Abstract
Biodiversity is known to promote ecosystem multifunctionality (EMF), but how grassland degradation influences the relationship between biodiversity and EMF remains unclear. Here, using paired observations at 44 sites (a total of 792 sampling quadrats) along a 2,600 km transect, we test how moderate grassland degradation influences 20 surrogates of ecosystem functions, EMF, plant richness, soil bacterial, fungal and protist richness, and biodiversity–EMF relationships in Tibetan alpine grasslands. Our results reveal significant declines in individual ecosystem functions and EMF with moderate grassland degradation. By contrast, both plant richness and integrated soil biodiversity exhibit significant increases. The structural equation models analyses show that following degradation, the effect of soil biodiversity on EMF strengthens, whereas that of plant richness weakens. These findings offer large-scale empirical evidence that moderate grassland degradation can amplify both soil biodiversity and its functional importance, emphasizing the key role of below-ground biodiversity in supporting ecosystem functioning in degraded grasslands.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data used in this study are available from the figshare data repository https://doi.org/10.6084/m9.figshare.30119233 (ref. 93). The sequence data generated in this study have been deposited in the Genome Sequence Archive in National Genomics Data Center, China National Center for Bioinformation https://ngdc.cncb.ac.cn/bioproject/browse/PRJCA046361 (ref. 94). Source data are provided with this paper.
References
Keck, F. et al. The global human impact on biodiversity. Nature 641, 395–400 (2025).
Blowes, S. A. et al. The geography of biodiversity change in marine and terrestrial assemblages. Science 366, 339–345 (2019).
van der Plas, F. Biodiversity and ecosystem functioning in naturally assembled communities. Biol. Rev. 94, 1220–1245 (2019).
Weiskopf, S. R. et al. Biodiversity loss reduces global terrestrial carbon storage. Nat. Commun. 15, 4354 (2024).
Hautier, Y. et al. Local loss and spatial homogenization of plant diversity reduce ecosystem multifunctionality. Nat. Ecol. Evol. 2, 50–56 (2018).
Hooper, D. U. et al. A global synthesis reveals biodiversity loss as a major driver of ecosystem change. Nature 486, 105–108 (2012).
Tilman, D., Reich, P. B. & Isbell, F. Biodiversity impacts ecosystem productivity as much as resources, disturbance, or herbivory. Proc. Natl Acad. Sci. USA 109, 10394–10397 (2012).
Pichon, N. A. et al. Nitrogen availability and plant functional composition modify biodiversity-multifunctionality relationships. Ecol. Lett. 27, e14361 (2024).
Hong, P. et al. Biodiversity promotes ecosystem functioning despite environmental change. Ecol. Lett. 25, 555–569 (2022).
Garcia, F. C., Bestion, E., Warfield, R. & Yvon-Durocher, G. Changes in temperature alter the relationship between biodiversity and ecosystem functioning. Proc. Natl Acad. Sci. USA 115, 10989–10994 (2018).
McIntire, E. J. & Fajardo, A. Facilitation as a ubiquitous driver of biodiversity. New Phytol. 201, 403–416 (2014).
Stein, A., Gerstner, K. & Kreft, H. Environmental heterogeneity as a universal driver of species richness across taxa, biomes and spatial scales. Ecol. Lett. 17, 866–880 (2014).
Koerner, S. E. et al. Change in dominance determines herbivore effects on plant biodiversity. Nat. Ecol. Evol. 2, 1925–1932 (2018).
Armstrong, R. A. & McGehee, R. Competitive exclusion. Am. Nat. 115, 151–170 (1980).
Horváth, Z., Ptacnik, R., Vad, C. F. & Chase, J. M. Habitat loss over six decades accelerates regional and local biodiversity loss via changing landscape connectance. Ecol. Lett. 22, 1019–1027 (2019).
Moi, D. A. et al. Human pressure drives biodiversity–multifunctionality relationships in large Neotropical wetlands. Nat. Ecol. Evol. 6, 1279–1289 (2022).
Maestre, F. T., Castillo-Monroy, A. P., Bowker, M. A. & Ochoa-Hueso, R. Species richness effects on ecosystem multifunctionality depend on evenness, composition and spatial pattern. J. Ecol. 100, 317–330 (2012).
Mori, A. S., Isbell, F. & Cadotte, M. W. Assessing the importance of species and their assemblages for the biodiversity ecosystem multifunctionality relationship. Ecology 104, e4104 (2023).
Bardgett, R. D. & van der Putten, W. H. Belowground biodiversity and ecosystem functioning. Nature 515, 505–511 (2014).
Tibbett, M., Fraser, T. D. & Duddigan, S. Identifying potential threats to soil biodiversity. PeerJ 8, e9271 (2020).
Bardgett, R. D. et al. Combatting global grassland degradation. Nat. Rev. Earth Environ. 2, 720–735 (2021).
Li, C., de Jong, R., Schmid, B., Wulf, H. & Schaepman, M. E. Changes in grassland cover and in its spatial heterogeneity indicate degradation on the Qinghai-Tibetan Plateau. Ecol. Indic. 119, 106641 (2020).
Du, Y. et al. Core microbes regulate plant-soil resilience by maintaining network resilience during long-term restoration of alpine grasslands. Nat. Commun. 16, 3116 (2025).
Zhou, H. et al. Alpine grassland degradation and its restoration in the Qinghai-Tibet Plateau. Grasses 2, 31–46 (2023).
Zhang, W., Xue, X., Peng, F., You, Q. & Hao, A. Meta-analysis of the effects of grassland degradation on plant and soil properties in the alpine meadows of the Qinghai-Tibetan Plateau. Glob. Ecol. Conserv. 20, e00774 (2019).
Zhang, J., Wang, J., Chen, W., Li, B. & Zhao, K. Vegetation of Xizang (Tibet) (Science, 1988).
Song, M., Dong, M. & Jiang, G. Importance of clonal plants and plant species diversity in the Northeast China Transect. Ecol. Res. 17, 705–716 (2002).
Manning, P. et al. Redefining ecosystem multifunctionality. Nat. Ecol. Evol. 2, 427–436 (2018).
Byrnes, J. E. K. et al. Investigating the relationship between biodiversity and ecosystem multifunctionality: challenges and solutions. Methods Ecol. Evol. 5, 111–124 (2014).
Byrnes, J. E. K., Roger, F. & Bagchi, R. Understandable multifunctionality measures using Hill numbers. Oikos 2023, e09402 (2023).
Stone, B. W. G. et al. Life history strategies among soil bacteria-dichotomy for few, continuum for many. ISME J. 17, 611–619 (2023).
Zhang, G., Yang, F. & Long, H. Save the life-sustaining mattic layer on the Qinghai-Tibetan Plateau. Innov 4, 100418 (2023).
Wagg, C., Bender, S. F., Widmer, F. & van der Heijden, M. G. Soil biodiversity and soil community composition determine ecosystem multifunctionality. Proc. Natl Acad. Sci. USA 111, 5266–5270 (2014).
Lange, M. et al. Plant diversity increases soil microbial activity and soil carbon storage. Nat. Commun. 6, 6707 (2015).
Fierer, N. & Jackson, R. B. The diversity and biogeography of soil bacterial communities. Proc. Natl Acad. Sci. USA 103, 626–631 (2006).
Kamble, P. N., Gaikwad, V. B., Kuchekar, S. R. & Bååth, E. Microbial growth, biomass, community structure and nutrient limitation in high pH and salinity soils from Pravaranagar (India). Eur. J. Soil Biol. 65, 87–95 (2014).
Tibbett, M. et al. Long-term acidification of pH neutral grasslands affects soil biodiversity, fertility and function in a heathland restoration. Catena 180, 401–415 (2019).
Tian, Q. et al. A novel soil manganese mechanism drives plant species loss with increased nitrogen deposition in a temperate steppe. Ecology 97, 65–74 (2016).
Li, Y. et al. Environmental heterogeneity and dispersal limitation simultaneously determine the spatial scaling of different microbial functional groups. Sci. Total Environ. 885, 163854 (2023).
Jing, X. et al. The links between ecosystem multifunctionality and above- and belowground biodiversity are mediated by climate. Nat. Commun. 6, 8159 (2015).
Dee, L. E. et al. Clarifying the effect of biodiversity on productivity in natural ecosystems with longitudinal data and methods for causal inference. Nat. Commun. 14, 2607 (2023).
Hu, W. et al. Aridity-driven shift in biodiversity-soil multifunctionality relationships. Nat. Commun. 12, 5350 (2021).
Zhang, M. et al. Experimental impacts of grazing on grassland biodiversity and function are explained by aridity. Nat. Commun. 14, 1–8 (2023).
Qin, S. et al. Temperature sensitivity of permafrost carbon release mediated by mineral and microbial properties. Sci. Adv. 7, eabe3596 (2021).
Zhang, Z. et al. Don’t judge toxic weeds on whether they are native but on their ecological effects. Ecol. Evol. 10, 9014–9025 (2020).
Loreau, M. & Hector, A. Partitioning selection and complementarity in biodiversity experiments. Nature 412, 72–76 (2001).
Herren, C. M. & McMahon, K. D. Cohesion: a method for quantifying the connectivity of microbial communities. ISME J. 11, 2426–2438 (2017).
Xun, W. B. et al. Specialized metabolic functions of keystone taxa sustain soil microbiome stability. Microbiome 9, 1–15 (2021).
Wright, A. J., Wardle, D. A., Callaway, R. & Gaxiola, A. The overlooked role of facilitation in biodiversity experiments. Trends Ecol. Evol. 32, 383–390 (2017).
Filstrup, C. T., King, K. B. & McCullough, I. M. Evenness effects mask richness effects on ecosystem functioning at macro-scales in lakes. Ecol. Lett. 22, 2120–2129 (2019).
Noulèkoun, F. et al. Contrasting ecological mechanisms mediate the impact of land conversion on ecosystem multifunctionality. Funct. Ecol. 39, 783–798 (2025).
Gu, S. et al. Microbial and chemical fertilizers for restoring degraded alpine grassland. Biol. Fertil. Soils 59, 911–926 (2023).
Editorial Committee for Vegetation Map of China. Vegetation Atlas of China (Science, 2001).
Ding, J. et al. Decadal soil carbon accumulation across Tibetan permafrost regions. Nat. Geosci. 10, 420–424 (2017).
Wang, Y. F. et al. Vegetation structural shift tells environmental changes on the Tibetan Plateau over 40 years. Sci. Bull. 68, 1928–1937 (2023).
Dong, S., Zhang, Y., Shen, H., Li, S. & Xu, Y. Overview of the third pole’s grasslands. in Grasslands on the Third Pole of the World: Structure, Function, Process, and Resilience of Social-Ecological Systems 31–72 (Springer, 2023).
Sun, J. et al. Nature-based solutions can help restore degraded grasslands and increase carbon sequestration in the Tibetan Plateau. Commun. Earth Environ. 5, 154 (2024).
IUSS Working Group. International Soil Classification System for Naming Soils and Creating Legends for Soil Maps. World Soil Resources Reports (Food and Agriculture Organization of the United Nations, 2015).
Liu, B. Actual Livestock Carrying Capacity Estimation Product in Qinghai-Tibet Plateau (2000–2019) (National Tibetan Plateau/Third Pole Environment Data Center, 2021); https://doi.org/10.11888/Ecolo.tpdc.271513
Che, R. et al. Degraded patch formation significantly changed microbial community composition in alpine meadow soils. Soil. Till. Res. 195, 104426 (2019).
Wen, L. et al. The construction of grassland degradation index for alpine meadow in Qinghai-Tibetan Plateau. Procedia Environ. Sci. 2, 1966–1969 (2010).
Zhang, H., Chen, W., Dong, L. & Wang, W. Grassland degradation amplifies the negative effect of nitrogen enrichment on soil microbial community stability. Glob. Change Biol. 30, e17217 (2024).
Zheng, H. et al. The implementation of ecological protection in Inner Mongolia has slowed down grassland degradation. Fundam. Res. https://doi.org/10.1016/j.fmre.2024.10.006 (2024).
Bastida, F., Luis Moreno, J., Teresa, H. & García, C. Microbiological degradation index of soils in a semiarid climate. Soil Biol. Biochem. 38, 3463–3473 (2006).
Li, X. J. et al. Root biomass distribution in alpine ecosystems of the northern Tibetan Plateau. Environ. Earth Sci. 64, 1911–1919 (2011).
Yang, Y., Fang, J., Ji, C. & Han, W. Above- and belowground biomass allocation in Tibetan grasslands. J. Veg. Sci. 20, 177–184 (2009).
Olsen, S. R. Estimation of Available Phosphorus in Soils by Extraction with Sodium Bicarbonate (US Department of Agriculture, 1954).
Vance, E. D., Brookes, P. C. & Jenkinson, D. S. An extraction method for measuring soil microbial biomass C. Soil Biol. Biochem. 19, 703–707 (1987).
Hedley, M. J., Stewart, J. W. B. & Chauhan, B. S. Changes in inorganic and organic soil phosphorus fractions induced by cultivation practices and by laboratory incubations. Soil Sci. Soc. Am. J. 46, 970–976 (1982).
Brookes, P. C., Landman, A., Pruden, G. & Jenkinson, D. S. Chloroform fumigation and the release of soil nitrogen: a rapid direct extraction method to measure microbial biomass nitrogen in soil. Soil Biol. Biochem. 17, 837–842 (1985).
German, D. P. et al. Optimization of hydrolytic and oxidative enzyme methods for ecosystem studies. Soil Biol. Biochem. 43, 1387–1397 (2011).
Oksanen, J. et al. vegan: Community ecology package. R package version 2.6-4 https://CRAN.R-project.org/package=vegan (2010).
Delgado-Baquerizo, M. et al. Changes in belowground biodiversity during ecosystem development. Proc. Natl Acad. Sci. USA 116, 6891–6896 (2019).
Finkel, O. M. et al. A single bacterial genus maintains root growth in a complex microbiome. Nature 587, 103–108 (2020).
Zhang, Y. et al. Temperature fluctuation promotes the thermal adaptation of soil microbial respiration. Nat. Ecol. Evol. 7, 205–213 (2023).
Ramirez, K. S. et al. Biogeographic patterns in below-ground diversity in New York City’s Central Park are similar to those observed globally. Proc. R. Soc. B 281, 20141988 (2014).
Amaral-Zettler, L. A., McCliment, E. A., Ducklow, H. W. & Huse, S. M. A method for studying protistan diversity using massively parallel sequencing of V9 hypervariable regions of small-subunit ribosomal RNA genes. PLoS ONE 4, e6372 (2009).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Callahan, B. J., McMurdie, P. J. & Holmes, S. P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J. 11, 2639–2643 (2017).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, 590–596 (2013).
Federhen, S. The NCBI Taxonomy database. Nucleic Acids Res. 40, 136–143 (2012).
Delgado-Baquerizo, M. et al. Multiple elements of soil biodiversity drive ecosystem functions across biomes. Nat. Ecol. Evol. 4, 210–220 (2020).
Yang, K. et al. A High-Resolution Near-Surface Meteorological Forcing Dataset for the Third Pole Region (TPMFD, 1979–2022) (National Tibetan Plateau/Third Pole Environment Data Center, 2023); https://doi.org/10.11888/Atmos.tpdc.300398
Delgado-Baquerizo, M. et al. Global homogenization of the structure and function in the soil microbiome of urban greenspaces. Sci. Adv. 7, eabg5809 (2021).
Cheng, C. et al. Plant species richness on the Tibetan Plateau: patterns and determinants. Ecography 1, e06265 (2023).
Lefcheck, J. S. piecewiseSEM: piecewise structural equation modelling in R for ecology, evolution, and systematics. Methods Ecol. Evol. 7, 573–579 (2016).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2023).
Gao, X. et al. Grassland degradation alters plant and soil biodiversity–multifunctionality relationships. figshare https://doi.org/10.6084/m9.figshare.30119233 (2025).
Gao, X. Grassland degradation alters plant and soil biodiversity-multifunctionality relationships. National Genomics Data Center, China National Center for Bioinformation https://ngdc.cncb.ac.cn/bioproject/browse/PRJCA046361 (2025).
Jarvis, A., Reuter, H. I., Nelson, A. & Guevara, E. Hole-filled SRTM for the globe Version 4. CGIAR-CSI SRTM 90m Database http://srtm.csi.cgiar.org (2008).
Acknowledgements
The authors thank Z. Tang (Institute of Ecology, College of Urban and Environmental Science, Peking University), B. Schmid (Remote Sensing Laboratories, Department of Geography, University of Zurich) and N. A. Pichon (Swiss Federal Institute for Forest, Snow and Landscape Research WSL, Birmensdorf, Switzerland) for their comments and suggestions on the manuscript. This study was financially supported by the Strategic Priority Research Program of Chinese Academy of Sciences (XDA26020201, Y.Y.), the National Natural Science Foundation of China (32588202, Y.Y.; 32425004, Y.Y.; 32301501, X.G.) and the New Cornerstone Science Foundation through the XPLORER PRIZE (Y.Y.).
Author information
Authors and Affiliations
Contributions
Y.Y. conceived the idea. Y.Y., D.Z. and X.G. designed the research. Y.P. and Y.Y. designed the field sampling. Y.P., X.G., Q.L., Z.W., Y.N., S.Y. and X.L. performed field samplings. X.G., Z.W., L.Z., Y.N., Y.L. and S.Y. performed the experiments. X.G. and D.Z. analysed the data. X.G., D.Z. and Y.Y. wrote the paper with inputs from J.P., Y.H., M.L., S.W., L.J., B.W., S.Q., Y.S. and L.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Eric Allan, Ling Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
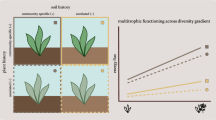
Extended Data Fig. 1 Schematic diagram of the sampling design.
At a specific site, three blocks were selected, and within each block a 30 × 30 m plot was established in both non-degraded and degraded grasslands. Within each plot, three 1 × 1 m quadrats were randomly chosen to conduct vegetation surveys and collect plant and soil samples (9 soil cores at 0-10 cm depth). Altitude map derived from the Shuttle Radar Topography Mission (SRTM)-Digital Elevation Model (DEML)95. Drone images of contrasting non-degraded and degraded grasslands show that plant productivity and the proportion of grasses and sedges (for example, S. purpurea and K. humilis) decreases while the proportion of forbs (for example, L. nanum and S. chamaejasme) increases upon degradation. Photos were captured by Qinlu Li in Babao Town (37.86°N, 100.44°E, altitude 3663 m), July 2021. ND, non-degraded grasslands; DG, degraded grasslands.
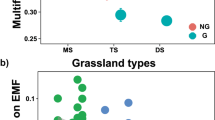
Extended Data Fig. 2 Comparisons of ecosystem multifunctionality (EMF) between non-degraded (ND) and degraded grasslands (DG).
a, Averaged approach; b, multiple-threshold approach; c, d, hill-number based approaches when q = 1 (c) and q = 2 (d). The open circles shown are the site-level values. The ends of a box denote the 25th and 75th quartiles, and whiskers extend to 1.5 times the standard deviation (n = 44 independent sites). The horizontal line within each box represents the median value. Linear mixed models with a correction for environmental covariates were employed to conduct statistical comparisons between non-degraded and degraded grasslands across the 44 sites. Climate and soil factors were treated as covariates, site and grassland type (alpine steppe, alpine meadow or swamp meadow) were treated as random factors. All the reported two-sided P values were not corrected for multiple comparisons, and detailed F-statistics were presented in Supplementary table 1.
Extended Data Fig. 3 Comparisons of plant community in non-degraded (ND) and degraded grasslands (DG).
a, Cover of grasses and sedges; b, cover of forbs. The dashed lines indicate median values; solid lines represent the values for each site (n = 44 independent sites). Linear mixed models with a correction for environmental covariates were employed to conduct statistical comparisons between non-degraded and degraded grasslands across the 44 sites (cover of grasses and sedges: F1,46.72 = 276.01, P = 3.10 × 10−21; cover of forbs: F1, 44.72 = 52.66, P = 4.40 × 10−9; the two-sided P values were not corrected for multiple comparisons). Climate and soil factors were treated as covariates, site and grassland type (alpine steppe, alpine meadow or swamp meadow) were treated as random factors.
Extended Data Fig. 4 Differences in the relative abundance of copiotrophic and oligotrophic bacteria between non-degraded (ND) and degraded grasslands (DG).
a, Copiotrophic bacteria; b, oligotrophic bacteria. The ends of a box denote the 25th and 75th quartiles, and whiskers extend to 1.5 times the standard deviation (n = 44 independent sites). The horizontal line within each box represents the median value. The open circles represent the values for each site. Linear mixed models with a correction for environmental covariates were employed to conduct statistical comparisons between non-degraded and degraded grasslands across the 44 sites (copiotrophic bacteria: F1,51.14 = 5.69, P = 0.0209; oligotrophic bacteria: F1, 51.01 = 6.46, P = 0.0141; the two-sided P values were not corrected for multiple comparisons). Climate and soil factors were treated as covariates, site and grassland type (alpine steppe, alpine meadow or swamp meadow) were treated as random factors. The demarcations of copiotrophic and oligotrophic bacteria were as follows31, oligotrophs: Acidobacteriota, Actinobacteriota, Planctomycetota, Chloroflexi, Verrucomicrobiota, Cyanobacteria; copiotrophs: Firmicutes, Gemmatimonadota, Bacteroidota, Proteobacteria. Copiotrophic bacteria are generally fast-slowing and resource acquisitive, while oligotrophic bacteria tend to be slow-growing and resource conservative31.
Extended Data Fig. 5 Differences in cover of clonal and non-clonal plants in non-degraded (ND) and degraded grasslands (DG).
a, Clonal plants; b, non-clonal plants. The horizontal line within each box represents the median value; the open circles represent the values for each site (n = 44 independent sites). Linear mixed models with a correction for environmental covariates were employed to conduct statistical comparisons between non-degraded and degraded grasslands across the 44 sites (cover of clonal plants: F1,47.77 = 265.38, P = 3.89 × 10−21; cover of non-clonal plants: F1, 44.67 = 39.42, P = 1.24 × 10−7; the two-sided P values were not corrected for multiple comparisons). Climate and soil factors were treated as covariates, site and grassland type (alpine steppe, alpine meadow or swamp meadow) were treated as random factors.
Extended Data Fig. 6 Relationships between soil biodiversity and plant richness in non-degraded (ND) and degraded grasslands (DG).
Soil biodiversity was calculated by averaging biodiversity across the three taxa (soil bacteria, fungi and protists) after standardization33,73. Linear mixed models (n = 44 independent sites) were used to determine the linkages between soil biodiversity and plant richness, with grassland type (alpine steppe, alpine meadow or swamp meadow) being treated as the random factor (non-degraded grasslands: F1,42 = 27.68, P = 4.55 × 10−6; degraded grasslands: F1,41.71 = 24.97, P = 1.09 × 10−5; the two-sided P values were not corrected for multiple comparisons). Shaded areas are the 95% confidence intervals; the open circles are the values for each site. Soil biodiversity still exhibited significant linkages with plant richness when considering the environmental covariates (climate and soil factors).
Extended Data Fig. 7 Differences in cover of poisonous weeds in non-degraded (ND) and degraded grasslands (DG).
The ends of a box denote the 25th and 75th quartiles, and whiskers extend to 1.5 times the standard deviation. The open circles represent the values for each site (n = 44 independent sites). Linear mixed models with a correction for environmental covariates were employed to conduct statistical comparisons between non-degraded and degraded grasslands across the 44 sites (F1,48.06 = 34.34, P = 4.08 × 10−7; the two-sided P value was not corrected for multiple comparisons). Climate and soil factors were treated as covariates, site and grassland type (alpine steppe, alpine meadow or swamp meadow) were treated as random factors.
Extended Data Fig. 8 Comparisons of plant and microbial community evenness between non-degraded (ND) and degraded grasslands (DG).
a, Plant evenness; b, bacteria evenness; c, fungi evenness; d, protist evenness. The ends of a box denote the 25th and 75th quartiles, and whiskers extend to 1.5 times the standard deviation (n = 44 independent sites). The horizontal line within each box represents the median value. The open circles represent the values for each site. Linear mixed models with a correction for environmental covariates were employed to conduct statistical comparisons between non-degraded and degraded grasslands across the 44 sites (plant: F1,48.52 = 143.60, P = 4.15 × 10−16; bacteria: F1, 42.24 = 11.05, P = 0.0018; fungi: F1, 48.86 = 4.45, P = 0.0400; protist: F1, 50.95 = 0.25, P = 0.6211; the two-sided P values were not corrected for multiple comparisons). Climate and soil factors were treated as covariates, site and grassland type (alpine steppe, alpine meadow or swamp meadow) were treated as random factors.
Extended Data Fig. 9 Comparisons of microbial interspecific cohesion between non-degraded (ND) and degraded grasslands (DG).
a, Microbial positive cohesion; b, microbial negative cohesion. The ends of a box denote the 25th and 75th quartiles, and whiskers extend to 1.5 times the standard deviation (n = 44 independent sites). The open circles represent the values for each site, the horizontal line within each box represents the median value. Linear mixed models with a correction for environmental covariates were employed to conduct statistical comparisons between non-degraded and degraded grasslands across the 44 sites (positive cohesion: F1,49.67 = 24.93, P = 7.71 × 10−6; negative cohesion: F1, 41.20 = 0.11, P = 0.7464; the two-sided P values were not corrected for multiple comparisons). Climate and soil factors were treated as covariates, site and grassland type (alpine steppe, alpine meadow or swamp meadow) were treated as random factors.
Extended Data Fig. 10 Comparisons of microbial community composition between non-degraded (ND) and degraded grasslands (DG).
a, Bacterial community composition; b, fungal community composition; c, protist community composition. Data shown in this figure are the relative abundance of microbes at the level of phylum. Community composition of bacteria, fungi and protist all exhibited significant changes following degradation according to permutational multivariable analysis of variance (all P < 0.05). Linear mixed models with a correction for environmental covariates were employed to conduct statistical comparisons of the relative abundance of each microbial group between non-degraded and degraded grasslands across the 44 sites. Climate and soil factors were treated as covariates, site and grassland type (alpine steppe, alpine meadow or swamp meadow) were treated as random factors. The plus (+) and minus (-) signs denote that microbial taxa significantly increase or decrease, respectively, following grassland degradation. All the reported two-sided P values were not corrected for multiple comparisons.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1 and 2, Figs. 1–9, Tables 1–4 and Refs. 1–17.
Supplementary Data 1 (download XLSX )
Statistical source data.
Source data
Source Data Figs. 2–4 and Extended Data Figs. 2–10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gao, X., Zhang, D., Peng, Y. et al. Grassland degradation alters plant and soil biodiversity–multifunctionality relationships. Nat. Plants 11, 2487–2497 (2025). https://doi.org/10.1038/s41477-025-02147-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02147-x