Abstract
The plant hormone auxin regulates a wide variety of transcriptional responses depending on the cell type, environment and species. How this diversity is achieved may be related to the specific complement of auxin-signalling components in each cell. The levels of activators (class-A AUXIN RESPONSE FACTORS) and repressors (class-B ARFs) are particularly important. Tight regulation of ARF protein levels is probably key in determining this balance. Through comparative analysis of novel, dominant mutants in maize and the moss Physcomitrium patens, we have discovered a ~500-million-year-old mechanism of class-B ARF protein-level regulation mediated by proteasome degradation, important in determining cell fate decisions across land plants. Thus, our results add a key piece to the puzzle of how auxin regulates plant development.
Similar content being viewed by others
Main
Plants develop organs continuously through the activity of stem cells. Pattern formation, differentiation and tissue deformation define the shape of complex organs, while iterative development of these organs builds the plant body. Modulation of these developmental processes gives rise to a huge diversity in organ shapes and plant architectures. Despite this complexity, a single phytohormone, auxin, has a major conserved role across land plants1,2,3,4, underpinning cell fate decisions, organ initiation, shape and plant architecture. How auxin precisely regulates so many distinct processes is one of the biggest unresolved questions in plant biology.
Canonical auxin signalling occurs through the activity of a suite of deeply conserved transcriptional regulators found in all land plants3,5,6,7. The AUXIN/INDOLE-3-ACETIC-ACID repressors (AUX/IAAs), AUXIN RESPONSE FACTORS (ARFs) and the TRANSPORT INHIBITOR RESPONSE 1/AUXIN-SIGNALING F-BOX (TIR1/AFB) co-receptors are components of the best-characterized auxin-mediated transcriptional regulation pathway8,9,10. All three components are in multigene families that have greatly expanded in flowering plants3. When auxin levels are low, AUX/IAAs are bound to ARFs and recruit transcriptional co-repressors, preventing ARF activation of auxin-responsive genes. In elevated auxin levels, the TIR1/AFB co-receptor forms a complex with auxin and AUX/IAA repressors, resulting in ubiquitylation and degradation of AUX/IAAs via the 26S proteasome11,12,13. Degradation of AUX/IAAs relieves ARF repression, facilitating auxin transcriptional responses14. Recent studies have elaborated on this core pathway, showing that the TIR1/AFB proteins possess adenyl cyclase activity activated upon auxin binding and required for ARF activity15. In addition, the ABP1-TMK pathway, responsible for perceiving extracellular auxin, can influence ARF function by phosphorylating the non-canonical Aux/IAA proteins IAA32 and IAA34 (ref. 16).
Land plant ARFs are divided into three phylogenetic lineages; class-A, -B and -C, each associated with functions based on protoplast and Marchantia studies. Class-A ARFs are activators, while class-B and -C are potential repressors4,17,18. All ARFs share a DNA-binding domain (DBD), a middle domain and a C-terminal Phox and Bem1 (PB1) domain19,20. Differences in domain sequences lead to variation in DNA-binding activity21,22,23,24,25, interacting partners (including other ARFs, AUX/IAAs, other transcription factors, chromatin remodellers19) and oligomerization26. Modulation of auxin response is believed to be determined by the complement of ARF and AUX/IAA proteins in the cell, and in vitro analysis has revealed different activation levels of ARF–AUX/IAA protein combinations, indicating potential subfunctionalization mechanisms27. However, the elegant model of auxin-dependent ARF de-repression is not able to account for all auxin responses. For example, only 5 of the 22 Arabidopsis thaliana (Arabidopsis hereafter) ARFs functionally interact with AUX/IAAs in vivo19.
Although genetic studies implicate class-B function in diverse developmental processes28,29,30, their mode of action and regulation is unclear. Studies in non-vascular plants such as Marchantia and Physcomitrium indicate that class-B ARFs can directly compete with class-A ARFs to bind to promoter sequences13,31. In maize, class-A and -B ARFs exhibit clear DNA-binding site differences. However, a proportion of the sites (16%) are bound by both classes, thus supporting the hypothesis that competition between class-A and -B ARFs may be important for the regulation of some auxin-regulated genes across land plants24,32,33. In addition, class-B ARFs can function independently of AUX/IAAs, directly recruiting the TOPLESS (TPL) repressor to chromatin34,35,36,37,38, providing an AUX/IAA-independent way to repress gene expression. Regardless of mechanism, the level of class-B ARFs is a key determinant of auxin-regulated transcription. Without a clear understanding of class-B ARF protein regulation and how it has evolved, we lack a key piece of the auxin regulatory system puzzle.
To address this deficiency, we leveraged a comparative approach by investigating dominant class-B ARF mutants in the distantly related species Zea mays (maize) and the moss Physcomitrium patens (P. patens), which diverged ~500 million years ago (Ma)39. The mutations result in amino acid substitutions in a loop region in the DBD. We demonstrate that this contains a novel regulatory region that is required for degradation of class-B ARF proteins, thus revealing a deeply conserved mechanism of class-B ARF regulation in land plants.
Truffula has pleiotropic defects
Maize is a classic model genetic system with a rich history of developmental mutant analyses40. We found the striking, dominant Truffula (Trf) mutant in an ethyl methanesulfonate (EMS)-mutagenized population and analysed it in two inbred backgrounds: W22 and Mo17. In both inbreds, when compared to normal siblings, Trf mutants had reduced plant height, increased leaf number, and tassels and tassel flowers with ear traits (Fig. 1a–d and Supplementary Fig. 1). The mutation did not transmit through the ovule parent in Mo17, but did at a reduced rate in W22, permitting analysis of homozygotes. Mutant plants were half the height of normal siblings in W22 (Fig. 1a) and two-thirds the size in Mo17 (Fig. 1b). The reduced height results from shorter internodes, the stem section between leaves (Supplementary Fig. 1a,b). On average in Mo17, non-mutant siblings produced 15 leaves, while heterozygous mutants produced 19. In W22, non-mutants produced 15 leaves, heterozygotes produced 29 and homozygotes produced 54 (Fig. 1e). Mutants flowered at the same time as normal siblings (Supplementary Fig. 1c), hence the increase in leaf number is due to a shorter plastochron (time between leaf initiations) in Trf. Leaf phyllotaxy was also altered in the upper nodes of Trf, switching from distichous to spiral-like (Supplementary Fig. 1d). Additional Trf phenotypes were inbred dependent such as defects in midrib formation (15% of leaves), ranging from midribless leaves to multiple midribs (Fig. 1c, arrowheads), and leaf shape, forming very rare tube leaves in W22 (<1% of leaves). In W22, only a few kernels formed at the base of the normally staminate tassel, while in Mo17 many tassel florets were pistillate and tassel branches were often transformed into ears wrapped in husk leaves (Fig. 1d).
a–d, Normal (+/+) and Trf siblings (Trf/+, Trf/Trf) in the W22 (a,c) and Mo17 (b,d) inbred backgrounds. a,b, Mature plants. c, Midrib phenotypes in Trf leaves: midribless (i), normal (ii) and multiple midribs (iii). Scale bars, 1 cm. d, Tassel phenotypes showing ear-like traits in Trf siblings. Scale bar, 5 cm. e, Leaf number in mature plants of normal (blue) and Trf siblings (orange). **P < 0.05 determined by Kruskal–Wallis non-parametric test followed by Wilcox pairwise tests, n ≥ 6 plants. Black dot and error bars, mean ± s.d.
Trf is associated with a mutation in ZmARF28
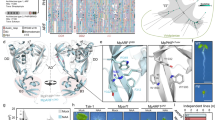
To identify the underlying causal mutation in Trf, we carried out whole-genome sequencing (WGS) bulk-segregant analysis41, identifying a region between 11.0 Mbp and 20.0 Mbp on chromosome 10 (Fig. 2a). Short sequence repeat (SSR) mapping (Fig. 2b and Supplementary Table 1) refined the region to between 13.5 Mbp and 14.0 Mbp. This region contains 8 annotated genes, of which 5 are expressed in the vegetative shoot apex according to RNA-seq (Fig. 2b, and Supplementary Fig. 2a and Table 2). Analysis of the WGS data in the mapping region reveals 188 EMS-type variants (G:C to A:T transitions) specific to Trf siblings. Of these SNPs, only one is in a conserved non-coding sequence, associated with Zm00001eb408820, a gene which is not significantly differentially expressed in Trf. SNPEff42 analysis predicts 7 moderate to high-effect EMS-type single nucleotide polymorphisms (SNPs) in the interval (Supplementary Table 3). Only a single SNP in exon 8 of ZmAUXIN RESPONSE FACTOR28 (ZmARF28, Zm00001eb408800), resulting in a Ser-to-Asn change in the DBD (Fig. 2c–f), is Trf specific when compared to other maize inbred backgrounds (Supplementary Table 3), suggesting that this mutation in ZmARF28 is probably responsible for the Trf phenotype.
a, Plotting WGS-BSA variant mean allelic distances in 0.5-Mbp windows across the maize NAM5.0 reference genome reveals a region on chromosome 10 (Chr10) which is more associated with Trf mutants than normal siblings (red box). n = 80 per genotype. Dot and error bars, mean ± s.e.m. Colour scale: mean allelic distance −0.25:dark blue to 0.45:red. b, PCR mapping narrows the region to between 13.5 Mbp and 14.0 Mbp. Mapping primer positions are indicated (AD_# and umc#), numbers represent Mbp position on Chr10, and recombinant numbers are shown in brackets. There are 8 genes in the interval, of which 5 have an FPKM > 1 in our shoot apical meristem RNA-seq dataset (green) and 3 have an FPKM < 1 (purple). A moderate effect, Trf-specific, EMS-type SNP (*) is present in Zm00001eb408800 (ZmARF28). c, Locations of the Trf, narD72, narD120, narC29 and narC18 mutations (red arrows). The exons (blue), 5’ and 3’ UTRs (grey), CRISPR sgRNA targets for deletions (magenta arrowheads), and small RNA regulatory sites (cyan) are indicated. d, Generalized cartoon depiction of the protein domains in class-B ARFs, with the DBD, B3 and PB1 domains indicated; the relative location of the mutations identified in this study are indicated in red. e,f, The maize Trf mutation and the moss narD72, narD120, narC29 and narC18 mutations map to the same loop region in the ARF protein. e, Multiple sequence alignments of ARFs from maize, Arabidopsis and moss. f, Homology modelling of the ZmARF28, PpARFb2 and PpARFb4 DBDs for both wild-type and mutant proteins (metrics in Supplementary Table 10). (i)–(iii), Zoomed-in images of the green-boxed regions. e,f, Mutation locations (red) and amino acids previously shown to be important for ARF function (blue) are indicated.
The Trf phenotype, although distinct from other reported ARF mutants in maize43, is consistent with the defects observed in maize seedlings treated with auxin transport inhibitors44 and other auxin-signalling mutants43 in which changes to organ initiation and medial-lateral leaf patterning are observed. To investigate the link between Trf and auxin response further, we carried out an RNA-seq analysis of vegetative shoot apices, at a time consistent with the formation of the Trf vegetative phenotypes (Supplementary Fig. 2a–d and Table 4). A total of 1,804 genes were significantly differentially expressed (sigDE), of which 455 were downregulated (Padj < 0.05, log2fold change (FC) < −0.5). Interestingly, when compared to published class-B ARF DNA affinity purification sequencing (DAP-seq) data24, potential class-B ARF target genes were both up- and downregulated (19% and 16%, respectively, Supplementary Fig. 2d), suggesting that ZmARF28’s role may not be only as a straightforward repressor. Consistent with a defect in an ARF, sigDE genes were enriched for Gene Ontology (GO) terms associated with auxin signalling and response (Padj = 0.00002, Supplementary Table 5). In line with the Trf phenotype, we also observed enrichments for GO terms associated with meristem development (Supplementary Fig. 2c). Crucially, this RNA-seq data showed that ZmARF28 is not significantly differentially expressed in Trf (Supplementary Fig. 2b), suggesting that the Trf mutation affects protein activity or post-translational regulation of ZmARF28. To further evaluate the auxin response in Trf, we treated seedlings with indole-acetic-acid (IAA) and evaluated transcriptional response using RNA-seq and quantitative (q)PCR (Supplementary Fig. 2e–h and Tables 6 and 7). At 30 min post-IAA treatment, both normal and Trf siblings showed significant differential gene expression in response to auxin, although the overlap is small (Supplementary Fig. 2f), suggesting differential responses potentially arising from the existing dysregulation of auxin-signalling components in Trf seedlings before treatment. Of the 1,130 sigDE genes, 831 genes were downregulated in Trf siblings treated with IAA versus mock (Padj < 0.05, log2FC < −0.5), consistent with a role as a repressor of auxin-responsive gene expression. Of the 299 significantly upregulated genes (Padj < 0.05, log2FC > 0.5), 58 were potential class-B ARF targets based on overlap with published DAP-seq data24 (Supplementary Fig. 2g), and 5 of these were AUX/IAA genes, highlighting a potential role of ZmARF28 in promoting the expression of other repressors of auxin response. Mapping fragments per kilobase of transcript per million mapped reads (FPKM) of all genes associated with auxin GO terms and expressed in both Trf and normal siblings showed clear auxin transcriptional response differences (Supplementary Fig. 2h). Combined, this evidence supports the hypothesis that the Trf phenotype is at least in part caused by altered auxin transcriptional response due to the identified mutation in ZmARF28.
As ZmARF28 is a co-orthologue of Arabidopsis AtARF2 (Supplementary Fig. 3), we carried out sequence alignments (Fig. 2e) to see whether the mutation was in an already characterized residue. We found that although the mutation is in the DBD, the Trf mutation is in the dimerization domain, not the B3 domain involved in DNA binding, and is outside of the core residues previously shown to be required for functionality in AtARF1 and AtARF5 (Fig. 2d–f, blue, ref. 21). Homology modelling revealed that the Trf residue is in an externally facing loop region and is not predicted to cause notable structural changes (Fig. 2f). These findings suggest that the novel Trf phenotype is not due to a mutation in an already characterized ARF residue, potentially revealing a novel regulatory module.
Although protein structure is predicted to be conserved, amino acid sequence is variable in this region of plant class-B ARFs (Fig. 2e). This suggests that the Trf phenotype could be due to the disruption of a grass-specific regulatory pathway in the expanded family of class-B ARFs. Alternatively, the Trf mutation could highlight the role of a functionally conserved region of class-B ARFs that has yet to be described due to lack of sequence conservation.
Mutations in P. patens class-B ARFs cause auxin resistance
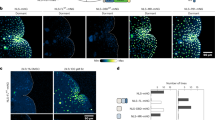
Evidence that the Trf mutation revealed an ancient mechanism of class-B ARF regulation emerged from parallel forward-genetic analyses in the moss P. patens. We found that the genomes of four newly isolated P. patens NAA-RESISTANT (nar) mutants45 had missense mutations in the same Trf region in two of the four class-B ARF genes (PpARFb2 and PpARFb4, Fig. 2c–f). narC29 and narC18 mutants have missense mutations, causing Glu-to-Lys substitutions (E263K) in PpARFb4/Pp6c6_11530, while narD72 has the same substitution in PpARFb2/Pp6c16_6840 (E266K). narD120 has a Gln-to-Arg substitution in PpARFb2 (R269Q). When the DBD are aligned, the residues affected in the moss mutants flank the position altered by the maize Trf mutation (Fig. 2e,f). These class-B ARF moss nar mutants had clear defects in the auxin-regulated transition from highly photosynthetic chloronemal cells to ‘nutrient foraging’ spreading caulonemal cells, and a subsequent delay in the production of 3D-leafy shoot (gametophore) growth forms. This effect was more pronounced on media containing ammonium which delays transitions promoted by auxin46. Unlike wild type, the nar mutants produce gametophores even when grown on media containing micromolar amounts of 1-naphthaleneacetic acid (NAA) (Fig. 3a). Further supporting the reduction in auxin response in these nar mutants, expression of the proDR5:DsRed2 auxin-signalling reporter was also dramatically reduced in narC29, narC18, narD72 and to a lesser extent in narD120 (Fig. 3a and Supplementary Fig. 4a).
a, Comparison of wild type, nar and ∆pparfb4 deletion mutants after 21 days of growth on standard medium (BCD) and BCD supplemented with 5 mM ammonium tartrate or 5 µM NAA. Plants grown on NAA were imaged for RFP fluorescence from the proDR5:DsRed auxin-response reporter. On NAA-containing media, wild-type and ∆pparfb4 strains produce ectopic rhizoids and high levels of RFP fluorescence, whereas the nar mutants produce green leafy gametophores. Arrowheads indicate leafy shoots in narD120 and ∆pparfb4. Scale bar, 5 mm. b, Comparison of wild type and gene-edited point mutants grown for 21 days, as in a. c, Expression of select auxin-responsive genes (PpRSL6, ROOT HAIR DEFECTIVE SIX-LIKE 6; PpIAA2, Aux/IAA2; PpHSL1-2, HOOKLESS1-LIKE 2) in wild type, pparfb2E266K and auxin-response mutant ppiaa2G325S grown for 1 h on media containing 10 µM IAA or 0.01% ethanol (3 technical replicates). Bars represent mean ± s.e.m. Transcript levels were normalized to the APT gene. d, Growth comparisons of wild type, the pparfb2E266K mutant and the pparfb2E266K mutant with second-site mutations, as in a. brd–, L614A + F615A substitutions in the B3 repression domain; pb1K→A, K682A substitution in the PB1 domain; ∆pb1, a 5-bp frameshift-causing deletion starting at G680 codon; dbd–, P194A + R196A substitutions affecting DNA-interacting residues. n ≥ 4. Scale bars, 5 mm.
To independently verify that pparfb mutations are responsible for the above nar phenotypes, we used oligo-mediated gene editing47 to introduce the E-to-K substitution to all four PpARFb loci. The pparfb2E266K and pparfb4E263K lines recapitulated the nar phenotypes of narD72 and narC29, respectively, and the pparfb1E266K and pparfb3E266K lines were also appreciably resistant to NAA (Fig. 3b and Supplementary Fig. 4b). pparfb2R269Q and pparfb4R266Q mutations conferred weak nar phenotypes similar to narD120 (Fig. 3b and Supplementary Fig. 4c). Interestingly, combining both the E-to-K and R-to-Q mutations in the same gene resulted in dramatically stronger phenotypes approaching those of quadruple mutants lacking all four TIR1/AFB genes which are completely auxin insensitive (Fig. 3b and Supplementary Fig. 4c). In-frame deletions and insertions in this region of PpARFb2 also conferred either intermediate phenotypes similar to those of pparfb2E266K or strong phenotypes similar to those of pparfb2E266K+R269Q (Supplementary Table 8). Reduced expression of proDR5:DsRed and reduced auxin induction of auxin-regulated genes in pparfb2E266K confirmed that the phenotypes are due to reduced auxin response (Fig. 3c and Supplementary Fig. 4d–e). Interestingly, Trf-like substitutions in PpARFb2 (M268N) and PpARFb4 (L265N) caused no phenotypic changes, but both Trf-like and ZmARF28-like 9-amino-acid swaps in the loop caused nar phenotypes (Supplementary Fig. 5).
The phenotypes of the four nar mutants are similar to those of lines that over-accumulate PpARFb protein either by expression with a strong heterologous promoter13 or by disruption of the miR1219 and tasiARF target sequences46 (Supplementary Fig. 4f). The nar mutations do not affect any known small RNA regulatory elements, but the phenotypic similarities suggest that the nar lesions represent dominant mutations. To test this, we used CRISPR/Cas9 to delete the mutated genes in the mutant lines. Deletion of either the wild-type PpARFb2 or PpARFb4 genes in moss has minor phenotypic effects13, but deletion of the mutated genes in the nar mutants restored NAA sensitivity indicating that, similar to the maize Trf mutant, the P. patens pparfb2 and pparfb4 variants are dominant mutations (Fig. 3a and Supplementary Fig. 4a,g) with defects in auxin signalling.
All major domains are required for the dominant phenotype
ARF domains confer specific functionality: the B3 repression domain (BRD) recruits the TPL repressor34,48, the PB1 domain mediates oligomerization49, and the DBD binds target DNA sequences and can facilitate binding to other proteins20,21. To better understand how the above mutations confer gain-of-function auxin phenotypes in moss, we introduced second-site mutations affecting key residues of the known functional domains into the pparfb2E266K mutant (Fig. 3d and Supplementary Fig. 6). We anticipated that if the functionality of a domain was required for the pparfb2E266K phenotype, we would restore the wild-type phenotype. Mutating the BRD (L614S + F615S) eliminates the interaction with a TPL homologue (TPL2/Pp6c9_10690) and reverts the pparfb2E266K phenotype to wild type (Fig. 3d and Supplementary Fig. 6a–c). Similarly, mutating a conserved lysine residue in the positive face of the PB1 domain (K682A) partially suppresses the arfb2E266K phenotype, and a frameshift mutation near the start of the domain results in near-complete suppression. Introducing a pair of substitutions (P194A + R196A) shown to abolish DNA binding21 restores NAA response, although additional phenotypes appear, suggesting dominant-negative effects. None of these functional-domain mutations caused phenotypic effects in an otherwise wild-type PpARFb2 (Supplementary Fig. 6a). These data show that the gain-of-function phenotypes depend on all known ARF protein functions: DNA binding, TPL recruitment and PB1-mediated oligomerization. Interestingly, the requirements of both the BRD and PB1 domains indicate that TPL recruitment through PB1 interactions with AUX/IAAs is insufficient and that oligomerization with other ARFs is required. Indeed, PpARFb4 overexpression phenotypes were previously shown to be independent of AUX/IAAs13.
Mutations increase class-B ARF stability
On the basis of the phenotypes and lack of ZmARF28 and PpARFb expression changes in the mutants, we hypothesized that the maize Trf and P. patens arfb mutant phenotypes were due to accumulation of ZmARF28 and PpARFb protein, respectively. To test whether the PpARFb proteins are more abundant in the moss mutants, we knocked in mYPet yellow fluorescent protein (YFP) tags to the four endogenous PpARFb loci and the mutants to quantify protein abundance. The YFP signal dramatically increased in all nuclei of the mutants compared with the weak and sporadic YFP signal detected in nuclei in the wild type (Fig. 4a,b and Supplementary Fig. 7a,b). Protein localization did not change in the mutants (Fig. 4a). Since the transcript levels are similar in the mutant and wild-type lines (Supplementary Fig. 7c), the increased YFP signal suggests that the mutations affect PpARFb stability. Consistent with accumulation of PpARFb protein resulting in the mutant phenotypes, mutations with higher YFP signal had stronger mutant phenotypes and less auxin response (Fig. 3a, and Supplementary Figs. 4 and 5).
a,b, PpARFb2-YFP signal in P. patens wild type, pparfb2R269Q, pparfb2E266K and the pparfb2E226K,R269Q double mutants. a, Representative confocal microscopy images. Yellow, PpARFb2-YFP; magenta, chlorophyll autofluorescence. Scale bar, 25 µm. b, Nuclear fluorescence quantification. Centre black dot and error bars, mean ± s.d.; **P < 0.01, determined using Kruskal–Wallis non-parametric test followed by Wilcox pairwise tests. c, Western blot of ZmARF28 in +/+ and Trf/+ mutant siblings from a W22 backcross population. Lanes are different biological replicates. Anti-tubulin was used as a loading control. d, Quantification of ZmARF28 abundance relative to TUBULIN. **P < 0.01, Welch 2-sample t-test. Centre black dot and error bars, mean ± s.d.; 3 technical replicates, 2 biological replicates in each. e, Western blots of ZmARF28-GFP accumulation in N. bethamiana, with (+) and without (−) MG132 treatment, n = 3. f, Mean ± s.e.m. ratiometric signal of AtARF2-mNeonGreen (blue) and AtARF2-Trf-mNeonGreen (orange) in Arabidopsis protoplasts, relative to mScarlet after cycloheximide (CHX) treatment (≥2,166 cells across 6 technical replicates analysed). g, Western blots of anti-GFP immunoprecipitated samples (αGFP IP) and cleared extracts (input) using anti-GFP and anti-ubiquitin antibodies. Extracts were from bortezomib-treated untransformed control (‘C’) and stable lines expressing YFP-fused ARFb2’s DBD with and without the E266K + R269Q substitutions. Mutant IP samples were diluted 5-fold to normalize DBD-YFP amounts loaded. n = 3.
To determine whether the wild-type moss proteins are subject to degradation via the 26S proteasome, we treated the wild-type YFP lines overnight with the proteasome inhibitor, bortezomib50. Quantification of the YFP signal revealed significant increases in protein level for all four wild-type lines in the presence of the inhibitor (Supplementary Fig. 7d–f), supporting the hypothesis that PpARFb proteins are subject to proteasome-dependent degradation.
To determine whether protein levels were also elevated in the maize Trf mutants, we developed an antibody against ZmARF28. Semi-quantitative western blot analysis of whole-protein extract from vegetative seedlings revealed a statistically significant increase in ZmARF28 protein abundance in Trf compared with normal siblings (Fig. 4c,d), supporting the hypothesis that ZmARF28 is stabilized in Trf. To test whether ZmARF28 was also subject to proteasome-dependent degradation, we expressed wild-type and Trf versions of ZmARF28-GFP in Nicotiana benthamiana and maize protoplasts treated with the proteasome inhibitor MG132 (ref. 51). Western blots revealed accumulation of ZmARF28 protein in the presence of MG132, which was more marked in the wild-type ZmARF28-GFP samples (Fig. 4e and Supplementary Fig. 2i). This suggests that both the Trf and moss mutants disrupt a deeply functionally conserved class-B ARF region required for proteasome-dependent degradation.
To further test the effect of the Trf mutation on protein stability in flowering plants, we expressed the Arabidopsis orthologue of ZmARF28, AtARF2, and a Trf variant of AtARF2 (arf2T298N) in Arabidopsis mesophyll protoplasts using a ratiometric reporter system. Protoplasts expressing AtARF2 or arf2T298N were treated with the transcriptional inhibitor cycloheximide (CHX). Similar to AtARF1 (ref. 52), AtARF2 shows a half-life of ~4 h. Conversely, arf2T298N only lost 23% of its initial protein level after 4 h of CHX treatment, suggesting that its half-life is longer than that of wild-type AtARF2 (Fig. 4f). These results show that the Trf mutation in AtARF2 results in increased protein accumulation not due to increased rates of translation but more likely due to increases in protein stability, further supporting functional conservation of this region.
To test whether the mutations disrupt ubiquitylation of the domain, we immunoprecipitated wild-type and mutant versions of the PpARFb2 DBD from bortezomib-treated proPpARFb2:DBDWT/mut-YFP moss lines and probed with an anti-ubiquitin antibody. Polyubiquitylated proteins were detected for the wild-type DBD but not for the E266K/R269Q version (Fig. 4g). Combined with protein stability findings, these data indicate that the dominant mutations in moss and maize uncovered a region required for class-B ARF degradation by the 26S proteasome.
Discussion
The precise complement and abundance of auxin-signalling components in each cell determines auxin sensitivity and how the auxin signal is decoded to produce distinct transcriptional responses. Data across land plants show that the relative levels of class-A and -B ARFs have a key role in regulating auxin response4,13. We leveraged a comparative analysis of species that diverged ~500 Ma and identified an ancient regulatory mechanism that controls class-B ARF accumulation. By combining our findings with an emerging understanding of differential class-A and -B binding preferences24,32,33, we can start to build a model of how class-A and -B ARF function may be regulated.
Our comparative mutational analysis revealed that the level of class-B ARFs is essential for dictating cell fate decisions in land plants. Elevated class-B ARFs, as in the dominant Pparfb mutations, block or delay differentiation towards a leafy gametophore fate. Similarly, in maize, class-B ARF ZmARF28 accumulation probably influences meristem cell fate, leaf medio-lateral patterning and sex determination. Defects in these auxin-determined decisions lead to changes in phyllotaxy, node elongation and inflorescence architecture, while fate decisions such as the transition to flowering remain unaffected. Comparisons of the Trf phenotype with other known maize ARF mutants (reviewed in ref. 43) highlight potential diversification of ARF function in organ-specific developmental programmes in different plant lineages.
It is remarkable that, despite a lack of sequence-level conservation, the same short region within the DBD of class-B ARFs has an essential role in determining protein level across land plants. Assuming that this region interacts with a second protein, perhaps a ubiquitin protein ligase, it is possible that the degradation-regulating region and interacting protein have co-evolved during the ~500 Myr that separate maize and P. patens, explaining differences in sequence and response, and the inability of the maize regulatory region to be recognized in moss. This further emphasizes that, similar to class-B ARFs regulation by the TAS3 trans-acting small interfering RNA (tasiRNA), some aspects of ARF regulation are ancient, although the outcomes can be different: in flowering plants, TAS3 regulation of ARF3 and ARF4 is required for adaxial–abaxial patterning, while in P. patens the pathway regulates protonemal development46,53,54. These differences further highlight lineage-specific changes to auxin-regulated cell fate decisions.
Our work underscores the importance of protein-level regulation of ARF levels in tuning auxin response. Previous work found that proteasome regulation of class-B AtARF1 and AtARF2 stability in Arabidopsis, is dependent on AtHOOKLESS1 activity55. More recently, the F-box protein AtAFF1 was shown to regulate the levels and behaviour of the class-A ARFs, AtARF7 and AtARF19 (ref. 56). Proteasomal degradation of MpARF1 and MpARF2 in Marchantia polymorpha has also recently been reported57,58. In addition, AtARF2 has recently been identified by proximity labelling with AtCUL1, part of the E3 Ligase SKP1–Cullin–F-box complex59, further supporting our findings here. Additional instances of proteasome regulation of ARFs, potentially conserved across evolution, as in the case of the Trf and nar mutants, may exist. It will be interesting to see whether similar non-sequence conserved regulatory mechanisms are common in the ARF lineages.
The TIR1/AFB–Aux/IAA pathway is subject to complex regulation consistent with its essential role in diverse processes. These parallel regulatory pathways enhance auxin response robustness, and modulation probably facilitated the expansion and fine-tuning of auxin-mediated processes across land plant lineages. Comparative analyses between species, such as the work presented here, is key to the discovery of the auxin puzzle pieces and how they fit together.
Methods
Maize plants
The Truffula seed was provided by Gerry Neuffer, originally called Ts-2620 and found in an EMS-mutagenized Mo17 population. The phenotype of Trf was analysed in backcrosses into W22 and Mo17 advanced to generation 10. Field-grown plants were grown at UC Berkeley Gill Tract, UC Davis and UMass Amherst during summer field seasons. All plants used for molecular analysis were backcrossed into W22 10 times. Seedlings grown for RNA and DNA extractions were grown under greenhouse conditions in 30 × 45 × 2.5 cm trays with peat-free soils plus osmocote. Seedlings grown for auxin treatment tests and western blots were grown in temperature-controlled greenhouses with supplemental light in 30 × 45 × 2.5 cm trays with peat-reduced osmocote compost with added fertilizer under 16 h 25 °C/8 h 21 °C day/night cycle.
DNA extraction
Leaf discs were collected from 80 Trf and N siblings in the W22 backcross population (at least 10 introgressions). High-quality gDNA was extracted using urea as described previously60. In brief, genomic DNA was extracted using a urea-based extraction method which is as follows. Tissue was ground in urea extraction buffer (4 M urea, 4 M NaCl, 1 M Tris pH 8, 0.5 M EDTA, 34 mM n-lauroyl sarcosine). Then an equal volume of phenol:chloroform:isoamyl alcohol (25:24:1), was added and vortexed to mix. After centrifugation, the supernatant was decanted and mixed with an equal volume of chloroform. After another centrifugation, the supernatant was decanted and mixed with 1/10th volume of 4.4 M NH4OAc pH 5.2 and 0.7 volume of isopropanol. Strands of DNA were then collected and washed with 70% ethanol in a separate tube. The dried pellet was resuspended in 100 µl of Tris-EDTA buffer (TE). DNA integrity was checked using gel electrophoresis and quantified using Qubit before sending for library preparation and 150-bp paired-end sequencing by Novogene. Sequencing was for 10× coverage of the maize genome. DNA for SSR mapping and genotyping was extracted from 1.5 cm of leaf tissue using the protocol described previously61.
Whole-genome sequencing bulk-segregant analysis
Sequence data were downloaded via ftp from Novogene servers and passed through the following analysis pipeline: FASTPQ62 > Bowtie2 (ref. 63) alignment to the B73_Reference_Nam5.0 genome > samtools mpileup variant calling64 > SNPeff42 and variant filtering in R, restricting to EMS-type SNPs. Jupyter notebooks containing the code for WGS-BSA analysis is available on GitHub at https://github.com/ThePlantShapeLab.
SSR mapping and genotyping maize plants
SSR markers were identified using the Maize GDB database and used to genotype Trf phenotype plants from successive generations (see Fig. 2 for position, and Supplementary Table 1 for primers). Custom SSR markers were identified and designed to refine the region further (Supplementary Table 1). Genotyping PCRs used AD_4 and AD_5. All PCRs were carried out in 20 µl reactions using GoTaq Green Master Mix (Promega), following manufacturer instructions. PCR reactions were carried out on a Bio-Rad T1000 thermocycler using the following conditions: 98 °C for 5 min, then 35 cycles of 98 °C for 2 min, 58 °C for 30 s, 72 °C for 30 s, then 72 °C for 10 min. The PCR product (10 µl) was analysed using thin metaphor agarose (3.5% in 1× TBE, 1:10,000 SYBR safe, 90 V, 1 h) electrophoresis and imaged on a BioRad GelDoc GO imaging system.
Auxin treatment of maize
A segregating population of 5-week-old Trf siblings was genotyped using primers AD_4 and AD_5. Plants were cut at the root–shoot node and submerged in either 100 µm indole-3-acetic acid (Sigma, I5148) in 1% ethanol, or 1% ethanol only, ensuring that at least 10 cm of the shoot base was completely submerged. Shoot apices were harvested (~1 cm encompassing the base of the meristem and young leaf tissue, outer leaf removed), dried briefly and flash frozen in liquid nitrogen at 0 min and 30 min. Two replicate pools for each treatment and genotype group were collected, containing 3 individuals in each pool.
Maize RNA extraction
For the Trf versus normal (N) sibling RNA-seq experiment, a segregating population of 3-week-old maize seedlings were first genotyped for Trf (see Supplementary Table 1 for primers). The outer 2 leaves were removed and 0.5 cm of the shoot apex was dissected and flash frozen in liquid nitrogen. Three replicate pools were created for Trf and N siblings, with 10 individuals per pool. For the auxin treatment of Trf versus N siblings, tissues were treated as described above. RNA for all experiments was extracted from each pool using TRIzol (Invitrogen) as described in the manual. RNA was DNAse treated to remove gDNA contamination using Invitrogen DNaseI (18068015) digestion. RNA quality was checked using gel electrophoresis (1× Agarose in 1× TBE) and quantified using Qubit before sending for library preparation and 150-bp paired-end sequencing by Novogene, or generating cDNA libraries using the iScript gDNA clear cDNA synthesis kit (Bio-Rad).
RNA-seq analysis
Sequence data were downloaded via ftp from Novogene servers and passed through the following analysis pipeline: FASTPQ62 > HiSAT2 (ref. 65) alignment to the Zea mays Reference NAM5.0 genome > FeatureCounts66 > DeSeq2 (ref. 67) analysis of differential gene expression using genes with more than 5 counts in 3 or more samples. All analysis was carried out locally (3.6 GHz 8-Core Intel Core i9 processor, 32 GB RAM) using R packages combined with Jupyter notebooks. Jupyter notebooks containing the code for RNA-seq analysis are available on GitHub at https://github.com/ThePlantShapeLab. GO term analysis was carried out using the GAMER Maize annotations68 and the GOSeq R package69; significantly enriched GO terms had a Benjamini and Hochberg cut-off Padj value of <0.05. Comparisons of significantly differentially expressed genes (Padj < 0.05, log2FC < −0.5 or >0.5) with potential class-B ARF regulatory targets were carried out using the DAP-seq data published in ref. 24 aligned to the maize B73 NAM5.0 genome (provided by Andrea Gallavotti and Mary Galli, Rutgers University). DAP-seq peak data for three class-B ARFs in the same subclade as ZmARF28: ZmARF13 Zm00001eb170540, ZmARF10 Zm00001eb142540 and ZmARF25 Zm00001eb363810 were used (ZmARF28 does not have any published DAP-seq data). DAP-seq target genes were filtered and considered ‘high-confidence’ targets if the DAP-seq peak was within 1 kb of the transcriptional start site, generating a list of 4,654 class-B target genes, of which 61% are shared.
cDNA synthesis and qPCR analysis
cDNA was synthesized from 750 ng RNA using the iScript gDNA clear cDNA synthesis kit (BioRad) according to the manufacturer’s manual. cDNA was then diluted 1:10 for subsequent analysis. Primers for qPCR were designed spanning intron/exon boundaries for auxin-responsive gene ZmSAUR27 (Zm00001eb104110, primers AER639:CTGGAGGGTGGAAAGGGAAG, AER640:AGCATCAAAAGGCTCCATGGA) and the GAPDH housekeeping gene (Zm00001eb173410, primers AER647:CATCATTCCTAGCAGCACCG, AER648:TACACAAGCAGCAACCATCC) using Benchling and checked for specificity using the Maize GDB BLAST tool (https://www.maizegdb.org/). Primers were then tested using standard RT–PCR conditions (1 µl 1:10 cDNA, 2 µl of each 5 µM primer, 10 µl GoTaq Green Master Mix, 5 µl nuclease-free H2O, run on BioRad ThermoCycler T2100 30 cycles of 95 °C/58 °C/72 °C) and evaluated using gel electrophoresis (1% agarose in TBE, 90 V, 40 min). All primers generated single, specific bands at the expected size (~500 bp). To confirm that identity bands were excised, the DNA was extracted using Qiagen Gel Extraction kit and sent for Sanger sequencing. qPCR was performed with the optimized primers using PowerUp SYBR Green Master Mix (Applied Biosystems) according to manufacturer instructions, and the Applied Biosystems StepOnePlus Real-Time PCR System. Optimum conditions were determined as follows: first, primer specificity was tested by a melting curve analysis, yielding a single peak per primer pair. Next, serial dilutions of each sample were used to determine the optimum cDNA concentration, which was found to be the same 1:10 dilution as above, with a 20 µl final volume for each reaction. Reactions were carried out as follows: a hold step (2 min, 50 °C) and initial denaturation (2 min, 95 °C); 40 cycles of 95 °C (15 s) followed by 60 °C (1 min); ending with a melt curve (60 to 95 °C). The \({{2}{^{-{\Delta\Delta}{C}_{T}}}}\) method was used to calculate relative quantification70, with values normalized to the average of the mock treatment for the WT pools.
Antibody production
Antibody was developed from the ZmARF28 full-length coding sequence cloned into pDEST17 bacterial destination vector carrying the 6His tag for antigen production. Recombinant protein expression was carried out in E. coli Rosetta strain (BL21). After purification, a total of 200 µg of protein was sent to Cocalico Biologicals where two guinea pig immunizations were performed. To improve specificity of this antibody, purification from antiserum was carried out using a synthetic 20-amino-acid peptide from position 505 to 524 which is a highly specific region to ZmARF28 (amino acids 505–524: RPFPNKISGTRSSTWVTADA) using magnetic beads from Invitrogen. A cysteine (C) residue was added at the N-terminal end of the peptide to enable covalent binding to the beads used for antibody purification. To validate specificity of the purified antibody, a western blot was carried out following the described protocol in the next section, using 20 mg of soluble total protein from vegetative B73 maize shoot apical meristem.
ZmARF28 protein extraction from maize and western blots
Maize seedling (3-week-old) shoot apices were harvested and flash frozen in liquid nitrogen before grinding to a fine powder. Powdered tissue (100 mg) was then thawed in 1× SDS + dithiothreitol loading buffer and vortexed. Samples were heated at 95 °C for 5 min before loading on a 10% acrylamide gel for electrophoresis. Semi-dry transfer to a methanol-activated PVDF membrane in 1× Tris-glycine/20% methanol was carried out. Membranes were air dried, reactivated in methanol, washed in TBS/0.05% Tween for 5 min and then blocked for 1 h with 6% skimmed milk in TBS/0.05% Tween at room temperature. Membranes were incubated overnight in the primary anti-ARF28 antibody diluted 1:250 in 1% milk/TBS/0.05% Tween. The following day, membranes were washed (3× 5 min in TBS/0.05% Tween) and incubated with 1:5,000 anti-guinea pig-HRP secondary antibody (Thermo Fisher, A18769) at room temperature with rocking for 2 h. After washing (4× 5 min in TBS/0.05% Tween), membranes were incubated with Clarity Western ECL substrate (Bio-rad, 1705060) and exposed to X-ray film. Film was exposed to the membrane for 10 min before developing. This experiment was repeated four times with similar results.
Expression of ZmARF28-GFP in N. benthamiana and western blots
The CDS of ZmARF28 and ZmARF28-Trf was cloned into pEARLYGATE103 using gateway cloning. Sequence was confirmed using plasmid nanopore sequencing (Source Biosciences). The ZmARF28-pEARLYGATE103 constructs were transformed into Agrobacterium strain AGL0, and successful transformants were confirmed using colony PCR. Glycerol stocks of colonies were created and used for all subsequent experiments. Overnight cultures of Agrobacterium strains containing ZmARF28-pEARLYGATE103, ZmARF28-Trf-pEARLYGATE103 or pEARLYGATE103 empty vector were grown in Luria–Bertani (LB) media with 10 µg ml−1 rifampicin and 50 µg ml−1 kanamycin at 28 °C with 200 r.p.m. shaking. The next day, the optical density at 600 nm (OD600) of the cultures was measured and the cultures diluted in infiltration buffer (10 mM MES, 10 mM MgCl2, 0.2 mM acetosyringone) to OD600 = 0.3 for pEARLYGATE103 constructs. Leaves of N. benthamiana plants at 2 weeks post transplantation to soil were infiltrated with a pEARLYGATE103 strain and then left to grow for a further 2 days (16 h 22 °C/8 h 22 °C day/night cycle). Infiltrated leaves were then infiltrated with 5 µl 100 µM MG132 in DMSO, or DMSO only, diluted in 1 ml infiltration buffer and incubated for 4 h before harvest and snap freezing in liquid nitrogen. Leaves were ground to a very fine powder in liquid nitrogen before adding extraction buffer (50 mM NaCl, 1% Igepal CA-630, 0.5% sodium deoxycholate, 0.1% SDS, 50 mM Tris-HCl (pH 8.0), 1 mM EDTA, 50 mM NEM, 1× cOmplete EDTA-free protease inhibitor cocktail (Roche, 11836170001), 1× PhosSTOP phosphatase inhibitor (Roche, PHOSS-RO)) and continuing to grind until defrosted. Samples were transferred to Eppendorf tubes and centrifuged at 6,500 × g at 4 °C for 12 min. The supernatant was transferred to a new Eppendorf tube and centrifuged at 9,800 × g at 4 °C for 3 min. Protein was quantified using a Qubit Broad Range Protein kit and the Qubit fluorometer 4 (Invitrogen). Total protein extract was diluted to 0.5 µg ml−1 in extraction buffer. A volume of 10 µl of extract was mixed with 4× SDS + dithiothreitol (5× buffer stock: 0.25% bromophenol blue, 0.5 M dithiothreitol, 50% glycerol, 10% sodium dodecyl sulfate, 0.25 M Tris-HCl pH6.8) loading buffer and heated at 95 °C for 5 min. Sample (20 µl) was loaded onto a 12% SDS–PAGE gel and ran in 1× Tris-glycine running buffer with 0.1% SDS at 70 V for 30 min, then 100 V for 90 min at room temperature. Proteins were transferred overnight at 4 °C using the Bio-Rad wet-transfer mini-blot system to a PVDF membrane. Membranes were washed in dH2O and dried before reactivating in methanol for 1 min and washing in dH2O. Membranes were then stained for 5 min in Ponceau stain, and blots were imaged after 3 rinses in dH2O. Ponceau stain was washed off the blot using 1× PBS-T (1× PBS, 0.05% Tween-20). Membranes were blocked at room temperature for 2 h in 5% milk/PBS-T with shaking, then incubated with 1:3,000 mouse anti-GFP (Roche, 11814460001) for 90 min, washed 3 times in 1× PBS-T, then incubated with 1:3,000 anti-mouse-HRP (Cell Signaling Technology, 7076) for 1 h at room temperature. Blots were washed three times in 1× PBS-T before incubating with SuperSignal West Pico PLUS Chemiluminescent Substrate (Thermo Scientific, 34577) for 1 min according to manufacturer instructions and imaged on an Azure 300 Chemiluminescent imager. Experiments were repeated 3 times.
Expression of ZmARF28-GFP in maize protoplasts
Protoplasts were extracted as follows. A 3–4-week-old seedling was harvested and the outer unexpanded leaves removed before being cut into 2-cm sections and surface sterilized in 0.5% bleach/1% Tween-20 for 3 min. Tissue was then rinsed in sterile water four times and patted dry using kimwipes. Tissue was finely sliced (<1 mm) in a sterile glass Petri dish and 10 ml of sterile enzyme solution added (10 mM KCl, 8 mM MES, 1 mM CaCl2, 0.5 M mannitol, 0.6% cellulase RS, 0.1% macroenzyme R-10, 0.1% bovine serum albumin, 0.1% polyvinylpyrrolidone K30) before incubating on an orbital shaker (20 mm orbit, 40 r.p.m.) for 2 h at room temperature in the dark. A volume of 10 ml sterile W5 solution (154 mM NaCl, 125 mM CaCl2, 5 mM KCl, 2 mM MES) was then added and incubated for 1 h at room temperature on an orbital shaker (20 mm orbit, 80 r.p.m.) in the dark. This was then filtered through a 70-µm cell strainer before centrifugation at 100 × g for 5 min. Protoplasts were resuspended in 1 ml suspension solution (0.4 M mannitol, 20 mM CalCl2, 5 mM MES) and viability checked using fluorescein diacetate (FDA) staining and confocal microscopy. Protoplasts (200 µl) at a concentration of 400,000 per ml were then incubated with 20 µg of plasmid DNA (full-length ZmARF28 or ZmARF28-Trf in pEARLYGATE103) for 5 min at room temperature. A volume of 220 µl PEG solution (40% PEG 4000, 0.1 M CalCl2, 0,4 M mannitol pH 5.7, 50 mM MES) was added and mixed by inversion before incubation at 28 °C for 15 min. PEG was diluted with 800 µl of W5 solution, protoplasts were pelleted (100 × g, 3 min), washed with 500 µl W5 solution and then resuspended in 300 µl incubation buffer (0.5 M mannitol, 4 mM KCl, 4 mM MES, 1% BSA, 50 µg ml−1 kanamycin) and incubated overnight at room temperature in the dark on an orbital shaker (20 mm orbit, 20 r.p.m.). Protoplasts were checked for GFP expression using confocal microscopy before proceeding with MG132 treatment (20 µM MG132 or DMSO mock treatment) and incubated for a further 3 h. Once complete, protoplasts were pelleted and washed with W5 solution before proceeding with protein extraction and western blots as described above for the N. benthamiana protein expression. Experiments were repeated 3 times.
Image analysis and figure assembly
All images were analysed in FIJI71. Figures were assembled using Adobe Photoshop.
Statistical analyses
All data were tested for normality using the Shapiro–Wilk test. On the basis of this analysis, data with a normal distribution were analysed using a Student’s t-test or analysis of variance (ANOVA) where appropriate. Data that were not normally distributed were analysed using the non-parametric Kruskal–Wallis test, followed by pairwise Wilcox tests. A Jupyter notebook containing the code for data analysis is available on GitHub at https://github.com/ThePlantShapeLab.
Moss growth
Moss was grown on BCD minimal medium or BCD supplemented with either 5 mM ammonium tartrate (BCDAT) or 5 µM NAA (BCD + NAA)72. BCDAT was used for routine growth. Ammonium directly or indirectly dampens auxin responses, which enhances the phenotypes of mutants with reduced auxin response. For growth assays, ~1-mm2 pieces of tissue from chloronema-rich cultures of each genotype were spotted in 4–6 different positions on the plates containing all three media, with each genotype represented by at least two independent lines. The nar mutants were isolated in the DR5:DsRed/NLS-4 background in the Gransden ecotype13, and all subsequent analyses were carried out in either the Reute ecotype or a strain with the DR5:DsRed transgene introgressed from the semi-fertile Gransden strain into Reute through three crosses. During the course of this work, it was discovered that the DR5:DsRed expression is often silenced upon retransformation, possibly due to the locus containing approximately 60 repeats of the transgene45.
Moss mutant screen
The screen for nar mutants and mutant genome sequencing was described previously39. Briefly, 30 plates of week-old lawns of DR5:DsRed/NLS-4 on cellophane-overlain BCDAT media were irradiated with 150 mJ cm−2 of UV light in a Stratalinker UV Crosslinker (Stratagene). After 24 h in darkness, the cellophanes were moved to new BCD + NAA plates, and the mutants were identified after 2–4 weeks. Confirmed mutants were screened for mutations in Aux/IAA and DIAGEOTROPICA genes by sequencing PCR products, and genomes of 12 of the strongest remaining mutants were sequenced. The full set of variants in the mutant genomes were filtered for those causing protein-altering changes in the same gene in two or more of the mutants. While both narC29 and narC18 had the E263K substitutions in ARFb4, narC18 also had a G608S substitution in class D ARF, ARFd1, which probably accounts for the stronger phenotype. For simplicity, only narC29 is shown.
CRISPR/Cas9
Plasmids for expressing one or two single guide RNAs (sgRNAs) and Cas9 were assembled either as described in ref. 73 or after modification to bypass the Gateway cloning step: the empty pENTR-L1L2-U6:BsaI insert was recombined into the pMH-Cas9-Gateway vector with LR Clonase II (Invitrogen), then the resulting plasmid backbone was replaced with that of pUK21 which lacks BsaI restriction sites and thus allows GoldenGate assembly of protospacers between the BsaI sites after the PpU6 promoter. A fragment of GFP was inserted between the BsaI sites to help distinguish recombinant clones from the parental vector. The hph selection gene was swapped for nptII, aacC1 or ble to allow selection with kanamycin/G418, gentamicin/G418 and Zeocin, respectively, in addition to Hyg. Protospacers were selected on the basis of predicted specificity and efficiency using the CRISPOR website (http://crispor.gi.ucsc.edu/)74. sgRNAs starting with G were preferred. Plasmids for making deletions contained dual PpU6:sgRNA genes targeting shortly after the start codons and before the stop codons. Oligo-mediated gene editing was achieved by designing double-stranded oligos matching the target but with the intended missense mutation(s) and silent mutations to create a restriction site and multiple mismatches in the PAM-proximal half of the sgRNA target. The oligos extended 21 bases before and after the mismatched bases.
Moss transformations
Protoplasts were prepared from week-old tissue and transformed as previously described72, except that the post-heat-shock dilution was done by slowly adding 10 ml 8.5% mannitol all at once rather than 6.5 ml in 10 steps. Transformations included 15–25 µg of each plasmid and 250 pmol of annealed oligos. Deletion lines were identified by dramatically truncated PCR products and Sanger sequencing. Genome-edited lines were identified by PCR across the edited site, followed by digestion with the restriction enzyme whose site was added to the template oligos. PCR products with the expected digestion pattern were sequenced. In several instances, lines exhibiting the same mutant phenotype but lacking the restriction site change or lacking the phenotype but having the restriction site change were found to have incorporated either the missense mutation or the restriction site, respectively, without the other. In the first editing experiment to re-create the E266K and R269Q mutations in PpARFb2, it was discovered that the proDR5:DsRed transgene’s gentamicin selection cassette also confers resistance to the G418 selection agent and this precluded selection of the CRISPR plasmid. We instead screened the transformants for resistance to NAA. Some of the NAA-resistant lines had the intended mutation but many had small in-frame insertions and deletions in the region.
YFP-tagging of endogenous loci was achieved by co-transforming a CRISPR plasmid targeting near the stop codon and a plasmid that included: 600–1,500 bp homology arm of the coding region preceding the stop codon fused in-frame to a linker (SRGGGGA) and YFP (encoding a bright, monomeric yellow fluorescent protein), followed by the Pisum rbcS terminator, a pro35S:Hyg:ter35S selection cassette flanked by loxP sites and a downstream homology arm consisting of the 3′ UTR and downstream sequences. Stable lines were identified by selecting for the CRISPR plasmid and the knock-in construct for 7 days, followed by 10–14 days of no selection and 7 days of Hyg selection. Clean insertion lines were identified by (1) PCR amplification with primers matching before the upstream homology arm and downstream from the downstream homology arm with primers to YFP and ter35S, and (2) lack of PCR amplification both across the insertion site and of the Cas9 gene. In the case of the pparfb1E266K and pparfb2E266K, the strength of the phenotypes increased upon integrating the YFP/Hyg sequences. Removal of the pro35S:Hyg:ter35S selection cassette by CRE/lox-mediated excision reverted the phenotype to that before insertion, suggesting that pro35S enhancers may act at a distance on the nearby promoters in moss as in Arabidopsis75. All YFP lines shown had their selection cassettes excised.
The proARFb2:DBD-YFP constructs with and without the E266K + R269Q substitutions and the overexpression constructs were constructed by GoldenGate assembly. The DBD-YFP assembly included plasmids containing mYPet and 3 kb of sequence upstream from ARFb2’s start codon, PCR products containing the DBD sequences before and after the loop sequence, double-stranded oligos with the wild-type or mutant loop sequences and the pMP2133 destination vector. The pMP2133 vector contained PIG1-locus-homology arms for targeting, AtHsp18.2 terminator sequences and a pro35S:nptII:ter35S selection cassette. The overexpression construct assembly included the pMP2280 destination vector, plasmids containing the maize Ubiquitin promoter, the PpARFb2 or ZmARF28 cDNAs (without stop codons), and a double-stranded oligo with a C-terminal 1× Myc epitope and a stop codon. The pMP2280 vector contained Pp108 homology arms flanking the cloning site, a nos terminator and a pro35S:Hyg:ter35S selection cassette. The constructs were co-transformed into Reute WT along with a CRISPR plasmid targeting the corresponding native neutral targeting locus. Transformants were selected for 7 days with hygromycin and G418, followed by 10 days off selection. Stable lines were identified by PCR after screening lines resistant to G418 and sensitive to hygromycin (DBD lines), or resistant to hygromycin (overexpression lines).
Moss RT–PCR
Week-old pparfb2E266K, ppiaa2G325S and the corresponding wild-type (DR5:DsRed/3×Re) tissues were homogenized, plated on BCDAT plates overlain with cellophanes and grown for 6 days. At 18 h before treatment, the cultures on cellophanes were moved to BCD plates to alleviate the antagonism on auxin response by ammonium. Approximately 100 mg of tissue were scraped from the cellophane and transferred in triplicate to wells of 6-well plates with BCD-agar media containing either 10 µM IAA or 0.01% ethanol with 1 ml liquid BCD media, with the same concentrations of IAA and ethanol. After 1 h, tissue was patted dry between paper towels, and RNA was extracted using the Quick-RNA Plant Miniprep kit (Zymo Research) and a Mini-Beadbeater-16 (BioSpec Products) according to manufacturer instructions. Total RNA (1.3 µg) was reverse transcribed using the Maxima H Minus cDNA Synthesis Master Mix (Thermo Fisher), diluted 8-fold with 0.1× TE, and 3 µl used per 20 µl SYBR-qPCR reaction using the CFX Connect-96 Real-Time PCR Detection System (Bio-Rad). Data represent two technical replicates for each of the three biological replicates for each genotype and treatment.
Yeast two-hybrid assay
Yeast interaction assays were carried out using the Matchmaker LexA Two-Hybrid System (Clontech). Interactions were indicated by β-galactosidase detection on SD/Gal/Raf/X-Gal plates. For testing BRD function, a C-terminally Myc-tagged full-length TPL2/Pp6c9_10690 cDNA was inserted into pGilda (LexA DBD), and full-length ARFb2 cDNA with and without the L614S and F615S substitutions were inserted into pB42AD. No dimerization was detected between DBD/DD domains using pGilda+ARFb2[aa1–375] and pB42AD + ARFb2[full-length].
Moss imaging
For growth assays, 3-week-old moss were photographed with a Nikon SMZ1500 and a DS-Ri1 camera. DsRed fluorescence was captured using a TRITC filter set. Confocal microscopy was performed using a Zeiss LSM880. Tissue was mounted in water or liquid BCD medium, and images were captured for differential interference contrast (DIC), YFP fluorescence and chlorophyll autofluorescence. FIJI was used for processing images and for measuring DsRed and YFP signals71. DsRed signal was measured from oval selections of whole plants and regions of background and calculated as IntDen[sample]−(IntDen[background]×Area[sample]/Area[Background]). For YFP, nuclei were identified in the YFP or DIC channels, circled as freehand selections and their integrated density measured. Note that not all nuclei in wild type, unlike the mutants, have detectable YFP signal and thus could not be measured, hence their quantification is probably an overestimate and the difference between wild type and the mutants is an underestimate. The proARFb2:DBD-YFP lines were mounted in water and imaged with a Keyence BZ-X810 microscope with an EYFP filter cube and a ×4 objective to capture brightfield and YFP (2 s exposure time) images.
Detection of ubiquitylated protein
Tissue from untransformed Reute WT (Re), proARFb2:DBD-YFP (T215-5, DBDY) and proARFb2:dbd-YFP (E266KR269Q, T216-2, dbdY) strains were homogenized and split into 10 plates of BCDAT medium overlain with cellophane sheets and grown for 9 days. Moss tissues (200–250 mg) were added to wells of 6-well plates containing solid BDCAT + 100 µM bortezomib (LC Laboratories, B-1408) plus 1 ml liquid BCDAT + 200 µM bortezomib. After 18 h, tissues from 3 wells (Re and DBDY) or 1 well (dbdY) were patted dry between paper towels and frozen in liquid nitrogen in triplicate. Frozen tissue was ground to a powder while frozen, then mixed with 1,350 µl (Re, DBDY) or 450 µl (dbdY) chilled lysis buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1% Ipegal CA-630, 50 mM N-ethylmaleimide, 1.75× plant protease inhibitor cocktail (Millipore, P9599), 50 mM NaF, 10 mM β-glycerophosphate) and transferred to microfuge tubes in ice. After pelleting debris at 17,000 g for 10 min, ‘input’ samples were removed and 25 µl of thrice-washed GFP-Trap magnetic beads (Bulldog Bio, GMA020) were added to each tube and rotated for 2 h at 4 °C. The beads were gathered using a magnetic stand, and the liquid was removed and replaced with 1 ml wash buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 0.05% Ipegal CA-630). Three washes in total were performed, with the beads being transferred to clean tubes in the second wash. The third wash was completely removed and replaced with 70 µl of 2× SDS sample buffer, mixed and heated to 95 °C for 10 min before removing immunoprecipitated (IP) samples from the beads. Samples were separated using 7.5% SDS–PAGE gels transferred to nitrocellulose membranes using the Bio-Rad wet-transfer mini-blot system. Membranes were rinsed with TBST and blocked for 2 h in 5% milk, then incubated overnight at 4 °C with the primary antibodies in 5% BSA. GFP was detected using the rabbit polyclonal antibody (Invitrogen, A-11122, 1:2,000 dilution), and ubiquitin was detected with the P4D1 mouse monoclonal antibody (Cell Signaling Technology, 3936T, 1:1,000). Blots were washed 3× in TBST and incubated for 2 h in 5% milk with the goat-anti-rabbit (Millipore, A6154) or goat-anti-mouse (Bio-Rad, 1706516) peroxidase-conjugated secondary antibodies. After 3 washes, the blots were incubated with SuperSignal West Pico PLUS Chemiluminescent Substrate for 5 min and imaged using an ImageQuant LAS 4000 mini-imager. For the IP samples, equivalent amounts of Re and DBDY elution samples were loaded along with a 1:5 dilution of the dbdY samples to give equivalent amounts of YFP fusion protein. This ratio was estimated from preliminary blot of the IP samples.
Cloning ARF2 and arf2T98N constructs
DNA encoding 4 GS repeats, a porcine teschovirus-1 2A (P2A) cleavage peptide76 and mScarlet3 were synthesized using TWIST biosciences fragment synthesis. The P2A-mScarlet3 insert was amplified by PCR using Q5 2× master mix (NEB). The protoplast expression vector pLCS107 was linearized by PCR using Q5 2× master mix (NEB). The insert was cloned into linearized pLCS107 using NEBuilder Hifi cloning reactions to generate pUBQ-mNEON-P2A-mScarlet. The arf2T298N variant was introduced into the ARF2 CDS in pENTR by PCR amplification and infusion cloning (Takara BioSciences). Both wild type and T298N ARF2 were amplified from a pENTR backbone by PCR and cloned in-frame with mNEON and the GS linker using NEBuilder Hifi cloning. The pUBQ-mNEON-2A-mScarlet backbone was linearized by PCR.
Arabidopsis protoplast expression and quantification
Arabidopsis mesophyll protoplasts were isolated from 14-day-old Col-0 leaves. Cells (100,000) were transformed with 20–30 μg of plasmid DNA carrying mNEON-ARF2-P2A-mScarlet3 or mNEON-arf2T298N-P2A-mScarlet3 constructs using the tape-sandwich method and incubated for 16 h in the dark77. Transformed cell populations were scored using the Beckman Coulter Cytoflex S flow cytometer and CytExpert software. A back gating strategy was taken to identify the population of intact protoplasts. Cells expressing mScarlet3 reporters were first identified by comparing transformed mNEON-ARF2-P2A-mScarlet3 cells to untransformed cells using the Y610 channel (561 nm excitation, 610 ± 20 nm emission, 1,000 gain) and then back gated on FSCvSSC. Cells were treated with 100 µM cycloheximide diluted in protoplast media for the indicated times. A minimum of 162 mScarlet3 positive cells from 6 independent transformations were used to collect mScarlet3 reporter levels. The mNEON levels were collected using the B525 channel (488 nm excitation, 525 ± 40 nm emission, 69 gain). FCS files for mScarlet3 positive cell populations were generated by Cytoflex and analysed using FlowKit78, NumPy79 and Pandas80 (https://zenodo.org/record/7979740) packages in Python with custom scripts. Ratios were determined by taking the mNEON signal divided by the mScarlet3 signal on a per cell basis. The mean ratio of each replicate at timepoint 0 was generated and all cells from that replicate were normalized to this value. A minimum of 2,166 cells across all 6 replicates were collected. Graphs were generated using seaborn81 and Matplotlib82 in Python.
ARF DNA-binding domain structural predictions
Protein structure predictions of ARF DNA-binding domains as defined in ref. 21 from wild-type and mutant ZmARF28, PpARFb2 and PpARFb4 were generated using CollabFold v.1.5.2 (https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/AlphaFold2.ipynb)83 accessed on 8 August 2023 using default conditions. The top model from each prediction was visualized with PyMol 2.5 (http://www.pymol.org). AlphaFold pTM values are shown in Supplementary Table 10.
Phylogenetic tree construction
Protein sequences for class-B ARFs from 13 species were identified by reciprocal BLAST searches and aligned using T-COFFEE (v.11.00.8cbe486)84. Non-conserved regions were trimmed using Mesquite (v.3.70)85, and a tree was inferred using MrBayes (v.3.2.7)86 with the settings ‘aamodelpr=mixed, rates=invgamma, 2 runs of 4 chains for 1,000,000 generations’.
Databases used were for the following species: Alsophila spinulosa87, Amborella trichopoda88, Arabidopsis thaliana89, Ceratodon purpureus90, Ceratopteris richardii91, Diphasiastrum complanatum92, Marchantia polymorpha93, Physcomitrium patens94, Selaginella moellendorffii95, Solanum lycopersicum96, Takakia lepidozioides97, Thuja plicata98 and Zea mays99. Class-B ARF sequences were found neither in the three published genomes nor in the 1KP transcriptomes for hornwort species100,101,102.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All original numerical, sequence and image data are included in the manuscript or freely available via the University of Edinburgh DataShare service at https://doi.org/10.7488/ds/7903 (ref. 103); mutant and parental P. patens raw sequence reads are available at BioProject (PRJNA1210789); raw sequence reads for the Zea mays work is available for download at NCBI GEO accession numbers GSE293375, GSE293431, GSE293432. Seeds and plant material are available on request from A.E.R. (maize), M.E. (P. patens) and L.C.S. (Arabidopsis). Source data are provided with this paper.
Code availability
All code used for the maize sequence analyses is available on GitHub at https://github.com/ThePlantShapeLab/Truffula (ref. 104).
References
Matthes, M. S. et al. Auxin EvoDevo: conservation and diversification of genes regulating auxin biosynthesis, transport, and signaling. Mol. Plant 12, 298–320 (2019).
Thelander, M., Landberg, K. & Sundberg, E. Auxin-mediated developmental control in the moss Physcomitrella patens. J. Exp. Bot. 69, 277–290 (2018).
Mutte, S. K. et al. Origin and evolution of the nuclear auxin response system. Elife 7, e33399 (2018).
Kato, H. et al. Auxin-mediated transcriptional system with a minimal set of components is critical for morphogenesis through the life cycle in Marchantia polymorpha. PLoS Genet. 11, e1005084 (2015).
Powers, S. K. & Strader, L. C. Regulation of auxin transcriptional responses. Dev. Dyn. 249, 483–495 (2020).
Jing, H. & Strader, L. C. AUXIN RESPONSE FACTOR protein accumulation and function. BioEssays 45, 2300018 (2023).
Hernández-García, J. et al. Evolutionary origins and functional diversification of Auxin Response Factors. Nat. Commun. 15, 10909 (2024).
Ramos Báez, R. & Nemhauser, J. L. Expansion and innovation in auxin signaling: where do we grow from here? Development 148, dev187120 (2021).
Leyser, O. Auxin signaling. Plant Physiol. 176, 465–479 (2018).
Nemhauser, J. L. Back to basics: what is the function of an Aux/IAA in auxin response? New Phytol. 218, 1295–1297 (2018).
de Figueiredo, M. R. A. & Strader, L. C. Intrinsic and extrinsic regulators of Aux/IAA protein degradation dynamics. Trends Biochem. Sci. 47, 865–874 (2022).
Tao, S. & Estelle, M. Mutational studies of the Aux/IAA proteins in Physcomitrella reveal novel insights into their function. New Phytol. 218, 1534–1542 (2018).
Lavy, M. et al. Constitutive auxin response in Physcomitrella reveals complex interactions between Aux/IAA and ARF proteins. Elife 5, e13325 (2016).
Weijers, D. & Wagner, D. Transcriptional responses to the auxin hormone. Annu. Rev. Plant Biol. 67, 539–574 (2016).
Qi, L. et al. Adenylate cyclase activity of TIR1/AFB auxin receptors in plants. Nature 611, 133–138 (2022).
Cao, M. et al. TMK1-mediated auxin signalling regulates differential growth of the apical hook. Nature 568, 240–243 (2019).
Tiwari, S. B., Hagen, G. & Guilfoyle, T. The roles of auxin response factor domains in auxin-responsive transcription. Plant Cell 15, 533–543 (2003).
Ulmasov, T., Hagen, G. & Guilfoyle, T. J. Activation and repression of transcription by auxin-response factors. Proc. Natl Acad. Sci. USA 96, 5844–5849 (1999).
Cancé, C., Martin‐Arevalillo, R., Boubekeur, K. & Dumas, R. Auxin response factors are keys to the many auxin doors. New Phytol. 235, 402–419 (2022).
Smit, M. E. et al. Specification and regulation of vascular tissue identity in the Arabidopsis embryo. Development 147, dev186130 (2020).
Boer, D. R. et al. Structural basis for DNA binding specificity by the auxin-dependent ARF transcription factors. Cell 156, 577–589 (2014).
Franco-Zorrilla, J. M. et al. DNA-binding specificities of plant transcription factors and their potential to define target genes. Proc. Natl Acad. Sci. USA 111, 2367–2372 (2014).
O’Malley, R. C. et al. Cistrome and epicistrome features shape the regulatory DNA landscape. Cell 165, 1280–1292 (2016).
Galli, M. et al. The DNA binding landscape of the maize AUXIN RESPONSE FACTOR family. Nat. Commun. 9, 4526 (2018).
Stigliani, A. et al. Capturing auxin response factors syntax using DNA binding models. Mol. Plant 12, 822–832 (2019).
Powers, S. K. et al. Nucleo-cytoplasmic partitioning of ARF proteins controls auxin responses in Arabidopsis thaliana. Mol. Cell 76, 177–190 (2019).
Weijers, D. et al. Developmental specificity of auxin response by pairs of ARF and Aux/IAA transcriptional regulators. EMBO J. 24, 1874–1885 (2005).
Ellis, C. M. et al. AUXIN RESPONSE FACTOR1 and AUXIN RESPONSE FACTOR2 regulate senescence and floral organ abscission in Arabidopsis thaliana. Development 132, 4563–4574 (2005).
Sessions, A. et al. ETTIN patterns the Arabidopsis floral meristem and reproductive organs. Development 124, 4481–4491 (1997).
Pekker, I., Alvarez, J. P. & Eshed, Y. Auxin response factors mediate Arabidopsis organ asymmetry via modulation of KANADI activity. Plant Cell 17, 2899–2910 (2005).
Kato, H. et al. Design principles of a minimal auxin response system. Nat. Plants 6, 473–482 (2020).
Freire-Rios, A. et al. Architecture of DNA elements mediating ARF transcription factor binding and auxin-responsive gene expression in Arabidopsis. Proc. Natl Acad. Sci. USA 117, 24557–24566 (2020).
Rienstra, J., Hernández-García, J. & Weijers, D. To bind or not to bind: how Auxin Response Factors select their target genes. J. Exp. Bot. 74, 6922–6932 (2023).
Ikeda, M. & Ohme-Takagi, M. A novel group of transcriptional repressors in Arabidopsis. Plant Cell Physiol. 50, 970–975 (2009).
Piya, S., Shrestha, S. K., Binder, B., Stewart, C. N. Jr & Hewezi, T. Protein–protein interaction and gene co-expression maps of ARFs and Aux/IAAs in Arabidopsis. Front. Plant Sci. 5, 744 (2014).
Choi, H.-S., Seo, M. & Cho, H.-T. Two TPL-binding motifs of ARF2 are involved in repression of auxin responses. Front. Plant Sci. 9, 372 (2018).
Liu, X., Galli, M., Camehl, I. & Gallavotti, A. RAMOSA1 ENHANCER LOCUS2-mediated transcriptional repression regulates vegetative and reproductive architecture. Plant Physiol. 179, 348–363 (2019).
Kuhn, A. et al. Direct ETTIN–auxin interaction controls chromatin states in gynoecium development. Elife 9, e51787 (2020).
Morris, J. L. et al. The timescale of early land plant evolution. Proc. Natl Acad. Sci. USA 115, E2274–E2283 (2018).
Richardson, A. E. & Hake, S. The power of classic maize mutants: driving forward our fundamental understanding of plants. Plant Cell 34, 2505–2517 (2022).
Klein, H. et al. Bulked-segregant analysis coupled to whole genome sequencing (BSA-Seq) for rapid gene cloning in maize. G3 8, 3583–3592 (2018).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92 (2012).
Cowling, C. L., Dash, L. & Kelley, D. R. Roles of auxin pathways in maize biology. J. Exp. Bot. 74, 6989–6999 (2023).
Scanlon, M. J. The polar auxin transport inhibitor N-1-naphthylphthalamic acid disrupts leaf initiation, KNOX protein regulation, and formation of leaf margins in maize. Plant Physiol. 133, 597–605 (2003).
Bascom, C. Jr et al. Clade-D auxin response factors regulate auxin signaling and development in the moss Physcomitrium patens. PLoS Biol. 21, e3002163 (2023).
Plavskin, Y. et al. Ancient trans-acting siRNAs confer robustness and sensitivity onto the auxin response. Dev. Cell 36, 276–289 (2016).
Yi, P. & Goshima, G. Transient cotransformation of CRISPR/Cas9 and oligonucleotide templates enables efficient editing of target loci in Physcomitrella patens. Plant Biotechnol. J. 18, 599 (2020).
Causier, B., Ashworth, M., Guo, W. & Davies, B. The TOPLESS interactome: a framework for gene repression in Arabidopsis. Plant Physiol. 158, 423–438 (2012).
Nanao, M. H. et al. Structural basis for oligomerization of auxin transcriptional regulators. Nat. Commun. 5, 3617 (2014).
Beathard, C., Mooney, S., Al-Saharin, R., Goyer, A. & Hellmann, H. Characterization of Arabidopsis thaliana R2R3 S23 MYB transcription factors as novel targets of the ubiquitin proteasome-pathway and regulators of salt stress and abscisic acid response. Front. Plant Sci. 12, 629208 (2021).
Smalle, J. & Vierstra, R. D. The ubiquitin 26s proteasome proteolytic pathway. Annu. Rev. Plant Biol. 55, 555–590 (2004).
Salmon, J., Ramos, J. & Callis, J. Degradation of the auxin response factor ARF1. Plant J. 54, 118–128 (2008).
Garcia, D., Collier, S. A., Byrne, M. E. & Martienssen, R. A. Specification of leaf polarity in Arabidopsis via the trans-acting siRNA pathway. Curr. Biol. 16, 933–938 (2006).
Fahlgren, N. et al. Regulation of AUXIN RESPONSE FACTOR3 by TAS3 ta-siRNA affects developmental timing and patterning in Arabidopsis. Curr. Biol. 16, 939–944 (2006).
Li, H., Johnson, P., Stepanova, A., Alonso, J. M. & Ecker, J. R. Convergence of signaling pathways in the control of differential cell growth in Arabidopsis. Dev. Cell 7, 193–204 (2004).
Jing, H. et al. Regulation of AUXIN RESPONSE FACTOR condensation and nucleo-cytoplasmic partitioning. Nat. Commun. 13, 4015 (2022).
Das, S. et al. Quantitative imaging reveals the role of MpARF proteasomal degradation during gemma germination. Plant Commun. 5, 101039 (2024).
de Roij, M., Hernández García, J., Das, S., Borst, J. W. & Weijers, D. ARF degradation defines a deeply conserved step in auxin response. Preprint at. bioRxiv https://doi.org/10.1101/2024.09.12.612472 (2024).
Sun, F. et al. TurboID-based proteomic profiling reveals proxitome of ASK1 and CUL1 of the SCF ubiquitin ligase in plants. New Phytol. 244, 2127–2136 (2024).
Chen, J. & Dellaporta, S. in The Maize Handbook (eds Freeling, M. & Walbot, V.) 526–527 (Springer, 1994).
Lunde, C. Small-scale DNA extraction method for maize and other plants. Bio Protoc. https://doi.org/10.21769/BioProtoc.2782 (2018).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wimalanathan, K., Friedberg, I., Andorf, C. M. & Lawrence-Dill, C. J. Maize GO Annotation—Methods, Evaluation, and Review (maize-GAMER). Plant Direct 2, e00052 (2018).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, R14 (2010).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the \({{2}{^{-{\Delta\Delta}{C}_{T}}}}\) method. Methods 25, 402–408 (2001).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Cove, D. J. et al. The moss Physcomitrella patens: a novel model system for plant development and genomic studies. Cold Spring Harb. Protoc. 2009, pdb-emo115 (2009).
Mallett, D. R., Chang, M., Cheng, X. & Bezanilla, M. Efficient and modular CRISPR-Cas9 vector system for Physcomitrella patens. Plant Direct 3, e00168 (2019).
Concordet, J.-P. & Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 46, W242–W245 (2018).
Yoo, S. Y. et al. The 35S promoter used in a selectable marker gene of a plant transformation vector affects the expression of the transgene. Planta 221, 523–530 (2005).
Kim, J. H. et al. High cleavage efficiency of a 2A peptide derived from porcine teschovirus-1 in human cell lines, zebrafish and mice. PLoS One 6, e18556 (2011).
Wu, F.-H. et al. Tape-Arabidopsis Sandwich - a simpler Arabidopsis protoplast isolation method. Plant Methods 5, 16 (2009).
White, S. et al. FlowKit: a Python toolkit for integrated manual and automated cytometry analysis workflows. Front. Immunol. 12, 768541 (2021).
Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362 (2020).
Pandas Development Team. pandas-dev/pandas. GitHub https://github.com/pandas-dev/pandas (2023).
Waskom, M. L. Seaborn: statistical data visualization. J. Open Source Softw. 6, 3021 (2021).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Notredame, C., Higgins, D. G. & Heringa, J. T-Coffee: a novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 302, 205–217 (2000).
Maddison, W. P. & Maddison, D. R. M. Mesquite: a modular system for evolutionary analysis. v.3.70 http://www.mesquiteproject.org (2023).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Huang, X. et al. The flying spider-monkey tree fern genome provides insights into fern evolution and arborescence. Nat. Plants 8, 500–512 (2022).
DOE-JGI. Amborella trichopoda var. SantaCruz_75 HAP1 v2.1. http://phytozome-next.jgi.doe.gov/ (2022).
Cheng, C.-Y. et al. Araport11: a complete reannotation of the Arabidopsis thaliana reference genome. Plant J. 89, 789–804 (2017).
Carey, S., Yu, Q. & Harkess, A. The diversity of plant sex chromosomes highlighted through advances in genome sequencing. Genes 12, 381 (2021).
Marchant, D. B. et al. Dynamic genome evolution in a model fern. Nat. Plants 8, 1038–1051 (2022).
DOE-JGI. Diphasiastrum complanatum v3.1. http://phytozome-next.jgi.doe.gov/ (2022).
Bowman, J. L. et al. Insights into land plant evolution garnered from the Marchantia polymorpha genome. Cell 171, 287–304.e15 (2017).
Lang, D. et al. The Physcomitrella patens chromosome-scale assembly reveals moss genome structure and evolution. Plant J. 93, 515–533 (2018).
Banks, J. A. et al. The Selaginella genome identifies genetic changes associated with the evolution of vascular plants. Science 332, 960–963 (2011).
Zhou, Y. et al. Graph pangenome captures missing heritability and empowers tomato breeding. Nature 606, 527–534 (2022).
Hu, R. et al. Adaptive evolution of the enigmatic Takakia now facing climate change in Tibet. Cell 186, 3558–3576.e17 (2023).
Shalev, T. J. et al. The western redcedar genome reveals low genetic diversity in a self-compatible conifer. Genome Res. 32, 1952–1964 (2022).
Hufford, M. B. et al. De novo assembly, annotation, and comparative analysis of 26 diverse maize genomes. Science 373, 655–662 (2021).
Zhang, J. et al. The hornwort genome and early land plant evolution. Nat. Plants 6, 107–118 (2020).
Li, F.-W. et al. Anthoceros genomes illuminate the origin of land plants and the unique biology of hornworts. Nat. Plants 6, 259–272 (2020).
Leebens-Mack, J. H. et al. One thousand plant transcriptomes and the phylogenomics of green plants. Nature 574, 679–685 (2019).
Richardson, A. et al. Comparative mutant analyses reveal a novel mechanism of ARF regulation in land plants [dataset]. DataShare https://doi.org/10.7488/ds/7903 (2025).
ThePlantShapeLab. ThePlantShapeLab/Truffula: V1.0. Zenodo https://doi.org/10.5281/zenodo.15115846 (2025).
Acknowledgements
We thank G. Neuffer for the kind donation of the original Trf seed; S. Leiboff (Oregon State University), M. Bartlett (UMass Amherst), D. Weijers (Wageningen University) and S. Hetherington (University of Edinburgh) for discussions and comments; Z. Ban and M. Narasimhan (UCSD), Orosa and Spoel Labs (University of Edinburgh), and C. H. Hurst (University of Dundee) for advice on immunoprecipitation; A. Gallavotti and M. Galli (Rutgers University) for kindly sharing the class-B ARF DAP-seq peaks aligned to the B73 NAM5.0 genome; the ARS/USDA Plant Gene Expression Center plant growth support staff, the University of Edinburgh Plant Growth Facility, and CYVERSE (work supported by the National Science Foundation under Award Numbers DBI-0735191, DBI-1265383 and DBI-1743442; www.cyverse.org) for technical support and resources. Funding was provided by the University of Edinburgh Start-Up Fund (A.E.R.), UKRI EP/Y010116/1 (A.E.R.) and Royal Society IES\R2\212032 (A.E.R.); National Science Foundation IOS1922543 (S.H.), PGRP BIO-2112056 (L.C.S.); National Institutes of Health R01 GM043644 (M.E.), R35GM141892 (M.E.) and R35 GM136338 (L.C.S.); and the Ledell Family Research Scholarship for Science and Engineering (W.S.).
Author information
Authors and Affiliations
Contributions
A.E.R., L.C.S., M.E., S.H., N.M. and M.J.P. conceptualized the project. S.H., A.d.N., K.J. and A.E.R. cloned and phenotyped Trf; A.E.R. carried out the WGS, RNA-seq and auxin treatment analyses; N.M. carried out the protoplast stability experiments and the AlphaFold analysis; M.J.A.-J. purified the ZmARF28 antibody and carried out the maize western blots; A.E.R., H.J. and T.M. carried out plasmid cloning and the RT–qPCR analysis in maize; T.M. and A.E.R. carried out the MG132 test of ZmARF28 and associated western blots; W.S. carried out the nar mutant screen and the initial mutant characterization with help from M.J.P.; M.J.P. generated and characterized all moss deletion, genome-edited and reporter lines with support from W.S.; N.D. carried out the yeast two-hybrid experiments. M.L. provided reagents. A.E.R., N.M. and M.J.P. performed visualization. A.E.R., L.C.S., M.E. and S.H. acquired funding. A.E.R., L.C.S., M.E. and S.H. administered the project. A.E.R. and M.J.P. curated data. A.E.R., L.C.S., M.E. and S.H. supervised the project. A.E.R., L.C.S., M.E., S.H., N.M. and M.J.P. wrote the original draft. A.E.R., L.C.S., M.E., S.H., N.M. and M.J.P. reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
L.C.S. is on the scientific advisory board of Prose Foods. All other authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Yanhua Qi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–7, legends for Supplementary Tables 1–10 and unprocessed western blots for Supplementary Fig. 2I.
Supplementary Tables (download XLSX )
Supplementary Tables 1–10.
Source data
Source Data Fig. 4 (download PDF )
Unprocessed western blot images for Fig. 4c,e,g.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Prigge, M.J., Morffy, N., de Neve, A. et al. Comparative mutant analyses reveal a novel mechanism of ARF regulation in land plants. Nat. Plants 11, 821–835 (2025). https://doi.org/10.1038/s41477-025-01973-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-01973-3
This article is cited by
-
ARF degradation fine-tunes auxin response in land plants
Nature Plants (2025)