Abstract
The receptor-like cytoplasmic kinase BIK1 and its close homologue PBL1 have been widely recognized as central components of plant immunity. However, most genetic studies of BIK1 and PBL1 functions were carried out with single transfer DNA (T-DNA) insertional mutant alleles. Some phenotypes observed in these mutants, for example autoimmunity, have been difficult to reconcile with the proposed role of BIK1 and PBL1 in pattern-triggered immunity. In this study, we generated several new alleles of bik1 and pbl1 by CRISPR–Cas9-based gene editing and systematically analysed these mutants alongside existing T-DNA insertional lines. These analyses reinforced the central role of BIK1 and PBL1 in pattern-triggered immunity mediated by both receptor kinases and receptor-like proteins. At the same time, however, we revealed several pleiotropic phenotypes associated with T-DNA insertions that are not necessarily linked to loss of BIK1 or PBL1 function. Further analyses of newly generated bik1 pbl1 double mutants uncovered an even greater contribution of these kinases to immune signalling and disease resistance than previously appreciated. These findings clarify longstanding ambiguities surrounding BIK1 and PBL1 functions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The raw sequencing data of Col-0, bik1-1 and pbl1-1 generated by the DNBSEQ platform have been deposited in the National Center for Biotechnology Information database under the accession number PRJNA1255789. The third-generation raw data of bik1-1 have been submitted to the Genome Sequence Archive in the National Genomics Data Center with the record number CRA030180. The assembled genome is available via Figshare at https://doi.org/10.6084/m9.figshare.30148612.v1 (ref. 58). Other data are provided in Supplementary Table 1. Materials created in this study are available from the laboratories of the corresponding authors. Source data are provided with this paper.
Change history
04 March 2026
A Correction to this paper has been published: https://doi.org/10.1038/s41477-026-02259-y
References
Veronese, P. et al. The membrane-anchored Botrytis-induced kinase1 plays distinct roles in Arabidopsis resistance to necrotrophic and biotrophic pathogens. Plant Cell 18, 257–273 (2006).
Lu, D. et al. A receptor-like cytoplasmic kinase, BIK1, associates with a flagellin receptor complex to initiate plant innate immunity. Proc. Natl Acad. Sci. USA 107, 496–501 (2009).
Zhang, J. et al. Receptor-like cytoplasmic kinases integrate signaling from multiple plant immune receptors and are targeted by a Pseudomonas syringae effector. Cell Host Microbe 7, 290–301 (2010).
Liu, Z. et al. BIK1 interacts with PEPRs to mediate ethylene-induced immunity. Proc. Natl Acad. Sci. USA 110, 6205–6210 (2013).
Kadota, Y., Shirasu, K. & Zipfel, C. Regulation of the NADPH oxidase RBOHD during plant immunity. Plant Cell Physiol. 56, 1472–1480 (2015).
Li, L. et al. The FLS2-associated kinase BIK1 directly phosphorylates the NADPH oxidase RbohD to control plant immunity. Cell Host Microbe 15, 329–338 (2014).
Liang, X. & Zhou, J. M. Receptor-like cytoplasmic kinases: central players in plant receptor kinase-mediated Signaling. Annu. Rev. Plant Biol. 69, 267–299 (2018).
Ranf, S. et al. Microbe-associated molecular pattern-induced calcium signaling requires the receptor-like cytoplasmic kinases, PBL1 and BIK1. BMC Plant Biol. 14, 374 (2014).
Monaghan, J., Matschi, S., Romeis, T. & Zipfel, C. The calcium-dependent protein kinase CPK28 negatively regulates the BIK1-mediated PAMP-induced calcium burst. Plant Signal. Behav. 10, e1018497 (2015).
Hou, S. et al. The Arabidopsis MIK2 receptor elicits immunity by sensing a conserved signature from phytocytokines and microbes. Nat. Commun. 12, 5494 (2021).
Chen, X. et al. Arabidopsis perceives caterpillar oral secretion to increase resistance by reactive oxygen species-enhanced glucosinolate hydrolysis. New Phytol. 246, 1304–1318 (2025).
Yuan, M. et al. Pattern-recognition receptors are required for NLR-mediated plant immunity. Nature 592, 105–109 (2021).
Rao, S. et al. Roles of receptor-like cytoplasmic kinase VII members in pattern-triggered immune signaling. Plant Physiol. 177, 1679–1690 (2018).
Pruitt, R. N. et al. The EDS1-PAD4-ADR1 node mediates Arabidopsis pattern-triggered immunity. Nature 598, 495–499 (2021).
Kadota, Y. et al. Direct regulation of the NADPH oxidase RBOHD by the PRR-associated kinase BIK1 during plant immunity. Mol. Cell 54, 43–55 (2014).
Bi, G. et al. Receptor-like cytoplasmic kinases directly link diverse pattern recognition receptors to the activation of mitogen-activated protein kinase cascades in Arabidopsis. Plant Cell 30, 1543–1561 (2018).
Thor, K. et al. The calcium-permeable channel OSCA1.3 regulates plant stomatal immunity. Nature 585, 569–573 (2020).
Ma, M. et al. A surface-receptor-coupled G protein regulates plant immunity through nuclear protein kinases. Cell Host Microbe 30, 1602–1614 (2022).
Li, Y. et al. Plasma membrane-nucleo-cytoplasmic coordination of a receptor-like cytoplasmic kinase promotes EDS1-dependent plant immunity. Nat. Plants 8, 802–816 (2022).
Kong, L. et al. Dual phosphorylation of DGK5-mediated PA burst regulates ROS in plant immunity. Cell 187, 609–623 (2024).
Feng, F. et al. A Xanthomonas uridine 5’-monophosphate transferase inhibits plant immune kinases. Nature 485, 114–118 (2012).
Liang, X. et al. A Phytophthora capsici RXLR effector targets and inhibits the central immune kinases to suppress plant immunity. New Phytol. 232, 264–278 (2021).
Yu, G. et al. The Arabidopsis E3 ubiquitin ligase PUB4 regulates BIK1 and is targeted by a bacterial type-III effector. EMBO J. 41, e107257 (2022).
Sun, Z. M. et al. The Ralstonia solanacearum Type III effector RipAW targets the immune receptor complex to suppress PAMP-triggered immunity. Int. J. Mol. Sci. 25, 183 (2023).
Blekemolen, M. C. et al. The PTI-suppressing Avr2 effector from Fusarium oxysporum suppresses mono-ubiquitination and plasma membrane dissociation of BIK1. Mol. Plant Pathol. 24, 1273–1286 (2023).
Wan, W. L. et al. Comparing Arabidopsis receptor kinase and receptor protein-mediated immune signaling reveals BIK1-dependent differences. New Phytol. 221, 2080–2095 (2019).
Liang, X. et al. Arabidopsis heterotrimeric G proteins regulate immunity by directly coupling to the FLS2 receptor. eLife 5, e13568 (2016).
Wang, J. et al. A regulatory module controlling homeostasis of a plant immune kinase. Mol. Cell 69, 493–504 (2018).
Alonso, J. M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003).
Li, G. J., Chen, K., Sun, S. & Zhao, Y. Osmotic signaling releases PP2C-mediated inhibition of Arabidopsis SnRK2s via the receptor-like cytoplasmic kinase BIK1. EMBO J. 43, 6076–6103 (2024).
Zhang, H., Huang, L., Hong, Y. & Song, F. BOTRYTIS-INDUCED KINASE1, a plasma membrane-localized receptor-like protein kinase, is a negative regulator of phosphate homeostasis in Arabidopsis thaliana. BMC Plant Biol. 16, 152 (2016).
Luo, J. et al. Intraorganellar calcium imaging in Arabidopsis seedling roots using the GCaMP variants GCaMP6m and R-CEPIA1er. J. Plant Physiol. 246-247, 153127 (2020).
Tian, W. et al. A calmodulin-gated calcium channel links pathogen patterns to plant immunity. Nature 572, 131–135 (2019).
Melotto, M., Underwood, W., Koczan, J., Nomura, K. & He, S. Y. Plant stomata function in innate immunity against bacterial invasion. Cell 126, 969–980 (2006).
Görlach, J. et al. Benzothiadiazole, a novel class of inducers of systemic acquired resistance, activates gene expression and disease resistance in wheat. Plant Cell 8, 629–643 (1996).
Gheysen, G., Montagu, M. V. & Zambryski, P. Integration of Agrobacterium tumefaciens transfer DNA (T-DNA) involves rearrangements of target plant DNA sequences. Proc. Natl Acad. Sci. USA 84, 6169–6173 (1987).
Nacry, P., Camilleri, C., Courtial, B., Caboche, M. & Bouchez, D. Major chromosomal rearrangements induced by T-DNA transformation in Arabidopsis. Genetics 149, 641–650 (1998).
Pucker, B., Kleinbölting, N. & Weisshaar, B. Large scale genomic rearrangements in selected Arabidopsis thaliana T-DNA lines are caused by T-DNA insertion mutagenesis. BMC Genomics 22, 599 (2021).
Ulker, B. et al. Getting the most out of publicly available T-DNA insertion lines. Plant J. 56, 665–677 (2008).
Jupe, F. et al. The complex architecture and epigenomic impact of plant T-DNA insertions. PLoS Genet. 15, e1007819 (2019).
van Butselaar, T. et al. The role of salicylic acid in the expression of receptor-like protein 23 and other immunity-related genes. Phytopathology 114, 1097–1105 (2024).
Lin, W. et al. Tyrosine phosphorylation of protein kinase complex BAK1/BIK1 mediates Arabidopsis innate immunity. Proc. Natl Acad. Sci. USA 111, 3632–3637 (2014).
Wang, C. et al. Mechanisms of calcium homeostasis orchestrate plant growth and immunity. Nature 627, 382–388 (2024).
Dindas, J. et al. Direct inhibition of phosphate transport by immune signaling in Arabidopsis. Curr. Biol. 32, 488–495 (2022).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Roach, M. J., Schmidt, S. A. & Borneman, A. R. Purge haplotigs: allelic contig reassignment for third-gen diploid genome assemblies. BMC Bioinf. 19, 460 (2018).
Alonge, M. et al. Automated assembly scaffolding using RagTag elevates a new tomato system for high-throughput genome editing. Genome Biol. 23, 258 (2022).
Marçais, G. et al. MUMmer4: a fast and versatile genome alignment system. PLoS Comput. Biol. 14, e1005944 (2018).
He, W. et al. NGenomeSyn: an easy-to-use and flexible tool for publication-ready visualization of syntenic relationships across multiple genomes. Bioinformatics 39, btad121 (2023).
Goel, M., Sun, H., Jiao, W. B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 277 (2019).
Wang, Z. P. et al. Egg cell-specific promoter-controlled CRISPR/Cas9 efficiently generates homozygous mutants for multiple target genes in Arabidopsis in a single generation. Genome Biol. 16, 144 (2015).
Cuppels, D. A. Generation and characterization of Tn5 insertion mutations in Pseudomonas syringae pv. tomato. Appl. Environ. Micro. 51, 323–327 (1986).
Ma, S. W., Morris, V. L. & Cuppels, D. A. Characterization of a DNA region required for production of the phytotoxin coronatine by Pseudomonas syringae pv. tomato. J. Bacteriol. 4, 69–77 (1991).
Su, J. et al. Regulation of stomatal immunity by interdependent functions of a pathogenresponsive MPK3/MPK6 cascade and abscisic acid. Plant Cell 29, 526–542 (2017).
Bi, G. et al. The cytosolic thiol peroxidase PRXIIB is an intracellular sensor for H2O2 that regulates plant immunity through a redox relay. Nat. Plants 8, 1160–1175 (2022).
Kim, S. I. et al. Global profiling of CPL3-mediated alternative splicing reveals regulatory mechanisms of DGK5 in plant immunity and phosphatidic acid homeostasis. Genome Biol. 26, 65 (2025).
Ngou, B. P. M., Ahn, H. K., Ding, P. & Jones, J. D. G. Mutual potentiation of plant immunity by cell-surface and intracellular receptors. Nature 592, 110–115 (2021).
Song, B. bik1 genome. figshare https://doi.org/10.6084/m9.figshare.30148612.v1 (2025).
Acknowledgements
We thank S. Han for sharing GCaMP6m transgenic line and S. Yang for original Col-0. The study was supported by Biological Breeding—National Science and Technology Major Projects (2023ZD04070) to M.H., National Natural Science Foundation of China (31761143017), National Key R&D Program of China (2021YFA1300700) and Yazhouwan National Laboratory Project (2310JM01) to J.-M.Z., funding from University of Zurich, European Research Council under the grant agreement (nos. 309858 and 773153, grants ‘PHOSPHOinnATE’ and ‘IMMUNO-PEPTALK’), Swiss National Science Foundation (grants nos. 31003A_182625 and 310030_212382) to C.Z., Collaborative Research Center funding from the German Research Foundation (grant no. CRC1101-D10) to T.N. and funding from University of Michigan to P.H. and L.S.
Author information
Authors and Affiliations
Contributions
Conceptualization: J.-M.Z., C.Z. and P.H. Methodology: B.S., S.C., L.K., Y.G. and S.M. Investigation: B.S., S.C., L.K., S.-I.K., J.F., X.L., M.L., Y.Z., H.W. and M.H. Visualization: B.S., S.C., H.W., Y.G. and S.M. Funding acquisition: J.-M.Z., C.Z., P.H., L.S. and T.N. Project administration: J.-M.Z., C.Z. and P.H. Supervision: J.-M.Z., C.Z., P.H., L.S. and T.N. Writing: J.-M.Z., C.Z., P.H., T.N., T.A.D., H.W., B.S. and S.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Alleles of bik1 generated by CRISPR and T-DNA insertions.
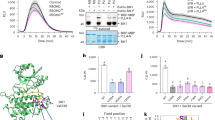
a, Diagram shows the nature of mutations of various bik1 alleles. Note that the bik1-1 allele carries a 4.3 Mb chr 1 fragment immediately before the T-DNA insertion. For CRISPR-Cas9 alleles, red numbers in parentheses indicate number of nucleotides deleted or inserted into BIK1. Graph on the left depicts predicted BIK1 polypeptide length of each allele. Note each of the bik1-1Δ/2 and bik1-6/8 alleles was obtained from two independent events. b, Deletion of the truncated coding region from bik1-1 (bik1-1Δ) failed to rescue the growth phenotype (recorded in Beijing). Scale bar: 1 cm. c, Immunoblot of selected bik1 alleles using anti-BIK1 antibodies (Ann Arbor). Total protein isolated from indicated seedlings was examined with anti-BIK1 immunoblot. Ponceau (Ponc) staining of Rubisco (RBC) indicates equal loading of protein. d, Growth phenotypes of different bik1 alleles recorded in Tübingen. Scale bar: 2 cm. e, Primary root length of bik1 alleles under normal and Pi deficient conditions (Ann Arbor). Seedlings were grown in 1/2 (Mock) for 7 days then transferred to 1/2 MS without phosphate (- Pi) or 250 μM KH2PO4 (+ Pi) for another 7 days. f, Rosette width of bik1 alleles relative to Col-0 on 1/2 MS (mock) or 1/2 MS supplemented with mannitol (Ann Arbor). Seedlings were grown in 1/2 (Mock) or 1/2 MS + 100 mM mannitol for 15 days. Violin plots in (e) and (f) show individual data points (e, n = 12-26, f, n = 29-33) and median values (centre lines). **P ≤ 0.01, ****P ≤ 0.0001 (two-way ANOVA, Tuckey’s post-test). ns, not significant. Each experiment was repeated twice with consistent results, and data from one representative experiment are shown.
Extended Data Fig. 2 BIK1 positively regulates immune responses to different immunogenic patterns.
a-c, The bik1-1, bik1-2 and bik1-3 mutants all show impaired ROS burst in response to flg22, Pep1, and chitin. 3 × pbl, pbl30/31/32 triple mutant. d-g, The BIK1 transgene fully restores Pep1- and flg22-induced ROS burst to bik1-3. Two independent transgenic lines (#21 and #32) of bik1-3 complemented with BIK1 under the control of the native promoter were tested. h, bik1-1 shows elevated whereas bik1-2 and bik1-3 show reduced ROS burst in response to nlp20. i, bik1-1, but not bik1-2 and bik1-3, shows enhanced resistance to Pst infiltrated into the leaf. The experiments in (a, h) were performed in Tübingen. All other experiments were performed in Beijing and Zurich. Data are total photo counts during the 40 min recording (a and h, n = 11-70), mean ± s.e.m. for real-time luminescence (n = 12 for b, 8 for c, g, 6 for d-f), or mean ± s.d. for bacterial titer (i, n = 8). Nine independent experiments were performed for (a and h), and data from all experiments are included, with different letters indicating significant differences at p ≤ 0.05 (Kruskal–Wallis test, Dunn’s multiple comparison test). The centre line indicates the median, the bounds of the box show the 25th and the 75th percentiles, and the whiskers indicate 1.5 × IQR. Experiments in (b-g) were repeated ≥ three times with similar results, and data from one experiment are shown. (i) shows representative data from three experiments in which bik1-2, bik1-3 and Col-0 did not differ. See Fig. 3b. **P ≤ 0.01 (one-way ANOVA, Tukey’s post-test). ns, not significant.
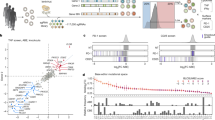
Extended Data Fig. 3 Analyses of the pbl1-1 line and generation of gene-edited pbl1 alleles.
a, b, Lack of PR1 induction in response to Pst (a) and BTH (b) in the pbl1-1 mutant. c, The lack of PR1 induction was not restored in pbl1-1 stable transgenic lines complemented with PBL1 under the control of the 35S promoter. d, Ethylmethane sulfonate mutant alleles of pbl1-2 and pbl1-5 show normal PR1 induction in response to BTH. e, The pbl1-1 line carries a previously unknown T-DNA insertion in the PR1 promoter. f, Diagram shows the nature of mutations of various pbl1 alleles. Red numbers in parentheses indicate number of nucleotides deleted or inserted into PBL1. Graph on the left depicts predicted PBL1 polypeptide length of each allele. The original pbl1-1 line contains a second T-DNA insertion in the promoter of PR1. Note we named the ethylmethane sulfonate alleles of pbl1 (cce5-1 through cce5-4)8, pbl1-2 through pbl1-6. g, Growth phenotype of bik1-9, pbl1-9, and bik1-10 pbl1-10 (recorded in Zurich). Scale bar: 2 cm. Data are mean ± s.d. ****P ≤ 0.0001 (one-way ANOVA, Tukey’s post-test, n = 3). ns, not significant. Each experiment was repeated at least three times with similar results, and data from one representative experiment are shown.
Supplementary information
Supplementary Table 1 (download XLSX )
Primers used in this study.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download PDF )
Unprocessed western blots and/or gels.
Source Data Fig. 3 (download PDF )
Unprocessed western blots and/or gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Song, B., Choi, S., Kong, L. et al. New alleles of Arabidopsis BIK1 reinforce its predominant role in pattern-triggered immunity and caution interpretations of other reported functions. Nat. Plants 12, 284–293 (2026). https://doi.org/10.1038/s41477-025-02187-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02187-3