Abstract
Quorum sensing (QS) orchestrates collective microbial behaviors and functional acclimatization through chemical communication. However, QS in natural waters is challenged by dilution, alkaline hydrolysis, and enzymatic degradation of freely dissolved autoinducers. Here, we demonstrate that extracellular vesicles (EVs) act as selective, durable, and protective vectors for QS signal molecules under environmental stresses. Specifically, EVs preferentially package hydrophobic acyl‑homoserine lactones, concentrate them locally, and shield them from alkaline hydrolysis, and exhibiting long-distance transport. In addition, EVs possess specific affinity to recipients, thus influencing microbial community. Field investigation via multi-omics showed that EV abundance covaried with salinity, nutrients, chlorophyll a, and biomass, which were validated by culture experiments. Our statistical framework demonstrated that organisms producing moderate EV levels contributed significantly to maintaining community stability and ecosystem functions. Distinctively within this group, QS-active species (including Burkholderiaceae, Pseudomonadaceae, Rhodobacteraceae, Roseobacteraceae, Flavobacteriaceae etc.) emerge as key drivers facilitating these crucial ecological roles. Furthermore, metaproteomics of field EVs reveal QS receptor and synthesis proteins, suggesting coordinated transport of signals and proteins, which indicate new routes for QS crosstalk, particularly for taxa bearing luxR/I solos. Our results show that moderately generated EVs are the potentially important QS signal carriers and ecological regulation hubs in natural waters.
Similar content being viewed by others
Introduction
Quorum sensing (QS) is a sophisticated communication mechanism that relies on the density of signal molecules called autoinducers1,2. In QS, cells produce and release autoinducers to the extracellular environment, and they are perceived by themselves or other cells3. After sensing their cell numbers via monitoring changes in the concentration of autoinducers, cells will collectively alter gene expression4. Generally, there are diverse kinds of QS signals, such as N-Acyl-homoserine lactones (AHLs), 4-hydroxy-2-alkylquinolines (HAQs), diffusible signal factors (DSFs), auto-inducing peptides (AIPs), Cholerae autoinducer-1 (CAI-1), indole, and autoinducer 2 (AI-2) etc. Much laboratory-based information exists on these QS languages, yet little is known about how these systems operate in nature5. In addition, the extensive hydrophobicity of QS signal molecules makes it difficult for them to exist in natural waters6. It is apparent that many factors, including chemical, physical, and biological variables, can affect signaling in complex environments7.
AHLs produced by gram-negative bacteria are one of the most well-characterized signal molecules controlled by LuxIR-type QS system8,9,10,11,12,13,14. Bioinformatic analysis shows that the communication via AHLs occupied the main position in the activated sludge of wastewater treatment plants15 and human gut microbiota16. Some research on AHL signals has predominantly been conducted under controlled laboratory conditions using pure cultures17,18. However, their behavior and functions can be greatly affected by environmental stresses in natural open water, such as dilution of water6, fluid flow6, temperature9, pH9, and protease19. At the same time, the genetic architectures of luxI and luxR genes are frequently incomplete, and luxR solos lacking LuxI are prevalent18,20, as well as taxa with luxI uncoupled from receptors17,21. Such modularity implies a community context where reception and emission can be decoupled, enabling eavesdropping22, crosstalk11, and interspecies coordination. We are therefore far from fully capturing and understanding the AHL signaling routes in aquatic ecosystems.
Extracellular vesicles (EVs) with membrane-bound structures can be produced by cells across all domains of life and are ubiquitous in natural environments23,24,25,26,27,28,29,30,31. Studies with model microbes have shown that EVs can be crucial mediators of biotic and abiotic interaction via their specific cargoes (metabolites, nucleic acids, proteins, lipids)2,23,32. Notably, ~86% of N-hexadecanoyl-L-homoserine lactone (C16-HSL) is packaged into EVs33, whereas less than 1% of the 3-oxo-dodecanoyl homoserine lactone (3-oxo-C12-HSL) and butyryl homoserine lactone (C4-HSL) are encapsulated34. Prior studies indicated that EVs concentrate33 and traffic34 those signals with significant hydrophobic character. However, we lack knowledge that connects EV cargoes selectivity to the chemistry of natural QS repertoires and whether EV-mediated transport extends signal range and stability in real scenarios. It is still an open question whether microbial species can release EVs at what levels to adapt to the environment, and whether the uptake of EVs can causally reconfigure the microbial community’s structure and function in situ. Addressing these gaps is essential to understanding how EV-mediated molecular dialog scales to the organization and robustness of aquatic microbial ecosystems.
Here, we hypothesize that EVs provide protective and effective signaling routes in the field, and their producers have irreplaceable advantages in species interactions and functions. Therefore, after we found that the free AHL concentrations did not reach the effect threshold in a subtropical urban reservoir, we explored the characteristics and advantages of EVs carrying AHLs through AHL-positive strains isolated from sampling sites. We also studied the specific interactive cells of EVs using a microcosm experiment. We investigated the spatiotemporal distribution and influencing factors of EVs, and then used metagenome coupling with metaproteome to explore the ecological importance of species with different EV levels and QS proteins in EVs. We further compiled a QS species inventory and identified QS-related genes in metagenome-assembled genomes (MAGs) to elucidate the potential mechanisms of QS crosstalk mediated by EVs. Building on watershed-scale surveys, isolate-based assays, metaproteomic and metagenomic profiling of field EVs, and microcosm experiments, we evaluated EVs as regulated vehicles for QS and as potential important drivers of ecological functions.
Results
EVs selectively package and protect hydrophobic AHLs
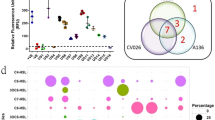
To evaluate the effective concentration of free signals capable of mediating QS in natural aquatic environments, we employed high-sensitivity liquid chromatography-tandem mass spectrometry (LC-MS/MS) to quantify the abundance of 13 representative AHLs in the surface water samples of Xinglinwan Reservoir (XLR) (Fig. 1a). Results showed that the occurrence of C4-HSL, C12-HSL (N-Dodecanoyl-L-homoserine lactone), 3-oxo-C16-HSL (N-(3-Oxohexadecanoyl)-L-homoserine lactone), and C14-HSL (N-Tetradecanoyl-L-homoserine lactone) were at the range between not detected and 0.1 ng/mL (Fig. 1a, Table S1). Only 16.7% (6/36) of measurements exceeded published activity thresholds (0.01–10 ng/mL)33,35,36,37. These results suggested that free AHLs in open water were difficult to prevent from dilution and hydrolysis to accumulate to the effective activation thresholds, and non-free AHL pools may underpin effective.
a The concentrations of 13 types of AHLs in water samples from XLR across three seasons. b The biosensor assays of 24 AHLs-active strains isolated from XLR. We used Chromobacterium violaceum CVO26 (CVO26), Agrobacterium tumefaciens A136 (A136), and Agrobacterium tumefaciens KYC55 (KYC55) as biosensors. The red boxes indicated the AHL positive results (turning purple or blue) for their corresponding biosensor. The clustering represents a phylogenetic analysis based on 16S rRNA gene sequences. c Preferential packaging of hydrophobic AHLs within EVs from 12 isolated strains. The diagram showed that the proportion of EV-associated AHLs (y-axis) across 13 different kinds of AHLs (x-axis). The blue arrow indicates that the hydrophobicity of the 13 AHLs molecules increases from left to right. d The three-dimensional (3D) confocal laser scanning microscopy (CLSM) image showed the agglomerates of EVs attached to the biofilm matrices formed in XLR (white arrows). Red, FM4-64-labeled EVs derived from XL10. Green, biofilm stained with SYTO 9. e The hydrolysis course of C14-HSL in EVs at pH8 and pH10 monitored by LC-MS/MS.
We hypothesize that EVs can be non-free component carriers of AHLs. To verify this, the partition of AHLs between EV-free supernatants and EV fractions was calculated in pure cultured strains isolated from the XLR. First, we isolated 95 bacterial strains from XLR spanning Proteobacteria (n = 68), Firmicutes (n = 19), Bacteroidetes (n = 4), Actinobacteria (n = 1), and unassigned (n = 3), representing the dominant phyla observed in the reservoir (Table S2). Then, we screened typical AHL types using three biosensor strains targeting complementary acyl-chain lengths: Chromobacterium violaceum CVO26 (CVO26), Agrobacterium tumefaciens A136 (A136), and Agrobacterium tumefaciens KYC55 (KYC55). Specifically, CVO26 produces the purple pigment violacein selectively in response to short-chain molecules (C4- to C6-HSL), whereas A136 produces blue pigmentation in the presence of 3-oxo-C4- to 3-oxo-C12-HSL and C5- to C10-HSL38. KYC55 exhibits blue coloration in response to a broad spectrum of 3-oxo-C4- to 3-oxo-C18-HSL and C4- to C18-HSL38. Biosensor assays revealed that 25.3% (24/95) of isolated strains were AHL-positive (Fig. 1b, Fig. S2). To further validate the signaling molecules, we applied the LC-MS/MS methods against a standard panel of 13 kinds of typical AHLs. The analysis successfully resolved specific AHLs profiles in 12 isolates (Fig. S3, Table S3). The remaining biosensor positive strains lacked spectral matches likely due to the production of non-targeted or higher molecular weight variates beyond the 13 AHLs standards. For bioassay-positive strains, we calculated the partition of AHLs between EV-free supernatants and EV fractions. The percentage of the molecules associated with EVs ranged from 10% to 49%, and exhibited increasing trend with the hydrophobicity of AHLs (Fig. 1c). Notably, AHLs exhibit a broad spectrum of hydrophobicity ranging from −1.96 to 6.91 (collected from PubChem and ChemSpider), quantified by the octanol-water partition coefficient (Kow), where higher Kow values indicate greater hydrophobicity (Fig. S4b). A marked hydrophobicity bias was observed that AHLs with Kow >3.50 being preferentially packaged within EVs.
We then analyzed the concentrated effects, stability, and protection of EVs on AHLs using a representative isolated strain XL10 (uncultured Pseudomonas sp.) from the repository of isolated strains from XLR (full list provided in Table S2). XL10 generated up to 24.2% of C14-HSL associated with EVs (Table S3). In late stationary phase, monocultures of strain XL10 contained 1.53 × 109 EV particles·mL−1, and the EV-associated C14-HSL pool reached ~0.0285 ng/mL which has exceeded the reported biologically active thresholds range (>0.01 ng/mL, depending on receptor sensitivity)33,35,36,37. In addition, the computational results showed that each EV could carry 36 C14-HSL molecules, and the average central distance of EV-associated C14-HSL was 17.1 nm (Fig. S5), implying local concentration by particle clustering. We also found that EVs of XL10 displayed a zeta potential of −7.25 ± 0.44 mV, consistent with aggregation/coagulation propensity39, and were frequently observed as agglomerates attached to biofilm matrices (Fig. 1d).
Given the susceptibility of AHLs to hydrolysis, we next investigated whether vesicular packaging protects these signals. We found that EV-associated C14-HSL maintained more than half of concentration up to 48 h at both pH8 and pH10 (Fig. 1e, Table S4-1). In contrast, the half-life of free C14-HSL was 35.3 h at pH8 and 34.1 h at pH10 (Table S4-2), which was similar to previously reported (36 h)14. Using the isolated strain XL10 as the representative model, the results demonstrated the potential capacity of EVs to selectively package hydrophobic AHLs, locally concentrate signals, and improve stability against alkaline hydrolysis, which may display significant inter-specific heterogeneity.
EVs show species-selective binding and influence the microbial community
The finding that EVs act as protective vectors solves the puzzle of signal instability, yet these stabilized EVs must interact with target recipients to exert ecological impact. Therefore, the role of EVs in establishing a functional framework were analyzed using the isolated strain XL92 (Flavobacterium sp.), which was a potential keystone species significantly correlated with crucial functional pathways in our metagenomic data of XLR (Fig. S6). Confocal microscopy showed uptake/adhesion of DiO-labeled EVs from XL92 by pure cultured Synechococcus sp. FACHB-1061 and Microcystis aeruginosa FACHB-905 and by their biofilms (Fig. 2a, Fig. S7a). In samples from field incubations, EVs were taken up by or adhered to specific algae, bacteria, and biofilm matrices (Fig. 2b, Fig. S7b, c). We tracked EV-recipient interactions in field communities using fluorescence-activated cell sorting (FACS) coupled to 16S/18S rRNA gene sequencing (Fig. 2c, Fig. S8). Prokaryotic genera with higher EV interaction included Aeromonas, Cyanobium PCC-6307, Acinetobacter, Flavobacterium, and Vibrio, jointly representing >40% (±4.22%) of total prokaryotic abundance. Among eukaryotes, Ciliophora, Cryptomonadales, Rotifera, and Ochrophyta dominated the labeled community (Fig. 2d). Our results indicated that naturally occurring EV uptake was not restricted to a single exclusive target but rather widely distributed. EVs may act as a broad-spectrum yet amenable medium for community-wide communication.
a Representative confocal microscopy images of Synechococcus sp. FACHB-1061 (up) and Microcystis aeruginosa FACHB-905 (down) co-incubation with DiO-labeled EVs derived from XL92 (Flavobacterium sp.) (scale bar, 2 μm). b Representative confocal microscopy images of field species from XLR co-incubation with DiO-labeled EVs derived from XL92 (Flavobacterium sp.) (scale bar, 5 μm). c The scheme for tracking DiO-labeled EVs uptake or conglutination by aquatic organisms. d The bar graph showed the results of 16S (left) and 18S (right) rRNA gene sequencing for species with a preference for taking up or attaching EVs (group P, n = 3) compared with the control group (group N, n = 3). The data were representative of three independent experiments. e The scheme of the microcosmic experiment setup. The four experimental groups and one control group lasted for 7 days. The additional amounts of EVs and bacteria were determined according to the environmental concentration. f The clustering tree (Bray-Curtis dissimilarity) showed their similarity of community structure. Group P represents the sorted microbial fraction that successfully takes up DiO-labeled EVs, while group N represents the fraction that did not exhibit EV fluorescence. In the microcosmic experiments, group A, B, C, D, and E represent the different group which were treated with strain XL92 bacterial cells, intact EVs derived from XL92, ruptured EVs derived from XL92, XL92 bacterial supernatant depleted of EVs, and control with PBS, respectively. g The bubble diagram showed the pathway enrichment analysis from functional predictions for prokaryotes by PICRUSt2. The color and the size of the bubbles represented the P value and the abundance of pathways, respectively.
To determine whether these targeted EV recipients impact on the community structure, we conducted 7-day microcosms assays to assess biological structure and functions under environmentally relevant conditions (Fig. 2e). Treatments included intact EVs, the source XL92 cells, ruptured EVs, EV-depleted supernatants, and a vehicle control. The amplicon sequencing results of microcosm experiment showed that group treated with intact EVs (group B) clustered closely with those supplemented with source XL92 cells (group A), indicating convergent community composition under these two treatments. In contrast, groups added ruptured EVs (group C) and EV-depleted supernatant (group D) clustered together, supporting the specific impact of intact EVs on microbial community structure (Fig. 2f). To further understand the potential physiological consequences of these structural shifts, we extrapolated the functional profiles of the bacterial communities using PICRUSt2 based on 16S rRNA gene amplicons. The prediction analysis revealed that intact EVs increased predicted metabolic capacities relative to controls, particularly starch/sucrose metabolism, glycolysis/gluconeogenesis, peptidoglycan biosynthesis, carbon metabolism, and two-component systems with mirroring the effects of source-cell addition (Fig. 2g). By contrast, ruptured EVs and EV-depleted supernatants resembled controls. These results indicated that EV particle integrity is critical for ecological efficacy. By acting as structured vectors, EVs likely affect community composition and functions through protected bioavailability and simultaneous delivery of multiple bioactive cues to specific recipients, a capacity lost upon vesicle rupture.
EVs are widespread and influenced by biotic and abiotic drivers in natural waters
To investigate whether these EV-based mechanisms operate under realistic environmental constraints, and not just in discrete isolates, it is crucial to assess their stability and occurrence in the field. Spanning varying levels of urbanization and nutritional status, the Houxi River watershed (HRW), in which the XLR is located downstream, provides an ideal environmental gradient for this purpose (Fig. 3a). The average EV particle concentrations from HRW varied between 3.22 × 105 and 3.05 × 107 particles·mL−1 with the particle sizes ranging from 68 to 79 nm (Fig. 3b, Fig. S9), which were measured using NanoFCM. The particle and protein concentrations of field EVs are the highest in the downstream of HRW. Along the gradient, EV particle counts and EV-associated protein concentrations covaried with salinity, chl-a, total phosphorus, nitrite, and total organismal abundance (Fig. 3c). Structural equation modeling indicated that environmental variables (especially pH, DO, and chl-a) and species jointly exerted direct and indirect effects on EV abundance (Fig. 3d). Consistently, controlled cultures of the isolated strain XL92 showed that EV production was highest at early exponential growth and sensitive to temperature, pH, and salinity, ranging from 0.9 to 18.3 EVs·cell−1·generation−1 at the early stage of the logarithmic growth phase (Log 1) and converging to 0.25–2.1 EVs·cell−1·generation−1 at the late logarithmic phase (Log 3) (Fig. 3e, Fig. S10). To sum up, EVs are widespread in natural waters and their abundance is influenced by biotic and abiotic factors. Source-tracking (fast expectation-maximization microbial source tracking, FEAST) suggested limited upstream inputs to the downstream EV pool in HRW (2.7% from midstream, 0.99% from upstream in Fig. 3f), indicating broad spatiotemporal connectivity and transport potential.
a Map of the sampling sites in HRW (n = 13). b The box plots showed the particle number (up) and protein content (down) of EVs in HRW, in which Wilcoxon tests were performed. US: upstream; MS: midstream; DS: downstream. c Heatmaps showed pairwise correlations between the characteristics of EVs and biotic as well as abiotic factors in HRW. The color gradient indicated Spearman’s correlation coefficients with an asterisk (see notes below). d Structural equation modeling analysis showed the influence of biotic and abiotic factors on EV particle number. e The EVs production rates during the logarithmic growth phase of XL92 (Flavobacterium sp.) strain (three repetitions) at different pH, salinity, and temperature when a single variable was controlled. Log 1, Log 2, and Log 3 represent the first, second, and third stages of logarithmic growth, respectively. The significant asterisk markers were represented as *P < 0.05, **P < 0.01, ***P < 0.001. f The proportions of the EVs taxonomic sources in HRW using FEAST (fast expectation-maximization microbial source tracking) algorithm. g Scanning electron microscopy (SEM) images (scale bar, 100 nm) of the XL92 (Flavobacterium sp.) strain isolated from sampling sites. The red arrows pointed to the EVs which were bubbling. Notes: APS average particle size, HRW Houxi River watershed, Tem water temperature, Tur turbidity, chl-a chlorophyl a, DO dissolved oxygen, ORP oxidation reduction potential, Con conductivity, Sal salinity, Trans transparency, TN total nitrogen, NH4_N ammonium nitrogen, NO3_N nitrate nitrogen, NO2_N nitrite nitrogen, TP total phosphorus, PO4_P phosphate phosphorus, TDS total dissolved solids, ACD algae cell density.
Moderate EV producers orchestrate species interactions and ecological functions in situ
Metabarcoding of EV-associated DNA traced EV origins to diverse taxa and revealed spatiotemporal heterogeneity in producers. To assess EV-associated dynamics at the community level, we cross-examined the taxonomic overlap between aquatic organism DNA (whole community) and purified EV-associated DNA (EV-released species) datasets (Figs. S11, 12). Organisms identified in both datasets were defined as aquatic organisms with detected EVs (DAs), which indicated confirmable in situ EV release. Conversely, taxa present in the community but consistently absent from the EV dataset were classified as aquatic organisms with undetected EVs (UDAs). DAs were further subdivided as aquatic organisms with high EV levels (H-DAs) and aquatic organisms with low EV levels (L-DAs) based on comparing the abundance of EV-associated DNA with that of its source species.
DAs comprised >90% of total abundance across three months, with L-DAs >70% (Fig. S13). The downstream sites (XLR) exhibited the lowest Shannon diversity in the spatial continuum, which may be attributable to the mild eutrophication status of it. Among aquatic organisms with different EV levels, UDAs exhibited higher diversity than DAs, and H-DAs exceeded L-DAs (Fig. 4a). Community dissimilarities were significant across all sampling sites (Fig. S14), and DAs, especially L-DAs, contributed most to β-diversity (Fig. 4b). L-DAs were also subject to stronger environmental filtering (Figs. S15, 16).
a The Shannon diversity of aquatic organisms with varied EV levels. US, MS, and DS meant the upstream, midstream, and downstream of HRW. b The cumulative contribution of overall differences in microbial communities caused by different types of aquatic organisms using the similarity percentage analysis (SIMPER). c The niche breadths in UDAs, DAs, H-DAs, and L-DAs in three seasons of XLR. d The robustness of aquatic organisms with different EV levels measured as the proportion of taxa remained with 90% of the taxa targeted removed from prokaryotes and eukaryotes co-occurrence networks. Each error bar corresponded to the standard deviation of 1000 repetitions of simulation. e Network vulnerability (the maximum node vulnerability) measured by nodes vulnerability in prokaryotes and eukaryotes co-occurrence networks. f The degree of the nodes from the co-occurrence networks (*P < 0.05, **P < 0.01, ***P < 0.001; Wilcoxon test). g Ternary plots of pathway enrichment analysis of aquatic organisms with different EV levels in three seasons of XLR. Position was determined by the Reporter Score of organisms. Colors of circles correspond to different pathways. h The cumulative contribution percentage of aquatic organisms with different EV levels to the selected functions. i The functional diversity in the top 6 MAGs taxa (family level). The heatmap showed the gene number of MAGs within each family containing the corresponding genes. CAZymes Carbohydrate-Active Enzymes, AA Auxiliary Activities, CBM Carbohydrate-Binding Modules, CE Carbohydrate Esterases, GH Glycoside Hydrolases, GT Glycosyltransferases, PL Polysaccharide Lyases, SLH S-Layer Homology.
Niche analyses across HRW and XLR indicated broader niche breadths for DAs than UDAs, with L-DAs exceeding H-DAs (Fig. 4c, Fig. S17). By mapping the different EV levels of aquatic organisms onto network nodes and conducting targeted removal simulations, we assessed the importance of distinct species types. The analysis revealed that network robustness declined more upon removal of DAs than UDAs, and more upon removal of L-DAs than H-DAs (Fig. 4d, Fig. S18a, b), implying that moderate EV producers occupy the topological centers. Network vulnerability was lower for DAs than UDAs and lower for L-DAs than H-DAs (Fig. 4e). DAs exhibited higher positive and negative cohesion (with L-DAs > H-DAs, Fig. S18c, d) and higher node degree (Fig. 4f), consistent with more keystone-like roles. Furthermore, we applied the LEfSe (Linear Discriminant Analysis Effect Size) algorithm to screen for seasonal indicator taxa (biomarker). Notably, a cross-referenced analysis revealed that the majority of these seasonal biomarkers aligned with DAs (Fig. S19a-c), suggesting that prominent EV producers are effectively the drivers of seasonal fluctuations.
Pathway enrichment analysis showed the most genes in DAs (including L-DAs and H-DAs) were significantly related to diverse metabolism followed by cellular processes, environmental information processing, and genetic information processing, which were similar to those of its L-DAs component, underscoring the ecological prominence of L-DAs (Fig. 4g). For 11 focal functional genes, DAs contributed >97% across seasons (Fig. 4h). Intriguingly, we identified 389 MAGs from XLR. The metabolic profile analysis demonstrated that these MAGs encompassed diverse C/N/P metabolism and signaling (Fig. 4i). By aligning the taxonomy of these MAGs with the family-level aquatic organisms monitored in parallel studies (with different EV levels detailed in Fig. S19d), we confirmed that they were more abundant in DAs.
QS species occupy key ecological positions and their EVs facilitate crosstalk
To investigate QS species and their EV levels, we compiled a complete QS species inventory for bacterial taxa of XLR. This was constructed by integrating known QS databases (SigMol, Quorumpeps, and QSDB) with taxon-specific potentials identified from metagenomic data via local BLASTP (Fig. S20). We then constructed an undirected QS communication network spanning eight kinds of QS languages. This analysis highlighted dense cross-species connectivity, with AIPs and AHLs acting as particularly prominent mediators (Fig. 5a, Fig. S21). In the QS species inventory, those taxa (e.g., Burkholderiaceae, Pseudomonadaceae, Rhodobacteraceae, Roseobacteraceae, Flavobacteriaceae) were prominent DAs and enriched as keystones (Fig. S22a). Across three seasons, the average relative abundance of QS species was 40.1% in spring, 40.2% in summer, and 35.0% in winter (Fig. S22b). QS species assigned to L-DAs contributed most to community structure, particularly in summer (Fig. S23a). Contributions of QS species to key ecological functions were 42.9% (±8.22%) in spring, 45.0% (±8.19%) in summer, and 39.0% (±16.7%) in winter (Fig. 5b). QS species also exhibited wide niche breadths (Fig. 5c). To sum up, based on functional profiling, data analysis indicated that QS community members had impact on ecological functions.
a QS database of organisms from XLR. The phylogenetic tree of species with quorum-sensing genes found in XLR based on eight QS languages (AHLs, DSFs, HAQs, CAI-1, AIPs, AI-2, Indole, others). Heatmap on the outermost layer indicated QS languages distribution in each cluster, existence was colored in blue; non-existence was colored in gray. b The cumulative contribution percentage of QS aquatic organisms with different EV levels to the selected functions. The x-axis represents the percentage scale (0-100%). c The niche breadths in different types of QS organisms in three seasons of XLR. d The luxR solos in MAGs of XLR. The color of each MAG represents their Family taxon and the AHL gene means their luxR genes. e Heatmap showing the log-transformed abundance of identified QS synthases and receptor proteins from field EVs of XLR.
Notably, while the vast majority of species were restricted to a single QS language, we identified taxa with “multilingual” capacities, in which 87, 13, and 2 species possessed two, three, and four QS languages, respectively (Fig. S21). This multi-signal process underscores complex crosstalk communication in the open water. Among these QS systems, AHLs are produced by LuxI family synthases and specifically interact with cytoplasmic LuxR family transcription factors to influence gene expression22. Commonly, the luxR and luxI genes appear in pairs, while 27 of 389 MAGs harbored AHL-related genes exclusively as luxR solos, spanning five phyla and dominated by Rhodobacteraceae (18/27, Fig. 5d). Given that Synechococcus was the ubiquitous and highly abundant genus in the XLR, we extended our analysis to this lineage by downloading their genomes from NCBI. Results showed that Synechococcus also chiefly carries LuxR-type receptors without adjacent luxI, suggesting broad potential for eavesdropping on exogenous AHLs (Fig. S23b). While suspecting eavesdropping on exogenous AHLs, we also speculate whether EVs can transmit LuxR homologs. Metaproteomics of field EVs detected QS synthesis and receptor proteins, with AHL receptor homologs peaking in summer and primarily derived from Proteobacteria (notably Rhodobacteraceae and Syntrophorhabdaceae, Fig. 5e, Fig. S23c).
Discussion
Natural aquatic environments impose strong hydrolysis, dilution, and enzymatic pressures on freely dissolved QS signals, leaving open the question of how recipients reliably perceive these cues in the wild. Here, we demonstrate that EVs provide a regulated, selective, and ecologically consequential route for QS in nature. Our results established a coherent mechanism-to-ecosystem framework in which EVs (i) preferentially load and stabilize hydrophobic AHLs, (ii) targeted and reprogrammed recipient taxa to produce causal shifts in community structure and function, and (iii) were produced under biotic and abiotic control, facilitating QS species as key roles in community structure and ecological functions.
As is well documented, AHLs play key roles in species communications, but their functions can be largely affected by environmental stresses40. Short-chain acyl-HSLs diffuse freely across the cell membrane41, whereas long-chain acyl-HSLs with higher hydrophobicity require transporters to active export42,43. EVs demonstrated a tendency to traffic hydrophobic AHLs with a Kow greater than 3.50. If this result applies to all types of QS languages, EVs preferentially traffic a range of signals that encompasses over half of the QS signals (55.3%, Fig. S4). By shielding AHLs from alkaline hydrolysis, EVs extended signal persistence and enabled transport over long distances, well beyond canonical biological calling distance44. Field and experimental evidence further indicated local concentration via vesicle coagulation, collectively lowering the effective activation threshold for AHLs in noisy, fluctuating waters. As for the targeting of EVs to the recipients, it can be influenced by surface hydrophobicity45, cell envelope structure45, surface charge46, and surface lipids28,47. The specific interactions between EVs and their targeted cells were selective (Fig. 2d). As evidenced by our microcosm experiments, the uptake of EVs led to influence in community structure (Fig. 2f) and regulation of metabolic potential (Fig. 2g). EVs properties, like selective loading, physicochemical protection, and spatial aggregation, define a robust conduit for maintaining QS competence where free signals rarely reach activity levels.
The QS communication network in natural aquatic ecosystem is extremely complex. In previous studies, AHLs exist only in Proteobacteria48, and AIPs are found mostly in Firmicutes49. Surprisingly, there were both AHLs and AIPs in Actinobacteria, Bacteroidota, Planctomycetota, and Cyanobacteria in XLR (Fig. 5a; Supplementary Data-QS datasets). Studies have shown that some of organisms contain multiple QS operons, but others only have “orphan” LuxR or LuxI homologs, which are termed as luxR solos or luxI solos17,18,20. There are ~76% of annotated LuxR proteins belong to the luxR solos in prokaryotes20, which regulate gene expression by binding to AHLs produced by luxI genes encoded elsewhere50,51, or independently of AHLs binding52. In our study, the only identified MAGs with LuxI/R system were all luxR solos (Fig. 5d), and the ubiquitous genus Synechococcus was almost all luxR solos (Fig. S23b). While incomplete MAGs may preclude absolute confirmation of gene absence, the consistency of recurrence of the luxR solos across multiple MAGs and high-quality NCBI genomes points to a widespread biological strategy for QS. The variety of commensal and competitive interactions among bacteria in natural communities is expected to create many situations in which reception or emission of a signal may be independently favored. Those luxR/I solos may be actively lost or obtained during evolution, which conforms to the Black Queen hypothesis53. EVs provide a mechanism to pair orphan receptors with exogenous AHLs, or conversely to deliver receptor proteins to luxI solos. These results predict broad eavesdropping and interspecific crosstalk in situ, consistent with the observed diversity of QS systems in the XLR. The nature of the autoinducers varies among species and among systems in the same species, and each pathway responds to its own autoinducer signal54. However, more and more evidence revealed the crosstalk of signal molecules among various species2,11,12,17,20,21,54, in which the specificity is difficult to adapt to complex environments. In this view, EVs offer a new way for crosstalk coordination in mixed communities.
Critically, EV production in the field was jointly shaped by both biotic and abiotic factors. At the ecosystem scale, moderate producers were found to disproportionately contribute to β-diversity, niche breadth, network robustness, and core ecological functions, suggesting a strategic role for mid-intensity vesiculation in stabilizing interactions under environmental variability. Taken together, these findings suggest that the environmental modulation of EV fluxes can serve as a potential indicator that effectively mirrors the intricate dynamic status of aquatic ecosystems.
In sum, EVs provide a field-validated solution to the central problem of QS under environmental stresses: they load, protect, concentrate, and deliver hydrophobic signals to the right recipients over relevant space-time scales. EVs and their environmentally moderate producers function as hubs of ecological regulation. These findings elevate EVs to active architects of microbial communication and ecosystem functions in aquatic ecosystems.
Methods
Study area and sampling strategy
The sampling strategy was designed to encompass both spatial heterogeneity across urbanization gradients in a watershed and temporal dynamics within a reservoir with a high level of urbanization located in the downstream of the watershed. The study proceeded in two distinct phases:
Watershed-scale survey
The microbiome and field EV analysis were conducted in the Houxi River watershed (HRW) (24°34′-24°46′N, 117°55′-118°7′E) in December 2021, which was located in Xiamen City, Fujian Province, southeast China. The land cover data with 10-meter resolution was obtained from ESRI’s comprehensive datasets as of June 24, 2021 (https://livingatlas.arcgis.com/landcover) and classified the watershed into five primary categories: cultivated land, forestland, grassland, water bodies, and artificial surfaces (Fig. S2). HRW had different urbanization levels from upstream to downstream, and surface water samples (0–50 cm) were collected.
Seasonal dynamics in urban reservoir
Based on the survey, the downstream of HRW, XLR (24°34′-24°36′N, 118°03′-118°04′E), was selected for intensive temporal monitoring. The region is characterized by a subtropical monsoon climate, and the autumn is notably brief and transitions rapidly into the winter. Consequently, to capture representative seasonal changes, surface water samples were collected in April, August, and December of 2022, which represent spring, summer, and winter, respectively. This temporal coverage allows effective investigating microbiota and their EVs dynamics across the full range of representative environmental gradients characterizing this aquatic ecosystem.
Determination of environmental factors
A total of 16 environmental factors were measured. Water temperature (Tem), pH, dissolved oxygen (DO), turbidity (Tur), electrical conductivity (Con), salinity (Sal), chlorophyll a (chl-a), and oxidation-reduction potential (ORP) were measured in situ with a Hydrolab DS5 multiparameter water quality analyzer (Hach Company, Loveland, CO, USA). Total carbon (TC), total organic carbon (TOC), total nitrogen (TN), ammonium nitrogen (NH4_N), nitrate nitrogen (NO3_N), nitrite nitrogen (NO2_N), total phosphorus (TP) and phosphate phosphorus (PO4_P) were measured following a standard method55. Water transparency (Trans) was assessed with a Secchi disk. For algae counting, 1 L water samples were immediately fixed with Lugol’s solution and kept in the dark at room temperature until further processing56.
Isolation, identification, and culture of bacterial strains from Xinglinwan Reservoir
Bacteria were isolated from XLR by spreading 100 μL of water sample onto LB or 2216E agar plates. The plates were incubated at 28 °C (water temperature when sampling) and single colony with unique morphologies was re-streaked three times to eliminated cross-contamination. Colony PCR by universal primers (27F, 1492R) for 16S rRNA gene was performed, and the PCR products were purified and sequenced by Sanger sequencing (Sangon Biotech, Shanghai). The sequences were aligned using the BLAST tool on the NCBI to determine the closest match. The isolated bacteria were cultured in LB or 2216E media at 28 °C, 180 rpm. Microcystis aeruginosa FACHB-905 and Synechococcus sp. FACHB-1061 were from Freshwater Algae Culture Collection at the Institute of Hydrobiology (FACHB) and cultured them in BG-11 media under 12 h constant light flux and 12 h dark at 25 °C.
Biosensor-based quorum-sensing assays
All the 95 isolated bacteria were checked with the ability of producing AHLs signal molecules using Chromobacterium violaceum CVO26 (CVO26), Agrobacterium tumefaciens A136 (A136), and Agrobacterium tumefaciens KYC55 (KYC55) as the biosensors. CVO26 shows purple color for the positive detection of C4- and C6-HSL. A136 shows blue for 3-oxo-C4- to 3-oxo-C12-HSL and C5- to C10-HSL. KYC55 shows blue for 3-oxo-C4- to 3-oxo-C18-HSL and C4- to C18-HSL38.
Extraction and determination of AHLs from field and strains
We used ethyl acetate to extract AHLs in the liquid and then detected them using LC-MS/MS57. Briefly, the supernatants from field samples and bacterial culture were extracted twice with equal volumes of ethyl acetate, and the mixture was vortex-mixed for 1 h each time. The organic phase was collected after standing for 15 min and was dried under nitrogen gas. The dried samples were re-suspended in methyl alcohol with 0.1% formic acid and passed through 0.22 μm pinhole membrane filters (PTFE, JINTENG) to obtain the crude extract and were stored at −20 °C until detection.
Chromatographic separation was performed using a Kinetex C-18 column (2.6 μm, 2.1 × 100 mm). The eluate was subsequently analyzed by an ABI 6500 mass spectrometer (Applied Biosystems, USA) operating in multiple reaction monitoring mode, with mobile phase A of 0.1% FA in pure water and mobile phase B of 100% methyl alcohol, respectively. The AHLs were eluted at a flow rate of 300 µL/min by gradient elution of mobile phase increasing to 20% B in 0.01 min; then increased to 90% B in 4 min and held for 5 min; then decreased to 20% B at 6 min, and held for 3 min, for a total run of 9 min. The injection volume was 10 µL and the column was maintained at 40 °C. We conducted Recovery rate experiments to validate our extraction efficiency. Specifically, standard mixtures of AHLs were spiked into the field-sterile water matrix and EV samples at defined concentrations and processed using the identical ethyl acetate liquid-liquid extraction workflow as the field and EV samples. The results showed that the Recovery rates were between 57% and 94%.
We chose 13 different kinds of AHLs, which are representative and well studied7,57. The AHLs we used as standards are C4-HSL (N-Butyryl-L-homoserine lactone), C6-HSL (N-Hexanoyl-L-homoserine lactone), C8-HSL (N-Octanoyl-L-homoserine lactone), C10-HSL (N-Decanoyl-L-homoserine lactone), C12-HSL (N-Dodecanoyl-L-homoserine lactone), C14-HSL (N-Tetradecanoyl-L-homoserine lactone), C16-HSL (N-Hexadecanoyl-L-homoserine lactone), 3-oxo-C6-HSL (N-(3-Oxohexanoyl)-L-homoserine lactone), 3-oxo-C8-HSL (N-(3-Oxooctanoyl)-L-homoserine lactone), 3-oxo-C10-HSL (N-(3-Oxodecanoyl)-L-homoserine lactone), 3-oxo-C12-HSL (N-(3-Oxododecanoyl)-L-homoserine lactone), 3-oxo-C14-HSL (N-(3-Oxotetradecanoyl)-L-homoserine lactone), and 3-oxo-C16-HSL (N-(3-Oxohexadecanoyl)-L-homoserine lactone). Specifically, for each identified AHL structure, the octanol-water partition coefficient (Kow) was retrieved from standard chemical property databases (PubChem, ChemSpider).
For quantification, standard curves were constructed using synthetic AHL standards at concentrations of 0.002, 0.005, 0.01, 0.02, and 0.1 µg/mL. The concentrations of AHLs in water and EV samples were calculated based on the linear regression equations derived from the peak areas of these external standards (R2 > 0.99).
Extraction of crude EVs
EVs from field water samples (~25–30 L) were isolated via fractional filtration through 3, 0.8, 0.45, and 0.2 μm polycarbonate membranes, followed by concentration to ~200 mL using tangential flow filtration (TFF, Millipore membrane, 100 kDa cutoff). The concentrated filtrate was passed through a 0.2 μm syringe (Millipore) filter to remove residual cells and pelleted by ultracentrifugation at 100,000 × g for 2 h at 4 °C (Beckman-counter SW32 Ti rotor). The EV pellet was resuspended in PBS and stored at –20 °C.
For bacterial cultures, EVs were harvested from late-log phase supernatants after centrifugation at 10,000 × g to remove cells, followed by 0.45 and 0.2 μm filtration, concentration, and ultracentrifugation using the same protocol.
Purification of EVs
Purification of EVs employed Optiprep (Iodixanol; SigmaAldrich) density gradients tailored to the sample origin. Field EV pellets were layered over continuous 5–55% OptiPrep gradients in buffer matching reservoir salinity (4.2% NaCl, 10 mM HEPES, pH8), whereas cultured EVs were applied to stepped gradients (5–40%) adjusted to culture salinity (1% NaCl, 10 mM HEPES, pH7.4). Gradients were centrifuged at 100,000 × g for 16 h at 4 °C. EV-containing fractions, identified by particle abundance and purity metrics (Fig. S1), were collected, diluted >5-fold with PBS, and pelleted by repeated ultracentrifugation. This protocol ensured high-purity EV preparations for downstream analyses.
Extraction and determination of AHLs from EVs
To accurately quantify EV-associated AHLs, EVs were resuspended in 3–5 times volume of methanol and thoroughly vortexed to ensure complete membrane disruption and cargo release prior to LC-MS/MS measurement, and then dried under nitrogen gas58. The other detection procedures are the same as those for isolated strains.
The stability assays of EV-associated AHLs
To investigate the protection of EVs on AHLs at different pH, we used field water from XLR to incubate EVs of strain XL10. Specifically, the water samples from XLR were passed through 0.2 µm filter membrane and ultracentrifuged at 16 h (100,000 × g, 4 °C) to remove the EVs in it. The pH of the supernatant was adjusted to 8 and 10, respectively. After extracting EVs of XL10, we used the field water from XLR to wash and dissolve EVs. Finally, we regularly monitor the concentration of C14-HSL in EVs.
The kinetics of AHLs hydrolysis
AHLs hydrolysis follows pseudo-first order kinetics59 using Eq. (1) below:
\({{\text{C}}}_{0}\): The initial concentration of AHLs \({{\text{C}}}_{{\text{t}}}\): The concentration at time t k: the decay rate constant (hydrolysis rates)
Pseudo-first order regression analysis of R2
The AHLs half-lives t1/2 is calculated using Eq. (2) below:
Quantitation and physicochemical characterization of EVs
The EV particle concentration and size distribution were measured using a nanoflow cytometry (N30, NanoFCM, Xiamen, China). Protein content of EV samples was determined using the Micro BCA Protein Assay Kit (Pierce). The concentrations were then normalized to the total volume of the original water samples.
The zeta potential of EVs was tested using Zetasizer Nano (Malvern, UK).
Electron microscopy of EVs
Transmission electron microscopy (TEM) was employed by placing EVs onto copper grids for 3 min, negatively staining with 2% uranyl acetate for 50 s, and air drying. Imaging was performed using a Hitachi H-7650 electron microscope (100 kV) equipped with an Advanced Microscopy Techniques digital camera. For scanning electron microscopy (SEM), cells were fixed in 2.5% glutaraldehyde (4 °C, 12 h), rinsed in 0.1 M PBS, sequentially dehydrated in an ethanol series (30–100%), freeze-dried, sputter-coated, and examined with a Hitachi S4800 SEM.
Microcosmic experiments setup
Field water from XLR was treated with PBS (group E), XL92 (Flavobacterium sp.) (group A), intact EVs from XL92 (group B), ruptured EVs (High temperature (105 °C, 30 min)) (group C), XL92 supernatant depleted of EVs (group D). Triplicates were performed for each treatment. The microcosms were run for 7 days at room temperature under conditions consistent with natural circadian rhythms. To simulate wave-induced mixing, the microcosms were gently shaken every 12 h to ensure homogenization and oxygenation. Amplicon sequencing was then performed to determine the water microbial composition.
Labeling and tracking EV interactions with organisms and biofilm
EVs (derived from XL92 (Flavobacterium sp.)) and microorganisms (Microcystis aeruginosa FACHB-905, Synechococcus sp. FACHB-1061, and aquatic organisms from XLR) were labeled with DiO (VybranTM), FM4-64 (MCE) or Hoechst 33342 (MCE) in accordance with the manufacturer’s instructions. Briefly, 1×107 cells were incubated for 1 h at 28 °C with DiO-labeled EVs. After two times of washing with PBS, the cells were visualized using a confocal microscope (Carl Zeiss AG, Germany). Cells with positive fluorescence (group P) were sorted via flow cytometry, and then applied to DNA extraction and 16S rRNA and 18S rRNA gene sequencing for taxonomic identification. At the same time, the biological negative controls were performed, in which microorganisms were incubated with PBS (buffer) without EVs (Fig. S7d).
Biofilm was cultivated on sterile slides, which were placed in XLR until they matured (at least 6 days)60. These slides were washed by pipetting PBS gently to remove all attached and free aquatic organisms. The biofilm was stained using SYTO 9 (Life Technologies-Invitrogen) according to the manufacturer’s instructions. Afterwards, the biofilm was incubated with FM4-64-labeled EVs for 1 h at 28 °C. After two times of washing with PBS, the slide was visualized using a confocal microscope, and the 3D images were obtained by scanning biofilm at 0.5 μm intervals along the Z axis.
Microbiota 16S and 18S rRNA gene sequencing
To identify bacterial taxa that preferentially take up EVs, DiO-labeled EVs derived from the isolated XL92 (Flavobacterium sp.) were incubated with the natural microbiota collected from XLR. After washing two times with PBS, the DiO-positive microbiota from field water samples were sorted using a Moflo-XDP cell sorter (Beckman, USA). The total DNA was extracted specifically from the sorted cell populations with QIAamp DNA kits (QIAGEN, Germany), following the manufacture’s process. Prokaryotes and eukaryotes were investigated using 16S (V3-V4 region, primer: 341F: CCTACGGGAGGCAGCAG; 806 R: GGACTACHVGGGTWTCTAAT61) and 18S (V9 region, primer: 1380F: CCCTGCCHTTTGTACACAC; 1510 R: CCTTCYGCAGGTTCACCTAC61) rRNA gene sequencing. The resulting PCR amplicons were purified using 1% agarose gel electrophoresis. Sequencing libraries were established using the NEBNext® Ultra™ II DNA Library Prep Kit for Illumina® (New England Biolabs, USA) and sequenced on the Illumina Nova 6000 (Guangdong Magigene Biotechnology Co., Ltd., Guangzhou, China).
Quantification of EVs during the bacterial growth cycle
To quantify EV production rates in bacterial cultures under different environmental conditions, triplicate bacterial cultures were prepared as described previously. Samples (1 mL) were collected every two hours and fixed with 0.125% (final concentration) glutaraldehyde for 10 min in the dark for flow cytometry analysis. To isolate EVs, 1 mL of culture was gently filtered through a 0.2 μm syringe filter (Millipore) to remove bacterial cells. The EV particle number in filtrates was examined by NanoFCM immediately.
Extraction and sequencing of metagenomic DNA
Total DNA was extracted from water sample filters and EVs using Qiagen kits (Qiagen, Germany) according to the manufacturer’s protocols. The EV-associated DNA was pretreated with DNase as previously24. DNA yield and quality were quantified via a Qubit HS dsDNA kit using a Qubit 4.0 Fluorometer (Thermo Fisher Scientific, USA).
For microbial community profiling (HRW samples), the V3-V4 regions of 16S rRNA genes (341 F/806 R) and the V9 region of 18S rRNA genes (1380 F/1510 R)61 were sequenced using the Illumina NovaSeq 6000 platform (Illumina Inc., San Diego, CA, USA).
For samples from XLR, the artificial spike-in sequences were added to the sample DNA pools for absolute quantitative amplicon sequencing62, and the Illumina NovaSeq 6000 sequencer was used for sequencing. The shotgun metagenomic sequencing was used for samples of XLR. Illumina NovaSeq 6000 platform was adopted to construct Illumina shotgun DNA library and sequencing. Briefly, DNA extraction was fragmented to an average size of about 350 bp for paired-end library construction. Paired-end library was constructed using NEXTFLEX Rapid DNA-Seq (Bioo Scientific, Austin, TX, USA). For the processing of shotgun metagenome sequencing data, the Transcripts Per Million (TPM) method was used to normalize the abundance. Taxonomic annotations and other functions were aligned to NR database, KEGG database, and CAZy database.
Metagenome binning
High-quality contigs (>1000 bp) from metagenomic assemblies were binned into MAGs using MetaBAT (v2.12.1), CONCOCT (v0.5.0), and MaxBin (v2.2.5)63,64. Finally, 389 MAGs were dereplicated (completeness ≥50% and contamination < 10%). MAG abundance within samples was calculated using CoverM (v0.6.1). Phylogenetic reconstruction was performed using GTDB-Tk (v2.3.0), with resultant trees visualized in iTOL. Comprehensive functional annotation of MAGs was achieved by comparison to public reference databases (NR, KEGG, CAZy). Functional, phylogenetic, and abundance statistics were integrated for downstream analyses.
Real-time quantitative PCR (qPCR) was used to quantify the number of 18S rRNA and 16S rRNA gene copies from HRW using a LightCycler 480 instrument (Roche, Basel, Switzerland)61.
Classification of aquatic organisms with different EV levels
We observed significant differences in the reads number obtained from DNA sequencing of aquatic organisms compared to those associated with EVs from the same species (Fig. S11). Specifically, based on the presence of species predicted by the sequencing data, we found that some species were exclusive to EV-associated DNA, some species were exclusive to aquatic organisms DNA, and some species were present in both datasets.
For the species that were exclusively identified in EV-associated DNA, we propose four potential explanations. First, EVs could effectively preserve the DNA of deceased organisms, as well as species that were not easily collected via filtration methods. Second, EVs could be introduced from organisms locating on elsewhere. Third, discrepancies may arise due to sequencing errors or instability. Lastly, the EVs from some species do not contain DNA.
In our study, we designated species identified solely in aquatic organism DNA sequencing as “aquatic organisms with undetected EVs” (UDAs), while those identified in both datasets were termed “aquatic organisms with detected EVs” (DAs). Additionally, the Spearman correlation between EV reads and species OTU reads corresponding to each OTU at each sampling point in prokaryotes and eukaryotes were both significantly positive (Fig. S12). This result indicated that, to a certain extent, the more of species, the higher the EV levels they have. Therefore, among the DAs, we further categorized them based on the comparative abundance of each species in the two datasets. Specifically, the abundance of species i in EV-associated DNA sequencing is denoted as (EVs-reads)i, while in aquatic organism sequencing was named as (Aquatic organisms-reads)i. When (EVs-reads)i ≥ (Aquatic organism-reads)i, it indicates that species i produces a higher quantity of EVs, and these are classified as “aquatic organisms with high EV levels” (H-DAs). Conversely, when (EVs-reads)i < (Aquatic organism-reads)i, it means lower EV production by species i, and these are classified as “aquatic organisms with low EV levels” (L-DAs). It is important to note that the EV production is affected by various biotic and abiotic factors. The terminology used here is specific to the spatiotemporal context of our sampling and may not be applicable under all conditions.
Co-occurrence network construction
Prior to network construction, the OTU (Operational Taxonomic Unit) abundance profiles obtained from the 16S and 18S rRNA gene amplicon sequencing of the HRW samples were filtered based on their frequency and abundance. OTUs with a presence rate below 18% were removed. Additionally, OTUs with ≤10 sequences were excluded. Subsequently, all possible pairwise Spearman’s correlation (r) between OTUs were calculated using the psych R package. Only robust correlations (|r| > 0.8, P-value < 0.05, adjusted by fdr method) were incorporated into the network analyses. Network visualization was performed using Gephi version 0.9.7 (https://gephi.org). Network topological properties were calculated using the igraph R package. The stability of the co-occurrence networks was indicated by robustness and vulnerability. For network stability analysis, robustness was calculated to simulate targeted removing65. We measured the robustness when 90% of target nodes were removed, and the vulnerability was indicated by the maximal vulnerability of nodes in the network which were measured using the relative contribution of the node to the global efficiency.
Metaproteomics analysis of EVs
For metaproteomic profiling of the field EVs derived from XLR, 100 μg of EV protein was first assessed through sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) to verify sample quality. EV proteins were enzymatically digested with trypsin, and the resulting peptides were fractionated by high-pH reversed-phase ultra-high performance liquid chromatography (UHPLC) using a Vanquish Flx UHPLC system (Thermo Scientific). Peptides were separated using a gradient of 0% buffer B (80% acetonitrile) for 16 min, 3.8%-43% buffer B for 21 min, and reaching 100% buffer B for 1 min. After separation, peptide concentrations were measured using the NANO DROP ONE (Thermo Scientific), and equal quantities from each sample were combined for tandem mass spectrometry (LC-MS/MS) analysis on a Vanquish Neo system (Thermo Scientific), operating with data-dependent acquisition (DDA) mode under Thermo Xcalibur 4.7. Peptide libraries were generated using Spectronaut™ software to facilitate downstream data-independent acquisition (DIA) quantification. The DIA mass spectrometric raw data were searched and processed with the Spectronaut software (Version 19) software against the non-redundant metagenomic-constructed database from XLR metagenome. The parameters are as follows up: The peptide length range was set to 7-52; Enzyme cutting site was trypsin/P; The maximum missed cleavage site was 2; Carbamidomethylation of cysteines as fixed modification, and oxidation of methionines and protein N-terminal acetylation as variable modifications; Protein FDR ≤ 0.01, Peptide FDR ≤ 0.01, Peptide Confidence ≥99%, XIC width ≤75ppm. The protein quantification method was MaxLFQ.
Construction of quorum quorum-sensing database of Xinglinwan Reservoir
The overall workflow of the Quorum sensing (QS) gene, species collection and database construction were summarized in Fig. S20. The autoinducer synthases and receptors are from SigMol (http://bioinfo.imtech.res.in/manojk/sigmol), Quorumpeps (http://quorumpeps.ugent.be), and QSDB (https://qsdb.uni-konstanz.de/). We integrated the species in the three QS databases and the results of local BLASTP using the metagenomic sequencing data (identity >70%, E-value < 1e-5). We compiled the QS species from BLASTP and species from three public QS database (SigMol, Quorumpeps, and QSDB) to form the QS species inventory of XLR.
Structural equation modeling
To investigate the relationship among the environmental factors, EV particle numbers, and the aquatic organisms, structural equation modeling analysis was performed using lavaan package66. We filtered the environmental factors which had strong collinearity (variance inflation factor >10), using vif function in car package (version, 3.1-3). The filtered environmental factors included water temperature, pH, DO, chl-a, salinity, and TN:TP. To evaluate the model’s performance, we employed three commonly used indicators: standardized root mean squared residual (SRMR < 0.08 represents a qualified model, SRMR = 0.016 in our model) and goodness-of-fit index (GIF > 0.95 for satisfactory performance, GFI = 0.993 in our model)67.
Niche breadths calculating
We assessed the ecological adaptation potential on the community level regarding niche breadth. Levins’ niche breadth (Bj) was calculated using the formula:
Bj represents the habitat suitability for species j; N is the total number of species; Pij is the proportion of species j in sample i relative to its total abundance across all sites. Niche breadths were calculated using niche.width function from the spaa R package68.
Statistical analyses
The phylogenetic analysis of the isolated strains was based on 16S rRNA gene sequences and used MEGA11 (Version 11.0.13). Briefly, we calculated the clustering using the Neighbor-Joining (NJ) method for the distance matrix. Bootstrap values (based on 1000 replicates) were calculated to assess the robustness of the branches. The differentially abundant taxa among seasons (biomarkers) were discovered by way of linear discriminant analysis effect size analysis (LEfSe)69 in R. Taxa with an LDA score >5.0 and P < 0.05 were considered significant discriminative features. Alpha diversity, beta diversity, and the similarity percentage analysis (SIMPER) were calculated using the diversity function in the “vegan” package (https://github.com/vegandevs/vegan). Hierarchical clustering was performed continuously based on Bray–Curtis distances using vegan package in R. Student’s t test and Wilcoxon test was used to evaluate the significance of the results. We performed the redundancy analysis (RDA) to show the community trait response to environmental gradients70. We used the hierarchical partitioning analysis (HPA) to estimate the importance of each environmental factors. Hierarchical partitioning calculates the variable importance from all subset models, leading to an unordered assessment of importance71. The pathway enrichment analysis was performed using the R package ReporterScore72. We used the microbial source-tracking method fast expectation-maximization microbial source tracking (FEAST)73 to determine the potential origin of EVs. The relative contribution of taxa to functions using the method described by Ofek-Lalzar et al.74. Phylogenetic Investigation of Communities by Reconstruction of Unobserved States 2 (PICRUSt2) was performed for predicting function of prokaryotes in microcosmic experiment (https://github.com/picrust/picrust2) using R software. All statistical analyses were performed in R software (v4.2.3) (https://www.r-project.org). We download the genome of Synechococcus from NCBI (https://www.ncbi.nlm.nih.gov/datasets/genome/). To identify if they have QS related genes, we compared the genomes with the collected autoinducer synthases and receptors using local BLASTP (identity >70%, E-value < 1e-5).
Data availability
All the DNA sequences generated during the current study are available in the BIG Submission75, BIG Sub: https://ngdc.cncb.ac.cn/gsa/browse/CRA027247. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (https://proteomecentral.proteomexchange.org) via the iProX partner repository76,77 with the dataset identifier PXD065377.
References
Mukherjee, S. & Bassler, B. L. Bacterial quorum sensing in complex and dynamically changing environments. Nat. Rev. Microbiol. 17, 371–382 (2019).
Coolahan, M. & Whalen, K. E. A review of quorum-sensing and its role in mediating interkingdom interactions in the ocean. Commun. Biol. 8, 179 (2025).
Decho, A. W., Norman, R. S. & Visscher, P. T. Quorum sensing in natural environments: emerging views from microbial mats. Trends Microbiol. 18, 73–80 (2010).
Papenfort, K. & Bassler, B. L. Quorum-sensing signal-response systems in Gram-negative bacteria. Nat. Rev. Microbiol. 14, 576–588 (2016).
Abisado R.G. et al. Bacterial quorum sensing and microbial community interactions. mBio. 9, (2018).
Hmelo, L. R. Quorum sensing in marine microbial environments. Ann. Rev. Mar. Sci. 9, 257–281 (2017).
Horswill, A. R., Stoodley, P., Stewart, P. S. & Parsek, M. R. The effect of the chemical, biological, and physical environment on quorum sensing in structured microbial communities. Anal. Bioanal. Chem. 387, 371–380 (2007).
Wei, J. R. et al. A mobile quorum-sensing system in Serratia marcescens. J. Bacteriol. 188, 1518–1525 (2006).
Yates, E. A. et al. N-acylhomoserine lactones undergo lactonolysis in a pH-, temperature-, and acyl chain length-dependent manner during growth of Yersinia pseudotuberculosis and Pseudomonas aeruginosa. Infect. Immun. 70, 5635–5646 (2002).
Wellington, S. & Greenberg, E. P. Quorum sensing signal selectivity and the potential for interspecies cross talk. mBio 10, e00146–00119 (2019).
Slinger, B. L., Banerjee, S., Chandler, J. R. & Blackwell, H. E. Interspecies crosstalk via LuxI/LuxR-type quorum sensing pathways contributes to decreased nematode survival in coinfections of Pseudomonas aeruginosa and Burkholderia multivorans. ACS Chem. Biol. 19, 2557–2568 (2024).
Wu, F., Menn, D. J. & Wang, X. Quorum-sensing crosstalk-driven synthetic circuits: from unimodality to trimodality. Chem. Biol. 21, 1629–1638 (2014).
Arashida, N. et al. Identification of novel long chain N-acylhomoserine lactones of chain length C(20) from the marine phototrophic bacterium Rhodovulum sulfidophilum. Biosci. Biotechnol. Biochem. 82, 1683–1693 (2018).
Decho, A. W. et al. Autoinducers extracted from microbial mats reveal a surprising diversity of N-acylhomoserine lactones (AHLs) and abundance changes that may relate to diel pH. Environ. Microbiol. 11, 409–420 (2009).
Wang, Y. C. et al. Diversity, influential factor, and communication network construction of quorum-sensing bacteria in global wastewater treatment plants. Water Res. 279, 123437 (2025).
Wu, S. et al. Machine learning aided construction of the quorum-sensing communication network for human gut microbiota. Nat. Commun. 13, 3079 (2022).
Albataineh, H., Duke, M., Misra, S. K., Sharp, J. S. & Stevens, D. C. Identification of a solo acylhomoserine lactone synthase from the myxobacterium Archangium gephyra. Sci. Rep. 11, 3018 (2021).
Fuqua, C. The QscR quorum-sensing regulon of Pseudomonas aeruginosa: an orphan claims its identity. J. Bacteriol. 188, 3169–3171 (2006).
Roche, D. M. et al. Communications blackout? Do N-acylhomoserine-lactone-degrading enzymes have any role in quorum sensing? Microbiology 150, 2023–2028 (2004).
Hudaiberdiev, S. et al. Census of solo LuxR genes in prokaryotic genomes. Front. Cell Infect. Microbiol. 5, 20 (2015).
Zan, J. et al. A solo luxI-type gene directs acylhomoserine lactone synthesis and contributes to motility control in the marine sponge symbiont Ruegeria sp. KLH11. Microbiology 161, 50–56 (2015).
Chandler, J. R., Heilmann, S., Mittler, J. E. & Greenberg, E. P. Acyl-homoserine lactone-dependent eavesdropping promotes competition in a laboratory co-culture model. ISME J. 6, 2219–2228 (2012).
Toyofuku, M., Schild, S., Kaparakis-Liaskos, M. & Eberl, L. Composition and functions of bacterial membrane vesicles. Nat. Rev. Microbiol. 21, 415–430 (2023).
Zhu, L. T. et al. Diverse functional genes harboured in extracellular vesicles from environmental and human microbiota. J. Extracell. Vesicles 11, e12292 (2022).
Biller, S. J. et al. Bacterial vesicles in marine ecosystems. Science 343, 183–186 (2014).
Qin, Y. et al. Widespread of potential pathogen-derived extracellular vesicles carrying antibiotic resistance genes in indoor dust. Environ. Sci. Technol. 56, 5653–5663 (2022).
Long, L. et al. Extracellular vesicles are prevalent and effective carriers of environmental allergens in indoor dust. Environ. Sci. Technol. 59, 1969–1983 (2025).
Teng, Y. et al. Plant-derived exosomal micrornas shape the gut microbiota. Cell Host Microbe 24, 637–652.e638 (2018).
Gaudin, M. et al. Hyperthermophilic archaea produce membrane vesicles that can transfer DNA. Environ. Microbiol. Rep. 5, 109–116 (2013).
Wang, Z. et al. Fungal extracellular vesicles mediate cross-kingdom trafficking of virulence effectors into plant cells to promote infection. Mol. Plant 18, 1369–1389 (2025).
Biller, S. J. et al. Prochlorococcus extracellular vesicles: molecular composition and adsorption to diverse microbes. Environ. Microbiol. 24, 420–435 (2022).
Hackl, T. et al. Novel integrative elements and genomic plasticity in ocean ecosystems. Cell 186, 47–62.e16 (2023).
Toyofuku, M. et al. Membrane vesicle-mediated bacterial communication. ISME J. 11, 1504–1509 (2017).
Mashburn, L. M. & Whiteley, M. Membrane vesicles traffic signals and facilitate group activities in a prokaryote. Nature 437, 422–425 (2005).
Emge, P. et al. Resilience of bacterial quorum sensing against fluid flow. Sci. Rep. 6, 33115 (2016).
Wang, J. et al. AHLs-mediated quorum sensing threshold and its response towards initial adhesion of wastewater biofilms. Water Res. 194, 116925 (2021).
Zhu, Y. L., Hou, H. M., Zhang, G. L., Wang, Y. F. & Hao, H. S. AHLs regulate biofilm formation and swimming motility of Hafnia alvei H4. Front Microbiol. 10, 1330 (2019).
Chu, W., Vattem, D. A., Maitin, V., Barnes, M. B. & McLean, R. J. Bioassays of quorum sensing compounds using Agrobacterium tumefaciens and Chromobacterium violaceum. Methods Mol. Biol. 692, 3–19 (2011).
Agmo Hernández V. An overview of surface forces and the DLVO theory. ChemTexts. 9 (2023).
Hmelo, L. & Van Mooy, B. A. S. Kinetic constraints on acylated homoserine lactone-based quorum sensing in marine environments. Aquat. Microb. Ecol. 54, 127–133 (2009).
Pearson, J. P., Delden, C. V. & Iglewski, B. H. Active efflux and diffusion are involved in transport of Pseudomonas aeruginosa cell-to-cell signals. J. Bacteriol. 181, 1203–1210 (1999).
Chan, Y. Y. et al. Control of quorum sensing by a Burkholderia pseudomallei multidrug efflux pump. J. Bacteriol. 189, 4320–4324 (2007).
Buroni, S. et al. Assessment of three Resistance-Nodulation-Cell Division drug efflux transporters of Burkholderia cenocepacia in intrinsic antibiotic resistance. BMC Microbiol. 9, 200 (2009).
Gantner, S. et al. In situ quantitation of the spatial scale of calling distances and population density-independent N-acylhomoserine lactone-mediated communication by rhizobacteria colonized on plant roots. FEMS Microbiol. Ecol. 56, 188–194 (2006).
MacDonald, K. L. & Beveridge, T. J. Bactericidal effect of gentamicin-induced membrane vesicles derived from Pseudomonas aeruginosa PAO1 on gram-positive bacteria. Can. J. Microbiol. 48, 810–820 (2002).
Tashiro, Y. et al. Interaction of bacterial membrane vesicles with specific species and their potential for delivery to target cells. Front. Microbiol. 8, 571 (2017).
Schatz, D. et al. Communication via extracellular vesicles enhances viral infection of a cosmopolitan alga. Nat. Microbiol. 2, 1485–1492 (2017).
Case, R. J., Labbate, M. & Kjelleberg, S. AHL-driven quorum-sensing circuits: their frequency and function among the Proteobacteria. ISME J. 2, 345–349 (2008).
Monnet, V. & Gardan, R. Quorum-sensing regulators in Gram-positive bacteria: ‘cherchez le peptide. Mol. Microbiol. 97, 181–184 (2015).
Lequette, Y., Lee, J. H., Ledgham, F., Lazdunski, A. & Greenberg, E. P. A distinct QscR regulon in the Pseudomonas aeruginosa quorum-sensing circuit. J. Bacteriol. 188, 3365–3370 (2006).
McIntosh, M., Krol, E. & Becker, A. Competitive and cooperative effects in quorum-sensing-regulated galactoglucan biosynthesis in Sinorhizobium meliloti. J. Bacteriol. 190, 5308–5317 (2008).
Subramoni, S. & Venturi, V. LuxR-family ‘solos’: bachelor sensors/regulators of signalling molecules. Microbiology 155, 1377–1385 (2009).
Morris JJ, Lenski RE, Zinser ER. The Black Queen Hypothesis: evolution of dependencies through adaptive gene loss. mBio. 3, (2012).
Lerat, E. & Moran, N. A. The evolutionary history of quorum-sensing systems in bacteria. Mol. Biol. Evol. 21, 903–913 (2004).
Rice EW, Baird RB, Eaton AD. Standard Methods for the Examination of Water and Wastewater, 23rd Edition (2017).
Lv, H. et al. Temperature and nutrients are significant drivers of seasonal shift in phytoplankton community from a drinking water reservoir, subtropical China. Environ. Sci. Pollut. Res. Int. 21, 5917–5928 (2014).
He, W., Yang, H., Wang, X., Li, H. & Dong, Q. Growth of Salmonella Enteritidis in the presence of quorum sensing signaling compounds produced by Pseudomonas aeruginosa. Int. J. Food Eng. 17, 959–968 (2021).
Fletcher, M. P., Diggle, S. P., Camara, M. & Williams, P. Biosensor-based assays for PQS, HHQ and related 2-alkyl-4-quinolone quorum-sensing signal molecules. Nat. Protoc. 2, 1254–1262 (2007).
Ziegler, E. W., Brown, A. B., Nesnas, N. & Palmer, A. G. Abiotic hydrolysis kinetics of N-Acyl-L-homoserine Lactones: natural silencing of bacterial quorum sensing signals. Eur. J. Org. Chem. 2019, 2850–2856 (2019).
Huang, Y. L., Ki, J. S., Lee, O. O. & Qian, P. Y. Evidence for the dynamics of Acyl homoserine lactone and AHL-producing bacteria during subtidal biofilm formation. ISME J. 3, 296–304 (2009).
Mo, Y. et al. Low shifts in salinity determined assembly processes and network stability of microeukaryotic plankton communities in a subtropical urban reservoir. Microbiome 9, 128 (2021).
Tkacz, A., Hortala, M. & Poole, P. S. Absolute quantitation of microbiota abundance in environmental samples. Microbiome 6, 110 (2018).
Nurk, S., Meleshko, D., Korobeynikov, A. & Pevzner, P. A. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 27, 824–834 (2017).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP-a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158 (2018).
Yuan, M. M. et al. Climate warming enhances microbial network complexity and stability. Nat. Clim. Change 11, 343–348 (2021).
Rosseel, Y. Lavaan: an R package for structural equation modeling. J. Stat. Software 48, 1–36 (2012).
Kenny, D. A., Kaniskan, B. & McCoach, D. B. The performance of RMSEA in models with small degrees of freedom. Sociol. Methods Res. 44, 486–507 (2014).
JL Z, KP M. spaa: an R package for computing species association and niche overlap. Research Progress of Biodiversity Conservation in China. 165–174 (2014).
Segata N, et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, (2011).
Kleyer, M. et al. Assessing species and community functional responses to environmental gradients: which multivariate methods? J. Veg. Sci. 23, 805–821 (2012).
Lai, J., Zou, Y., Zhang, J. & Peres-Neto, P. R. Generalizing hierarchical and variation partitioning in multiple regression and canonical analyses using the rdacca.hp R package. Methods Ecol. Evol. 13, 782–788 (2022).
Peng C, Chen Q, Tan S, Shen X, Jiang C. Generalized reporter score-based enrichment analysis for omics data. Brief Bioinform. 25, bbae116 (2024).
Shenhav, L. et al. FEAST: fast expectation-maximization for microbial source tracking. Nat. Methods 16, 627–632 (2019).
Ofek-Lalzar, M. et al. Niche and host-associated functional signatures of the root surface microbiome. Nat. Commun. 5, 4950 (2014).
Members & Partners, C.-N. Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2025. Nucleic Acids Res. 53, D30–D44 (2025).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2022).
Acknowledgements
This work was supported by National Natural Science Foundation of China (32161143016, 42177362), and National Basic Science Data Center “Environment Health DataBase” (NO. NBSDC-DB-21). We thank Prof. Weihua Chu from China Pharmaceutical University for his help in QS biosensor experiments. We thank Prof. Steven Biller for his advice and assistance on this paper.
Author information
Authors and Affiliations
Contributions
X.L.X. conducted the experiments, generated and analyzed the data, and wrote the manuscript. J.J.L. performed the microcosmic experiments. L.T.Z. helped with the data analysis and revised the manuscript. L.L. and Y.F.D. provided the suggestions for computational adjustments. L.F.L. and J.H. contributed to the AHLs testing experiments. H.H.C. helped with measuring environmental factors. Q.S.H. designed the experiments, wrote the manuscript, and got the funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, X., Lin, J., Zhu, LT. et al. Extracellular vesicles as structured vectors of quorum sensing signals influence aquatic microbial communities. npj Biofilms Microbiomes 12, 57 (2026). https://doi.org/10.1038/s41522-026-00924-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41522-026-00924-w