Abstract
Metabolism is essential for plant growth and has become a major target for crop improvement by enhancing nutrient use efficiency. Metabolic engineering is also the basis for producing high-value plant products such as pharmaceuticals, biofuels and industrial biochemicals. An inherent problem for such engineering endeavours is the tendency to view metabolism as a series of distinct metabolic pathways—glycolysis, the tricarboxylic acid cycle, the Calvin–Benson cycle and so on. While these canonical pathways may represent a dominant or frequently occurring flux mode, systematic analyses of metabolism via computational modelling have emphasized the inherent flexibility of the metabolic network to carry flux distributions that are distinct from the canonical pathways. Recent experimental estimates of metabolic network fluxes using 13C-labelling approaches have revealed numerous instances in which non-canonical pathways occur under different conditions and in different tissues. In this Review, we bring these non-canonical pathways to the fore, summarizing the evidence for their occurrence and the context in which they operate. We also emphasize the importance of non-canonical pathways for metabolic engineering. We argue that the introduction of a high-flux pathway to a desired metabolic product will, by necessity, require non-canonical supporting fluxes in central metabolism to provide the necessary carbon skeletons, energy and reducing power. We illustrate this using the overproduction of isoprenoids and fatty acids as case studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
da Fonseca‐Pereira, P., de Cássia Monteiro-Batista, R., Araújo, W. L. & Nunes-Nesi, A. Harnessing enzyme cofactors and plant metabolism: an essential partnership. Plant J. 114, 1014–1036 (2023).
Mira, M. M. et al. Plant stem cells under low oxygen: metabolic rewiring by phytoglobin underlies stem cell functionality. Plant Physiol. 193, 1416–1432 (2023).
Simons, M., Misra, A. & Sriram, G. in Plant Metabolism: Methods and Protocols Vol. 1083 (ed. Sriram, G.) 213–230 (Humana, 2014).
de Oliveira Dal’Molin, C. G. & Nielsen, L. K. Plant genome-scale reconstruction: from single cell to multi-tissue modelling and omics analyses. Curr. Opin. Biotechnol. 49, 42–48 (2018).
Gerlin, L., Frainay, C., Jourdan, F., Baroukh, C. & Prigent, S. Plant genome-scale metabolic networks. Adv. Bot. Res. 98, 237–270 (2021).
Allen, D. K. & Young, J. D. Tracing metabolic flux through time and space with isotope labeling experiments. Curr. Opin. Biotechnol. 64, 92–100 (2020).
Clark, T. J., Guo, L., Morgan, J. & Schwender, J. Modeling plant metabolism: from network reconstruction to mechanistic models. Annu. Rev. Plant Biol. 71, 303–326 (2020).
Kaste, J. A. M. & Shachar-Hill, Y. Accurate flux predictions using tissue-specific gene expression in plant metabolic modeling. Bioinformatics 39, btad186 (2023).
Ma, H. & Zeng, A. P. Reconstruction of metabolic networks from genome data and analysis of their global structure for various organisms. Bioinformatics 19, 270–277 (2003).
Fabregas, N. & Fernie, A. R. The metabolic response to drought. J. Exp. Bot. 70, 1077–1085 (2019).
Botha, F. C. & Marquardt, A. Metabolic control of sugarcane internode elongation and sucrose accumulation. Agronomy 14, 1487 (2024).
de Vries, S. & Feussner, I. Biotic interactions, evolutionary forces and the pan-plant specialized metabolism. Phil. Trans. R. Soc. B 379, 20230362 (2024).
Kruger, N. J. & Ratcliffe, R. G. Pathways and fluxes: exploring the plant metabolic network. J. Exp. Bot. 63, 2243–2246 (2012).
Judge, A. & Dodd, M. S. Metabolism. Essays Biochem. 64, 607–647 (2020).
Dixon, R. A., Chen, F., Guo, D. & Parvathi, K. The biosynthesis of monolignols: a ‘metabolic grid’, or independent pathways to guaiacyl and syringyl units? Phytochemistry 57, 1069–1084 (2001).
Lanier, E. R., Andersen, T. B. & Hamberger, B. Plant terpene specialized metabolism: complex networks or simple linear pathways? Plant J. 114, 1178–1201 (2023).
Bergman, M. E., Kortbeek, R. W., Gutensohn, M. & Dudareva, N. Plant terpenoid biosynthetic network and its multiple layers of regulation. Prog. Lipid Res. 95, 101287 (2024).
Palsson, B. Ø. Systems Biology: Constraint-Based Reconstruction and Analysis Ch. 12 (Cambridge Univ. Press, 2015).
Sauer, U. Metabolic networks in motion: 13C-based flux analysis. Mol. Syst. Biol. 2, 62 (2006).
Morandini, P. Rethinking metabolic control. Plant Sci. 176, 441–451 (2009).
Saa, P. A. & Nielsen, L. K. Formulation, construction and analysis of kinetic models of metabolism: a review of modelling frameworks. Biotechnol. Adv. 35, 981–1003 (2017).
Orth, J. D., Thiele, I. & Palsson, B. Ø. What is flux balance analysis? Nat. Biotechnol. 28, 245–248 (2010).
Zhu, X. G., Wang, Y., Ort, D. R. & Long, S. P. e-Photosynthesis: a comprehensive dynamic mechanistic model of C3 photosynthesis: from light capture to sucrose synthesis. Plant Cell Environ. 36, 1711–1727 (2013).
Feldman-Salit, A., Veith, N., Wirtz, M., Hell, R. & Kummer, U. Distribution of control in the sulfur assimilation in Arabidopsis thaliana depends on environmental conditions. New Phytol. 222, 1392–1404 (2019).
Guo, L. et al. Dynamic modeling of subcellular phenylpropanoid metabolism in Arabidopsis lignifying cells. Metab. Eng. 49, 36–46 (2018).
Wang, J. P. et al. Flux modelling for monolignol biosynthesis. Curr. Opin. Biotechnol. 56C, 187–192 (2019).
Rao, X. & Barros, J. Modeling lignin biosynthesis: a pathway to renewable chemicals. Trends Plant Sci. 29, 546–559 (2024).
Töpfer, N., Braam, T., Shameer, S., Ratcliffe, R. G. & Sweetlove, L. J. Alternative Crassulacean acid metabolism modes provide environment-specific water-saving benefits in a leaf metabolic model. Plant Cell 32, 3689–3705 (2020).
Schuster, S., Pfeiffer, T. & Fell, D. A. Is maximization of molar yield in metabolic networks favoured by evolution? J. Theor. Biol. 252, 497–504 (2008).
Sarkar, D. & Kundu, S. Systems biology of plant metabolic interactions. J. Biosci. 49, 56 (2024).
Mahadevan, R. & Schilling, C. H. The effects of alternate optimal solutions in constraint-based genome-scale metabolic models. Metab. Eng. 5, 264–276 (2003).
Tong, H., Kuken, A., Razaghi-Moghadam, Z. & Nikoloski, Z. Characterisation of effects of genetic variants via genome-scale metabolic modelling. Cell. Mol. Life Sci. 78, 5123–5138 (2021).
Töpfer, N. Environment-coupled models of leaf metabolism. Biochem. Soc. Trans. 49, 119–129 (2021).
Sampaio, M., Rocha, M. & Dias, O. Exploring synergies between plant metabolic modelling and machine learning. Comput. Struct. Biotechnol. J. 20, 1885–1900 (2022).
Cheung, C. Y. M. et al. A method for accounting for maintenance costs in flux balance analysis improves the prediction of plant cell metabolic phenotypes under stress conditions. Plant J. 75, 1050–1061 (2013).
Sweetlove, L. J., Williams, T. C. R., Cheung, C. Y. M. & Ratcliffe, R. G. Modelling metabolic CO2 evolution—a fresh perspective on respiration. Plant Cell Environ. 36, 1631–1640 (2013).
Poolman, M. G., Miguet, L., Sweetlove, L. J. & Fell, D. A. A genome-scale metabolic model of Arabidopsis and some of its properties. Plant Physiol. 151, 1570–1581 (2009).
Poolman, M. G., Kundu, S., Shaw, R. & Fell, D. A. Responses to light intensity in a genome-scale model of rice metabolism. Plant Physiol. 162, 1060–1072 (2013).
Cheung, C. Y. M., Poolman, M. G., Fell, D. A., Ratcliffe, R. G. & Sweetlove, L. J. A diel flux balance model captures interactions between light and dark metabolism during day–night cycles in C3 and Crassulacean acid metabolism leaves. Plant Physiol. 165, 917–929 (2014).
Yuan, H., Cheung, C. Y. M., Poolman, M. G., Hilbers, P. A. J. & van Riel, N. A. W. A genome-scale metabolic network reconstruction of tomato (Solanum lycopersicum L.) and its application to photorespiratory metabolism. Plant J. 85, 289–304 (2016).
Moreira, T. B. et al. A genome-scale metabolic model of soybean (Glycine max) highlights metabolic fluxes in seedlings. Plant Physiol. 180, 1912–1929 (2019).
Bender, M. L., Zhu, X. G., Falkowski, P., Ma, F. & Griffin, K. On the rate of phytoplankton respiration in the light. Plant Physiol. 190, 267–279 (2022).
Tan, X. L. J. & Cheung, C. Y. M. A multiphase flux balance model reveals flexibility of central carbon metabolism in guard cells of C3 plants. Plant J. 104, 1648–1656 (2020).
Sprent, N. et al. Metabolic modeling reveals distinct roles of sugars and carboxylic acids in stomatal opening as well as unexpected carbon fluxes. Plant Cell 37, koae252 (2025).
Hunt, H. et al. Analysis of companion cell and phloem metabolism using a transcriptome-guided model of Arabidopsis metabolism. Plant Physiol. 192, 1359–1377 (2023).
de Oliveira Dal’Molin, C. G. et al. Metabolic reconstruction of Setaria italica: a systems biology approach for integrating tissue-specific omics and pathway analysis of bioenergy grasses. Front. Plant Sci. 7, 1138 (2016).
Moreno-Villena, J. J. et al. Spatial resolution of an integrated C4+CAM photosynthetic metabolism. Sci. Adv. 8, eabn2349 (2022).
von Bismarck, T. et al. Growth in fluctuating light buffers plants against photorespiratory perturbations. Nat. Commun. 14, 7052 (2023).
Fu, X., Schlüter, U., Smith, K., Weber, A. P. M. & Walker, B. J. Metabolomics of related C3 and C4 Flaveria species indicate differences in the operation of photorespiration under fluctuating light. Plant Direct 8, e70012 (2024).
Shaw, R. & Kundu, S. Metabolic plasticity and inter-compartmental interactions in rice metabolism: an analysis from reaction deletion study. PLoS ONE 10, e0133899 (2015).
Cheung, C. Y. M., Ratcliffe, R. G. & Sweetlove, L. J. A method of accounting for enzyme costs in flux balance analysis reveals alternative pathways and metabolite stores in an illuminated Arabidopsis leaf. Plant Physiol. 169, 1671–1682 (2015).
Hay, J. & Schwender, J. Computational analysis of storage synthesis in developing Brassica napus L. (oilseed rape) embryos: flux variability analysis in relation to 13C metabolic flux analysis. Plant J. 67, 513–525 (2011).
Chatterjee, A. & Kundu, S. Revisiting the chlorophyll biosynthesis pathway using genome scale metabolic model of Oryza sativa japonica. Sci. Rep. 5, 14975 (2015).
Schuster, S., Dandekar, T. & Fell, D. Detection of elementary flux modes in biochemical networks: a promising tool for pathway analysis and metabolic engineering. Trends Biotechnol. 17, 53–60 (1999).
Huma, B., Kundu, S., Poolman, M. G., Kruger, N. J. & Fell, D. A. Stoichiometric analysis of the energetics and metabolic impact of photorespiration in C3 plants. Plant J. 96, 1228–1241 (2018).
Hill, S. A. & ap Rees, T. Fluxes of carbohydrate metabolism in ripening bananas. Planta 192, 52–60 (1994).
Ratcliffe, R. G. & Shachar-Hill, Y. Measuring multiple fluxes through plant metabolic networks. Plant J. 45, 490–511 (2006).
Kruger, N. J. & Ratcliffe, R. G. Fluxes through plant metabolic networks: measurements, predictions, insights and challenges. Biochem. J 465, 27–38 (2015).
Moreira, T. B., Lima, J. M., Coca, G. C. & Williams, T. C. R. Insights into the spatial and temporal organisation of plant metabolism from network flux analysis. Theor. Exp. Plant Physiol. 31, 215–226 (2019).
Koley, S., Jyoti, P., Lingwan, M. & Allen, D. K. Isotopically nonstationary metabolic flux analysis of plants: recent progress and future opportunities. New Phytol. 242, 1911–1918 (2024).
Williams, T. C. R. et al. Metabolic network fluxes in heterotrophic Arabidopsis cells: stability of the flux distribution under different oxygenation conditions. Plant Physiol. 148, 704–718 (2008).
Masakapalli, S. K. et al. Subcellular flux analysis of central metabolism in a heterotrophic Arabidopsis cell suspension using steady-state stable isotope labeling. Plant Physiol. 152, 602–619 (2010).
Le, X. H. & Millar, A. H. The diversity of substrates for plant respiration and how to optimize their use. Plant Physiol. 191, 2133–2149 (2023).
Alonso, A. P., Goffman, F. D., Ohlrogge, J. B. & Shachar-Hill, Y. Carbon conversion efficiency and central metabolic fluxes in developing sunflower (Helianthus annuus L.) embryos. Plant J. 52, 296–308 (2007).
Allen, D. K., Ohlrogge, J. B. & Shachar-Hill, Y. The role of light in soybean seed filling metabolism. Plant J. 58, 220–234 (2009).
Alonso, A. P., Val, D. L. & Shachar-Hill, Y. Central metabolic fluxes in the endosperm of developing maize seeds and their implications for metabolic engineering. Metab. Eng. 13, 96–107 (2011).
Lonien, J. & Schwender, J. Analysis of metabolic flux phenotypes for two Arabidopsis mutants with severe impairment in seed storage lipid synthesis. Plant Physiol. 151, 1617–1634 (2009).
Schwender, J., Shachar-Hill, Y. & Ohlrogge, J. B. Mitochondrial metabolism in developing embryos of Brassica napus. J. Biol. Chem. 281, 34040–34047 (2006).
Tcherkez, G. et al. In folio respiratory fluxomics revealed by 13C isotopic labeling and H/D isotope effects highlight the noncyclic nature of the tricarboxylic acid ‘cycle’ in illuminated leaves. Plant Physiol. 151, 620–630 (2009).
Gauthier, P. P. G. et al. In folio isotopic tracing demonstrates that nitrogen assimilation into glutamate is mostly independent from current CO2 assimilation in illuminated leaves of Brassica napus. New Phytol. 185, 988–999 (2010).
Shameer, S., Ratcliffe, R. G. & Sweetlove, L. J. Leaf energy balance requires mitochondrial respiration and export of chloroplast NADPH in the light. Plant Physiol. 180, 1947–1961 (2019).
Lim, S. L. et al. In planta study of photosynthesis and photorespiration using NADPH and NADH/NAD+ fluorescent protein sensors. Nat. Commun. 11, 3238 (2020).
Igamberdiev, A. U. & Bykova, N. V. Mitochondria in photosynthetic cells: coordinating redox control and energy balance. Plant Physiol. 191, 2104–2119 (2023).
Szecowka, M. et al. Metabolic fluxes in an illuminated Arabidopsis rosette. Plant Cell 25, 694–714 (2013).
Arrivault, S. et al. Metabolite pools and carbon flow during C4 photosynthesis in maize: 13CO2 labeling kinetics and cell type fractionation. J. Exp. Bot. 68, 283–298 (2017).
Medeiros, D. B. et al. 13CO2 labeling kinetics in maize reveal impaired efficiency of C4 photosynthesis under low irradiance. Plant Physiol. 190, 280–304 (2022).
Wieloch, T. & Sharkey, T. D. Compartment-specific energy requirements of photosynthetic carbon metabolism in Camelina sativa leaves. Planta 255, 103 (2022).
Xu, Y., Wieloch, T., Kaste, J. A. M. & Sharkey, T. D. Reimport of carbon from cytosolic and vacuolar sugar pools into the Calvin–Benson cycle explains photosynthesis labeling anomalies. Proc. Natl Acad. Sci. USA 119, e2121531119 (2022).
Treves, H. et al. Carbon flux through photosynthesis and central carbon metabolism show distinct patterns between algae, C3 and C4 plants. Nat. Plants 8, 78–91 (2022).
Hasunuma, T. et al. Metabolic turnover analysis by a combination of in vivo 13C-labelling from 13CO2 and metabolic profiling with CE-MS/MS reveals rate-limiting steps of the C3 photosynthetic pathway in Nicotiana tabacum leaves. J. Exp. Bot. 61, 1041–1051 (2010).
Ma, F., Jazmin, L. J., Young, J. D. & Allen, D. K. Isotopically nonstationary 13C flux analysis of changes in Arabidopsis thaliana leaf metabolism due to high light acclimation. Proc. Natl Acad. Sci. USA 111, 16967–16972 (2014).
Young, J. D., Shastri, A. A., Stephanopoulos, G. & Morgan, J. A. Mapping photoautotrophic metabolism with isotopically nonstationary 13C flux analysis. Metab. Eng. 13, 656–665 (2011).
Sharkey, T. D. The end game(s) of photosynthetic carbon metabolism. Plant Physiol. 195, 67–78 (2024).
Makowka, A. et al. Glycolytic shunts replenish the Calvin–Benson–Bassham cycle as anaplerotic reactions in cyanobacteria. Mol. Plant 13, 471–482 (2020).
Wieloch, T., Augusti, A. & Schleucher, J. Anaplerotic flux into the Calvin–Benson cycle: hydrogen isotope evidence for in vivo occurrence in C3 metabolism. New Phytol. 234, 405–411 (2022).
Evans, S. E. et al. Rubisco supplies pyruvate for the 2-C-methyl-d-erythritol-4-phosphate pathway. Nat. Plants 10, 1453–1463 (2024).
Xu, Y., Schmiege, S. C. & Sharkey, T. D. The oxidative pentose phosphate pathway in photosynthesis: a tale of two shunts. New Phytol. 242, 2453–2463 (2024).
Wieloch, T., Augusti, A. & Schleucher, J. A model of photosynthetic CO2 assimilation in C3 leaves accounting for respiration and energy recycling by the plastidial oxidative pentose phosphate pathway. New Phytol. 239, 518–532 (2023).
Xu, Y., Fu, X., Sharkey, T. D., Shachar-Hill, Y. & Walker, A. B. The metabolic origins of non-photorespiratory CO2 release during photosynthesis: a metabolic flux analysis. Plant Physiol. 186, 297–314 (2021).
Schwender, J., Goffman, F., Ohlrogge, J. B. & Shachar-Hill, Y. Rubisco without the Calvin cycle improves the carbon efficiency of developing green seeds. Nature 432, 779–782 (2004).
Schwender, J. et al. Quantitative multilevel analysis of central metabolism in developing oilseeds of oilseed rape during in vitro culture. Plant Physiol. 168, 828–848 (2015).
Tsogtbaatar, E., Cocuron, J. C. & Alonso, A. P. Non-conventional pathways enable pennycress (Thlaspi arvense L.) embryos to achieve high efficiency of oil biosynthesis. J. Exp. Bot. 71, 3037–3051 (2020).
Deslandes-Hérold, G. et al. The PRK/Rubisco shunt strongly influences Arabidopsis seed metabolism and oil accumulation, affecting more than carbon recycling. Plant Cell 35, 808–826 (2023).
Bauwe, H. Photorespiration—Rubisco’s repair crew. J. Plant Physiol. 280, 153899 (2023).
Rachmilevitch, S., Cousins, A. B. & Bloom, A. J. Nitrate assimilation in plant shoots depends on photorespiration. Proc. Natl Acad. Sci. USA 101, 11506–11510 (2004).
Rosa-Téllez, S. et al. The serine–glycine–one-carbon metabolic network orchestrates changes in nitrogen and sulfur metabolism and shapes plant development. Plant Cell 36, 404–426 (2024).
Walker, B., Schmiege, S. C. & Sharkey, T. D. Re-evaluating the energy balance of the many routes of carbon flow through and from photorespiration. Plant Cell Environ. 47, 3365–3374 (2024).
Farquhar, G. D., von Caemmerer, S. & Berry, J. A. A biochemical model of photosynthetic CO2 assimilation in leaves of C3 species. Planta 149, 78–90 (1980).
Abadie, C., Boex-Fontvieille, E. R. A., Carroll, A. J. & Tcherkez, G. In vivo stoichiometry of photorespiratory metabolism. Nat. Plants 2, 15220 (2016).
Abadie, C. & Tcherkez, G. 13C isotope labelling to follow the flux of photorespiratory intermediates. Plants 10, 427 (2021).
Busch, F. A., Sage, R. F. & Farquhar, G. D. Plants increase CO2 uptake by assimilating nitrogen via the photorespiratory pathway. Nat. Plants 4, 46–54 (2018).
Fu, X., Gregory, L. M., Weise, S. E. & Walker, B. J. Integrated flux and pool size analysis in plant central metabolism reveals unique roles of glycine and serine during photorespiration. Nat. Plants 9, 169–178 (2023).
Timm, S. et al. Glycine decarboxylase controls photosynthesis and plant growth. FEBS Lett. 586, 3692–3697 (2012).
Timm, S. et al. Mitochondrial dihydrolipoyl dehydrogenase activity shapes photosynthesis and photorespiration of Arabidopsis thaliana. Plant Cell 27, 1968–1984 (2015).
Kebeish, R. et al. Chloroplastic photorespiratory bypass increases photosynthesis and biomass production in Arabidopsis thaliana. Nat. Biotechnol. 25, 593–599 (2007).
Maier, A. et al. Transgenic introduction of a glycolate oxidative cycle into A. thaliana chloroplasts leads to growth improvement. Front. Plant Sci. 3, 38 (2012).
South, P. F., Cavanagh, A. P., Liu, H. W. & Ort, D. R. Synthetic glycolate metabolism pathways stimulate crop growth and productivity in the field. Science 363, eaat9077 (2019).
Eisenhut, M., Roell, M. S. & Weber, A. P. M. Mechanistic understanding of photorespiration paves the way to a new green revolution. New Phytol. 223, 1762–1769 (2019).
George, K. W., Alonso-Gutierrez, J., Keasling, J. D. & Lee, T. S. in Biotechnology of Isoprenoids (eds Schrader, J. & Bohlmann, J.) 355–389 (Springer, 2015).
Yang, W. et al. Advances in pharmacological activities of terpenoids. Nat. Prod. Commun. 15, 1934578X20903555 (2020).
Joshi, S. & Mishra, S. Recent advances in biofuel production through metabolic engineering. Bioresour. Technol. 352, 127037 (2022).
Jiang, H. & Wang, X. Biosynthesis of monoterpenoid and sesquiterpenoid as natural flavors and fragrances. Biotechnol. Adv. 65, 108151 (2023).
O’Neill, E. C. & Kelly, S. Engineering biosynthesis of high-value compounds in photosynthetic organisms. Crit. Rev. Biotechnol. 37, 779–802 (2017).
Dai, Z., Cui, G., Zhou, S. F., Zhang, X. & Huang, L. Cloning and characterization of a novel 3-hydroxy-3-methylglutaryl coenzyme A reductase gene from Salvia miltiorrhiza involved in diterpenoid tanshinone accumulation. J. Plant Physiol. 168, 148–157 (2011).
Kai, G. et al. Metabolic engineering tanshinone biosynthetic pathway in Salvia miltiorrhiza hairy root cultures. Metab. Eng. 13, 319–327 (2011).
Li, Y. et al. Advanced metabolic engineering strategies for increasing artemisinin yield in Artemisia annua. L. Hortic. Res. 11, uhad292 (2024).
Masakapalli, S. K. et al. Metabolic flux phenotype of tobacco hairy roots engineered for increased geraniol production. Phytochemistry 99, 73–85 (2014).
Rodriguez, S. et al. ATP citrate lyase mediated cytosolic acetyl-CoA biosynthesis increases mevalonate production in Saccharomyces cerevisiae. Microb. Cell Fact. 15, 48 (2016).
Chapman, K. D., Dyer, J. M. & Mullen, R. T. Commentary: why don’t plant leaves get fat? Plant Sci. 207, 128–134 (2013).
Wang, Y. Q. et al. Proteomic analysis of chromoplasts from six crop species reveals insights into chromoplast function and development. J. Exp. Bot. 64, 949–961 (2013).
Andrews, T. J. & Kane, H. J. Pyruvate is a by-product of catalysis by ribulosebisphosphate carboxylase/oxygenase. J. Biol. Chem. 266, 9447–9452 (1991).
Chu, K. L. et al. Metabolic flux analysis of the non-transitory starch tradeoff for lipid production in mature tobacco leaves. Metab. Eng. 69, 231–248 (2022).
Eastmond, P. J., Dennis, D. T. & Rawsthorne, S. Evidence that a malate/inorganic phosphate exchange translocator imports carbon across the leucoplast envelope for fatty acid synthesis in developing castor seed endosperm. Plant Physiol. 114, 851–856 (1997).
Morley, S. A. et al. Expression of malic enzyme reveals subcellular carbon partitioning for storage reserve production in soybeans. New Phytol. 239, 1834–1851 (2023).
Schwender, J. Walking the ‘design–build–test–learn’ cycle: flux analysis and genetic engineering reveal the pliability of plant central metabolism. New Phytol. 239, 1539–1554 (2023).
Walker, B. J., Kramer, D. M., Fisher, N. & Fu, X. Flexibility in the energy balancing network of photosynthesis enables safe operation under changing environmental conditions. Plants 9, 301 (2020).
Vanhercke, T. et al. Step changes in leaf oil accumulation via iterative metabolic engineering. Metab. Eng. 39, 237–246 (2017).
Strand, D. D. & Walker, B. J. Energetic considerations for engineering novel biochemistries in photosynthetic organisms. Front. Plant Sci. 14, 1116812 (2023).
Brunk, E. et al. Characterizing strain variation in engineered E. coli using a multi-omics-based workflow. Cell Syst. 2, P335–P346 (2016).
Brand, A. & Tissier, A. Control of resource allocation between primary and specialized metabolism in glandular trichomes. Curr. Opin. Plant Biol. 66, 102172 (2022).
Asadollahi, M. A. et al. Enhancing sesquiterpene production in Saccharomyces cerevisiae through in silico driven metabolic engineering. Metab. Eng. 11, 328–334 (2009).
Assil-Companioni, L. et al. Engineering of NADPH supply boosts photosynthesis-driven biotransformations. ACS Catal. 10, 11864–11877 (2020).
Ding, N., Yuan, Z., Sun, L. & Yin, L. Dynamic and static regulation of nicotinamide adenine dinucleotide phosphate: strategies, challenges, and future directions in metabolic engineering. Molecules 29, 3687 (2024).
Chen, R. et al. Engineering cofactor supply and recycling to drive phenolic acid biosynthesis in yeast. Nat. Chem. Biol. 18, 520–529 (2022).
Yu, T. et al. Metabolic reconfiguration enables synthetic reductive metabolism in yeast. Nat. Metab. 4, 1551–1559 (2022).
Clomburg, J. M., Qian, S., Tan, Z., Cheong, S. & Gonzalez, R. The isoprenoid alcohol pathway, a synthetic route for isoprenoid biosynthesis. Proc. Natl Acad. Sci. USA 116, 12810–12815 (2019).
Smith, E. N. et al. Improving photosynthetic efficiency toward food security: strategies, advances, and perspectives. Mol. Plant 6, 1547–1563 (2023).
Pasoreck, E. K. et al. Terpene metabolic engineering via nuclear or chloroplast genomes profoundly and globally impacts off‐target pathways through metabolite signalling. Plant Biotechnol. J. 14, 1862–1875 (2016).
Lynch, J. H., Huang, X. Q. & Dudareva, N. Silent constraints: the hidden challenges faced in plant metabolic engineering. Curr. Opin. Biotechnol. 69, 112–117 (2021).
Lynch, J. H. et al. Modulation of auxin formation by the cytosolic phenylalanine biosynthetic pathway. Nat. Chem. Biol. 16, 850–856 (2020).
Razaghi-Moghadam, Z. & Nikoloski, Z. GeneReg: a constraint-based approach for design of feasible metabolic engineering strategies at the gene level. Bioinformatics 37, 1717–1723 (2021).
Gurdo, N., Volke, D. C., McCloskey, D. & Nikel, P. I. Automating the design–build–test–learn cycle towards next-generation bacterial cell factories. New Biotechnol. 74, 1–15 (2023).
Yilmaz, S., Nyerges, A., van der Oost, J., Church, G. M. & Claassens, N. J. Towards next-generation cell factories by rational genome-scale engineering. Nat. Catal. 5, 751–765 (2022).
Sears, R. G., Lenaghan, S. C. & Stewart, C. N. AI to enable plant cell metabolic engineering. Trends Plant Sci. 29, 126–129 (2024).
Sahu, A., Blätke, M. A., Szymański, J. J. & Töpfer, N. Advances in flux balance analysis by integrating machine learning and mechanism-based models. Comput. Struct. Biotechnol. J. 19, 4626–4640 (2021).
Patané, A. et al. Multi-objective optimization of genome-scale metabolic models: the case of ethanol production. Ann. Oper. Res. 276, 211–227 (2019).
Li, F. et al. Deep learning-based kcat prediction enables improved enzyme-constrained model reconstruction. Nat. Catal. 5, 662–672 (2022).
Gollub, M. G., Backes, T., Kaltenbach, H. M. & Stelling, J. ENKIE: a package for predicting enzyme kinetic parameter values and their uncertainties. Bioinformatics 40, btae652 (2024).
Wang, T. et al. DeepEnzyme: a robust deep learning model for improved enzyme turnover number prediction by utilizing features of protein 3D-structures. Brief. Bioinform. 25, bbae409 (2024).
Kroll, A., Rousset, Y., Hu, X. P., Liebrand, N. A. & Lercher, M. J. Turnover number predictions for kinetically uncharacterized enzymes using machine and deep learning. Nat. Commun. 14, 4139 (2023).
Kroll, A., Engqvist, M. K. M., Heckmann, D. & Lercher, M. J. Deep learning allows genome-scale prediction of Michaelis constants from structural features. PLoS Biol. 19, e3001402 (2021).
Salas-Nuñez, L. F. et al. Machine learning to predict enzyme–substrate interactions in elucidation of synthesis pathways: a review. Metabolites 14, 154 (2024).
Erbe, R., Gore, J., Gemmill, K., Gaykalova, D. A. & Fertig, E. J. The use of machine learning to discover regulatory networks controlling biological systems. Mol. Cell 82, 260–273 (2022).
Acknowledgements
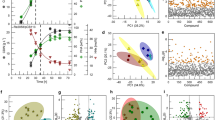
We thank T. C. R. Williams (Universidade de Brasília) for assistance with the data used for Fig. 2.
Author information
Authors and Affiliations
Contributions
L.J.S., R.G.R. and A.R.F. conceived and wrote the review.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Berkley Walker, John Morgan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sweetlove, L.J., Ratcliffe, R.G. & Fernie, A.R. Non-canonical plant metabolism. Nat. Plants 11, 696–708 (2025). https://doi.org/10.1038/s41477-025-01965-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-01965-3