Abstract
Glutamate dehydrogenase (GDH) resides at the crossroads of nitrogen and carbon metabolism, catalyzing the reversible conversion of L-glutamate to α-ketoglutarate and ammonium. GDH paralogs are ubiquitous across most species, presumably enabling functional specialization and genetic compensation in response to diverse conditions. Staphylococcus aureus harbors a single housekeeping GDH (GudB), whereas Bacillus subtilis encodes both a major and a minor GDH, GudB and RocG, respectively. In an unsuccessful attempt to identify an alternative GDH in S. aureus, we serendipitously discovered previously unrecognized GDH activity in two metabolic enzymes of B. subtilis. The hexameric Val/Leu/Ile dehydrogenase Bcd (formerly YqiT) catabolizes branched-chain amino acids and to a lesser extent glutamate using NAD+ as a cofactor. Removal of gudB and rocG unmasks the dual NAD(P)+-dependent GDH activity of RocA, which otherwise functions as a 3-hydroxy-1-pyrroline-5-carboxylate dehydrogenase. Bcd homologs are prevalent in free-living and obligate bacteria but are absent in most, if not all, staphylococci. Despite low sequence homology, Bcd structurally resembles the GudB/RocG family and can functionally compensate for the loss of GudB in S. aureus. Bcd is essential for the full maturation of biofilms. B. subtilis lacking GDHs exhibits severe impairments in rugose architecture and colony expansion of biofilms. This study underscores the importance of metabolic redundancy and highlights the critical role of substrate promiscuity in GDHs during biofilm development.
Similar content being viewed by others
Introduction
Glutamate dehydrogenases (GDHs) are housekeeping enzymes that connect nitrogen- and carbon-assimilation pathways by reversibly catalyzing the oxidative deamination of glutamate to produce α-ketoglutarate (α-KG) and ammonia using NAD+, NADP+ or dual NAD+-NADP+ as a cofactor. GDHs are widely distributed across eukaryotes and bacteria. They are classified by cofactor specificity1,2 and further sub-grouped by protein molecular weights3,4. NAD+-dependent GDHs, such as GudB enzymes from Staphylococcus aureus and Bacillus subtilis, are geared toward the catabolism of glutamate due to high Michaelis constant (Km) for ammonia (Fig. 1a)5. By contrast, NADP+-dependent GDHs predominantly proceed toward the reductive amination of ammonia. Small GDHs (S-GDH50 subfamily) in bacteria and eukaryotes generally adopt a homo-hexameric structure consisting of 45–50 kDa subunits, whereas large tetrameric GDHs (L-GDH115 and L-GDH180 subfamilies) in some bacteria and lower eukaryotes comprise 115–180 kDa protomers3,4.
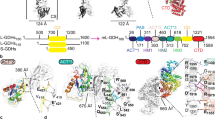
a Simplified diagram of nitrogen and carbon metabolism in S. aureus and B. subtilis. The glutamate synthases GltA-GltB convert α-ketoglutarate (αKG) and glutamine into two molecules of glutamate. The glutamine synthetase GlnA catalyzes the ATP-dependent synthesis of glutamine from glutamate and ammonia. GudB and RocG deaminate glutamate to produce αKG, which feeds into the tricarboxylic acid cycle (TCA), generating acetyl coenzyme A (acetyl-CoA) through pyruvate oxidation. B. subtilis carries both GudB and RocG, while S. aureus harbors only GudB. Additional glutamate degrading enzymes (indicated by question marks) remain unidentified. b B. subtilis produces two additional GDHs of unknown identity, as revealed by in-gel activity staining. Independent double knockouts (∆gudB∆rocG and ∆rocG∆gudB) show identical results. c Denaturing PAGE-immunoblot using the B. subtilis GudB antibody confirms proper gudB expression in the GudB-proficient strains. d Genetic knockouts confirm Bcd and RocA contributions to GDH activity. Both enzymes use NAD+ as the coenzyme (left) but only RocA utilizes NADP+ as the electron acceptor (right). e Zymogram analysis of a single bcd mutant and bcd-complemented strain rules out secondary mutations, substantiating Bcd as an alternative GDH. Zymography using NAD+ and glutamate substrate is shown. GDH activity of GudB and RocG was not detected in this assay, possibly due to the sequestration of GudB by GltAB and low transcription of rocG in LB and MSgg media.
GDHs have significant physiological importance and biotechnological applications. Like other amino acid dehydrogenases, GDHs are catalysts for high-value amino acids used in the pharmaceutical and food and feed additives industries6. The GDH enzyme immunoassay (EIA) is a first-line diagnostic test for Clostridioides difficile infections due to the high abundance of GDH in stool samples7. In bacteria, removing GDH leads to growth defects8,9,10,11,12, aberrant biofilm development13,14, attenuated virulence and host colonization15,16,17, and sensitivity to antibiotic and environmental stress17,18,19,20.
Enzyme promiscuity is central to metabolic diversity21. Duplication of GDH genes is well known in humans, yeast and bacteria. Human hGLUD1 and hGLUD2 differ significantly in tissue distribution and allosteric regulation22. In Firmicutes (synonym Bacillota), undomesticated B. subtilis strains, including NCIB3610, encode two functional GDH paralogs, GudB and RocG5,23,24, which are differentially regulated14,25. However, laboratory strains of B. subtilis, such as strain 168, synthesize a non-functional GudB due to the insertion of an 18-bp long direct repeat in gudB, leading to the duplication of 3 amino acids in the active center of the GDH5,23. By contrast, GudB is the only GDH documented in the opportunistic pathogen S. aureus (Fig. 1a).
We previously reported that deletion of gudB in S. aureus does not completely abolish cellular GDH activity, suggesting the existence of a second GDH. We found that a 20 kDa protein of unknown function, YwlG, compensates for the loss of gudB, possibly by activating a hitherto unknown GDH (Fig. S1a)26. YwlG lacks the N-terminal and C-terminal substrate- binding domains and does not appear to possess intrinsic GDH activity, despite folding into a hexamer and structurally resembling the B. subtilis GudB26,27. Moreover, YwlG-dependent stimulation of cellular GDH is physically inhibited by a ribosome hibernation factor (Hpf). YwlG-Hpf sequestration reciprocally prevents the formation of ribonuclease-resistant, hibernating 100S ribosomes (Fig. S1a)26. We hypothesize that YwlG promotes cellular GDH activity by interacting with an unidentified protein in the absence of Hpf. In the process of identifying this unknown protein in S. aureus, we fortuitously discovered that an annotated Val/Leu/Ile dehydrogenase, Bcd in B. subtilis NCIB3610 displays a previously unrecognized NAD+-specific GDH activity. Even more surprisingly, deletion of both gudB and rocG unmasks the NAD(P)+-dependent GDH activity of RocA, a 3-hydroxy-1-pyrroline-5-carboxylate dehydrogenase. Despite Bcd’s absence in S. aureus, B. subtilis Bcd is functionally interchangeable with S. aureus GudB. In B. subtilis, deletion of bcd in combination with either gudB or rocG deletion impairs biofilm maturation. This study reveals the substrate promiscuity of amino acid dehydrogenases and suggests that many more redundant GDH enzymes remain to be discovered in all organisms.
Results
Discovery of two new glutamate dehydrogenases (Bcd and RocA) in B. subtilis NCIB3610
We employed an established native gel activity staining-based zymography approach28,29 to identify the second GDH in total cell lysates of wild-type (WT) S. aureus and its ∆hpf null mutant. GDH activity was visualized by purple-stained protein bands, resulting from the dehydrogenase-mediated reduction of tetrazolium salts in the presence of glutamate substrate, a co-factor, and an electron carrier. In-gel zymography provides semi-quantitative and qualitative information about the native molecular weight of different GDH variants. Despite extensive experimental optimization and testing across multiple genetic backgrounds, we did not detect any GDH activity in S. aureus beyond the known GudB (Fig. S1b). Intriguingly, S. aureus GudB complexes migrate as a trimer ( ~ 135 kDa) on native PAGE, contrasting with the hexameric forms reported for other GudB homologs14. The identity of the trimeric GudB was confirmed by Western blots using a ∆gudB mutant as a control and a hexameric YwlG ( ~ 120 kDa) as the size marker (Fig. S1b). To rule out technical issues, we analyzed cell lysates from B. subtilis NCIB3610 LB or MSgg cultures as a control. On the zymogram, a GDH protein band corresponding to a hexameric GudB or RocG ( ~ 240 kDa) was detected in the zymogram. However, this multimer persisted upon gudB and rocG single deletion (Fig. 1b) and did not cross-react with the anti-GudB antibody, which recognizes both native and denatured GudB (Figs. S1c, and 1c). The absence of GudB activity in the WT B. subtilis is probably due to sequestration by GltA-GltB13. Further analysis of two independently constructed ∆gudB∆rocG double knockouts revealed that the 240-kDa species was still present, along with an unexpected appearance of a lower molecular weight GDH band. These observations raised questions about the identities of the two protein species detected (Fig. 1b).
In-gel mass spectrometry analysis identified the lower band as either RocA or PutC, while the upper band corresponded to Bcd (Fig. S2). None of these proteins are previously known to exhibit GDH activity. RocA is a 56-kDa 3-hydroxy-1-pyrroline-5-carboxylate dehydrogenase, and PutC is a 56-kDa 1-pyrroline-5-carboxylate dehydrogenase. Both proteins can functionally replace each other in the utilization of proline and arginine30. Bcd, on the other hand, is a 40-kDa Val/Leu/Ile dehydrogenase31, and its deletion triggers premature germination due to accumulation of valine32. Deletion of bcd, rocA and putC in the ∆gudB∆rocG double mutant confirmed Bcd as the ~240-kDa complex and RocA as the smaller oligomer. Both proteins were able to use NAD+ as the electron acceptor to degrade glutamate (Fig. 1d). RocA, but not Bcd, could also utilize NADP+ as a cofactor; however, its GDH activity was only apparent in the absence of both GudB and RocG (Fig. 1d). A ∆bcd deletion mutant and its chromosomally encoded Bcd complement driven by its native promoter further validated the identity of the native Bcd complex (Fig. 1e). In summary, Bcd is an alternative NAD+-dependent GDH enzyme in B. subtilis, and NAD(P)+-dependent GDH activity of RocA is manifested in the absence of both GudB and RocG through an as-yet-unknown mechanism.
Recombinant Bcd forms a hexamer and exhibits Glu/Val/Leu/Ile dehydrogenase activities
B. subtilis Bcd migrates as a hexamer on native PAGE (Fig. 1) whereas Lysinibacillus sphaericus Bcd assembles as an octamer33. To confirm the oligomeric state and active form of Bcd, we recombinantly purified the N-terminally His-tagged WT Bcd protein and its catalytically compromised mutants (K68A, K80A, D115A) from E. coli. Only the WT and K80A variants were amenable to native purification (Fig. 2a, b). Both proteins eluted as hexamers on a size-exclusion chromatography (Fig. 2c, Data S1). Bcd is annotated as a branched-chain amino acid dehydrogenase. Using zymography and various L-amino acids as substrates, we confirmed that the recombinant WT Bcd exhibits NAD+-dependent Val/Leu/Ile dehydrogenase activities and GDH activity in vitro. The K80A mutant showed a complete loss of dehydrogenase activity (Fig. 2d). Furthermore, in the presence of NAD+ and L-glutamate, measurable catabolic GDH activity was detected in the hexameric WT Bcd fractions collected from the size-exclusion chromatography, relative to a NAD(P)+-dependent bovine liver GDH control, whereas its Bcd(K80A) mutant had no activity (Fig. 2e). Similarly, untagged Bcd expressed in an S. aureus ∆gudB mutant demonstrated NAD+-specific GDH and valine dehydrogenase activities (Fig. S3). These results demonstrate that the active form of B. subtilis Bcd is predominantly a hexamer capable of catabolizing multiple substrates.
a Recombinant WT Bcd and its catalytically impaired mutants were purified by cobalt TALON affinity chromatography. b Structural prediction of Bcd monomer by Alphafold shows the catalytic sites. c WT Bcd and its K80A mutant form a hexameric assembly, as observed in size-exclusion chromatography. d Zymography shows that WT Bcd exhibits Glu/Val/Leu/Ile dehydrogenase activity, while the K80A mutant is catalytically inactive despite forming a hexamer. Zymograms followed by Coomassie Blue staining show equal loading and differing GDH activity. e In vitro GDH activity of purified Bcd hexamers collected from (c). WT Bcd exhibits measurable GDH activity, though significantly lower than that of the commercially available NAD(P)+-dependent bovine liver GDH control (CTRL). f Growth kinetics of B. subtilis mutants in CE minimal medium containing glucose and glutamate or glutamate alone as the carbon and nitrogen source. The ΔgudB∆rocG double and ΔgudB∆rocG∆bcd triple mutants grew poorly due to their inability to utilize glutamate. Overexpression of bcd partially rescues this growth defects. Color-coded traces represent the mean OD600 and shaded areas indicate the standard deviation from three independent biological replicates.
The growth of the B. subtilis∆gudB∆rocG mutant is severely impaired in CE minimal medium containing glutamate as the sole carbon source5,34. We found that overexpression of Bcd, driven by a constitutive rpsD promoter, could partially restore the growth of the ΔgudB∆rocG mutant to the level observed in the ∆gudB single mutant, although not to the levels of the WT or ∆rocG single mutant (Fig. 2f). This partial restoration of growth supports the involvement of Bcd in glutamate catabolism.
Bcd homologs are widespread across bacteria but absent in S. aureus
We next investigated the phylogeny and conservation of Bcd. Using OrthoFinder 2.5.5, we analyzed 136 bacterial genomes spanning a broad range of the bacterial clade within the tree of life. OrthoFinder identified hierarchical orthologous groups (HOGs), enabling the examination of orthology at various levels of the bacterial species tree. For example, a HOG at the root of the bacterial tree encompasses all genes descended from a single ancestral gene in the Last Bacterial Common Ancestor (LBCA). Conversely, a HOG at the Bacillus genus level includes all genes derived from the last common ancestor of all Bacillus species. At the root of the bacterial tree, RocG, GudB, and Bcd were grouped into a single HOG, suggesting they likely originated from a single ancestral gene near the LBCA. At the phylum level, Bcd was assigned to a distinct HOG (HOG_bcd) separate from GudB and RocG (HOG_gudB/rocG). GudB and RocG remained within the same HOG up to the species level, indicating a relatively recent divergence. Proteins from both HOG_bcd and HOG_gudB/rocG were identified in diverse bacterial phyla, with a notable absence of Bcd in most staphylococcal species, including S. aureus. The co-existence of gudB/rocG and bcd was common in Bacillus sp, Bdellovibrio sp., Halalkalibacterium sp., and Deinococcus sp. Additionally, bcd was prevalent in pseudomonads and obligate pathogens (Fig. 3a, Data S2).
a Phylogenetic tree shows that Bcd and GudB/RocG form two monophyletic clades. Multiple Bcd and GudB/RocG paralogs can be found in a single species, e.g., H. halodurans, which are color-coded accordingly. b Taxonomic distribution of Bcd homologs. The pie-chart size and black section represent the number of bacterial genomes and the frequency of Bcd presence, respectively. Terrabacteria mostly include Gram-positive bacteria such as Bacillota (Firmicutes); FCB phylum consists of Fibrobacteriota, Chloraobiota, and Bacteriodota. Less well-defined superphyla are loosely categorized by color. c Structural superposition of B. subtilis GudB1 structure (PDB 3k8z) with predicted structures of B. subtilis Bcd and S. aureus GudB reveals high resemblance among these GDHs. d Plasmid borne-Bcd partially restores GDH activity in a S. aureus ∆gudB mutant. EV, empty vector. Statistical significance was determined using Student’s t-test. *P < 0.05, ***P < 0.001, ns = not significant. e Growth defects of the S. aureus ∆gudB mutant are fully complemented by WT Bcd and its cognate GudB but not by the Bcd(K80A) mutant. Student’s t-test (n = 4), ***P < 0.001. f Western blot analysis shows endogenous GudB and plasmid-borne GudB expression in S. aureus. Anti-GudB does not cross-react with Bcd despite structural similarity. An asterisk indicates non-specific band.
To further explore Bcd orthology. We expanded our analysis to 26,373 genomes from the ProGenomes database35, which provides a non-redundant collection of high-quality genomes, each representing a single SpecI species cluster. A reciprocal BLAST approach was employed using Bcd from B. subtilis as the query in a forward BLASTp against all proteomes. The top hit from each target proteome was subsequently blasted back against the B. subtilis genome. If the Bcd query protein emerged as the best hit in the reciprocal BLAST, the corresponding target proteome hit was designated as a putative Bcd ortholog. Consistent with the initial analyses, Bcd homologs were predominantly found in the phyla Bacillota, Gemmatimondota, Bacterioidota, Pseudomonadota, Bdellovibrionota, Deinococcota, Myxococcota, and Chlamydiota, particularly in bacteria with free-living and obligate lifestyles (Fig. 3b).
B. subtilis Bcd rescues the growth defects of an S. aureus ∆gudB mutant
Bcd shares only 27–28% overall protein sequence identity with GudB and RocG, which is significantly lower than the identity between GudB and RocG (74–75%). At the structural levels, Alphafold modeling36 showed that B. subtilis Bcd is nearly identical to S. aureus GudB, with a root mean square deviation (r.m.s.d) of 1.49 Å (Fig. 3c). Plasmid-borne expression of Bcd restored the cellular GDH activity in the S. aureus ∆gudB mutant, although not to the levels achieved by expressing the cognate GudB (Fig. 3d). The S. aureus ∆gudB exhibits growth defects under various laboratory conditions, including lysogeny broth (LB) medium8,9. GudB is essential for growth following glucose consumption. A ∆gudB mutant prematurely enters stationary phase upon glucose depletion due to its inability to utilize glutamate as an alternative carbon source8. Indeed, we observed that deletion of gudB significantly hindered S. aureus growth in LB. These growth defects were fully complemented by plasmid-based expression of either the cognate GudB or heterologous B. subtilis Bcd (Fig. 3e). The modest difference observed between the GudB-complemented (blue trace) and Bcd-complemented (orange trace) strains during stationary phase may be partially due to the lower GDH activity of Bcd compared to GudB (Fig. 3d) to efficiently use glutamate as an alternative carbon source, leading to carbon limitation. In contrast, a catalytically inactive Bcd(K80A) failed to restore growth to the WT level (Fig. 3e, f). These findings confirm that B. subtilis Bcd can functionally replace S. aureus GudB in glutamate degradation.
Bcd is required for biofilm maturation in B. subtilis
The homeostasis of glutamate, glutamine and ammonium is known to regulate biofilm development11,13,37,38,39. Gradients of glutamate and ammonium influence the oscillatory dynamics of biofilm expansion through GDH-dependent ammonium synthesis within the interior of the colony, utilizing glutamate diffused from the periphery (Fig. 4a, left panel)37,38. To investigate the role of Bcd in biofilm formation, we examined the morphological and structural changes of colony biofilm on solid agar and floating pellicle biofilm at the air-liquid interface. Under biofilm-inducing conditions40, B. subtilis NCIB3610 forms rugose colony biofilms on LBGM agar, characterized by concentric rings of cell differentiation. Wrinkles form as a consequence of localized cell death, coupled with the stiffness and hydrophobicity provided by the exopolysaccharides- and proteins-containing extracellular matrix41. All mutant strains, with the exception of the ΔrocG∆bcd mutant, displayed comparable growth in LBGM medium relative to the WT (Fig. S4). GudB and RocG are considered major and minor GDH in strain NCIB3610, respectively. While the single ∆bcd mutant did not exhibit measurable defects in macrocolony formation, combining the ∆bcd deletion with ∆gudB deletion completely abolished the rugose architecture of the colonies. Additionally, combining ∆bcd deletion with ∆rocG deletion modestly altered the corrugated surface of the colonies. A triple mutant (∆gudB∆rocG∆bcd) not only abolished the wrinkly morphology but also exhibited dysregulated colony expansion. In general, biofilm phenotypes were more pronounced when both gudB and bcd knockouts were combined. The chromosomally encoded bcd from the sacA locus fully restored the rugose texture and corrected colony expansion of the triple mutant, but only partially restored radial patterns to WT levels (Fig. 4a). Frilled colony and appendage-like cell growth were observed at the edges of colonies in the ∆gudB∆rocG and ∆rocG∆bcd double mutants, suggesting the emergence of suppressors, which are commonly found in gudB and rocG mutants5,34,42. In line with the colony biofilm phenotype, the ∆gudB∆bcd and ∆gudB∆rocG∆bcd mutants were impaired in forming the corrugated architecture in the pellicle, instead producing a powdery texture that could be complemented by chromosomally encoded bcd (Fig. 4b). Bcd likely plays a secondary role in glutamate degradation (Fig. 2d-2f) and Val/Leu/Ile dehydrogenase activity of Bcd may partially contribute to the observed phenotypes. Nevertheless, the biofilm phenotypes confirm the functional redundancy of Bcd, GudB and RocG in glutamate degradation, highlighting the importance of GDH activity in biofilm development.
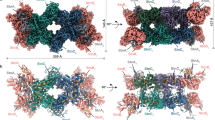
a Inactivation of the major GDH (GudB) and Bcd abolishes the wrinkled morphology of biofilm colonies. Removing gudB, rocG and bcd together results in aberrant biofilm expansion. (Left) A schematic of glutamate metabolism shows that biomass production requires external glutamate and ammonium generated internally via GDH-mediated glutamate degradation. b Pellicle biofilms at the air-liquid interface lose their rugose structures in the absence of GudB and Bcd. These defects are exacerbated in the ΔgudB∆rocG∆bcd triple mutant. Chromosomal expression of bcd from the sacA locus under its native promoter provides varying degrees of complementation in different mutants.
Discussion
Here we report a novel dehydrogenase activity for NAD+-specific Val/Leu/Ile dehydrogenase (Bcd/YqiT) in deaminating glutamate. We also identify a previously unrecognized NAD(P)+-dependent GDH activity of RocA, which is revealed when both gudB and rocG are deleted. The B. licheniformis RocA homolog (PDB 3RJL), a 3-hydroxy-1-pyrroline-5-carboxylate dehydrogenase, is structurally distinct from Bcd, GudB, and RocG. The additional GDH activity of RocA represents a new example of compensatory redundancy. Genetic compensation can also occur by establishing a new pathway to restore glutamate degradation through the overexpression of aspartase AnsB, allowing the ∆gudB∆rocG double mutant to feed glutamate into the TCA cycle25,43. Furthermore, a loss-of-function mutation in the glutamate synthase gltB of the ∆gudB∆rocG double mutant also prevents glutamate accumulation5.
Glutamate is one of the most abundant cytoplasmic anions and the major nitrogen reservoir in bacteria, with cellular concentrations reaching up to 150 mM44. Glutamate stabilizes interactions among biomolecules, acts as a counterion to potassium, buffers intracellular pH, maintains osmolarity, and serves as an amino group donor for at least 37 transamination reactions13,44,45,46. Glutamate overload is toxic for B. subtilis under conditions of GDH deficiency and excessive glutamate uptake and synthesis5,42,47. GudB and RocG, and to a lesser extent, Bcd and RocA are metabolically redundant in degrading glutamate, ensuring robustness against fluctuations in glutamate and gdh mutations. B. subtilis gudB is constitutively expressed, while rocG expression is repressed by glucose and activated by arginine14. gudB, rocG, bcd, and rocA are differentially expressed under various conditions14,31,48,49, allowing fine-tuned glutamate catabolism based on cellular needs.
GudB, RocG, Bcd and RocA are multi-tasking enzymes, performing regulatory role in addition to degrading multiple substrates. GudB and RocG act as trigger enzymes that not only carry GDH activity but also directly interact with the GltC transcription factor to regulate glutamate25,48,50,51 synthesis. RocA recognizes both L-glutamate 5-semialdehyde and L-glutamate substrates and uses NAD+ and NADP+ as electron acceptors. Bcd degrades branched-chain amino acids as well as glutamate (Fig. 2), with Bcd-mediated valine catabolism regulating the timing of germination in B. subtilis32. Our phylogenetic analysis suggests that the gudB/rocG and bcd clades may have diverged evolutionarily to expand their substrate repertoire (Fig. 3). Notably, combinations of point mutations in the substrate-binding pocket of leucine dehydrogenase and GDH can switch amino acids specificity in vitro52,53,54. The prevalence of Bcd in intracellular pathogens and bacteria inhabiting diverse aquatic and terrestrial environments suggests a selective growth advantage due to substrate ambiguity in these niches.
Niche specialization of GDH could partially explain our inability to identify a second GDH in S. aureus. The expression and activity of this alternative GDH in S. aureus may be too low14, inhibited by other factors13, or inactivated by post-translational modifications55 under the conditions we tested. For instance, B. subtilis GudB activity is repressed by the heterodimeric GltAB in non-fractionated samples, and rocG is normally transcriptionally repressed13, which may account for the absence of GudB- and RocG-mediated GDH activity on the zymograms (Fig. 1). Zymography has inherent limitations28, such as resolution and sensitivity issues, and our current electrophoresis protocols preclude the detection of multimer larger than 1200 kDa, such as the 1.6 MDa GltAB-GudB complex. Moreover, GltAB-GudB interaction inhibits GudB activity, potentially explaining a lack of GudB’s GDH activity on the zymogram from LB and Msgg cultures.
Mutations in gudB impair colony biofilm formation11,13,14. Consistent with previous reports, a ΔgudB mutant exhibits modest but measurable biofilm developmental defects. We find that inactivating both bcd and gudB severely diminishes the rugose phenotype, and inactivating all three GDHs produces rapidly growing smooth colonies (Fig. 4). These additive biofilm defects confirm the overlapping role of GudB, RocG and Bcd. GDH activity may influence biofilm maturation in multiple ways: (i) Biofilm oscillations are beneficial for cells to cope with nutrient deprivation and chemical stress. GDH insufficiency disrupts normal growth oscillations, which involve gradients of glutamate and ammonium37,38,56. (ii) Acidic and alkaline conditions hinder biofilm formation57. GDH deficiency can lead to high glutamate and low pH. (iii) Unlike strain B1, whose biofilm matrix is primarily composed of hydrophilic poly-γ-glutamate (γ-PGA), NCIB3610’s matrix contains very little γ-PGA58,59. However, the possibility that GDH deficiency increases γ-PGA levels and alters biofilm architecture cannot be entirely excluded.
Methods
Bacterial strains, plasmids, primers and chemicals
Strain USA300 JE2 is a community-associated methicillin-resistant Staphylococcus aureus (CA-MRSA, GenBank CP000255). The JE2 mutant derivatives carry a bursa aurealis transposon insertion were acquired from ATCC-managed BEI Resources Repository and were backcrossed to isogenic JE2 background by Φ11 phage transduction60. The in-frame ∆gudB::Km deletion mutant was created using standard allelic-exchange recombination61. A 2-kb flanking region of S. aureus gudB (locus SAUSA300_0861) was PCR amplified with the primer pairs P1747/P1748 and P1749/P1750 via 2-step PCR using JE2 genomic DNA as the template. The product was digested with SacI and SalI and cloned into the same sites of pBT2. The resulting pBT2∆gudB was digested with SmaI, dephosphorylated, and ligated to the blunt-ended 1.6-kb kanamycin (Km) resistance cassette that was released and end-repaired from pBTK by KpnI and PstI digestion. The resulting construct pBT2∆gudB::Km was propagated in E. coli DC10B, and the plasmid reisolated, electroporated into JE2 and selected at 30 °C on agar plates supplemented with 10 µg/mL chloramphenicol. The integrant was further selected by a 43 °C temperature upshift on chloramphenicol-containing agar plates. The homologous recombinant was resolved by 30 °C passages and cycloserine enrichment following the published procedures62. Mutant strains were verified by Sanger sequencing and Western blots.
Restriction-free cloning was used to construct pEPSA5_GudB, pEPSA5_Bcd, and pMCSG7 derivatives63. To construct N-terminally His-tagged bcd for protein purification, primers P2139/P2140 were used to assemble the DNA fragment onto pMCSG7. Primers used to construct pEPSA5_GudB and pEPSA5_Bcd were P1753/P1754 and P2130/P2131, respectively. Site-directed mutagenesis was performed using Quikchange mutagenesis kit (Agilent Genomics) using P2133/P2134 (K68A), P2135/P2136 (K80A), and P2137/P2138 (D115A).
B. subtilis NCIB3610 (ATCC 6051 or DSM10, GenBank CP020102) is an undomesticated biofilm former that is poorly competent24,64. Gene deletions mutants in NCIB3610 were made using phage transduction and a commercially available gene knockout library in B. subtilis 16865,66. SPP1 phage transduction was completed by moving gene knockouts from the donor 168 strain into the receiving NCIB3610 strain as described in detail previously67. Double mutants were made by utilizing the Cre recombinase recognition sites, loxP, built into the commercially available gene knockout library that allowed for removal or curing of the antibiotic cassette65. To remove the antibiotic cassette, the strain was transformed with a temperature sensitive plasmid that contained a constitutively expressed Cre recombinase using the previously described transformation method for NCIB3610 with all incubation steps carried out at 30 °C68. The curing of the antibiotic cassette was then completed using the described procedure65. Cured mutants were then used as the recipient strain for sequential phage transduction. All genotypes were confirmed with specific flanking primers for all genes and Sanger sequencing.
All strains, oligos and plasmids are listed in Tables S1-S2. Unless otherwise noted, S. aureus cells were grown aerobically at 37 °C in tryptic soy broth (TSB, BD Difco #211822), Msgg [5 mM potassium-phosphate buffer (pH 7) 100 mM MOPS, 2 mM MgCl2 ∙ 6H2O, 700 µM CaCl2, 50 µM MnCl2, 100 µM FeCl3 ∙ 6H2O, 1 µM ZnCl2, 2 µM thiamine-HCl, 0.5%(v/v) glycerol, 0.5% (w/v) monosodium glutamate]69 or Lysogeny Broth (LB, BD Difco #244610) at a 5:1-10:1 tube- or flask-to-medium ratio with a 1:100 dilution of an overnight seed culture. B. subtilis and E. coli were grown in LB. When necessary, erythromycin, chloramphenicol, kanamycin, ampicillin, xylose, IPTG were used at 5 µg/mL, 10 µg/mL (7.5 µg/mL in B. subtilis), 75 µg/mL (10 µg/mL in B. subtilis), 100 µg/mL, 10 mM, and 0.5 mM, respectively. Unless otherwise noted, all chemicals and oligonucleotides were purchased from Millipore Sigma-Aldrich and IDT DNA, respectively.
Chromosomal bcd complementation
The constitutively expressed rpsD promoter-driven Pc-bcd plasmid and native promoter-driven Pn-bcd plasmid were constructed by amplifying 500-bp of the rpsD or operonic bkdR-ptb-bcd promoters from the NCIB3610 genome and assembled onto pSac-Cm (or ECE174) using a restriction-free cloning method63,69. Primers used for the assembly were P2149/P2150 and P2154/P2155 (Table S2). The assembled plasmids were transformed into and integrated into NCIB3610 sacA locus using a natural competence protocol previously established68. Transformants were selected on LB agar containing 7.5 µg/mL chloramphenicol. Low efficiency of transformation was achievable despite the presence of ComI-containing plasmid pBS32 in the parental NCIB3610 strain68, which was verified by whole-genome sequencing. All genotypes were confirmed with specific flanking primers on sacA and bcd sequences.
His-tag affinity purification of Bcd recombinant proteins and size-exclusion chromatography
Overnight E. coli seed cultures were diluted 1/100-fold in 200 mL LB supplemented with 100 µg/mL ampicillin and incubated at 37 °C until to an OD600 = 0.5–0.6. Cells were induced with 0.5 mM IPTG for 3-4 hr. Then cultures were centrifuged at 3220×g for 15 min at 4 °C to pellet cells. Supernatant was discarded, and cells were resuspended in 5 mL of Buffer I (50 mM Tris-Cl (pH 7.5), 150 mM NaCl, 1 mM phenylmethylsulfonyl fluoride (PMSF)) containing 1 mM imidazole and lysed on a FastPrep-24 instrument using Lysing Matrix B beads (MP Biomedicals # 116911500). Following lysis, samples were centrifuged at 4 °C, 20,817×g for 5 min to remove cell debris. Approximately 5 mL of cell lysates was incubated with 0.5 ml TALON metal affinity beads (Takara, #635503) on a tube rotator at 4 °C for 1 hr. The column was washed extensively with Buffer I supplemented with 20 mM imidazole (7 ×1 mL), and proteins were eluted with Buffer I supplemented with 50–300 mM imidazole.
For size-exclusion chromatography, approximately 1.8 mg of Bcd(WT) and 2 mg of Bcd (K80A) proteins were loaded on HiPrepTM 16/60 Sephacryl S-100 HR column (Cytiva) on an ÄKTA Go chromatography system (Cytiva) and equilibrated with Buffer II [50 mM Tris-Cl (pH 7.5), 150 mM NaCl]70. To estimate the size of Bcd peaks a Gel Filtration Standard (BioRad #151-1901) was used.
Zymography and in-gel mass spectrometry
Zymography was performed following the published protocol29. Fifty microliters of overnight cultures were diluted in 25 ml of LB or Msgg (B. subtilis) and TSB (S. aureus). The cultures were grown for 24 h at 37 °C and 10 mL of culture were centrifuged at 3220×g for 15 min at 4 °C. Cell pellets were resuspended in 1 ml of sterile water and lysed on a FastPrep-24 instrument using Lysing Matrix B beads (MP Biomedicals). Following lysis, samples were centrifuged at 20,817×g at 4 °C for 5 min. Fifty microliters of the cell-free extracts were mixed with equal volume of 2×Native Sample Buffer (Invitrogen, catalog # LC2673) and 10 μl were analyzed on a 4–12% Tris-glycine native gel (Invitrogen #XP04122BOX) in Tris-Glycine Native Running Buffer (Invitrogen #LC2672) at constant 200 V for 80 min. The gels were rinsed in ddH2O for 5 min before incubation in 30 ml activity solution in the dark for 1 hr or overnight at 37 °C. The activity solution comprised of 0.5 mM NADP+ (Sigma #10128031001) or NAD+ (Sigma #NAD100-RO), 50 mM Tris-HCl (pH 8.0), 20 mM L-glutamate (Sigma #49601-100 G), 0.3 mg/ml Nitro Blue Tetrazolium (Sigma #N5514-10TAB), and 0.05 mg/mL phenazine methosulfate (Sigma #P9625-1G). For Leu/Val/Ile dehydrogenase activity, L-glutamate was substituted with 20 mM L-valine (Sigma #V0500-100G, L-leucine (Sigma #L8000-100G) or L-isoleucine (Sigma #I2752-100G). Dehydrogenase activity appeared as a dark purple against a clear background after extensive wash in ddH2O to remove residual yellow dyes. The gels were subsequently stained with Gelcode Blue Safe Stain (ThermoFisher #PI24956) to ensure equal loading. A HMW Native protein marker kit (Cytvia #17044501) was used to estimate Bcd or GudB oligomerization. For mass spectrometry, protein bands of interest were excised from the zymogram gel and subjected to in-gel trypsin digestion at 37 °C. Digested peptides were extracted with trifluoracetic acid (TFA) extraction buffer, desalted by passing through the C-18 Zip-tips71. The digested peptides were then separated on a nano-flow Ultimate 3000 and analyzed by Orbitrap Exploris high-resolution mass spectrometer. Protein identification using Nano LC-MS/MS was performed by Applied Biomics (Hayward, CA)
In vitro glutamate dehydrogenase (GDH) assays
GDH Activity Assay was performed as previously published26. Briefly, S. aureus and B. subtilis were grown aerobically in 25 ml of TSB and LB, respectively for 24 h at 37 °C. One milliliter of bacterial cultures was collected, resuspended in sterile water, and lysed on a FastPrep-24 instrument using Lysing Matrix B beads (MP Biomedicals), and 10 μL cell lysates were immediately used in the reactions. GDH activity was measured by the amount of NADH production upon glutamate consumption using a colorimetric (450 nm) kit according to manufacturer’s procedures (Abcam # ab102527 or Sigma #MAK099). To determine GDH activity of the purified proteins, 4.5 μM of Bcd proteins and 0.8 μM of positive control bovine liver GDH (Sigma #G2626) were used. One unit of GDH is defined as the amount of enzyme that generates 1.0 μmol of NADH per minute at pH 7.6 at 37 °C.
Native Gel immunoblots
Fifteen microliters of the cell-free extracts prepared as described in “Zymography” were analyzed on a 4–12% Tris-glycine native gel (Invitrogen). Proteins were transferred onto a 0.22 μm PVDF membrane (Bio-Rad #1704156) using a Trans-Blot Turbo Transfer System (Bio-Rad). Membranes were immunolabelled using epitope-specific rabbit polyclonal anti-GudBBs (1/1,000 dilution)13 and anti-YwlG (1/4000 dilution)26. HRP-conjugated anti-IgG secondary antibody (1/15,000 dilution) was from Cytiva (#NA9120) and SuperSignal™ West Dura chemiluminescence substrate (Thermo Scientific #34075) was used for signal detection. Images were acquired using iBright FL1500 system (ThermoFisher).
Phylogenetic analysis
To determine orthologous relationships between Bcd, RocG and GudB, we used OrthoFinder version 2.5.572 to analyze 136 bacterial genomes representing a wide span of the bacterial clade in the tree of life. A phylogenetic tree was constructed using all proteins from both bcd-HOG and gudB/rocG-HOG. Amino acid sequences were aligned with MUSCLE 5.1 using default settings, and an unrooted tree was inferred with FastTree version 2.1.11.
The analysis of bcd orthology was expanded to 26,373 genomes from the ProGenomes database. This dataset comprises a non-redundant collection of high-quality genomes, with one representative genome for each SpecI species cluster. A reciprocal BLAST approach was employed, using bcd from Bacillus subtilis 168 as the query in a forward BLASTp against all other proteomes. The top hit from each target proteome was subsequently blasted back against the Bacillus subtilis genome. If the bcd query protein was identified as the best hit in the reciprocal BLAST, the corresponding target proteome hit was designated as a putative bcd ortholog. Custom python scripts were used to run and analyze results from Orthofinder and reciprocal blast and are available on GitHub at https://github.com/JonWinkelman/reciprocal-blast (Trestle Biosciences, LLC). Interactive Tree of Life (iTol) tool and Adobe Illustrator were used for visualizing and annotating phylogenetic trees.
In silico analyses
The predicted structures for S. aureus GudB and B. subtilis Bcd were made using AlphaFold 373 and rendered with PyMOL Molecular Graphics System (version 3.0 Schrödinger, LLC), yielding a per-residue model confidence score (pLDDT) of >70–90 and >90 for GudB and Bcd, respectively. To discern relationships to known structures, the predicted structures were submitted to the DALI server (v5)74 to compare against available structures.
Biofilm assays and growth kinetics
Assays of biofilm formation were adapted from refs. 40,75. Single colonies of B. subtilis strains were cultured overnight in 3 mL of LB medium at 37 °C. The seed cultures were diluted in fresh 25 ml LB supplemented with 0.1 mM MnSO4•4H2O and 3% (v/v) glycerol (LBGM) to an OD600 of 0.1. To induce pellicle formation, 20 mL subculture were transferred to sterile 50-ml Pyrex bottles and incubated at 25 °C without agitation for 48 h before the images were recorded. For colony biofilm formation, 2 μL of the OD600 0.1 cultures were spotted onto the LBGM agar. Plates were incubated at 37 °C for 24 h prior to imaging. All experiments were repeated at least three times.
For growth kinetics, strains were grown to OD600 0.8-1.0 in LB, spun down, resuspended in 1×PBS and adjusted to OD600 0.8. Two microliters of cell suspension were seeded into 200 μL of CE minimal medium, LB, or LBGM40 in a 96-well microplate. Bacterial growth was recorded at 37 °C with 150 rpm shaking on a TECAN SPARK multimode plate reader equipped with a humidity cassette. The CE minimal medium comprises [70 mM K2HPO4, 30 mM KH2PO4, 25 mM (NH4)2SO4, 0.5 mM MgSO4, 10 μM MnSO4, 50 mg/L tryptophan] with and without 1 g/L glucose and 8 g/L potassium glutamate5.
Statistical analysis
Statistical analyses were performed using GraphPad Prism 10.1.1. The performed statistical tests are described in the figure legends.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Code availability
Custom python scripts used for phylogenetic analysis are publicly available on GitHub: https://github.com/JonWinkelman/reciprocal-blast.
References
Hudson, R. C. & Daniel, R. M. L-glutamate dehydrogenases: distribution, properties and mechanism. Comp. Biochem Physiol. B. 106, 767–792 (1993).
Brunhuber, N. M. & Blanchard, J. S. The biochemistry and enzymology of amino acid dehydrogenases. Crit. Rev. Biochem Mol. Biol. 29, 415–467 (1994).
Lázaro, M. et al. 3D architecture and structural flexibility revealed in the subfamily of large glutamate dehydrogenases by a mycobacterial enzyme. Commun. Biol. 4, 684 (2021).
Miñambres, B., Olivera, E. R., Jensen, R. A. & Luengo, J. M. A new class of glutamate dehydrogenases (GDH). Biochemical and genetic characterization of the first member, the AMP-requiring NAD-specific GDH of Streptomyces clavuligerus. J. Biol. Chem. 275, 39529–39542 (2000).
Commichau, F. M., Gunka, K., Landmann, J. J. & Stülke, J. Glutamate metabolism in Bacillus subtilis: gene expression and enzyme activities evolved to avoid futile cycles and to allow rapid responses to perturbations of the system. J. Bacteriol. 190, 3557–3564 (2008).
Seah, S. Y. Industrial Enzymes: Structure, Function and Applications (Dordrecht: Springer Netherlands, 2007).
Adamson, H. et al. Rapid Quantification of C. difficile Glutamate Dehydrogenase and Toxin B (TcdB) with a NanoBiT Split-Luciferase Assay. Anal. Chem. 94, 8156–8163 (2022).
DeMars, Z. & Bose, J. L. Redirection of Metabolism in Response to Fatty Acid Kinase in Staphylococcus aureus.J Bacteriol. 200, e00345–18 (2018).
Halsey, C. R. et al. Amino Acid Catabolism in Staphylococcus aureus and the Function of Carbon Catabolite Repression. mBio. 8, e01434–16 (2017).
Beaufay, F. et al. A NAD-dependent glutamate dehydrogenase coordinates metabolism with cell division in Caulobacter crescentus. EMBO J. 34, 1786–1800 (2015).
Noda-García, L. et al. Chance and pleiotropy dominate genetic diversity in complex bacterial environments. Nat. Microbiol. 4, 1221–1230 (2019).
Griffin, J. E. et al. High-resolution phenotypic profiling defines genes essential for mycobacterial growth and cholesterol catabolism. PLoS Pathog. 7, e1002251 (2011).
Jayaraman, V. et al. A counter-enzyme complex regulates glutamate metabolism in Bacillus subtilis. Nat. Chem. Biol. 18, 161–170 (2021).
Noda-Garcia, L., Romero Romero, M. L., Longo, L. M., Kolodkin-Gal, I. & Tawfik, D. S. Bacilli glutamate dehydrogenases diverged via coevolution of transcription and enzyme regulation. EMBO Rep. 18, 1139–1149 (2017).
Girinathan, B. P., Braun, S., Sirigireddy, A. R., Lopez, J. E. & Govind, R. Importance of Glutamate Dehydrogenase (GDH) in Clostridium difficile Colonization In Vivo. PLoS One 11, e0160107 (2016).
Chittick, L. & Okwumabua, O. Loss of expression of the glutamate dehydrogenase (gdh) of Streptococcus suis serotype 2 compromises growth and pathogenicity. Micro. Pathog. 188, 106565 (2024).
Huang, X., Lao, W., Zhou, Y., Sun, Y. & Wang, Q. Glutamate dehydrogenase enables Salmonella to survive under oxidative stress and escape from clearance in macrophages. FEBS Lett. 596, 81–94 (2022).
Gallant, J. L., Viljoen, A. J., van Helden, P. D. & Wiid, I. J. Glutamate Dehydrogenase Is Required by Mycobacterium bovis BCG for Resistance to Cellular Stress. PLoS One 11, e0147706 (2016).
Gazioglu, O. et al. Glutamate Dehydrogenase (GdhA) of Streptococcus pneumoniae Is Required for High Temperature Adaptation. Infect. Immun. 89, e0040021 (2021).
Lee, Y. H., Kingston, A. W. & Helmann, J. D. Glutamate dehydrogenase affects resistance to cell wall antibiotics in Bacillus subtilis. J. Bacteriol. 194, 993–1001 (2012).
Tawfik, D. S. Enzyme promiscuity and evolution in light of cellular metabolism. FEBS J. 287, 1260–1261 (2020).
Mathioudakis, L. et al. Localization of Human Glutamate Dehydrogenases Provides Insights into Their Metabolic Role and Their Involvement in Disease Processes. Neurochem Res. 44, 170–187 (2019).
Belitsky, B. R. & Sonenshein, A. L. Role and regulation of Bacillus subtilis glutamate dehydrogenase genes. J. Bacteriol. 180, 6298–6305 (1998).
Nye, T. M., Schroeder, J. W., Kearns, D. B. & Simmons, L. A. Complete Genome Sequence of Undomesticated Bacillus subtilis Strain NCIB 3610. Genome Announc 5 (2017).
Gunka, K. & Commichau, F. M. Control of glutamate homeostasis in Bacillus subtilis: a complex interplay between ammonium assimilation, glutamate biosynthesis and degradation. Mol. Microbiol 85, 213–224 (2012).
Ranava, D. et al. Bidirectional sequestration between a bacterial hibernation factor and a glutamate metabolizing protein. Proc. Natl Acad. Sci. USA 119, e2207257119 (2022).
Gunka, K. et al. Functional dissection of a trigger enzyme: mutations of the Bacillus subtilis glutamate dehydrogenase RocG that affect differentially its catalytic activity and regulatory properties. J. Mol. Biol. 400, 815–827 (2010).
Vandooren, J., Geurts, N., Martens, E., Van den Steen, P. E. & Opdenakker, G. Zymography methods for visualizing hydrolytic enzymes. Nat. Methods 10, 211–220 (2013).
Okwumabua, O., Persaud, J. S. & Reddy, P. G. Cloning and characterization of the gene encoding the glutamate dehydrogenase of Streptococcus suis serotype 2. Clin. Diagn. Lab Immunol. 8, 251–257 (2001).
Warneke, R. et al. Ornithine is the central intermediate in the arginine degradative pathway and its regulation in Bacillus subtilis. J. Biol. Chem. 299, 104944 (2023).
Debarbouille, M., Gardan, R., Arnaud, M. & Rapoport, G. Role of BkdR, a transcriptional activator of the SigL-dependent isoleucine and valine degradation pathway in Bacillus subtilis. J. Bacteriol. 181, 2059–2066 (1999).
Kasu, I. R., Reyes-Matte, O., Bonive-Boscan, A., Derman, A. I. & Lopez-Garrido, J. Catabolism of germinant amino acids is required to prevent premature spore germination in Bacillus subtilis. mBio 15, e0056224 (2024).
Baker, P. J., Turnbull, A. P., Sedelnikova, S. E., Stillman, T. J. & Rice, D. W. A role for quaternary structure in the substrate specificity of leucine dehydrogenase. Structure 3, 693–705 (1995).
Gunka, K., Stannek, L., Care, R. A. & Commichau, F. M. Selection-driven accumulation of suppressor mutants in Bacillus subtilis: the apparent high mutation frequency of the cryptic gudB gene and the rapid clonal expansion of gudB(+) suppressors are due to growth under selection. PLoS One 8, e66120 (2013).
Fullam, A. et al. proGenomes3: approaching one million accurately and consistently annotated high-quality prokaryotic genomes. Nucleic Acids Res. 51, D760–D766 (2023).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Liu, J. et al. Metabolic co-dependence gives rise to collective oscillations within biofilms. Nature 523, 550–554 (2015).
Bocci, F., Suzuki, Y., Lu, M. & Onuchic, J. N. Role of metabolic spatiotemporal dynamics in regulating biofilm colony expansion. Proc. Natl Acad. Sci. USA 115, 4288–4293 (2018).
Kimura, T. & Kobayashi, K. Role of Glutamate Synthase in Biofilm Formation by Bacillus subtilis. J. Bacteriol. 202, e00120–e00120 (2020).
Shemesh, M. & Chai, Y. A combination of glycerol and manganese promotes biofilm formation in Bacillus subtilis via histidine kinase KinD signaling. J. Bacteriol. 195, 2747–2754 (2013).
Vlamakis, H., Chai, Y., Beauregard, P., Losick, R. & Kolter, R. Sticking together: building a biofilm the Bacillus subtilis way. Nat. Rev. Microbiol 11, 157–168 (2013).
Gunka, K. et al. A high-frequency mutation in Bacillus subtilis: requirements for the decryptification of the gudB glutamate dehydrogenase gene. J. Bacteriol. 194, 1036–1044 (2012).
Flórez, L. A., Gunka, K., Polanía, R., Tholen, S. & Stülke, J. SPABBATS: A pathway-discovery method based on Boolean satisfiability that facilitates the characterization of suppressor mutants. BMC Syst. Biol. 5, 5 (2011).
Bennett, B. D. et al. Absolute metabolite concentrations and implied enzyme active site occupancy in Escherichia coli. Nat. Chem. Biol. 5, 593–599 (2009).
Young, V. R. & Ajami, A. M. Glutamate: an amino acid of particular distinction. J. Nutr. 130, 892S–900S (2000).
Oh, Y. K., Palsson, B. O., Park, S. M., Schilling, C. H. & Mahadevan, R. Genome-scale reconstruction of metabolic network in Bacillus subtilis based on high-throughput phenotyping and gene essentiality data. J. Biol. Chem. 282, 28791–28799 (2007).
Krüger, L. et al. Essentiality of c-di-AMP in Bacillus subtilis: Bypassing mutations converge in potassium and glutamate homeostasis. PLoS Genet 17, e1009092 (2021).
Stannek, L. et al. Evidence for synergistic control of glutamate biosynthesis by glutamate dehydrogenases and glutamate in Bacillus subtilis. Environ. Microbiol 17, 3379–3390 (2015).
Calogero, S. et al. RocR, a novel regulatory protein controlling arginine utilization in Bacillus subtilis, belongs to the NtrC/NifA family of transcriptional activators. J. Bacteriol. 176, 1234–1241 (1994).
Commichau, F. M., Herzberg, C., Tripal, P., Valerius, O. & Stülke, J. A regulatory protein-protein interaction governs glutamate biosynthesis in Bacillus subtilis: the glutamate dehydrogenase RocG moonlights in controlling the transcription factor GltC. Mol. Microbiol 65, 642–654 (2007).
Commichau, F. M. & Stülke, J. Trigger enzymes: bifunctional proteins active in metabolism and in controlling gene expression. Mol. Microbiol 67, 692–702 (2008).
Turnbull, A. P., Baker, P. J. & Rice, D. W. Analysis of the quaternary structure, substrate specificity, and catalytic mechanism of valine dehydrogenase. J. Biol. Chem. 272, 25105–25111 (1997).
Wang, X. G., Britton, K. L., Stillman, T. J., Rice, D. W. & Engel, P. C. Conversion of a glutamate dehydrogenase into methionine/norleucine dehydrogenase by site-directed mutagenesis. Eur. J. Biochem 268, 5791–5799 (2001).
Stillman, T. J. et al. Insights into the mechanism of domain closure and substrate specificity of glutamate dehydrogenase from Clostridium symbiosum. J. Mol. Biol. 285, 875–885 (1999).
Sun, Y. et al. Acetylation coordinates the crosstalk between carbon metabolism and ammonium assimilation in Salmonella enterica. EMBO J. 42, e112333 (2023).
Chou, K. T. et al. A segmentation clock patterns cellular differentiation in a bacterial biofilm. Cell 185, 145–157.e113 (2022).
Tran, P., Lander, S. M. & Prindle, A. Active pH regulation facilitates Bacillus subtilis biofilm development in a minimally buffered environment. mBio 15, e0338723 (2024).
Arnaouteli, S., Bamford, N. C., Stanley-Wall, N. R. & Kovács, ÁT. Bacillus subtilis biofilm formation and social interactions. Nat. Rev. Microbiol 19, 600–614 (2021).
Kesel, S. et al. Direct Comparison of Physical Properties of Bacillus subtilis NCIB 3610 and B-1 Biofilms. Appl Environ. Microbiol 82, 2424–2432 (2016).
Schneewind, O. & Missiakas, D. Genetic manipulation of Staphylococcus aureus. Curr. Protoc. Microbiol. 32, Unit 9C 3 (2014).
Bruckner, R. A series of shuttle vectors for Bacillus subtilis and Escherichia coli. Gene 122, 187–192 (1992).
Fuller, J. R. et al. Identification of a lactate-quinone oxidoreductase in Staphylococcus aureus that is essential for virulence. Front Cell Infect. Microbiol 1, 19 (2011).
Unger, T., Jacobovitch, Y., Dantes, A., Bernheim, R. & Peleg, Y. Applications of the Restriction Free (RF) cloning procedure for molecular manipulations and protein expression. J. Struct. Biol. 172, 34–44 (2010).
Branda, S. S., González-Pastor, J. E., Ben-Yehuda, S., Losick, R. & Kolter, R. Fruiting body formation by Bacillus subtilis. Proc. Natl Acad. Sci. USA 98, 11621–11626 (2001).
Koo, B. M. et al. Construction and Analysis of Two Genome-Scale Deletion Libraries for Bacillus subtilis. Cell Syst. 4, 291–305.e297 (2017).
Yasbin, R. E. & Young, F. E. Transduction in Bacillus subtilis by bacteriophage SPP1. J. Virol. 14, 1343–1348 (1974).
Kearns, D. B., Chu, F., Branda, S. S., Kolter, R. & Losick, R. A master regulator for biofilm formation by Bacillus subtilis. Mol. Microbiol 55, 739–749 (2005).
Konkol, M. A., Blair, K. M. & Kearns, D. B. Plasmid-encoded ComI inhibits competence in the ancestral 3610 strain of Bacillus subtilis. J. Bacteriol. 195, 4085–4093 (2013).
Lander, S. M., Fisher, G., Everett, B. A., Tran, P. & Prindle, A. Secreted nucleases reclaim extracellular DNA during biofilm development. NPJ Biofilms Microbiomes 10, 103 (2024).
Lipońska, A., Lee, H. & Yap, M. F. Staphylococcal exoribonuclease YhaM destabilizes ribosomes by targeting the mRNA of a hibernation factor. Nucleic Acids Res. 52, 8998–9013 (2024).
Yap, M. N. & Bernstein, H. D. Mutations in the Escherichia coli ribosomal protein L22 selectively suppress the expression of a secreted bacterial virulence factor. J. Bacteriol. 195, 2991–2999 (2013).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Holm, L. Using Dali for Protein Structure Comparison. Methods Mol. Biol. 2112, 29–42 (2020).
Mhatre, E. et al. The impact of manganese on biofilm development of Bacillus subtilis. Microbiology 162, 1468–1478 (2016).
Acknowledgements
We thank Jeffrey Bose for advice on gudB mutant complementation, Fitnat Yilidz for comments on biofilm experiments, the late Dan Tawfik for a generous gift of B. subtilis GudB antibody, and Vijay Jayaraman for feedback on GudB oligomerization. This work was supported by National Institutes of Health (NIH) grants R01GM121359, R01AI150986, and the Northwestern University Faculty Startup Fund (to MFY). A.P. received generous support from the National Science Foundation (NSF 2239567), NIH (R35GM147170), Pew Charitable Trusts (2019-A-06953), the Army Research office (W911NF-19-1-0136), and the David and Lucile Packard Foundation (2018-68055). S.M.L. was supported by an NIH predoctoral fellowship F31GM143907. Transposon mutants were obtained through Network on Antimicrobial Resistance in Staphylococcus aureus (NARSA) for distribution by BEI Resources, NIAID, NIH.
Author information
Authors and Affiliations
Contributions
D.R.—Conceptualization, investigation, validation, formal analysis, writing. S.M.L., S.Y.K.—Investigation, validation.JDW- Formal analysis, software, visualization. A.P.—Resources, funding acquisition, supervision. M.F.Y.—Conceptualization, project administration, investigation, funding acquisition, supervision, writing. D.R., S.M.L., S.Y.K., J.D.W., A.P., M.F.Y.—Review, edit and approve the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ranava, D., Lander, S.M., Kuan, SY. et al. A promiscuous Bcd amino acid dehydrogenase promotes biofilm development in Bacillus subtilis. npj Biofilms Microbiomes 11, 112 (2025). https://doi.org/10.1038/s41522-025-00750-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41522-025-00750-6