Abstract
Polyploidization is a common and important evolutionary process in the plant kingdom. Compared with diploid plant species, the intricate genome architecture of polyploid plant species presents substantial challenges in applying multi-omics approaches for crop breeding improvement. In this Review, we summarize the current techniques for analysing polyploid genomes, including constructing reference genomes and pan-genomes, and detecting variants. We also assess findings related to polyploid genome architecture, population genetics and breeding programmes, highlighting advanced techniques in the breeding of polyploid crops. Finally, we explore the challenges and demands posed by polyploid genome complexity during analysis with available biotechnological tools. This Review emphasizes the importance of a comprehensive understanding of polyploid genomic features for the further genetic improvement of polyploid crops.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Grant, V. Plant Speciation (Columbia Univ. Press, 1981).
Masterson, J. Stomatal size in fossil plants: evidence for polyploidy in majority of angiosperms. Science 264, 421–424 (1994).
Cui, L. et al. Widespread genome duplications throughout the history of flowering plants. Genome Res. 16, 738–749 (2006).
Dinesh Babu, K. S. et al. A short review on sugarcane: its domestication, molecular manipulations and future perspectives. Genet. Resour. Crop Evol. 69, 2623–2643 (2022).
Healey, A. L. et al. The complex polyploid genome architecture of sugarcane. Nature 628, 804–810 (2024).
The International Wheat Genome Sequencing Consortium (IWGSC). A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345, 1251788 (2014).
Stokstad, E. The new potato. Science 363, 574–577 (2019).
Sattler, M. C., Carvalho, C. R. & Clarindo, W. R. The polyploidy and its key role in plant breeding. Planta 243, 281–296 (2016).
Bird, K. A., VanBuren, R., Puzey, J. R. & Edger, P. P. The causes and consequences of subgenome dominance in hybrids and recent polyploids. N. Phytol. 220, 87–93 (2018).
Woodhouse, M. R. et al. Origin, inheritance, and gene regulatory consequences of genome dominance in polyploids. Proc. Natl Acad. Sci. USA 111, 5283–5288 (2014).
Lian, Q. et al. Acquisition of deleterious mutations during potato polyploidization. J. Integr. Plant Biol. 61, 7–11 (2019).
Conover, J. L. & Wendel, J. F. Deleterious mutations accumulate faster in allopolyploid than diploid cotton (Gossypium) and unequally between subgenomes. Mol. Biol. Evol. 39, msac024 (2022).
Stebbins, G. L. in Advances in Genetics Vol. 1 (ed. Demerec, M.) 403–429 (Elsevier, 1947).
Spoelhof, J. P., Soltis, P. S. & Soltis, D. E. Pure polyploidy: closing the gaps in autopolyploid research. J. Syst. Evol. 55, 340–352 (2017).
Song, Y. et al. Phased gap-free genome assembly of octoploid cultivated strawberry illustrates the genetic and epigenetic divergence among subgenomes. Hortic. Res. 11, uhad252 (2024).
Jin, X. et al. Haplotype-resolved genomes of wild octoploid progenitors illuminate genomic diversifications from wild relatives to cultivated strawberry. Nat. Plants 9, 1252–1266 (2023).
Chalhoub, B. et al. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345, 950–953 (2014).
Li, F. et al. Genome sequence of cultivated Upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat. Biotechnol. 33, 524–530 (2015).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Nurk, S. et al. HiCanu: accurate assembly of segmental duplications, satellites, and allelic variants from high-fidelity long reads. Genome Res. 30, 1291–1305 (2020).
Sun, H. et al. Chromosome-scale and haplotype-resolved genome assembly of a tetraploid potato cultivar. Nat. Genet. 54, 342–348 (2022).
Hoopes, G. et al. Phased, chromosome-scale genome assemblies of tetraploid potato reveal a complex genome, transcriptome, and predicted proteome landscape underpinning genetic diversity. Mol. Plant 15, 520–536 (2022).
Edger, P. P. et al. Origin and evolution of the octoploid strawberry genome. Nat. Genet. 51, 541–547 (2019).
Zheng, Y. et al. Allele-aware chromosome-scale assembly of the allopolyploid genome of hexaploid Ma bamboo (Dendrocalamus latiflorus Munro). J. Integr. Plant Biol. 64, 649–670 (2022).
Marcussen, T. et al. Ancient hybridizations among the ancestral genomes of bread wheat. Science 345, 1250092 (2014).
Zhang, T. et al. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat. Biotechnol. 33, 531–537 (2015).
Bertioli, D. J. et al. The genome sequence of segmental allotetraploid peanut Arachis hypogaea. Nat. Genet. 51, 877–884 (2019).
VanBuren, R. et al. Exceptional subgenome stability and functional divergence in the allotetraploid Ethiopian cereal teff. Nat. Commun. 11, 884 (2020).
Miao, J. et al. Chromosome-scale assembly and analysis of biomass crop Miscanthus lutarioriparius genome. Nat. Commun. 12, 2458 (2021).
Mitros, T. et al. Genome biology of the paleotetraploid perennial biomass crop Miscanthus. Nat. Commun. 11, 5442 (2020).
Session, A. M. & Rokhsar, D. S. Transposon signatures of allopolyploid genome evolution. Nat. Commun. 14, 3180 (2023).
Chen, H. et al. A chromosome-scale genome assembly of Artemisia argyi reveals unbiased subgenome evolution and key contributions of gene duplication to volatile terpenoid diversity. Plant Commun. 4, 100516 (2023).
Miao, Y. et al. Genome sequencing reveals chromosome fusion and extensive expansion of genes related to secondary metabolism in Artemisia argyi. Plant Biotechnol. J. 20, 1902–1915 (2022).
Song, A. et al. Analyses of a chromosome-scale genome assembly reveal the origin and evolution of cultivated chrysanthemum. Nat. Commun. 14, 2021 (2023).
Yang, J. et al. Haplotype-resolved sweet potato genome traces back its hexaploidization history. Nat. Plants 3, 696–703 (2017).
Zhang, X., Zhang, S., Zhao, Q., Ming, R. & Tang, H. Assembly of allele-aware, chromosomal-scale autopolyploid genomes based on Hi-C data. Nat. Plants 5, 833–845 (2019).
Zhang, J. et al. Allele-defined genome of the autopolyploid sugarcane Saccharum spontaneum L. Nat. Genet. 50, 1565–1573 (2018).
Chen, H. et al. Allele-aware chromosome-level genome assembly and efficient transgene-free genome editing for the autotetraploid cultivated alfalfa. Nat. Commun. 11, 2494 (2020).
Long, R. et al. Genome assembly of alfalfa cultivar Zhongmu-4 and identification of SNPs associated with agronomic traits. Genom. Proteom. Bioinform. 20, 14–28 (2022).
Wang, F. et al. The autotetraploid potato genome provides insights into highly heterozygous species. Plant Biotechnol. J. 20, 1996–2005 (2022).
Zhang, H. et al. The haplotype-resolved genome assembly of autotetraploid rhubarb Rheum officinale provides insights into its genome evolution and massive accumulation of anthraquinones. Plant Commun. 5, 100677 (2024).
Wang, S. et al. Haplotype-resolved chromosome-level genome of hexaploid Jerusalem artichoke provides insights into its origin, evolution, and inulin metabolism. Plant Commun. 5, 100767 (2024).
Zeng, X. et al. Chromosome-level scaffolding of haplotype-resolved assemblies using Hi-C data without reference genomes. Nat. Plants 10, 1184–1120 (2024).
Bao, Z. et al. Genome architecture and tetrasomic inheritance of autotetraploid potato. Mol. Plant 15, 1211–1226 (2022).
Serra Mari, R. et al. Haplotype-resolved assembly of a tetraploid potato genome using long reads and low-depth offspring data. Genome Biol. 25, 26 (2024).
Cheng, H., Asri, M., Lucas, J., Koren, S. & Li, H. Scalable telomere-to-telomere assembly for diploid and polyploid genomes with double graph. Nat. Methods 21, 967–970 (2024).
Bao, Y. et al. A chromosomal-scale genome assembly of modern cultivated hybrid sugarcane provides insights into origination and evolution. Nat. Commun. 15, 3041 (2024).
Deshpande, A. S. et al. Identifying synergistic high-order 3D chromatin conformations from genome-scale nanopore concatemer sequencing. Nat. Biotechnol. 40, 1488–1499 (2022).
Zhang, Z. et al. Haplotype-resolved genome assembly and resequencing provide insights into the origin and breeding of modern rose. Nat. Plants 10, 1659–1671 (2024).
Gabriel, L. et al. BRAKER3: fully automated genome annotation using RNA-seq and protein evidence with GeneMark-ETP, AUGUSTUS, and TSEBRA. Genome Res. 34, 769–777 (2024).
Zhang, R. et al. A high-resolution single-molecule sequencing-based Arabidopsis transcriptome using novel methods of Iso-seq analysis. Genome Biol. 23, 149 (2022).
Olson, N. D. et al. Variant calling and benchmarking in an era of complete human genome sequences. Nat. Rev. Genet. 24, 464–483 (2023).
Hook, P. W. & Timp, W. Beyond assembly: the increasing flexibility of single-molecule sequencing technology. Nat. Rev. Genet. 24, 627–641 (2023).
Wang, B. et al. Variant phasing and haplotypic expression from long-read sequencing in maize. Commun. Biol. 3, 78 (2020).
Wu, J., Mukhopadhyay, S. & Pérez-López Sr, E. Resistance gene enrichment sequencing refines the Brassica napus NLRome. Plant Physiol. 197, kiae631 (2025).
Holst, F. et al. Helixer—de novo prediction of primary eukaryotic gene models combining deep learning and a hidden Markov model. Preprint at bioRxiv https://doi.org/10.1101/2023.02.06.527280 (2023).
Gabriel, L., Becker, F., Hoff, K. J. & Stanke, M. Tiberius: end-to-end deep learning with an HMM for gene prediction. Bioinformatics 40, btae685 (2024).
Zhai, J. et al. Cross-species modeling of plant genomes at single nucleotide resolution using a pre-trained DNA language model. Proc. Natl Acad. Sci. USA 122, e2421738122 (2025).
Benegas, G., Batra, S. S. & Song, Y. S. DNA language models are powerful predictors of genome-wide variant effects. Proc. Natl Acad. Sci. USA 120, e2311219120 (2023).
Comai, L. The advantages and disadvantages of being polyploid. Nat. Rev. Genet. 6, 836–846 (2005).
Cheng, F. et al. Gene retention, fractionation and subgenome differences in polyploid plants. Nat. Plants 4, 258–268 (2018).
Yuan, J. et al. Dynamic and reversible DNA methylation changes induced by genome separation and merger of polyploid wheat. BMC Biol. 18, 171 (2020).
Ramírez-González, R. H. et al. The transcriptional landscape of polyploid wheat. Science 361, eaar6089 (2018).
Bourke, P. M., Voorrips, R. E., Visser, R. G. & Maliepaard, C. The double-reduction landscape in tetraploid potato as revealed by a high-density linkage map. Genetics 201, 853–863 (2015).
Julier, B. et al. Construction of two genetic linkage maps in cultivated tetraploid alfalfa (Medicago sativa) using microsatellite and AFLP markers. BMC Plant Biol. 3, 9 (2003).
Hurgobin, B. et al. Homoeologous exchange is a major cause of gene presence/absence variation in the amphidiploid Brassica napus. Plant Biotechnol. J. 16, 1265–1274 (2018).
Zhang, Z. et al. Homoeologous exchanges occur through intragenic recombination generating novel transcripts and proteins in wheat and other polyploids. Proc. Natl Acad. Sci. USA 117, 14561–14571 (2020).
Fan, C. et al. The KL system in wheat permits homoeologous crossing over between closely related chromosomes. Crop J. 11, 808–816 (2023).
Hu, G. et al. Evolutionary dynamics of chromatin structure and duplicate gene expression in diploid and allopolyploid cotton. Mol. Biol. Evol. 41, msae095 (2024).
Wu, Y. et al. Phylogenomic discovery of deleterious mutations facilitates hybrid potato breeding. Cell 186, 2313–2328.e15 (2023).
Herben, T., Suda, J. & Klimešová, J. Polyploid species rely on vegetative reproduction more than diploids: a re-examination of the old hypothesis. Ann. Bot. 120, 341–349 (2017).
Walkowiak, S. et al. Multiple wheat genomes reveal global variation in modern breeding. Nature 588, 277–283 (2020).
Li, J. et al. Convergence and divergence of diploid and tetraploid cotton genomes. Nat. Genet. 56, 2562–2573 (2024).
Hu, H. et al. Plant pangenomics, current practice and future direction. Agric. Commun. 2, 100039 (2024).
Schreiber, M., Jayakodi, M., Stein, N. & Mascher, M. Plant pangenomes for crop improvement, biodiversity and evolution. Nat. Rev. Genet. 25, 563–577 (2024).
Shi, J., Tian, Z., Lai, J. & Huang, X. Plant pan-genomics and its applications. Mol. Plant 16, 168–186 (2023).
Li, Y. et al. De novo assembly of soybean wild relatives for pan-genome analysis of diversity and agronomic traits. Nat. Biotechnol. 32, 1045–1052 (2014).
Zhao, Q. et al. Pan-genome analysis highlights the extent of genomic variation in cultivated and wild rice. Nat. Genet. 50, 278–284 (2018).
Hirsch, C. N. et al. Insights into the maize pan-genome and pan-transcriptome. Plant Cell 26, 121–135 (2014).
Jayakodi, M. et al. The barley pan-genome reveals the hidden legacy of mutation breeding. Nature 588, 284–289 (2020).
Jayakodi, M. et al. Structural variation in the pangenome of wild and domesticated barley. Nature 636, 654–662 (2024).
Chen, J. et al. Pangenome analysis reveals genomic variations associated with domestication traits in broomcorn millet. Nat. Genet. 55, 2243–2254 (2023).
Hickey, G. et al. Genotyping structural variants in pangenome graphs using the vg toolkit. Genome Biol. 21, 35 (2020).
MacNish, T. R. et al. Brassica Panache: a multi-species graph pangenome representing presence absence variation across forty-one Brassica genomes. Plant Genome 18, e20535 (2025).
Zhang, Y. et al. Structural variation reshapes population gene expression and trait variation in 2,105 Brassica napus accessions. Nat. Genet. 56, 2538–2550 (2024).
Sun, H. et al. The phased pan-genome of tetraploid European potato. Nature 642, 389–397 (2025).
Nieto Feliner, G., Casacuberta, J. & Wendel, J. F. Genomics of evolutionary novelty in hybrids and polyploids. Front. Genet. 11, 792 (2020).
Brown, M. R., Abbott, R. J. & Twyford, A. D. The emerging importance of cross‐ploidy hybridisation and introgression. Mol. Ecol. 33, e17315 (2024).
Douglas, G. M. et al. Hybrid origins and the earliest stages of diploidization in the highly successful recent polyploid Capsella bursa-pastoris. Proc. Natl Acad. Sci. USA 112, 2806–2811 (2015).
Wang, Z. et al. Dispersed emergence and protracted domestication of polyploid wheat uncovered by mosaic ancestral haploblock inference. Nat. Commun. 13, 3891 (2022).
Zhao, X. et al. Population genomics unravels the Holocene history of bread wheat and its relatives. Nat. Plants 9, 403–419 (2023).
Wang, T. et al. Interploidy introgression shaped adaptation during the origin and domestication history of Brassica napus. Mol. Biol. Evol. 40, msad199 (2023).
Hardigan, M. A. et al. Genome diversity of tuber-bearing Solanum uncovers complex evolutionary history and targets of domestication in the cultivated potato. Proc. Natl Acad. Sci. USA 114, E9999 (2017).
Shen, C. et al. The chromosome-level genome sequence of the autotetraploid alfalfa and resequencing of core germplasms provide genomic resources for alfalfa research. Mol. Plant 13, 1250–1261 (2020).
Monnahan, P. et al. Pervasive population genomic consequences of genome duplication in Arabidopsis arenosa. Nat. Ecol. Evol. 3, 457–468 (2019).
Zhou, Y. et al. Triticum population sequencing provides insights into wheat adaptation. Nat. Genet. 52, 1412–1422 (2020).
Kamal, N. et al. The mosaic oat genome gives insights into a uniquely healthy cereal crop. Nature 606, 113–119 (2022).
Lu, K. et al. Whole-genome resequencing reveals Brassica napus origin and genetic loci involved in its improvement. Nat. Commun. 10, 1154 (2019).
Wang, M. et al. Asymmetric subgenome selection and cis-regulatory divergence during cotton domestication. Nat. Genet. 49, 579–587 (2017).
Wang, S. et al. Characterization of polyploid wheat genomic diversity using a high‐density 90,000 single nucleotide polymorphism array. Plant Biotechnol. J. 12, 787–796 (2014).
Sun, C. et al. The Wheat 660K SNP array demonstrates great potential for marker‐assisted selection in polyploid wheat. Plant Biotechnol. J. 18, 1354–1360 (2020).
Winfield, M. O. et al. High‐density SNP genotyping array for hexaploid wheat and its secondary and tertiary gene pool. Plant Biotechnol. J. 14, 1195–1206 (2016).
Felcher, K. J. et al. Integration of two diploid potato linkage maps with the potato genome sequence. PLoS ONE 7, e36347 (2012).
You, Q. et al. Development of an Axiom Sugarcane100K SNP array for genetic map construction and QTL identification. Theor. Appl. Genet. 132, 2829–2845 (2019).
Meng, L., Li, H., Zhang, L. & Wang, J. QTL IciMapping: integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J. 3, 269–283 (2015).
Deb, S. K., Edger, P. P., Pires, J. C. & McKain, M. R. Patterns, mechanisms, and consequences of homoeologous exchange in allopolyploid angiosperms: a genomic and epigenomic perspective. N. Phytol. 238, 2284–2304 (2023).
Wu, Y. et al. Genomic mosaicism due to homoeologous exchange generates extensive phenotypic diversity in nascent allopolyploids. Natl Sci. Rev. 8, nwaa277 (2021).
Stein, A. et al. Mapping of homoeologous chromosome exchanges influencing quantitative trait variation in Brassica napus. Plant Biotechnol. J. 15, 1478–1489 (2017).
Jiang, L., Ren, X. & Wu, R. Computational characterization of double reduction in autotetraploid natural populations. Plant J. 105, 1703–1709 (2021).
Manrique-Carpintero, N. C. et al. Genome reduction in tetraploid potato reveals genetic load, haplotype variation, and loci associated with agronomic traits. Front. Plant Sci. 9, 944 (2018).
Otto, S. P. The evolutionary consequences of polyploidy. Cell 131, 452–462 (2007).
Bourke, P. M. et al. Partial preferential chromosome pairing is genotype dependent in tetraploid rose. Plant J. 90, 330–343 (2017).
Wadl, P. A. et al. Genetic diversity and population structure of the USDA sweetpotato (Ipomoea batatas) germplasm collections using GBSpoly. Front. Plant Sci. 9, 1166 (2018).
Mollinari, M. et al. Unraveling the hexaploid sweetpotato inheritance using ultra-dense multilocus mapping. G3 10, 281–292 (2020).
Hackett, C. A., Boskamp, B., Vogogias, A., Preedy, K. F. & Milne, I. TetraploidSNPMap: software for linkage analysis and QTL mapping in autotetraploid populations using SNP dosage data. J. Hered. 108, 438–442 (2017).
Takagi, H. et al. QTL ‐seq: rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations. Plant J. 74, 174–183 (2013).
Chen, J. et al. A tetrasomic inheritance model and likelihood‐based method for mapping quantitative trait loci in autotetraploid species. N. Phytol. 230, 387–398 (2021).
Ma, Z. et al. Resequencing a core collection of upland cotton identifies genomic variation and loci influencing fiber quality and yield. Nat. Genet. 50, 803–813 (2018).
Lu, Q. et al. A genomic variation map provides insights into peanut diversity in China and associations with 28 agronomic traits. Nat. Genet. 56, 530–540 (2024).
Song, J.-M. et al. Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat. Plants 6, 34–45 (2020).
Jin, S. et al. Structural variation (SV)-based pan-genome and GWAS reveal the impacts of SVs on the speciation and diversification of allotetraploid cottons. Mol. Plant 16, 678–693 (2023).
Dwivedi, S. L., Heslop‐Harrison, P., Amas, J., Ortiz, R. & Edwards, D. Epistasis and pleiotropy‐induced variation for plant breeding. Plant Biotechnol. J. 22, 2788–2807 (2024).
Jia, J. et al. Genome resources for the elite bread wheat cultivar Aikang 58 and mining of elite homeologous haplotypes for accelerating wheat improvement. Mol. Plant 16, 1893–1910 (2023).
Sharma, S. K. et al. Linkage disequilibrium and evaluation of genome-wide association mapping models in tetraploid potato. G3 8, 3185–3202 (2018).
Haque, E. et al. Polyploid GWAS reveals the basis of molecular marker development for complex breeding traits including starch content in the storage roots of sweet potato. Front. Plant Sci. 14, 1181909 (2023).
Chen, M. et al. Genome-wide identification of agronomically important genes in outcrossing crops using OutcrossSeq. Mol. Plant 14, 556–570 (2021).
Huang, B., Spooner, D. M. & Liang, Q. Genome diversity of the potato. Proc. Natl Acad. Sci. USA 115, E6392 (2018).
Cooke, D. P., Wedge, D. C. & Lunter, G. Benchmarking small-variant genotyping in polyploids. Genome Res. 32, 403–408 (2022).
Liao, W.-W. et al. A draft human pangenome reference. Nature 617, 312–324 (2023).
Groza, C. et al. Pangenome graphs improve the analysis of structural variants in rare genetic diseases. Nat. Commun. 15, 657 (2024).
Dong, Y. & Wang, Y.-Z. Seed shattering: from models to crops. Front. Plant Sci. 6, 476 (2015).
Van Dam, J., Kooman, P. L. & Struik, P. C. Effects of temperature and photoperiod on early growth and final number of tubers in potato (Solanum tuberosum L.). Potato Res. 39, 51–62 (1996).
Martínez-García, J. F., García-Martínez, J. L., Bou, J. & Prat, S. The interaction of gibberellins and photoperiod in the control of potato tuberization. J. Plant Growth Regul. 20, 377–386 (2001).
Johns, T. Detoxification function of geophagy and domestication of the potato. J. Chem. Ecol. 12, 635–646 (1986).
Shang, Y. et al. Biosynthesis, regulation, and domestication of bitterness in cucumber. Science 346, 1084–1088 (2014).
Avni, R. et al. Wild emmer genome architecture and diversity elucidate wheat evolution and domestication. Science 357, 93–97 (2017).
Jia, J. et al. Aegilops tauschii draft genome sequence reveals a gene repertoire for wheat adaptation. Nature 496, 91–95 (2013).
Zheng, Z. et al. Chloroplast and whole-genome sequencing shed light on the evolutionary history and phenotypic diversification of peanuts. Nat. Genet. 56, 1975–1984 (2024).
Desta, Z. A. & Ortiz, R. Genomic selection: genome-wide prediction in plant improvement. Trends Plant Sci. 19, 592–601 (2014).
Meuwissen, T., Hayes, B. & Goddard, M. Accelerating improvement of livestock with genomic selection. Annu. Rev. Anim. Biosci. 1, 221–237 (2013).
Batista, L. G., Mello, V. H., Souza, A. P. & Margarido, G. R. Genomic prediction with allele dosage information in highly polyploid species. Theor. Appl. Genet. 135, 723–739 (2022).
Yadav, S. et al. Accelerating genetic gain in sugarcane breeding using genomic selection. Agronomy 10, 585 (2020).
Gouy, M. et al. Experimental assessment of the accuracy of genomic selection in sugarcane. Theor. Appl. Genet. 126, 2575–2586 (2013).
Bonnett, D. et al. Response to early generation genomic selection for yield in wheat. Front. Plant Sci. 12, 718611 (2022).
Pandey, J., Scheuring, D. C., Koym, J. W., Endelman, J. B. & Vales, M. I. Genomic selection and genome-wide association studies in tetraploid chipping potatoes. Plant Genome 16, e20297 (2023).
Wang, Y. et al. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 32, 947–951 (2014).
Johansen, I. E. et al. High efficacy full allelic CRISPR/Cas9 gene editing in tetraploid potato. Sci. Rep. 9, 17715 (2019).
Li, J., Yu, X., Zhang, C., Li, N. & Zhao, J. The application of CRISPR/Cas technologies to Brassica crops: current progress and future perspectives. aBIOTECH 3, 146–161 (2022).
Mall, A. K. et al. CRISPR/Cas-mediated genome editing for sugarcane improvement. Sugar Tech. 27, 1–13 (2024).
Vondracek, K., Altpeter, F., Liu, T. & Lee, S. Advances in genomics and genome editing for improving strawberry (Fragaria × ananassa). Front. Genet. 15, 1382445 (2024).
Tripathi, L., Ntui, V. O. & Tripathi, J. N. CRISPR/Cas9-based genome editing of banana for disease resistance. Curr. Opin. Plant Biol. 56, 118–126 (2020).
Liu, Y. et al. Targeted mutagenesis in tetraploid switchgrass (Panicum virgatum L.) using CRISPR/Cas9. Plant Biotechnol. J. 16, 381–393 (2018).
Tang, H. et al. Fertility recovery of wheat male sterility controlled by Ms2 using CRISPR/Cas9. Plant Biotechnol. J. 19, 224–226 (2021).
Zhu, H., Li, C. & Gao, C. Applications of CRISPR–Cas in agriculture and plant biotechnology. Nat. Rev. Mol. Cell Biol. 21, 661–677 (2020).
Zhang, Y. et al. Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat. Commun. 7, 12617 (2016).
Li, J., Li, Y. & Ma, L. Recent advances in CRISPR/Cas9 and applications for wheat functional genomics and breeding. aBIOTECH 2, 375–385 (2021).
Zaman, Q. U., Li, C., Cheng, H. & Hu, Q. Genome editing opens a new era of genetic improvement in polyploid crops. Crop J. 7, 141–150 (2019).
Levy, A. A. & Feldman, M. Evolution and origin of bread wheat. Plant Cell 34, 2549–2567 (2022).
Rosyara, U. et al. Genetic contribution of synthetic hexaploid wheat to CIMMYT’s spring bread wheat breeding germplasm. Sci. Rep. 9, 12355 (2019).
Yu, H. et al. A route to de novo domestication of wild allotetraploid rice. Cell 5, 1156–1170 (2021).
Cheng, L. et al. Leveraging a phased pangenome for haplotype design of hybrid potato. Nature 640, 408–417 (2025).
Ye, M. et al. Generation of self-compatible diploid potato by knockout of S-RNase. Nat. Plants 4, 651–654 (2018).
Zhang, C. et al. The genetic basis of inbreeding depression in potato. Nat. Genet. 51, 374–378 (2019).
Zhang, C. et al. Genome design of hybrid potato. Cell 184, 3873–3883 (2021).
Washburn, J. D. & Birchler, J. A. Polyploids as a “model system” for the study of heterosis. Plant Reprod. 27, 1–5 (2014).
Mieulet, D. et al. Turning rice meiosis into mitosis. Cell Res. 26, 1242–1254 (2016).
Wang, Y. et al. Harnessing clonal gametes in hybrid crops to engineer polyploid genomes. Nat. Genet. 56, 1075–1079 (2024).
Liu, S. et al. A telomere-to-telomere genome assembly coupled with multi-omic data provides insights into the evolution of hexaploid bread wheat. Nat. Genet. 57, 1008–1020 (2025).
Wang, Z. et al. Near-complete assembly and comprehensive annotation of the wheat Chinese Spring genome. Mol. Plant 18, 892–907 (2025).
Athiyannan, N. et al. Long-read genome sequencing of bread wheat facilitates disease resistance gene cloning. Nat. Genet. 54, 227–231 (2022).
Jia, K.-H. et al. SubPhaser: a robust allopolyploid subgenome phasing method based on subgenome-specific k-mers. N. Phytol. 235, 801–809 (2022).
Lovell, J. T. et al. Genomic mechanisms of climate adaptation in polyploid bioenergy switchgrass. Nature 590, 438–444 (2021).
Yan, H. et al. Post-polyploidization centromere evolution in cotton. Nat. Genet. 57, 1021–1030 (2025).
Hu, G. et al. A telomere-to-telomere genome assembly of cotton provides insights into centromere evolution and short-season adaptation. Nat. Genet. 57, 1031–1043 (2025).
Sreedasyam, A. et al. Genome resources for three modern cotton lines guide future breeding efforts. Nat. Plants 10, 1039–1051 (2024).
Li, B. et al. A gap-free reference genome reveals structural variations associated with flowering time in rapeseed (Brassica napus). Hortic. Res. 10, uhad171 (2023).
Zhao, C. et al. Chromosome-scale genome assembly-assisted identification of Brassica napus BnDCPA1 for improvement of plant architecture and yield heterosis. Plant Commun. 5, 100854 (2024).
Gao, S. et al. The centromere landscapes of four karyotypically diverse Papaver species provide insights into chromosome evolution and speciation. Cell Genom. 4, 100626 (2024).
Yang, X. et al. Three chromosome-scale Papaver genomes reveal punctuated patchwork evolution of the morphinan and noscapine biosynthesis pathway. Nat. Commun. 12, 6030 (2021).
Kumar, S. et al. TimeTree 5: an expanded resource for species divergence times. Mol. Biol. Evol. 39, msac174 (2022).
Clark, J. W. & Donoghue, P. C. Whole-genome duplication and plant macroevolution. Trends Plant Sci. 23, 933–945 (2018).
Kreiner, J. M., Kron, P. & Husband, B. C. Evolutionary dynamics of unreduced gametes. Trends Genet. 33, 583–593 (2017).
Ramsey, J. & Schemske, D. W. Pathways, mechanisms, and rates of polyploid formation in flowering plants. Annu. Rev. Ecol. Syst. 29, 467–501 (1998).
Chen, Z. J. et al. Genomic diversifications of five Gossypium allopolyploid species and their impact on cotton improvement. Nat. Genet. 52, 525–533 (2020).
Dai, D. et al. Unzipped chromosome-level genomes reveal allopolyploid nematode origin pattern as unreduced gamete hybridization. Nat. Commun. 14, 7156 (2023).
Acknowledgements
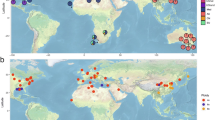
The assembled polyploid genomes in Fig. 1 are based on our own compilation from publicly available resources, primarily the PubPlant database, which may not include all reported species. We thank the developers and curators of PubPlant for their efforts in collecting and maintaining the data. We thank W. J. Lucas, J. Zhang, Z. Liu and N. Wang for project discussions and critical comments. This project was funded by a Guangdong Major Project of Basic and Applied Basic Research (2021B0301030004), the National Natural Science Foundation of China (grant numbers 32488302 and 31991181), the Fundamental Research Fund for the Central Universities (77000-12240018), the Agricultural Science and Technology Innovation Program (CAAS-ZDRW202404) and a Shenzhen Outstanding Talents Training Fund. Z.B. is supported by the grants to D. Weigel.
Author information
Authors and Affiliations
Contributions
Q.Z., Z.B. and S.H. co-designed the framework of the paper. L.C., Z.B. and Q.Z. wrote the draft of the paper and generated the figures. Q.K., L.L., N.S. and S.H. contributed to writing and revision of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Fay-Wei Li and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Tables 1 and 2 (download PDF )
Supplementary Tables 1 and 2.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cheng, L., Bao, Z., Kong, Q. et al. Genome analyses and breeding of polyploid crops. Nat. Plants 11, 1714–1728 (2025). https://doi.org/10.1038/s41477-025-02088-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02088-5