1;H1 is shown at contour level of 3.5 σ. SPX, purple; SPX’, yellow. Residues coordination with InsP6 are indicated as light-blue spheres. c, d, EM densities of substrate-binding site in AtPHO1;H1 (c) and AtPHO1;H1EXS (d), which are shown at the same contour levels of 5.5 σ. The relatively weak Pi density in AtPHO1;H1 might be due to insufficient resolution. e, EM densities of C-tail in AtPHO1;H1 (left) and AtPHO1;H1EXS (right), which are shown at the same contour levels of 5.5 σ.
Extended Data Fig. 3 Structural comparison of the SPX domains in different proteins.
a, SPX of AtPHO1;H1 is shown as ribbons colored from N to C terminus in blue to red. b-e, Structural superimposition of the AtPHO1;H1 SPX domain with other SPX domains from HsXPR1 (PDB:5IJH) (b), CtVTC4 (PDB:5IJP) (c), OsSPX1(PDB: 7E40) (d), and CtGDE1(PDB: 5IJJ) (e). All SPX domains are shown in the same orientations. The dashed box is represented to show the InsP6 binding site in c-e.
Extended Data Fig. 4 The structure and transport mechanism of Pi transporters.
a-d, The structure and transport mechanism of Pi transporters: Serendipita indica SiPT (a), Saccharomyces cerevisiae ScPHO90 (b), Galdieria sulphuraria GsGPT (c) and Thermotoga maritima TmPiT (d). e, Structural superimposition of the EXS domain of AtPHO1;H1 and the sodium-pumping rhodopsin KR2.
Extended Data Fig. 5 Sequence alignment of the C-subdomain of the EXS domain-containing proteins.
At: Arabidopsis thaliana; Os: Oryza sativa; Mt: Medicago truncatula; Zm: Zea mays; Hs, Homo sapiens. Dm: Drosophila melanogaster; Sc: Saccharomyces cerevisiae. The orange and blue dots indicate the residues critical for substrate phosphate binding and transport pathway in AtPHO1;H1, respectively.
Extended Data Fig. 6 Electrophysiological analysis of substrate-binding site residues.
a-i, Typical whole-cell recordings and average current–voltage curves for HEK293T cells expressing the wild type AtPHO1;H1 (WT), the mutants and a GFP control. Error bars mean ± s.e.m. Independent experiments were repeated for each construct with the numbers of the HEK293T cells tested: n = 6 (WT); n = 7 (D560A) (a); n = 7 (K635A) (b); n = 9 (Y636A) (c); n = 6 (Q672A) (d); n = 6 (D676A) (e); n = 7 (R716A) (f); n = 6 (R747A) (g); n = 6 (R748A) (h); n = 8 (I670A as an internal control) (i); n = 6 (GFP control). The same WT and mock control data were used for the comparison to the point mutated versions.
Extended Data Fig. 7 Substrate transport pathway of AtPHO1;H1W719A mutant and electrophysiological analysis of the gating residues.
a, Side view of the substrate transport pathway of AtPHO1;H1W719A mutant. The AtPHO1;H1W719A mutant structure was generated by simple mutation of Trp719 to Ala based on the AtPHO1;H1 structure. The transport pathway is shown as purple solid surface calculated by MOLE, and residues Y603, F607, Y610 and W719 in the pathway are shown as sticks and colored pink. The residues involving Pi binding are indicated by blue spheres. b, Top view of a. c, Typical whole-cell recordings and average current–voltage curves for HEK293T cells expressing the wild type AtPHO1;H1 (WT), the mutants and a GFP control. Error bars mean ± s.e.m. Independent experiments were repeated for each construct with the numbers of the HEK293T cells tested: n = 9 (WT); n = 8 (Y610A); n = 6 (W719A); n = 8 (GFP control). d, Statistic analysis of currents for the wild type AtPHO1;H1 (WT), the mutants and a GFP control measured at +80 mV in HEK293T cells. Error bars mean ± s.e.m. Independent experiments were repeated for each construct with the numbers of HEK293T cells tested: n = 9 (WT); n = 8 (Y610A); n = 6 (W719A); n = 8 (GFP control). Significances were determined using Two-tailed t-tests. ***P < 0.001, ****P < 0.0001 for WT versus mutants. Exact P values for all comparisons are provided in the Source data.
Extended Data Fig. 8 Electrophysiological analysis of InsP6-binding site.
a-e, Typical whole-cell recordings and average current–voltage curves for HEK293T cells expressing the wild type AtPHO1;H1 (WT), the mutants and a GFP control. Error bars mean ± s.e.m. Independent experiments were repeated for each construct with the numbers of HEK293T cells tested: n = 7 (WT); n = 7 (K31A) (a); n = 7 (K287A/290 A) (b); n = 9 (K370A/K373A/Y374A) (c); n = 6 (Y23F/K27A/K320A) (d); n = 9 (ΔSPX) (e); n = 6 (GFP control). The same WT and mock control data are used for a convenience of comparison to the point mutants.
Extended Data Fig. 9 Sequence alignment of the SPX domain of the SPX-EXS family proteins.
At: Arabidopsis thaliana; Os: Oryza sativa; Mt: Medicago truncatula; Zm: Zea mays; Hs, Homo sapiens. Dm: Drosophila melanogaster; Sc: Saccharomyces cerevisiae. The pink dots indicate the residues involved in InsP6 binding in AtPHO1;H1.
Extended Data Fig. 10 Sequence alignment of the C-tail of the SPX-EXS family proteins.
At: Arabidopsis thaliana; Os: Oryza sativa; Mt: Medicago truncatula; Zm: Zea mays; Hs, Homo sapiens. Dm: Drosophila melanogaster; Sc: Saccharomyces cerevisiae.
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1 and Table 1.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed gels.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fang, S., Yang, Y., Zhang, X. et al. Structural mechanism underlying PHO1;H1-mediated phosphate transport in Arabidopsis. Nat. Plants 11, 309–320 (2025). https://doi.org/10.1038/s41477-024-01895-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-024-01895-6